Abstract
In this paper, we demonstrate the use of two-mode clustering for genotype by trait and genotype by environment data. In contrast to two separate (one mode) clusterings on genotypes or traits/environments, two-mode clustering simultaneously produces homogeneous groups of genotypes and traits/environments. For two-mode clustering, we first scan all two-mode cluster solutions with all possible numbers of clusters using k-means. After deciding on the final numbers of clusters, we continue with a two-mode clustering algorithm based on a genetic algorithm. This ensures optimal solutions even for large data sets. We discuss the application of two-mode clustering to multiple trait data stemming from genomic research on tomatoes as well as an application to multi-environment data on barley.
Similar content being viewed by others
Introduction
Genotype by environment interaction is the phenomenon that occurs when genotypes respond differentially to changes in the environment. An attractive approach to model genotype by environment interaction is by the identification of groups of genotypes and groups of environments that internally exhibit a certain homogeneity, thereby relegating the genotype by environment interaction to differences between the genotypic and environmental groups. A well know example of this approach is the two-way (or two-mode) clustering method described in (Corsten and Denis 1990). The same strategy of reducing the complexity of genotype by environment two-way tables by grouping genotypes and environments can also be applied to genotype by trait data matrices, provided the traits are expressed on a scale that allows direct comparison, like, for example, when the traits are all metabolite concentrations.
Clustering methods order objects (genotypes) or variables (environments, traits) in groups that are similar with respect to some measure, e.g. Euclidean distance or the correlation coefficient (Vandeginste et al. 1998). Clustering is a popular technique due to its visualization probabilities and ease of use. Regular clustering, i.e. one-way clustering, aims at finding the best partitioning in one direction of a two-way table or data matrix. The best partitioning may be defined as the clustering that results in the minimum sum of squared distances across clusters between the data assigned to a cluster and the corresponding cluster center (in other words, the total within cluster distance is minimal). As opposed to regular, one-way clustering, two-way, or two-mode, clustering aims to find the best partitioning of the data in two directions (both genotypes and environments/traits). The added benefit in comparison with one-way clustering is that it becomes immediately clear why certain objects have been clustered together, since their variables have also been clustered simultaneously.
There are different algorithms available for two-mode clustering, one example is two-mode k-means (Vichi 2001; Rocci and Vichi 2008; van Rosmalen et al. 2009). Some methods have a tendency to get stuck in local optima. Other two-mode cluster algorithms are based on global optimization methods, such as Simulated Annealing, Tabu Search (van Rosmalen et al. 2009) and Genetic Algorithms (GA) (Hageman et al. 2008b; Cavill et al. 2009). Recently, we have introduced two-mode clustering using a Genetic Algorithm in metabolomics (Hageman et al. 2008a, b). GAs work on a group of solutions at the time, using biologically inspired operators such as mutation and crossover to explore the search space. It can take large steps in the search space thereby minimizing the risk of getting trapped in a local optimum.
Two-mode clustering has shown to be a valuable tool for the identification of biological meaningful clusters in metabolomics data (Hageman et al. 2008b). It can clearly identify genotypes that behave similarly and also show simultaneously in which environments or for which traits they behave similarly. After two-mode clustering, a careful scrutiny of the genotypes and corresponding molecular markers can possibly reveal which markers are responsible for a particular phenotypic response.
We will demonstrate this by performing a genotype by trait and a genotype by environment analysis using two-mode clustering on tomato and barley data.
Materials and methods
Data
The first dataset is on tomatoes and maintained by the Center for BioSystems Genomics (CBSG, http://www.cbsg.nl/). The CBSG is a joint venture in the field of plant genomics of breeding companies, biotech companies, research institutes and universities in the Netherlands. The goal of the tomato CBSG project is to develop a marker assisted strategy for quality traits. This dataset consisted of 94 genotypes, all cultivars provided by five companies involved with the project, which fell into three major categories; cherry, round and beef tomatoes. Almost all cultivars were F1 hybrids. For two-mode clustering, we used information on the metabolites, and sensory studies. Metabolic profiles were measured using GC-MS and LC-MS, more details on this dataset can be found here (Ursem et al. 2008; van Berloo et al. 2008; Gavai et al. 2009). Traits in this context mean metabolites and sensory attributes. The data was range scaled to get all metabolites and sensory attributes at the same level.
The second example dataset is the barley Steptoe × Morex doubled haploid population (Kleinhofs et al. 1993), a well known population from the North American Barley Genome Mapping Project (http://wheat.pw.usda.gov/ggpages/SxM/). The data matrix consisted of yield of 150 genotypes in 10 environments (Table 1). The genotype by environment matrix was first column and row-centered. This means that the focus of the attention of the two-mode clustering procedure is in describing the patterns in genotype by environment interaction.
Two-mode clustering
Two-mode clustering tries to find clusters in objects and variables simultaneously, as opposed to one-way or one-mode clustering, where either objects or variables are clustered. In this work, we aim to find the optimal two-mode cluster solution between genotypes and environments or genotypes and traits. There are several algorithms available for creating two-mode clusters. In this paper we will use two techniques for finding an optimal two-mode partitioning: two-mode k-means and genetic algorithm based two-mode clustering.
In general, two-mode clustering decomposes matrix Y (which contains for our purposes genotypes by environment or genotypes by traits information) into three parts, as shown in Fig. 1:
Schematic decomposition of matrix Y for three trial solutions. Each trial solution has its own decomposition of matrix Y and consequently has its own residuals. Some trial solution will have lower residuals and therefore perform better. Matrix R and C are filled with 0’s and 1’s to give an impression of their contents
where
-
Y (I × J): data matrix of I rows and J columns
-
R (I × P): membership matrix for I rows (genotypes) of matrix of Y, allowing for P row clusters.
-
M (P × Q): matrix containing cluster averages for P row and Q column clusters
-
C (J × Q): membership matrix for J columns (environments or metabolites/traits) of matrix Y.
-
E (I × J): matrix of residuals, containing the difference between each measurement and its cluster average from matrix M.
Membership or incidence matrices R and C contain only zeros and a single one on each row and uniquely assign each genotype by environment or genotype by trait element of Y to one of the P and Q clusters. The location of the one indicates membership to that particular cluster. The quality of the two-mode cluster algorithm is largely depending on its ability to find the best solution for the membership matrices R and C. The use of global optimizers reduces the risk of reaching sub optimal solutions and is the reason we used GAs for obtaining the final solution.
Two-mode k-means and genetic algorithms two-mode clustering both use the same decomposition of matrix Y. The difference between the two methods is how they come to their final solution. Two-mode k-means works on one single solution and iteratively recalculates cluster centers and adjusts row and column cluster memberships to the nearest cluster. For a detailed discussion of two-mode k-means see (van Rosmalen et al. 2009).
The inner workings of two-mode clustering with GAs are also described elsewhere, but repeated here in short for clarity. For a more detailed discussion on two-mode clustering, GAs or the combination of the two, the reader is referred to (Corsten and Denis 1990; Vichi 2001; Hageman et al. 2003; Madeira and Oliveira 2004; Van Mechelen et al. 2004; Turner et al. 2005; Hageman et al. 2008b; van Rosmalen et al. 2009).
Genetic algorithm
GAs are a special class of global optimization routines, based on the theory of evolution. GAs minimize a function by searching the search space for an optimal solution. For two-mode clustering, GAs try to find the optimal membership matrices for row and column objects that result in a minimal within cluster distance. The GA does not operate on the membership matrices R and C itself, but rather on a vector that represents R and C in a condensed form and that contains the cluster numbers for each data entry of Y. These data entries are often interaction residuals resulting from the fit of an additive two-way analysis of variance model to a two-way table of means. Figure 2 shows how the vector translates into the membership matrices R and C. Operations on a representation vector are more efficient within a GA than operations on sparse membership matrices.
The basic GA method consists of 6 steps that are being iterated.
-
1.
Initialization: GAs work on a group of trial solutions at a time (a group of trial solutions is called a population). At the start of the GA the population is filled with random solutions which are just random assignments of data elements to clusters.
-
2.
Evaluation: each trial solution in the population is evaluated. A trial solution is a vector with cluster number assignments. In GA terminology a trial solution is called a string. In this case, for each string the total within cluster sum of squares (SS res ) is calculated as shown in the Eq. (2).
Here, K r and K c are the numbers of row and column clusters, n rc is the number of data entry points in cluster identified by the row index r and column index c, y i(r)j(c) indicates data entry (i, j) within the cluster r,c, y rc indicates the mean for the cluster r,c.
-
3.
Stop: a stop criterion is checked, usually a minimal change in SS res for the last number of iterations (called generations), otherwise a predefined maximum number of generations.
-
4.
Selection: a fraction of the best strings, that is, the ones with the smallest SS res , are selected for the next generation.
-
5.
Recombination: the selected strings are recombined (called crossover) to yield new strings.
-
6.
Mutation: small random changes (called mutation) are applied to the new strings.
Figure 3 shows an example of the 2 point crossover (top part) and mutation (bottom part).
Examples for two point crossover (top part) and mutation (bottom part). The vertical lines in the top part indicate the cutting locations. At these cutting locations the strings will be disconnected and some parts will be exchanged (recombined) with another string. At the bottom part, the cross indicates the cluster assignment that will be randomly changed (mutated)
An important aspect of GAs is choosing adequate values for the parameters defining the GA itself (Hageman et al. 2003). This can be done with trial and error or with an experimental design. In this case we used the parameters from our previous study with metabolomics data (Hageman et al. 2008b).
Numbers of row and column clusters
The decomposition as shown in Eq. 1 requires a predefined number of row and column clusters (as indicated with P and Q), which is usually unknown beforehand. There are a number of methods available for estimating the optimal number of clusters (e.g. BIC, GAP statistic, knee/L/scree plots) (Milligan and Cooper 1985; Salvador and Chan 2004). We used the knee method, where the number of row and column clusters are plotted against SS res , the squared within cluster distance (Hageman et al. 2008b). The point where the increase in the number of clusters only marginally decreases SS res is evidenced in the graph by a knee or L shape. This point is regarded as the optimal numbers of clusters. Since the creation of the knee plot requires the calculation of SS res for all possible combinations of numbers of clusters, to save computation time, this stage was performed using two-mode k-means clustering. Although two mode k-means can get stuck in a local optimum, the global shape and trends of the knee plot will still show us how many clusters can be considered optimal. After the choice for a particular numbers of clusters has been made, the two-mode clustering is repeated from scratch with the GA based two-mode clustering using the cluster numbers obtained with two-mode k-means. The idea is that if two-mode k-means is stuck in a local optimum, the GA based two-mode clustering may overcome such a local optimum due to the nature of its optimization approach, and find a better two-mode partitioning.
Software
Two-mode k-means clustering and GA based two-mode clustering were programmed in Matlab 7.1 (Mathworks 2008), the latter using the Genetic Algorithm and Direct Search toolbox. All GA runs were performed in five fold to exclude any (un)lucky starting positions. The settings for the GA and k-means two-mode cluster algorithms can be found in Table 2. All calculations were performed on an Intel Core 2 CPU at 1.86 GHz.
To compare the results from two-mode clustering, the tomato data set is also analyzed using principal component analysis. For comparison, an AMMI model (van Eeuwijk et al. 2005) was fitted to the barley data. A mixed model multi-environment QTL mapping was performed following the methods as described by (Boer et al. 2007), and in a more basic form presented in (Malosetti et al. 2004). All these analyses were performed in GenStat 12th edition (Payne et al. 2009).
Results
Tomato data
To obtain an estimate for the correct number of clusters in each direction, all possible combinations of cluster numbers between two and eight were calculated using two-mode k-means. Figure 4 shows the knee plot for the tomato dataset (left part). Two-mode k-means is likely to find a local optimum, but will nevertheless provide a good idea on the correct numbers of clusters. When deciding on the numbers of clusters the biological interpretation of the resulting clustering has also been taken into account. The numbers of clusters for the tomato dataset were chosen as three clusters in the genotype direction and four clusters in the metabolites/traits direction.
The tomato data set has been clustered using the two-mode k-means algorithm using three genotype clusters and four metabolites/trait clusters. The GA was not able to find a solution with a lower residual error, indicating that two-mode k-means was also able to find the same solution. The relative small numbers of clusters make it probably not too difficult to find this solution. Figure 5 shows the two-mode clustering result for the tomato dataset.
The greatest effect in this data set is the difference between cherry tomatoes and the others varieties. The clustering on genotype clearly shows the distinction between the cherry tomatoes and the beef and round. One of the genotype clusters solely consists of cherry tomatoes (cluster three). The inspection of the trait direction can provide insights into what is causing this separation. Trait clusters three and four give a clear contrast between cherry tomatoes and the rest of tomato types. Cluster three shows a higher than average concentration of glucose, fructose and sucrose (and corresponding properties like brix and the sensory attribute taste ‘sweet’). Cluster four contains sensory attributes ‘mouth feel mealy’, ‘taste unripe’ and ‘taste watery’ which are all below average for the cherry tomatoes. Property ‘fruit weight’ is also below average for the cherry tomatoes, which is expected as they are smaller in comparison to the other types. Trait’s cluster one contains the sensory trait ‘taste tomato’ together with the metabolite isobutylthiazole, one of the odorants associated with the smell of tomatoes. None of the tomatoes types showed a higher concentration of isobutylthiazole, suggesting that all these tomatoes taste equally well like tomatoes.
To compare the two-mode cluster results with other data analyses techniques, we present a PCA plot in Fig. 6. This figure also clearly shows the distinction between cherry tomatoes (yellow circles) and the other ones. The PCA plot also indicates that cherry tomatoes are more ‘taste sweet’ and ‘scent sweet’. Beef and round tomatoes (red and blue circles) are also not separated in the plot. These tomatoes are more ‘taste watery’, ‘mouth feel mealy’ and ‘taste unripe’ in comparison with the cherry tomatoes.
Barley data
The right part of Fig. 4 shows the knee plot for the barley dataset. The numbers of clusters for the barley data were chosen to be six genotype clusters and four environment clusters. The GA was able to find a better solution compared to the k-means solution, the within cluster distance for the GA solution was 3.4% lower. Figure 7 shows the GA two-mode clustering result for the barley data set. The two-way clustering reflects combinations of genotypes and environments showing a positive genotype by environment interaction, that is, environments where the performance of genotypes deviates upwards from additivity of environmental and genotypic effects. For example, the genotypes in group one (upper left corner) had a positive interaction with environments ID92, MTd91, and MTi91 (environment group one). The clustering discriminates between sets of genotypes having positive interaction in one or more sets of environments. For example, while genotypes in cluster one showed a positive genotype by environment interaction with environment group one and two (MTd92, MTi92, and WA92), genotypes in group two showed a positive interaction with environment group one and three (ID91 and WA91). Positive interactions were observed between genotype’s group three and environment groups two (MTd92, MTi92, and WA92) and four (MAN92, and SKs92). Similar patterns can be observed for the other groups of genotypes. In summary, the two-mode clustering allowed to graphically display groups of genotypes and environments that had a positive interaction. Since the main effect is not included, a best performance combination of genotypes and environments can not be inferred from this graph, but it would point to favorable interaction patterns potentially pointing to specific adaptation patterns.
Results from two-mode clustering of Steptoe × Morex barley data. Red colors indicate interaction residuals above average, black colors around the average and green colors below average. See Table 3 for full contents of markers per genotype cluster
We can directly compare the results from the two-mode clustering with the AMMI biplot in Fig. 8. The AMMI biplot has been created by performing PCA on the interaction residuals after row and column centering of the tomato data. We can easily recognize groups of genotypes that are close to each other and clustered together with two-mode clustering. Examples are the cluster ID91, MTd91, MTi91 and WA92, MTi92, Mtd92. The only discrepancy between the AMMI biplot and the two-mode clustering is that SKs92 and WA92 are close in the biplot but not clustered together in the two-mode clustering. Perhaps SKs92 and WA92 are not that close when taking higher principal components into account.
Although molecular marker information was not used during the clustering, the examination of the correspondence between marker genotypes and genotype clusters can reveal some interesting patterns. An association between molecular marker genotype and genotypic groups can reveal chromosome regions linked to specific adaptation. Markers almost fixed within genotypic groups (81–100% homozygous for a particular allele) are given in Table 3.
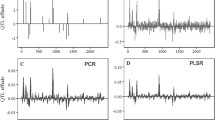
From Table 3, it is remarkable that most of the markers that were found fixed (or almost fixed) inside the genotypic groups map to chromosome two (around 30–60 cM), three (around 70–100 cM) and seven (around 50–70 cM). Chromosome two, three, and seven have been shown to harbor the most important QTLs explaining G × E, that is, QTL by environment interaction (QTL × E). This is also confirmed by the results of QTL analysis in Fig. 9 which shows the important QTLs being located on chromosome two, three and seven.
Results of a multi-environment QTL analysis on Steptoe × Morex barley data. The upper part of the figure presents the profile of the associated P value (on a log10 scale) of the H0 of no QTL effect in any of the environments (for environments abbreviations see Table 1). The lower plot gives for every chromosome position the magnitude of the QTL effects in each of the environments, the higher the intensity of the color the larger the QTL effect (white equals to no effect). The color indicates which of the two parents contributed the high value allele (blue = Steptoe, red = Morex)
The two-mode clustering could then be seen as a quick step to sample potentially interesting markers associated to the patterns of variation in the column direction (either environment or traits).
Concluding remarks
We have demonstrated the use of two-mode clustering to explore genotype by trait and genotype by environment data. We have first examined different numbers of clusters in the two-mode clustering using k-means and the knee plot. After deciding on the final numbers of clusters, a GA based two-mode clustering was used for finding a better solution than k-means. The GA only found a better solution for the genotype by environment data.
Two-mode clustering is able to extract relevant features from both data sets. It finds genotypes with a similar response in particular sets of environments (barley data set) or set of genotypes that share some common set of characteristics (tomato data set). In the interpretation of the two-mode clustering, external information can be useful. For example, in the barley data set, a closer analysis of the genotype clusters in relation to molecular markers provided information about relevant markers associated to the differential response of genotypes in particular environments.
Two-mode clustering as presented in this paper is an easy to use tool and gives a clear and straightforward to interpret graphical overview of a multi-dimensional data set, capturing the most relevant features of the data set under study. Our implementation of two-mode clustering is very similar in spirit to the two-way clustering algorithm presented by (Corsten and Denis 1990), but it seems easier to generalize to other contexts like three-mode cluster analysis or clustering procedures for generalized linear models in place of linear models. We are currently working on such types of extensions to the two-mode k-means and GA two means clustering algorithm.
References
Boer MP, Wright D, Feng LZ, Podlich DW, Luo L, Cooper M, van Eeuwijk FA (2007) A mixed-model quantitative trait loci (QTL) analysis for multiple-environment trial data using environmental covariables for QTL-by-environment interactions, with an example in maize. Genetics 177:1801–1813
Cavill R, Keun HC, Holmes E, Lindon JC, Nicholson JK, Ebbels TMD (2009) Genetic algorithms for simultaneous variable and sample selection in metabonomics. Bioinformatics 25:112–118
Corsten LCA, Denis JB (1990) Structuring interaction in 2-way tables by clustering. Biometrics 46:207–215
Gavai AK, Tikunov Y, Ursem R, Bovy A, van Eeuwijk F, Nijveen H, Lucas PJF, Leunissen JAM (2009) Constraint-based probabilistic learning of metabolic pathways from tomato volatiles. Metabolomics 5:419–428
Hageman JA, Streppel M, Wehrens R, Buydens LMC (2003) Wavelength selection with Tabu search. J Chemometr 17:427–437
Hageman JA, Hendriks M, Westerhuis JA, van der Werf MJ, Berger R, Smilde AK (2008a) Simplivariate models: ideas and first examples. Plos One 3(9):1–12
Hageman JA, van den Berg RA, Westerhuis JA, van der Werf MJ, Smilde AK (2008b) Genetic algorithm based two-mode clustering of metabolomics data. Metabolomics 4:141–149
Kleinhofs A, Kilian A, Maroof MAS, Biyashev RM, Hayes P, Chen FQ, Lapitan N, Fenwick A, Blake TK, Kanazin V, Ananiev E, Dahleen L, Kudrna D, Bollinger J, Knapp SJ, Liu B, Sorrells M, Heun M, Franckowiak JD, Hoffman D, Skadsen R, Steffenson BJ (1993) A molecular, isozyme and morphological map of the barley (Hordeum-Vulgare) genome. Theor Appl Genet 86:705–712
Madeira SC, Oliveira AL (2004) Biclustering algorithms for biological data analysis: a survey. IEEE/ACM Trans Comput Biol Bioinform 1:24–45
Malosetti M, Voltas J, Romagosa I, Ullrich SE, van Eeuwijk FA (2004) Mixed models including environmental covariables for studying QTL by environment interaction. Euphytica 137:139–145
Mathworks (2008) Matlab 7.1
Milligan GW, Cooper MC (1985) An examination of procedures for determining the number of clusters in a data set. Psychometrika 50:159–179
Payne R, Harding S, Murray D, Soutar D, Baird D, Glaser A, Channing I, Welham S, Gilmour A, Thompson R, Webster R (2009) The guide to GenStat release 12, part 2: statistics. VSN International, Hemel Hempstead
Rocci R, Vichi M (2008) Two-mode multi-partitioning. Comput Stat Data Anal 52:1984–2003
Salvador S, Chan P (2004) Determining the number of clusters/segments in hierarchical clustering/segmentation algorithms. Proceedings of the 16th IEEE international conference on tools with artificial intelligence (ICTAI 2004)
Turner HL, Bailey TC, Krzanowski WJ, Hemingway CA (2005) Biclustering models for structured microarray data. IEEE/ACM Trans Comput Biol Bioinform 2:316–329
Ursem R, Tikunov Y, Bovy A, van Berloo R, van Eeuwijk F (2008) A correlation network approach to metabolic data analysis for tomato fruits. Euphytica 161:181–193
van Berloo R, Zhu AG, Ursem R, Verbakel H, Gort G, van Eeuwijk FA (2008) Diversity and linkage disequilibrium analysis within a selected set of cultivated tomatoes. Theor Appl Genet 117:89–101
van Eeuwijk FA, Malosetti M, Yin XY, Struik PC, Stam P (2005) Statistical models for genotype by environment data: from conventional ANOVA models to eco-physiological QTL models. Aust J Agric Res 56:883–894
Van Mechelen I, Bock HH, De Boeck P (2004) Two-mode clustering methods: a structured overview. Stat Methods Med Res 13:363–394
van Rosmalen J, Groenen PJF, Trejos J, Castillo W (2009) Optimization strategies for two-mode partitioning. J Classif 26:155–181
Vandeginste BGM, Massart DL, Buydens LMC, Jong SD, Lewi PJ, Smeyers-Verbeke J (1998) Handbook of chemometrics. Elsevier, Amsterdam
Vichi M (2001) Double k-means clustering for simultaneous classification of objects and variables. Springer, Heidelberg
Acknowledgments
This project was (co)financed by the Centre for BioSystems Genomics (CBSG) which is part of the Netherlands Genomics Initiative/Netherlands Organisation for Scientific Research.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Hageman, J.A., Malosetti, M. & van Eeuwijk, F.A. Two-mode clustering of genotype by trait and genotype by environment data. Euphytica 183, 349–359 (2012). https://doi.org/10.1007/s10681-010-0236-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-010-0236-6