Abstract
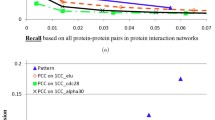
We develop a technique to validate large-scale gene regulatory networks (GRN) by comparing with corresponding protein-protein interaction (PPI) networks. The GRN are obtained with Bayesian networks while PPI networks are obtained from database of known PPI interactions. We look for exact matches and then reduced networks by skipping one or more genes in GRN. We demonstrate our technique on expression profiles of differentially expressed genes in the S. cerevisiae cell cycle. We validate GRNs against a merged database of 53235 genes. The precisions of GRN obtained over all genes were from 0.82 to 0.95 in all the phases. In particular we realized that one-skip and two-skip model significantly improved accuracy of the GRN of different phases of cell cycle.
Chapter PDF
Similar content being viewed by others
Keywords
References
Nikolsky, Y.: Biological networks and analysis of experimental data in drug discovery. Drug discovery today 10(9), 653 (2005)
Wyrick, J.J., Young, R.A.: Deciphering gene expression regulatory networks. Curr. Opin. Genet. Dev. 12(2), 130–136 (2002)
Fromont-Racine, M., Rain, J.C., Legrain, P.: Toward a functional analysis of the yeast genome through exhaustive two-hybrid screens. Nature Genetics 16, 277–282 (1997)
Duda, R.O.: Pattern classification [Book]
Friedman, N., Linial, M., Nachman, I., Pe’er, D.: Using bayesian networks to analyze expression data. J. Computational Biology 7(3), 601–620 (2000)
Lauritzen, S.: Graphical Models. Oxford University Press, Oxford (1996)
Luo, F., Tang, K., Khan, L.: Hierarchical clustering of gene expression data. Bioinformatics and Bioengineering, 2003. In: Proceedings. Third IEEE Symposium, 10-12 March, pp. 328–335 (2003)
Wei, W., Xin, L., Min, X., Jinrong, P., Setiono, R.: A hybrid SOM-SVM method for analyzing zebra fish gene expression. Pattern Recognition, 2004. In: ICPR 2004. Proceedings of the 17th International Conference, vol. 2, pp. 323–326 (2004)
Alter, O., Brown, P.O., Botstein, D.: Singular value decomposition for genome-wide expression data processing and modeling. Proc. Natl. Acad. Sci. USA 97, 10101–10106 (2000)
Lee, S., Batzoglou, S.: Application of Independent component analysis to microarrays. Genome Biology 4, R76 (2003)
Bar-Joseph, Z.: Analyzing Time Series Gene Expression Data. Bioinformatics 20(16), 2493–2503 (2004)
Fang-Xiang, et al.: A Genetic Algorithm for Inferring Time Delays in Gene Regulatory Networks, CSB (2004)
Wu, C.C., Huang, H., Juan, H., Chen, S.: Gene Networks: an interactive tool for reconstruction of genetic networks from microarray data. Bioinformatics advanced access (2004)
Liang, S., Fuhrman, S., Somogyi, R.: REVEAL: a general reverse engineering algorithm for inference of genetic network architectures. In: Pacific Symposium on Biocomputing, vol. 3, pp. 18–29 (1998)
Friedmann, N., Linial, M., Nachman, I., Pe’er, D.: Using Bayesian Networks to Analyze expression data. Journal of Computational Biology 7, 601–620 (2000)
Heckerman, D., Geiger, D., Chickering, D.M.: Learning Bayesian networks: The combination of knowledge and statistical data. Machine Learning 9, 309–347 (1999)
Friedman, N., Murphy, K., Russell, S.: Learning the structure of dynamic probabilistic networks. In: Proc. Fourteenth Conference on Uncertainty in Artificial Intelligence (UAI 1998), pp. 139–147 (1998)
Troyanskaya, O., et al.: Missing value estimation methods for DNA microarray. Bioinformatics 17(6), 520–525 (2001)
Davierwala, A.P., et al.: Synthetic genetic interaction spectrum of essential genes. Nature Genetics 37(10), 1147–1152 (2005)
Krogan, N.J., et al.: High-definition macromolecular composition of yeast RNA-processing complexes. Mol. Cell 13(2), 225–239 (2004)
Young, K.H.: Yeast Two-Hybrid: So Many Interactions (in) So Little Time. Biology of reproduction 58, 302–311 (1998)
Page, D., Ong, I.M.: Experimental design of time series data for learning from dynamic Bayesian networks. PSB 11, 267–278 (2006)
Bahler, J.: Cell-Cycle Control of Gene Expression in Budding and Fission Yeast. Annu. Rev. Genet. 39, 69–94 (2005)
Spellman, et al.: Comprehensive Identification of Cell Cycle–regulated Genes of the Yeast Saccharomyces cerevisiae by Microarray Hybridization. Molecular Biology of the Cell 9, 3273–3297 (1998)
Stark, C., Breitkreutz, B.J., Reguly, T., Boucher, L., Breitkreutz, A., Tyers, M.: BioGRID: a general repository for interaction datasets. Nucleic Acids Res. 34, D535–D539 (2006)
Author information
Authors and Affiliations
Editor information
Rights and permissions
Copyright information
© 2007 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Chaturvedi, I., Sakharkar, M.K., Rajapakse, J.C. (2007). Validation of Gene Regulatory Networks from Protein-Protein Interaction Data: Application to Cell-Cycle Regulation. In: Rajapakse, J.C., Schmidt, B., Volkert, G. (eds) Pattern Recognition in Bioinformatics. PRIB 2007. Lecture Notes in Computer Science(), vol 4774. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-540-75286-8_29
Download citation
DOI: https://doi.org/10.1007/978-3-540-75286-8_29
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-75285-1
Online ISBN: 978-3-540-75286-8
eBook Packages: Computer ScienceComputer Science (R0)