Abstract
Temperature is a key factor in determining plant growth and development, geographical distribution, and seasonal behavior. Plants accurately sense subtle changes in ambient temperature and alter their growth and development accordingly to improve their chances of survival and successful propagation. Thermomorphogenesis encompasses a variety of morphological changes that help plants acclimate to warm environmental temperatures. Revealing the molecular mechanism of thermomorphogenesis is important for breeding thermo-tolerant crops and ensuring food security under global climate change. Plant adaptation to elevated ambient temperature is regulated by multiple signaling pathways and epigenetic mechanisms such as histone modifications, histone variants, and non-coding RNAs. In this review, we summarize recent advances in the mechanism of epigenetic regulation during thermomorphogenesis with a focus on the model plant Arabidopsis thaliana and briefly discuss future prospects for this field.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Temperature plays an important role in plant growth and development and ambient temperature affects the entire plant life cycle (Ding et al. 2020; Lippmann et al. 2019). For example, high ambient temperatures accelerate plant growth, promote flowering, reduce seed production, and reduce disease resistance (Lee et al. 2020; Li et al. 2018). Therefore, changes in ambient temperature due to global climate change have major implications for agriculture and natural ecosystems. Indeed, the latest assessment from the UN’s Intergovernmental Panel on Climate Change (IPCC) indicates that global temperatures will continue to rise in the future (Tollefson 2021). Researchers have estimated that the yields of wheat, rice, maize, and soybean will decrease by 6%, 3.2%, 7.4%, and 3.1% for each degree-Celsius increase in global average temperature (Zhao et al. 2017). Therefore, exploring the mechanisms by which temperature regulates plant growth and development will facilitate the breeding of new crop varieties that tolerate high temperature to improve food security.
Plant growth and development are highly plastic, which helps plants survive the constantly changing external environmental temperatures. At high ambient temperature (below the heat-stress range), plants undergo adaptive growth, which leads to a variety of morphological changes termed thermomorphogenesis (Casal and Balasubramanian 2019; Quint et al. 2016). Typical thermomorphogenesis phenotypes of Arabidopsis thaliana include hypocotyl elongation, petiole lengthening, and hyponastic growth of petiole and leaf (Casal and Balasubramanian 2019; Quint et al. 2016).
Plants have evolved intricate and accurate systems for sensing temperature and transmission of temperature signals; these systems quickly sense changes in the surrounding environment and make corresponding adjustments, thus enhancing plant tolerance to ambient temperature. Thermomorphogenesis is regulated by the integration of signals from the ambient temperature, light, hormonal, and circadian clock pathways (Casal and Balasubramanian 2019; Ding et al. 2020). In addition, epigenetic regulatory mechanisms play an important role in plant temperature responses (Casal and Balasubramanian 2019; Chang et al. 2019; He et al. 2021a; Zhao et al. 2021).
Epigenetic regulatory mechanisms are complicated and diverse, including covalent modifications of DNA and histone tails, histone variants, and non-coding RNAs (Allis and Jenuwein 2016; Boquete et al. 2021). Epigenetic mechanisms regulate many crucial cellular processes including transcription and RNA metabolism, thus affecting growth and development (Gallusci et al. 2017; Martire and Banaszynski 2020; Ueda and Seki 2020). Many epigenetic regulatory factors and chromatin modifications are involved in the plant response to stress, which are important for plants to acclimate to harmful environmental conditions (Chang et al. 2019; He et al. 2021a; Meyer 2015; Ueda and Seki 2020). In this review, we discuss recent progress in understanding the epigenetic mechanisms that regulate thermomorphogenesis in Arabidopsis, potential research priorities, and their implications for crop breeding.
Histone modification
Various post-translational modifications of core histones, including methylation and acetylation, participate in the complex mechanisms regulating transcription in plants (He et al. 2021a; Ueda and Seki 2020). Histone methylation occurs primarily on the lysine (K) and arginine (R) residues of the core histone tail, including mono-, di, and tri-methylation of H3K4, K9, K27, and K36 (Grosselin et al. 2019; Ueda and Seki 2020). Histone methylation plays an important role in regulating the expression of temperature-sensitive genes, particularly methylation of H3K4, H3K36, and H3K27.
Histone methylation is associated with transcriptional activation in thermomorphogenesis
H3K4me3 and H3K4me2 are usually associated with gene activation (Zhang et al. 2009) and H3K4 methylation is dynamically regulated. JUMONJI C (JmjC) domain-containing proteins (JMJ) are a class of important histone demethylases (Lu et al. 2008). For example, the histone H3K4m3 demethylases JMJ14, JMJ15, and JMJ18 demethylate histone H3K4me3 associated with a cluster of genes, thus mediating their downregulation at warmer temperatures (Cui et al. 2021).
Other epigenetic regulators are also involved in regulating H3K4 methylation of specific genes in response to warm temperatures. The RNA-binding protein FLOWERING CONTROL LOCUS A (FCA) regulates H3K4me2 demethylation in FLOWERING LOCUS C (FLC) to promote flowering and restricts H3K4me2 on the auxin biosynthesis-related gene YUCCA8 (YUC8) promoter (Lee et al. 2014; Liu et al. 2007). PHYTOCHROME INTERACTING FACTOR 4 (PIF4), the basic helix-loop-helix transcription factor in Arabidopsis, plays a centre role in thermomorphogenesis (Koini et al. 2009). PIF4 promotes high temperature-induced hypocotyl elongation by binding to the promoter region of YUC8 to activate YUC8 expression (Sun et al. 2012). Moreover, FCA promotes the dissociation of PIF4 from the YUC8 promoter by modifying H3K4me2 level at YUC8 chromatin, thus limiting thermal acceleration of stem growth (Lee et al. 2014). The transcription factor SEUSS (SEU), a positive regulator of thermomorphogenesis, mediates the enrichment of H3K4me3 in the chromatin regions of YUC8 and INDOLE-3-ACETIC ACID INDUCIBLE 19 (IAA19) to promote hypocotyl elongation in warmer temperature (Huai et al. 2018). Chromatin remodeling factors such as INOSITOL REQUIRING 80 (INO80) have important roles in regulating H3K4 methylation response to temperature. The atino80 mutants displayed strong reductions of hypocotyl and petiole elongation under high ambient temperatures (Xue et al. 2021). The INO80 chromatin remodeling complex (INO80-C) promotes H3K4me3 deposition and transcript elongation at PIF4 targets to regulate gene expression in response to warm temperature (Xue et al. 2021).
Histone H3 lysine 36 trimethylation (H3K36me3) is an important marker for transcriptional elongation and positively regulates transcription rate (Wagner and Carpenter 2012). At warm temperatures, H3K36me3 levels tend to be higher and present in broader regions (Pajoro et al. 2017). H3K36me3 mediated by SET DOMAIN GROUP8 (SDG8) and SDG26 is required for temperature-induced alternative splicing, and mutations of SDG8 and SDG26 impair warmer temperature-induced flowering in Arabidopsis (Pajoro et al. 2017). MORF-RELATED GENE 1 (MRG1) and MRG2 are involved in H3K36me3 binding (Xu et al. 2014) and the mrg1-1mrg2-3 double mutants show reduced temperature-induced flowering, further demonstrating that H3K36 plays a regulatory role in plant responses to fluctuating ambient temperatures (Pajoro et al. 2017).
Histone methylation is associated with transcriptional inhibition in thermomorphogenesis
H3K27me3 marks are located in gene bodies of genes with high and low transcription rates and negatively regulate these genes under warm temperatures (Sidaway-Lee et al. 2014). JMJ13 and JMJ30/JMJ32 negatively regulate temperature-dependent flowering (Gan et al. 2014; Zheng et al. 2019). Warm temperature induces the expression of JMJ30 and JMJ13, thus facilitating their protein accumulation (Gan et al. 2014; Zheng et al. 2019). JMJ30 directly binds to the FLC locus, removing H3K27me3, promoting FLC expression, and thereby inhibiting warm temperature-induced flowering (Gan et al. 2014). JMJ13 has two homologs: EARLY FLOWERING 6 (ELF6/JMJ11) and RELATIVE OF EARLY FLOWERING 6 (REF6/JMJ12). REF6 recognizes a CTCTGYTY motif via its four Cys2His2 zinc fingers domains and is mainly involved in the demethylation of H3K27me2 and H3K27me3; moreover REF6 activity is affected by non-CG DNA methylation (Cui et al. 2016; Lu et al. 2011; Qiu et al. 2019). REF6 and HEAT SHOCK FACTOR A2 (HSFA2) form a heritable transcriptional feedback loop that regulates several processes, such as flowering and immunity, in response to heat stress (Liu et al. 2019). Recent studies have shown that hypocotyl elongation of ref6 mutants was inhibited under warm ambient temperatures and REF6 enzyme activity is essential for plant response to this process. REF6 and PIF4 synergistically activate the expression of thermo-responsive genes at warm ambient temperature, thereby participating in the regulation of thermomorphogenesis (He et al. 2021b). In addition, the ATP-dependent chromatin remodeling factor PICKLE (PKL) down regulates H3K27me3 at IAA19 and IAA29, promoting gene expression in a temperature-dependent manner (Zha et al. 2017). pkl mutations lead to reduced sensitivity of hypocotyl elongation to warm temperatures, suggesting that PKL plays an important role in regulating thermomorphogenesis (Zha et al. 2017). However, the relationship of REF6 and PKL is still unknown, needs to be further addressed.
Histone deacetylation in thermomorphogenesis
Histone acetylation and deacetylation play important roles in plant growth, development, and environmental adaptation (Chen et al. 2020a; Liu et al. 2014; Ma et al. 2013). Histone acetylation and deacetylation are reversible and dynamically regulated by histone acetyltransferases (HATs) and histone deacetylases (HDACs) (Chen et al. 2020a). HDACs are ubiquitous and relatively well conserved in eukaryotes. Recent studies have gradually begun to reveal the mechanisms by which histone deacetylation regulates gene expression and responds to environmental temperature changes.
Three histone deacetylases, HDA9, HDA15 and HDA19, are involved in thermomorphogenesis of Arabidopsis. At high ambient temperatures, the hda9 mutant has shorter hypocotyls compared to the wild type, suggesting that the deletion of HDA9 resulted in a weaker response to warm temperatures (Shen et al. 2019; van der Woude et al. 2019). Notably, the impaired thermomorphogenesis in had9 mutants is independent of light-quality signaling and phytochrome B (phyB) (van der Woude et al. 2019). The expression level of several warm-temperature marker genes was down-regulated in hda9-1 mutants compared with the wild type at 27 °C, which further confirmed that the hda9 mutant was insensitive to warm temperature (Shen et al. 2019). However, no direct binding or significant enrichment of HDA9 was detected on several warm-temperature marker genes (HSP20, HSP20-like, IAA19, IAA29, and bZIP63) (Shen et al. 2019), suggesting that HDA9 is not directly associated with these genes. HDA9 does not directly bind to these warm-temperature marker genes, but it is necessary for their expression, suggesting that HDA9 may regulate the expression of warm-temperature marker genes through other mechanisms at elevated ambient temperatures.
HDACs form histone deacetylase complexes (Chen et al. 2020a). For example, the SANT (SWI3/DAD2/N-CoR/TFIII-B) domain protein POWERDRESS (PWR) interacts with HDA9 and is required for HDA9 function (Chen et al. 2016; Kim et al. 2016). The pwr and had9 mutants showed highly similar phenotypes, including impaired thermomorphogenesis (Mayer et al. 2019; Tasset et al. 2018; van der Woude et al. 2019). Warm temperatures induce H3K9 deacetylation at the + 1 nucleosomes of PIF4 and YUC8, but in pwr mutants, these nucleosomes exhibit H3K9 hyperacetylation at warm temperatures (Tasset et al. 2018). Similarly, in had9 mutants, warm temperature induced an increase in H3K9 acetylation at the transcriptional start site and gene body of YUC8, demonstrating the importance of histone deacetylation in thermomorphogenesis (van der Woude et al. 2019).
Similar to the hda9-1 mutant, the hda19 mutants also had shorter hypocotyls than that of the wild type at high ambient temperatures and downregulated expression of several warm-temperature marker genes (HSP20, HSP20-like, IAA19, IAA29 and bZIP63), indicating a diminished response to warm temperatures (Shen et al. 2019). However, unlike HDA9, HDA19 can bind to the exon and/or promoter regions of several warm-temperature marker genes (HSP20, HSP20-like, IAA19, IAA29, and bZIP63), and it appears that HDA19 binding is related to the expression of these genes. HDA19 inhibits photomorphogenesis in Arabidopsis (Benhamed et al. 2006; Jing et al. 2021). Compared with the wild-type, the hda19-2 mutant showed an enhanced photomorphogenesis phenotype, with shorter hypocotyls under far-red (FR), red light (RL) and blue light (BL) conditions (Benhamed et al. 2006; Jing et al. 2021). Considering that HDA19 functions in both thermomorphogenesis and photomorphogenesis, it seems that HDA19 may be involved in the regulation of hypocotyl elongation through the integration of light and temperature signals.
Recent studies have shown that HDA15 is a direct repressor of Arabidopsis responses to elevated ambient temperature, and seedlings of hda15-1 mutants show enhanced high ambient temperature responses (Shen et al. 2019). Compared with wild-type Col-0 plants, hda15-1 seedlings showed a long hypocotyl phenotype at 20 °C and 27 °C; in particular, the hypocotyls of hda15-1 seedlings were significantly longer than those of Col-0 at high ambient temperatures, indicating that hda15-1 seedlings showed a hyper-response to elevated ambient temperature. Transcriptome analysis showed that hda15-1 mutants induced the expression of some warm temperature-induced genes at 20 °C, and the expression of these genes was more upregulated at high ambient temperature, indicating that HDA15 negatively regulates the expression of warm temperature-induced genes. Further analysis revealed that HDA15 interacts with the transcription factor LONG HYPOCOTYL IN FAR RED1 (HFR1) to synergistically inhibit temperature responses (Shen et al. 2019).
Three HDACs (HDA9, HDA15, and HDA19) are reported to be involved in thermomorphogenesis. Interestingly, different HDACs have different or even opposite functions in thermomorphogenesis (Shen et al. 2019). Analysis of the differentially expressed target genes revealed that different HDACs regulate hypocotyl elongation by distinct mechanisms, indicating the complexity and diversity of HDAC functions in thermomorphogenesis. The functions of HDACs depend to some extent on their interacting proteins, which may be one of the reasons for the differential response of HDACs to elevated ambient temperature. Screening and identifying more HDAC-interacting proteins and studying the role of the complexes formed by HDAC and HDAC-interacting proteins in the warm temperature response will help to reveal the specific mechanism by which HDACs participate in thermomorphogenesis. In addition, current research has mostly focused on revealing the roles of HDACs in warm temperature responses and analyzing the target genes regulated by HDACs, while the effects of HDACs on histone acetylation and chromatin state are not clear. Analyzing the effect of HDAC deletion on the acetylation level of specific histone sites and comparing the changes of histone acetylation in response to warm temperature will help to reveal the mechanism by which HDACs regulate the expression of temperature-responsive genes.
H2A.Z nucleosomal dynamics
H2A.Z is a conserved variant of histone H2A present in most eukaryotic organisms. In most cases, H2A.Z is enriched on the first nucleosome downstream of the transcription start site (TSS + 1 nucleosome) and in the gene body, causing nucleosomes to wrap DNA more tightly, which influences the ability of RNA polymerase (Pol) II to transcribe genes in plants (Cortijo et al. 2017; Kumar and Wigge 2010; Xue et al. 2021). H2A.Z is incorporated into chromatin to replace H2A by the SWI2/SNF2‐RELATED 1 complex (SWR1c), composed of ACTIN‐RELATED PROTEIN 6 (ARP6), ARP4, SWR1 COMPLEX 4 (SWC4), SERRATED LEAVES AND EARLY FLOWERING (SEF) and PHOTOPERIOD‐INDEPENDENT EARLY FLOWERING 1 (PIE1) in Arabidopsis (Aslam et al. 2019; Choi et al. 2007; Deal et al. 2005; Gómez-Zambrano et al. 2018; Kobor et al. 2004; Krogan et al. 2003; March-Díaz et al. 2007; Mizuguchi et al. 2004; Noh and Amasino 2003).
H2A.Z loss-of-function mutants (hta9hta11) and arp6 both have elevated HEAT SHOCK PROTEIN 70 (HSP70) expression along with early flowering, long hypocotyls, petiole elongation, and other warm temperature-related phenotypes (Kumar and Wigge 2010). Furthermore, temperature affects the occupancy of H2A.Z, with high occupancy of H2A.Z at lower temperatures and eviction of H2A.Z at higher temperatures (Kumar and Wigge 2010). High ambient temperature induce the eviction of H2A.Z-containing nucleosomes, which contributes to the formation of an open DNA structure, thus facilitating the binding of transcription factors to activate or inhibit gene expression at high temperature (Cortijo et al. 2017; Kumar and Wigge 2010; Xue et al. 2021). For example, temperature-induced H2A.Z nucleosome dynamics gate the accessibility of PIF4 to FLOWERING LOCUS T (FT) for transcriptional activation and control plants timing of flowering in response to temperature (Kumar et al. 2012).
However, the eviction of H2A.Z-containing nucleosomes is not directly controlled by temperature, as loss of H2A.Z in chromatin does not respond to temperature in vitro (Cortijo et al. 2017). Several factors regulate the substitution of H2A.Z in nucleosomes associated with temperature-responsive genes. One of these factors is the transcription factor HEAT SHOCK FACTOR A1 (HSFA1) (Cortijo et al. 2017). HSFA1 is part of a clade of Arabidopsis HEAT SHOCK FACTORs (HSFs), which are the pivotal transcriptional activators of the heat shock response in eukaryotes. As the temperature rises, HSFA1 binds to temperature-responsive genes and initiates transcription by dynamically facilitating H2A.Z loss, thus antagonizing the function of H2A.Z in response to warm temperature (Cortijo et al. 2017). Warm temperature selectively enhances the translation of another HSF protein, HSFA2, by affecting the formation of an RNA hairpin within the 5′-untranslated region of the HSFA2 mRNA (Chung et al. 2020). However, how HSFs regulate the eviction of H2A.Z-containing nucleosomes at temperature-responsive genes remains to be studied.
In addition, other epigenetic factors regulate the response of H2A.Z to temperature. HDA9 mediates histone deacetylation at YUCCA8 and regulates temperature-induced H2A.Z eviction from nucleosomes at PIF4 targets, but does not affect the expression of PIF4 and HSP70, suggesting that nucleosome dynamics and histone deacetylation are coupled during warm-temperature responses (van der Woude et al. 2019). Another important regulatory factor is the chromatin-remodeling factor INO80. The INO80 chromatin remodeling complex directly interacts with PIF4 to promote temperature-induced H2A.Z eviction at PIF4 targets, activating the expression of thermo-responsive and auxin-related genes (Xue et al. 2021). INO80 also interacts with H2A.Z, positively regulating H2A.Z-containing nucleosomes at the key flowering repressor gene FLC and regulating H2A.Z-mediated gene expression in response to light, thereby controlling plant growth and development (Yang et al. 2020; Zhang et al. 2015a, b). A recent study showed that PIFs (PIF4/PIF5/PIF7) undergo rapid regulation to reshape the H2A.Z and H3K9ac epigenetic landscape by interacting with EIN6 ENHANCER (EEN), a subunit of the INO80 chromatin-remodeling complex, in response to low R:FR conditions. The PIFs-INO80 module may be a hub allowing plants to regulate their growth by adapting to environmental changes at the epigenetic level (Willige et al. 2021). Light and temperature usually change in concert under natural conditions and the light and temperature signaling pathways usually share the same downstream regulatory elements.
Phytochrome B (phyB), the dominant red-light photoreceptor, exists in two photo-interconvertible forms, Pr (R light-absorbing, biologically inactive) and Pfr (FR light-absorbing, biologically active) (Li et al. 2011; Reed et al. 1993; Somers et al. 1991). PhyB also functions as a thermosensor, undergoing temperature-dependent reversion from the active Pfr form to the inactive Pr form (Jung et al. 2016; Legris et al. 2016). The Pfr form of phyB interacts with SWC6 and ARP6, positively regulating H2A.Z deposition at YUC9 and thus mediating the inhibition of hypocotyl growth under red light (Mao et al. 2021; Wei et al. 2021). Therefore, phyB may regulate H2A.Z eviction in response to warm temperature. In addition to phyB, the blue light (BL) photoreceptor cryptochrome 1 (CRY1) has been reported to act in a similar mechanism for H2A.Z regulation under blue light. CRY1 regulates H2A.Z deposition at ELONGATED HYPOCOTYL 5 (HY5) target genes by interacting with SWC6 and ARP6, thus positively regulating photomorphogenesis (Mao et al. 2021). CRY1 and CRY2 play important roles in temperature signal transduction. CRY1 directly interacts with PIF4 to inhibit its transcriptional activity in a blue light-dependent manner, thus regulating hypocotyl elongation (Ma et al. 2016) and CRY2 accumulates at higher temperature and is degraded at lower temperature (Li et al. 2021; Ma et al. 2021).
Since both phyB and CRY1 regulate the accumulation of H2A.Z at specific sites through similar mechanisms, and they play important roles in warm temperature signal transduction, it will be interesting to investigate the mechanism by which these photoreceptors regulate H2A.Z occupancy at ambient temperatures. In addition, the NUCLEAR FACTOR‐Y subunit C (NF-YC), also known as HISTONE‐ASSOCIATED PROTEIN5 (HAP5) or CCAAT BINDING FACTOR C (CBF‐C) has been reported to regulate light‐induced H2A.Z deposition associated with hypocotyl elongation‐related genes, repressing related gene expression in a light‐dependent manner (Zhang et al. 2021). Due to the close relationship between light and temperature signaling, NF-YC may also be a potential regulatory factor for the eviction and deposition of H2A.Z at warmer temperatures.
The important role of histone modification and histone variants in plant thermomorphogenesis is gradually emerging (See Fig. 1 and Table 1). Although many specific mechanisms need to be further elucidated, this emerging picture undoubtedly provides a new perspective on the mechanisms by which plants respond to environmental temperature. In addition to revealing the mechanism by which single epigenetic markers participate in thermomorphogenesis, we should also consider crosstalk between epigenetic markers in thermomorphogenesis. At warm temperatures, HDA9 mediates histone deacetylation of YUC8 and induces eviction of H2A.Z from nucleosomes at the YUC8 locus, suggesting that histone variants and histone deacetylation act in the temperature response (van der Woude et al. 2019). A recent report finds that INO80 is involved in promoting thermomorphogenesis by integrating H2A.Z eviction and H3K4me3 deposition (Xue et al. 2021). Recent studies have also shown that PIF transcription factors affect the H2A.Z and H3K9ac landscape in a light-quality-dependent manner (Willige et al. 2021). In addition, phyB has been reported to directly interact with VERNALIZATION INSENSITIVE 3-LIKE1/VERNALIZATION 5 (VIL1/VRN5), the core component of Polycomb Repressive Complex 2, to promote the enrichment of H3K27me3 in a light-dependent manner (Kim et al. 2021); furthermore, the direct interaction of phyB‐SWC6/ARP6 promotes the deposition of H2AZ at YUC9, thereby inhibiting hypocotyl elongation in red light (Wei et al. 2021). Considering that light signaling regulates thermomorphogenesis, the changes in H2A.Z and histone modifications may occur as part of crosstalk in response to ambient temperature. Such crosstalk between epigenetic markers may act as a fine-tuning mechanism to regulate plant responses to subtle changes in ambient temperature. Decoding how chromatin landscapes change in response to temperature changes, and thus rapidly fine-tune gene expression, will play a key role in uncovering the mechanisms by which plant morphogenesis responds to environmental challenges.
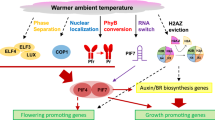
Roles of histone modifications and histone variants in regulating thermomorphogenesis in Arabidopsis thaliana. (i) Phytochrome B is not only the main red light sensor, but also a thermosensor. Under red light, phyB is transformed into a Pfr homodimer, enters the nucleus, and blocks PIF4 activity. Warm temperature promotes the reversion of phyB back to its inactive Pr form, releasing PIF4 to promote downstream temperature response gene expression. (ii) RELATIVE OF EARLY FLOWERING 6/JUMONJI DOMAIN-CONTAINING PROTEIN 12 (REF6/JMJ12) mediates H3K27me3 demethylation at several PIF4 target genes to positively regulate thermomorphogenesis. (iii) FLOWERING CONTROL LOCUS A (FCA)-mediated inhibition of H3K4me2 deposition results in inhibited SEUSS (SEU)-mediated enrichment of H3K4me3, which facilitates hypocotyl elongation at warmer ambient temperatures. (iv) At warmer ambient temperatures, PICKLE (PKL) mediates the downregulation of H3K27me3 levels, which promotes hypocotyl elongation. The INO80 chromatin remodeling complex (INO80-C) promotes H3K4me3 deposition and stimulates hypocotyl elongation in response to warm temperature. (v) The histone deacetylases HDA9 and HDA19 stimulate thermomorphogenesis but HDA15 directly represses plant responses to high ambient temperature. PWR, a SANT-domain containing protein known to interact with HDA9, can also stimulate thermomorphogenesis. (vi) INO80-C, HDA9 and HEAT SHOCK FACTOR A1 (HSFA1) accelerate warm temperature-induced eviction of the histone variant H2A.Z and stimulate hypocotyl elongation. (vii) FCA, SEU and INO80-C interact with PIF4, and regulate histone modifications and histone variants at PIF4 targets. The stars and triangles represent different histone methylations; the hexagons represent histone acetylation. Positive regulators of thermomorphogenesis are represented as ellipses and negative regulators as rectangles
Non-coding RNAs and RNA metabolism
Small RNAs (single-stranded 21–24 nucleotide RNAs) regulate the post-transcriptional expression of genes and include microRNAs (miRNAs) and small interfering RNAs (siRNAs). Small RNAs play key roles in regulating plant growth and development, signal transduction, stress responses, and genome stability (D'Ario et al. 2017; Deng et al. 2016; Si et al. 2020). In addition, the role of small RNAs in ambient temperature adaptation of plants is emerging (Gyula et al. 2018; Kim et al. 2012; Lee et al. 2010; Serivichyaswat et al. 2017). Temperature regulates the abundance of certain small RNAs and such ambient temperature-responsive miRNAs are involved in the regulation of flowering time (Gyula et al. 2018; Lee et al. 2010; Serivichyaswat et al. 2017). Although warm temperatures promoted the accumulation of miR172a, plants that overexpressed miR172a exhibited a temperature-independent early flowering phenotype, suggesting that changes in miR172a accumulation affected plant temperature perception (Lee et al. 2010). The accumulation of miR169 is down-regulated at warm temperatures, leading to the activation of NF‐YA‐dependent genes that promote flowering (Gyula et al. 2018).
Long non-coding RNAs (lncRNAs, > 200 nucleotide RNAs that are not translated into functional proteins) occur widely in eukaryotes (Statello et al. 2021; Wierzbicki et al. 2021) and regulate gene expression at the transcriptional and post-transcriptional levels (Statello et al. 2021; Wierzbicki et al. 2021). Ongoing work has identified lncRNAs whose expression responds to environmental temperature changes, suggesting that lncRNAs have potential regulatory roles in temperature-dependent plant development (Severing et al. 2018). For example, expression of the lncRNA FLOWERING LONG INTERGENIC NON-CODING RNA (FLINC) was down-regulated at warm temperatures and the flinc mutants flowered earlier than wild type, confirming the regulatory role of FLINC in thermomorphogenesis (Severing et al. 2018). However, the molecular mechanisms of the known ambient temperature-regulated lncRNAs is not clear, and their biological functions in plant thermomorphogenesis remain to be revealed.
The metabolism and processing of mRNAs is particularly important in environmental temperature responses. For example, alternative splicing modulates plant responses to high temperature (Ling et al. 2021). RNA transcription and processing are tightly coupled, providing an opportunity for epigenetic factors to regulate RNA processing (Soles and Shi 2021; Tellier et al. 2020). For example, H3K36me3 plays an important role in regulating temperature-induced alternative splicing and temperature-dependent flowering time (Pajoro et al. 2017). Indeed, the important regulatory role of epigenetic modification in RNA metabolism such as alternative polyadenylation and RNA modification is attracting increasing attention (Kan et al. 2021; Soles and Shi 2021). Moreover, temperature can affect translation: at higher temperatures, the RNA hairpin in the 5′-untranslated region of PIF7 adopts a loose conformation, leading to enhanced translation efficiency (Chung et al. 2020). Exploring the effects of epigenetic regulatory mechanisms on RNA metabolism of key genes involved in thermomorphogenesis will deepen our understanding of plant responses to warm temperature.
Concluding remarks and future prospects
Plant cell types have distinct functions and the synergies among different cell types determine the plasticity of plant development. Revealing the functions of cell types, even at the single-cell level, is crucial to understanding the environmental adaptation of plants (Rich-Griffin et al. 2020). High-throughput single-cell sequencing techniques developed in recent years have provided unprecedented insights into critical cellular processes. Cell type-specific transcriptome analysis has begun to reveal the basic cellular activities involved in plant development and stress adaptation (Rich-Griffin et al. 2020). It is also beginning to be possible to quantitatively measure the dynamics of transcription activity in single cells in living plants under changing environmental conditions (Alamos et al. 2021; Hani et al. 2021). However, the study of cell type-specific or single-cell epigenomics in plant species remains limited.
The role of epigenetic factors in defining individual cell types and the extent to which they vary during plant development or under environmental stress remains unknown. Epigenome studies based on single-cell behavior have tremendous potential for exploring the regulatory mechanisms of plant perception and responses to changes in environmental temperature. In addition, the explosion of single-cell genomics in recent years has created an opportunity to jointly reveal the variation and differential use of genomic information in different cell types of plant species during development, differentiation, and response to environmental stresses at multiple levels (Perkel 2021; Thibivilliers and Libault 2021). The combination of cellular epigenomics and other omics at the single-cell level will help us describe the association between cell behavior and plant thermomorphogenesis phenotypes under continuously changing ambient temperatures. Obtaining multiomic molecular information at the single-cell level will be critical to reveal the functions of individual genes and the molecular mechanisms that regulate their expression, and to develop new strategies to optimize temperature-responsive agronomic traits, ultimately achieving sustainable improvements in agricultural productivity.
Thermomorphogenesis of plants is closely related to their adaptation to global climate change, as global warming has a profound impact on plant morphology, development, and geographical distribution (Lee et al. 2020; Lippmann et al. 2019). In-depth analysis of the molecular mechanisms of thermomorphogenesis is helpful to explore ways to improve the adaptability of plants to climate change and produce climate-smart crops (Lippmann et al. 2019; Park et al. 2021). Epigenetic regulatory mechanisms are critical for plant development and phenotypic plasticity, including adaptive responses to ambient temperature. Epigenetic stability and diversity and their genetic characteristics are a new source of phenotypic variation to improve plant adaptation to changing global climate and to drive breeding to ensure crop yield and quality. Future studies on the variation characteristics of crop epigenetic markers and their association with gene expression and phenotype will expand the space for plant breeding. Considering epigenetic modification in response to environmental temperature changes will facilitate the exploration of breeding strategies needed to address food security challenges in the context of global warming.
References
Alamos S, Reimer A, Niyogi KK, Garcia HG (2021) Quantitative imaging of RNA polymerase II activity in plants reveals the single-cell basis of tissue-wide transcriptional dynamics. Nat Plants 7:1037–1049
Allis CD, Jenuwein T (2016) The molecular hallmarks of epigenetic control. Nat Rev Genet 17:487–500
Aslam M, Fakher B, Jakada BH, Cao S, Qin Y (2019) SWR1 chromatin remodeling complex: a key transcriptional regulator in plants. Cells 8:1621
Benhamed M, Bertrand C, Servet C, Zhou DX (2006) Arabidopsis GCN5, HD1, and TAF1/HAF2 interact to regulate histone acetylation required for light-responsive gene expression. Plant Cell 18:2893–2903
Boquete MT, Muyle A, Alonso C (2021) Plant epigenetics: phenotypic and functional diversity beyond the DNA sequence. Am J Bot 108:553–558
Casal JJ, Balasubramanian S (2019) Thermomorphogenesis. Annu Rev Plant Biol 70:321–346
Chang YN, Zhu C, Jiang J, Zhang H, Zhu JK, Duan CG (2019) Epigenetic regulation in plant abiotic stress responses. J Integr Plant Biol 62(5):563
Chen X (2004) A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303:2022–2025
Chen X, Lu L, Mayer KS, Scalf M, Qian S, Lomax A, Smith LM, Zhong X (2016) POWERDRESS interacts with HISTONE DEACETYLASE 9 to promote aging in Arabidopsis. eLife. https://doi.org/10.7554/eLife.17214
Chen X, Ding AB, Zhong X (2020a) Functions and mechanisms of plant histone deacetylases. Sci China Life Sci 63:206–216
Chen CY, Tu YT, Hsu JC, Hung HC, Liu TC, Lee YH, Chou CC, Cheng YS, Wu K (2020b) Structure of Arabidopsis HISTONE DEACETYLASE15. Plant Physiol 184:1585–1600
Choi K, Park C, Lee J, Oh M, Noh B, Lee I (2007) Arabidopsis homologs of components of the SWR1 complex regulate flowering and plant development. Development 134:1931–1941
Chung BYW, Balcerowicz M, Di Antonio M, Jaeger KE, Geng F, Franaszek K, Marriott P, Brierley I, Firth AE, Wigge PA (2020) An RNA thermoswitch regulates daytime growth in Arabidopsis. Nat Plants 6:522–532
Cortijo S, Charoensawan V, Brestovitsky A, Buning R, Ravarani C, Rhodes D, van Noort J, Jaeger KE, Wigge PA (2017) Transcriptional regulation of the ambient temperature response by H2A.Z nucleosomes and HSF1 transcription factors in Arabidopsis. Mol Plant 10:1258–1273
Cui X, Lu F, Qiu Q, Zhou B, Gu L, Zhang S, Kang Y, Cui X, Ma X, Yao Q et al (2016) REF6 recognizes a specific DNA sequence to demethylate H3K27me3 and regulate organ boundary formation in Arabidopsis. Nat Genet 48:694–699
Cui X, Zheng Y, Lu Y, Issakidis-Bourguet E, Zhou DX (2021) Metabolic control of histone demethylase activity involved in plant response to high temperature. Plant Physiol 185:1813–1828
D’Ario M, Griffiths-Jones S, Kim M (2017) Small RNAs: big impact on plant development. Trends Plant Sci 22:1056–1068
Deal RB, Kandasamy MK, McKinney EC, Meagher RB (2005) The nuclear actin-related protein ARP6 is a pleiotropic developmental regulator required for the maintenance of FLOWERING LOCUS C expression and repression of flowering in Arabidopsis. Plant Cell 17:2633–2646
Deng X, Song XW, Wei LY, Liu CY, Cao XF (2016) Epigenetic regulation and epigenomic landscape in rice. Natl Sci Rev 3:309–327
Ding Y, Shi Y, Yang S (2020) Molecular regulation of plant responses to environmental temperatures. Mol Plant 13:544–564
Franks RG, Wang C, Levin JZ, Liu Z (2002) SEUSS, a member of a novel family of plant regulatory proteins, represses floral homeotic gene expression with LEUNIG. Development 129:253–263
Gallusci P, Dai ZW, Genard M, Gauffretau A, Leblanc-Fournier N, Richard-Molard C, Vile D, Brunel-Muguet S (2017) Epigenetics for plant improvement: current knowledge and modeling avenues. Trends Plant Sci 22:610–623
Gan ES, Xu Y, Wong JY, Goh JG, Sun B, Wee WY, Huang J, Ito T (2014) Jumonji demethylases moderate precocious flowering at elevated temperature via regulation of FLC in Arabidopsis. Nat Commun 5:5098
Gómez-Zambrano Á, Crevillén P, Franco-Zorrilla JM, López JA, Moreno-Romero J, Roszak P, Santos-González J, Jurado S, Vázquez J, Köhler C et al (2018) Arabidopsis SWC4 binds DNA and recruits the SWR1 complex to modulate histone H2A.Z deposition at key regulatory genes. Mol Plant 11:815–832
Grosselin K, Durand A, Marsolier J, Poitou A, Marangoni E, Nemati F, Dahmani A, Lameiras S, Reyal F, Frenoy O et al (2019) High-throughput single-cell ChIP-seq identifies heterogeneity of chromatin states in breast cancer. Nat Genet 51:1060–1066
Gyula P, Baksa I, Toth T, Mohorianu I, Dalmay T, Szittya G (2018) Ambient temperature regulates the expression of a small set of sRNAs influencing plant development through NF-YA2 and YUC2. Plant Cell Environ 41:2404–2417
Hani S, Cuyas L, David P, Secco D, Whelan J, Thibaud MC, Merret R, Mueller F, Pochon N, Javot H et al (2021) Live single-cell transcriptional dynamics via RNA labelling during the phosphate response in plants. Nat Plants 7:1050–1064
He K, Cao X, Deng X (2021a) Histone methylation in epigenetic regulation and temperature responses. Curr Opin Plant Biol 61:102001
He K, Mei H, Zhu J, Qiu Q, Cao X, Deng X (2021b) The histone H3K27 demethylase REF6/JMJ12 promotes thermomorphogenesis in Arabidopsis. Natl Sci Rev. https://doi.org/10.1093/nsr/nwab213
Ho KK, Zhang H, Golden BL, Ogas J (2013) PICKLE is a CHD subfamily II ATP-dependent chromatin remodeling factor. Biochem Biophys Acta 1829:199–210
Huai J, Zhang X, Li J, Ma T, Zha P, Jing Y, Lin R (2018) SEUSS and PIF4 coordinately regulate light and temperature signaling pathways to control plant growth. Mol Plant 11:928–942
Jing Y, Guo Q, Lin R (2021) The SNL-HDA19 histone deacetylase complex antagonizes HY5 activity to repress photomorphogenesis in Arabidopsis. New Phytol 229:3221–3236
Jung JH, Domijan M, Klose C, Biswas S, Ezer D, Gao M, Khattak AK, Box MS, Charoensawan V, Cortijo S et al (2016) Phytochromes function as thermosensors in arabidopsis. Science 354:886–889
Kan RL, Chen J, Sallam T (2021) Crosstalk between epitranscriptomic and epigenetic mechanisms in gene regulation. Trends Genet. https://doi.org/10.1016/j.tig.2021.06.014
Kim JJ, Lee JH, Kim W, Jung HS, Huijser P, Ahn JH (2012) The microRNA156-SQUAMOSA PROMOTER BINDING PROTEIN-LIKE3 module regulates ambient temperature-responsive flowering via FLOWERING LOCUS T in Arabidopsis. Plant Physiol 159:461–478
Kim YJ, Wang R, Gao L, Li D, Xu C, Mang H, Jeon J, Chen X, Zhong X, Kwak JM et al (2016) POWERDRESS and HDA9 interact and promote histone H3 deacetylation at specific genomic sites in Arabidopsis. Proc Natl Acad Sci USA 113:14858–14863
Kim J, Bordiya Y, Kathare PK, Zhao B, Zong W, Huq E, Sung S (2021) Phytochrome B triggers light-dependent chromatin remodelling through the PRC2-associated PHD finger protein VIL1. Nat Plants 7:1213–1219
Kobor MS, Venkatasubrahmanyam S, Meneghini MD, Gin JW, Jennings JL, Link AJ, Madhani HD, Rine J (2004) A protein complex containing the conserved Swi2/Snf2-related ATPase Swr1p deposits histone variant H2A.Z into euchromatin. PLoS Biol 2:587–599
Koini MA, Alvey L, Allen T, Tilley CA, Harberd NP, Whitelam GC, Franklin KA (2009) High temperature-mediated adaptations in plant architecture require the bHLH transcription factor PIF4. Curr Biol 19:408–413
Krogan NJ, Keogh MC, Datta N, Sawa C, Ryan OW, Ding H, Haw RA, Pootoolal J, Tong A, Canadien V et al (2003) A Snf2 family ATPase complex required for recruitment of the histone H2A variant Htz1. Mol Cell 12:1565–1576
Kumar SV, Wigge PA (2010) H2A.Z-containing nucleosomes mediate the thermosensory response in Arabidopsis. Cell 140:136–147
Kumar SV, Lucyshyn D, Jaeger KE, Alos E, Alvey E, Harberd NP, Wigge PA (2012) Transcription factor PIF4 controls the thermosensory activation of flowering. Nature 484:242–245
Lee H, Yoo SJ, Lee JH, Kim W, Yoo SK, Fitzgerald H, Carrington JC, Ahn JH (2010) Genetic framework for flowering-time regulation by ambient temperature-responsive miRNAs in Arabidopsis. Nucleic Acids Res 38:3081–3093
Lee HJ, Jung JH, Cortes Llorca L, Kim SG, Lee S, Baldwin IT, Park CM (2014) FCA mediates thermal adaptation of stem growth by attenuating auxin action in Arabidopsis. Nat Commun 5:5473
Lee JH, Kim JY, Kim JI, Park YJ, Park CM (2020) Plant thermomorphogenic adaptation to global warming. J Plant Biol 63:1–9
Legris M, Klose C, Burgie ES, Rojas CC, Neme M, Hiltbrunner A, Wigge PA, Schafer E, Vierstra RD, Casal JJ (2016) Phytochrome B integrates light and temperature signals in Arabidopsis. Science 354:897–900
Li J, Li G, Wang H, Wang Deng X (2011) Phytochrome signaling mechanisms. Arabidopsis 9:e0148
Li C, Gu L, Gao L, Chen C, Wei CQ, Qiu Q, Chien CW, Wang S, Jiang L, Ai LF et al (2016) Concerted genomic targeting of H3K27 demethylase REF6 and chromatin-remodeling ATPase BRM in Arabidopsis. Nat Genet 48:687–693
Li B, Gao K, Ren H, Tang W (2018) Molecular mechanisms governing plant responses to high temperatures. J Integr Plant Biol 60:757–779
Li Y, Shi Y, Li M, Fu D, Wu S, Li J, Gong Z, Liu H, Yang S (2021) The CRY2-COP1-HY5-BBX7/8 module regulates blue light-dependent cold acclimation in Arabidopsis. Plant Cell. https://doi.org/10.1093/plcell/koab215
Ling Y, Mahfouz MM, Zhou S (2021) Pre-mRNA alternative splicing as a modulator for heat stress response in plants. Trends Plant Sci. https://doi.org/10.1016/j.tplants.2021.07.008
Lippmann R, Babben S, Menger A, Delker C, Quint M (2019) Development of wild and cultivated plants under global warming conditions. Curr Biol 29:R1326–R1338
Liu F, Quesada V, Crevillén P, Bäurle I, Swiezewski S, Dean C (2007) The Arabidopsis RNA-binding protein FCA requires a lysine-specific demethylase 1 homolog to downregulate FLC. Mol Cell 28:398–407
Liu X, Yang S, Zhao M, Luo M, Yu CW, Chen CY, Tai R, Wu K (2014) Transcriptional repression by histone deacetylases in plants. Mol Plant 7:764–772
Liu J, Feng L, Gu X, Deng X, Qiu Q, Li Q, Zhang Y, Wang M, Deng Y, Wang E et al (2019) An H3K27me3 demethylase-HSFA2 regulatory loop orchestrates transgenerational thermomemory in Arabidopsis. Cell Res 29:379–390
Lu F, Li G, Cui X, Liu C, Wang XJ, Cao X (2008) Comparative analysis of JmjC domain-containing proteins reveals the potential histone demethylases in Arabidopsis and rice. J Integr Plant Biol 50:886–896
Lu F, Cui X, Zhang S, Liu C, Cao X (2010) JMJ14 is an H3K4 demethylase regulating flowering time in Arabidopsis. Cell Res 20:387–390
Lu F, Cui X, Zhang S, Jenuwein T, Cao X (2011) Arabidopsis REF6 is a histone H3 lysine 27 demethylase. Nat Genet 43:715–719
Ma X, Lv S, Zhang C, Yang C (2013) Histone deacetylases and their functions in plants. Plant Cell Rep 32:465–478
Ma D, Li X, Guo Y, Chu J, Fang S, Yan C, Noel JP, Liu H (2016) Cryptochrome 1 interacts with PIF4 to regulate high temperature-mediated hypocotyl elongation in response to blue light. Proc Natl Acad Sci USA 113:224–229
Ma L, Li X, Zhao Z, Hao Y, Shang R, Zeng D, Liu H (2021) Light-response Bric-A-Brack/Tramtrack/broad proteins mediate cryptochrome 2 degradation in response to low ambient temperature. Plant Cell. https://doi.org/10.1093/plcell/koab219
Macknight R, Bancroft I, Page T, Lister C, Schmidt R, Love K, Westphal L, Murphy G, Sherson S, Cobbett C et al (1997) FCA, a gene controlling flowering time in Arabidopsis, encodes a protein containing RNA-binding domains. Cell 89:737–745
Mao Z, Wei X, Li L, Xu P, Zhang J, Wang W, Guo T, Kou S, Wang W, Miao L et al (2021) Arabidopsis cryptochrome 1 controls photomorphogenesis through regulation of H2A.Z deposition. Plant Cell 33:1961–1979
March-Díaz R, García-Domínguez M, Florencio FJ, Reyes JC (2007) SEF, a new protein required for flowering repression in Arabidopsis, interacts with PIE1 and ARP6. Plant Physiol 143:893–901
Martin-Trillo M, Lazaro A, Poethig RS, Gomez-Mena C, Pineiro MA, Martinez-Zapater JM, Jarillo JA (2006) EARLY IN SHORT DAYS 1 (ESD1) encodes ACTIN-RELATED PROTEIN 6 (AtARP6), a putative component of chromatin remodelling complexes that positively regulates FLC accumulation in Arabidopsis. Development 133:1241–1252
Martire S, Banaszynski LA (2020) The roles of histone variants in fine-tuning chromatin organization and function. Nat Rev Mol Cell Biol 21:522–541
Mayer KS, Chen X, Sanders D, Chen J, Jiang J, Nguyen P, Scalf M, Smith LM, Zhong X (2019) HDA9-PWR-HOS15 is a core histone deacetylase complex regulating transcription and development. Plant Physiol 180:342–355
Meyer P (2015) Epigenetic variation and environmental change. J Exp Bot 66:3541–3548
Mizuguchi G, Shen X, Landry J, Wu WH, Sen S, Wu C (2004) ATP-driven exchange of histone H2AZ variant catalyzed by SWR1 chromatin remodeling complex. Science 303:343–348
Noh YS, Amasino RM (2003) PIE1, an ISWI family gene, is required for FLC activation and floral repression in Arabidopsis. Plant Cell 15:1671–1682
Pajoro A, Severing E, Angenent GC, Immink RGH (2017) Histone H3 lysine 36 methylation affects temperature-induced alternative splicing and flowering in plants. Genome Biol 18:102
Pandey R, Muller A, Napoli CA, Selinger DA, Pikaard CS, Richards EJ, Bender J, Mount DW, Jorgensen RA (2002) Analysis of histone acetyltransferase and histone deacetylase families of Arabidopsis thaliana suggests functional diversification of chromatin modification among multicellular eukaryotes. Nucleic Acids Res 30:5036–5055
Park YJ, Kim JY, Lee JH, Han SH, Park CM (2021) External and internal reshaping of plant thermomorphogenesis. Trends Plant Sci 26:810–821
Perkel JM (2021) Single-cell analysis enters the multiomics age. Nature 595:614–616
Qiu Q, Mei H, Deng X, He K, Wu B, Yao Q, Zhang J, Lu F, Ma J, Cao X (2019) DNA methylation repels targeting of Arabidopsis REF6. Nat Commun 10:2063
Quint M, Delker C, Franklin KA, Wigge PA, Halliday KJ, van Zanten M (2016) Molecular and genetic control of plant thermomorphogenesis. Nat Plants 2:15190
Reed JW, Nagpal P, Poole DS, Furuya M, Chory J (1993) Mutations in the gene for the red/far-red light receptor phytochrome B alter cell elongation and physiological responses throughout Arabidopsis development. Plant Cell 5:147–157
Rich-Griffin C, Stechemesser A, Finch J, Lucas E, Ott S, Schafer P (2020) Single-cell transcriptomics: a high-resolution avenue for plant functional genomics. Trends Plant Sci 25:186–197
Serivichyaswat PT, Susila H, Ahn JH (2017) Elongated hypocotyl 5-homolog (HYH) negatively regulates expression of the ambient temperature-responsive microRNA gene MIR169. Front Plant Sci 8:2087
Severing E, Faino L, Jamge S, Busscher M, Kuijer-Zhang Y, Bellinazzo F, Busscher-Lange J, Fernandez V, Angenent GC, Immink RGH et al (2018) Arabidopsis thaliana ambient temperature responsive lncRNAs. BMC Plant Biol 18:145
Shen Y, Lei T, Cui X, Liu X, Zhou S, Zheng Y, Guerard F, Issakidis-Bourguet E, Zhou DX (2019) Arabidopsis histone deacetylase HDA15 directly represses plant response to elevated ambient temperature. The Plant J Cell Mol Biol 100:991–1006
Si F, Cao X, Song X, Deng X (2020) Processing of coding and non-coding RNAs in plant development and environmental responses. Essays Biochem 64:931–945
Sidaway-Lee K, Costa MJ, Rand DA, Finkenstadt B, Penfield S (2014) Direct measurement of transcription rates reveals multiple mechanisms for configuration of the Arabidopsis ambient temperature response. Genome Biol 15:R45
Soles LV, Shi YS (2021) Crosstalk between mRNA 3’-end processing and epigenetics. Front Genet. https://doi.org/10.3389/fgene.2021.637705
Somers DE, Sharrock RA, Tepperman JM, Quail PH (1991) The hy3 long hypocotyl mutant of Arabidopsis is deficient in phytochrome B. Plant Cell 3:1263–1274
Statello L, Guo CJ, Chen LL, Huarte M (2021) Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol 22:96–118
Sun J, Qi L, Li Y, Chu J, Li C (2012) PIF4-mediated activation of YUCCA8 expression integrates temperature into the auxin pathway in regulating arabidopsis hypocotyl growth. PLoS Genet 8:e1002594
Tasset C, Singh Yadav A, Sureshkumar S, Singh R, van der Woude L, Nekrasov M, Tremethick D, van Zanten M, Balasubramanian S (2018) POWERDRESS-mediated histone deacetylation is essential for thermomorphogenesis in Arabidopsis thaliana. PLoS Genet 14:e1007280
Tellier M, Maudlin I, Murphy S (2020) Transcription and splicing: a two-way street. Wiley Interdiscipl Rev RNA. https://doi.org/10.1002/wrna.1593
Thibivilliers S, Libault M (2021) Plant single-cell multiomics: cracking the molecular profiles of plant cells. Trends Plant Sci 26:662–663
Tian Z, Li X, Li M, Wu W, Zhang M, Tang C, Li Z, Liu Y, Chen Z, Yang M et al (2020) Crystal structures of REF6 and its complex with DNA reveal diverse recognition mechanisms. Cell Discov 6:17
Tollefson J (2021) IPCC climate report: earth is warmer than it’s been in 125,000 years. Nature 596:171–172
Ueda M, Seki M (2020) Histone modifications form epigenetic regulatory networks to regulate abiotic stress response. Plant Physiol 182:15–26
van der Woude LC, Perrella G, Snoek BL, van Hoogdalem M, Novak O, van Verk MC, van Kooten HN, Zorn LE, Tonckens R, Dongus JA et al (2019) HISTONE DEACETYLASE 9 stimulates auxin-dependent thermomorphogenesis in Arabidopsis thaliana by mediating H2A.Z depletion. Proc Natl Acad Sci USA 116:25343–25354
Wagner EJ, Carpenter PB (2012) Understanding the language of Lys36 methylation at histone H3. Nat Rev Mol Cell Biol 13:115–126
Wei X, Wang W, Xu P, Wang W, Guo T, Kou S, Liu M, Niu Y, Yang HQ, Mao Z (2021) Phytochrome B interacts with SWC6 and ARP6 to regulate H2A.Z deposition and photomorphogensis in Arabidopsis. J Integr Plant Biol 63:1133–1146
Wierzbicki AT, Blevins T, Swiezewski S (2021) Long noncoding RNAs in plants. Annu Rev Plant Biol 72:245–271
Willige BC, Zander M, Yoo CY, Phan A, Garza RM, Trigg SA, He Y, Nery JR, Chen H, Chen M et al (2021) PHYTOCHROME-INTERACTING FACTORs trigger environmentally responsive chromatin dynamics in plants. Nat Genet 53:955–961
Xu L, Zhao Z, Dong A, Soubigou-Taconnat L, Renou JP, Steinmetz A, Shen WH (2008) Di- and tri- but not monomethylation on histone H3 lysine 36 marks active transcription of genes involved in flowering time regulation and other processes in Arabidopsis thaliana. Mol Cell Biol 28:1348–1360
Xu Y, Gan ES, Zhou J, Wee WY, Zhang X, Ito T (2014) Arabidopsis MRG domain proteins bridge two histone modifications to elevate expression of flowering genes. Nucleic Acids Res 42:10960–10974
Xue M, Zhang H, Zhao F, Zhao T, Li H, Jiang D (2021) The INO80 chromatin remodeling complex promotes thermo-morphogenesis by connecting H2AZ eviction and active transcription in Arabidopsis. Mol Plant 14:1799
Yang Z, Qiu Q, Chen W, Jia B, Chen X, Hu H, He K, Deng X, Li S, Tao WA et al (2018) Structure of the Arabidopsis JMJ14-H3K4me3 complex provides insight into the substrate specificity of KDM5 subfamily histone demethylases. Plant Cell 30:167–177
Yang C, Yin L, Xie F, Ma M, Huang S, Zeng Y, Shen WH, Dong A, Li L (2020) AtINO80 represses photomorphogenesis by modulating nucleosome density and H2A.Z incorporation in light-related genes. Proc Natl Acad Sci USA 117:33679–33688
Zha P, Jing Y, Xu G, Lin R (2017) PICKLE chromatin-remodeling factor controls thermosensory hypocotyl growth of Arabidopsis. Plant Cell Environ 40:2426–2436
Zhang X, Bernatavichute YV, Cokus S, Pellegrini M, Jacobsen SE (2009) Genome-wide analysis of mono-, di- and trimethylation of histone H3 lysine 4 in Arabidopsis thaliana. Genome Biol 10:R62
Zhang C, Cao L, Rong L, An Z, Zhou W, Ma J, Shen WH, Zhu Y, Dong A (2015a) The chromatin-remodeling factor AtINO80 plays crucial roles in genome stability maintenance and in plant development. Plant J Cell Mol Biol 82:655–668
Zhang S, Zhou B, Kang Y, Cui X, Liu A, Deleris A, Greenberg MV, Cui X, Qiu Q, Lu F et al (2015b) C-terminal domains of a histone demethylase interact with a pair of transcription factors and mediate specific chromatin association. Cell Discov. https://doi.org/10.1038/celldisc.2015.3
Zhang C, Qian Q, Huang X, Zhang W, Liu X, Hou X (2021) NF-YCs modulate histone variant H2A.Z deposition to regulate photomorphogenic growth in Arabidopsis. J Integr Plant Biol 63:1120–1132
Zhao Z, Yu Y, Meyer D, Wu C, Shen WH (2005) Prevention of early flowering by expression of FLOWERING LOCUS C requires methylation of histone H3 K36. Nat Cell Biol 7:1256–1260
Zhao C, Liu B, Piao S, Wang X, Lobell DB, Huang Y, Huang M, Yao Y, Bassu S, Ciais P et al (2017) Temperature increase reduces global yields of major crops in four independent estimates. Proc Natl Acad Sci USA 114:9326–9331
Zhao JG, Lu ZG, Wang L, Jin B (2021) Plant responses to heat stress: physiology, transcription, noncoding RNAs, and epigenetics. Int J Mol Sci. https://doi.org/10.3390/ijms22010117
Zheng S, Hu H, Ren H, Yang Z, Qiu Q, Qi W, Liu X, Chen X, Cui X, Li S et al (2019) The Arabidopsis H3K27me3 demethylase JUMONJI 13 is a temperature and photoperiod dependent flowering repressor. Nat Commun 10:1303
Acknowledgements
We apologize to all colleagues whose work could not be cited in this review due to space limitations. We thank Mande Xue for helpful discussion and critical proofreading. This work was supported by grants from the National Natural Science Foundation of China (31788103 to X.C., 31801063 to Y.H.); the Chinese Academy of Sciences (Strategic Priority Research Program XDB27030201 and QYZDY-SSW-SMC022 to X.C.); the China Postdoctoral Science Foundation (2016M600143 to Y.H., 2020M680744 to Y.Y.), and the State Key Laboratory of Plant Genomics.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All the authors declare no conflict of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hou, Y., Yan, Y. & Cao, X. Epigenetic regulation of thermomorphogenesis in Arabidopsis thaliana. aBIOTECH 3, 12–24 (2022). https://doi.org/10.1007/s42994-022-00070-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42994-022-00070-9