Abstract
This paper presents a fully convolutional network-based segmentation method to create an eyeball model data for patient-specific ophthalmologic surgery simulation. In order to create an elaborate eyeball model for each patient, we need to accurately segment eye structures with different sizes and complex shapes from high-resolution images. Therefore, we aim to construct a fully convolutional network to enable accurate segmentation of anatomical structures in an eyeball from training on sparsely-annotated images, which can provide a user with all annotated slices if he or she annotates a few slices in each image volume data. In this study, we utilize a fully convolutional network with full-resolution residual units that effectively learns multi-scale image features for segmentation of eye macro- and microstructures by acting as a bridge between the two processing streams (residual and pooling streams). In addition, a weighted loss function and data augmentation are utilized for network training to accurately perform the semantic segmentation from only sparsely-annotated axial images. From the results of segmentation experiments using micro-CT images of pig eyeballs, we found that the proposed network provided better segmentation performance than conventional networks and achieved mean Dice similarity coefficient scores of 91.5% for segmentation of eye structures even from a small amount of training data.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Semantic segmentation of medical images is an essential technique for creating anatomical model data that are available for surgical planning, training, and simulation. In the field of ophthalmology, elaborate artificial eyeball models [1, 2] have been developed for training and simulation of eye surgeries, and it is desired to create realistic eyeball model data for patient-specific surgical simulation through the segmentation of detailed eye structures. Thus, we focus on segmenting not only the entire eyeball structure but also microstructures (e.g., Zinn’s zonule) in the eyeball, which conventional modalities such as computed tomography (CT) have difficulty capturing, by using higher-resolution modalities such as micro CT.
To efficiently create patient-specific eyeball model data from high-resolution images, we need to take into account the following three points: (a) full- or semi-automation of segmentation for reducing the burden of manual annotation, (b) accurate extraction of eye structures with different sizes and complex shapes, and (c) image processing at full resolution without downsampling. Therefore, we utilize a fully convolutional network (FCN) [3], which is one of the most powerful tools for end-to-end semantic segmentation, to construct a segmentation method to fulfill the key points.
For accurate segmentation of objects with different sizes and complex shapes in the images, it is important to construct a network architecture that can obtain image features for localization and recognition of the objects. In general, deep convolutional neural networks can obtain coarse image features for recognition on deep layers and fine image features for localization on shallow layers. Many studies [3,4,5,6] have proposed a network architecture to obtain multi-scale image features for semantic segmentation by residual units (RUs) or skip connections, which combine different feature maps output from different layers. U-net proposed by Ronneberger et al. [6] achieved good performance for semantic segmentation of biomedical images by effectively using long-range skip connections. Moreover, their research group showed that 3D U-net [7], which was developed as the extended version of U-net, could provide accurate volumetric image segmentation based on training from sparsely-annotated images on three orthogonal planes. However, such 3D FCNs have difficulty handling images at full resolution and obtaining full-resolution image features essential for strong localization performance because of the limitation of GPU memory.
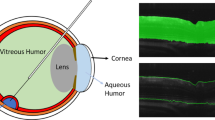
Therefore, we aim to construct a 2D network architecture that provides improved localization and recognition for semantic segmentation of high-resolution medical images by using advanced RUs instead of conventional skip connections found in FCN-8s [3] or U-net [6]. Moreover, we also aim to propose a training strategy in which the network can learn from sparsely-annotated images and provide accurate label propagation to the remaining images in volumetric image data, because it is not easy to collect a large amount of high-resolution image volumes for network training from different cases. The concept of our proposed method is shown in Fig. 1. The originality of this study lies in introducing a FCN with the advanced RUs and its training strategy to achieve accurate segmentation of eye structures in an end-to-end fashion even from sparsely-annotated volumetric images.
2 Methods
2.1 Network Architecture
In this study, we focus on full-resolution residual units (FRRUs) [8], which was designed to facilitate the combination of multi-scale image features while keeping similar training characteristics as ResNet [9]. We utilize the network architecture that consists of four pooling steps followed by four upsampling steps like U-net [6] as a base and construct a residual-based FCN incorporating FRRUs into the basal network architecture to enhance the localization and recognition performances for segmentation of eye structures. Figure 2 shows the architectures of U-net and the proposed network. The box in the figure represents a feature map output by each convolution layer or FRRU and the number of channels is denoted under the box. U-net fuses the same-size feature maps between pooling stages and upsampling stages with skip connections, while the proposed network jointly computes image features on two processing streams by using FRRUs. One stream (i.e., residual stream) conveys full-resolution fine image features for localization, which are obtained by adding successive residuals, and the other stream (i.e., pooling stream) conveys coarse image features for recognition, which are computed through convolution and pooling steps.
The detail of a FRRU structure is indicated in Fig. 3. Each classical RU [9] has one input and one output, while each FRRU computes two outputs from two inputs. Let \(x_n\) and \(y_n\) be the residual and the pooling inputs to n-th FRRU, respectively. Then, the outputs are computed as follows:
where \(W_n\) denote the parameters of the residual function \(\mathcal{G}\) and the pooling function \(\mathcal{H}\). As shown in Fig. 3, the FRRU concatenates the pooling input with the residual input operated by a pooling layer, and subsequently obtains the concatenated features (i.e., the output of the function \(\mathcal{H}\)) through two 3 \(\times \) 3 convolution layers. The output of \(\mathcal{H}\) is passed to the next layer as the pooling stream. Moreover, the output of \(\mathcal{H}\) are also resized by the function \(\mathcal{G}\) and reused as features added to the residual stream. This design of the FRRU makes it possible to combine and compute the two stream simultaneously and successively.
Therefore, the proposed network, which are composed of a sequence of FRRUs, gains the ability to precisely localize and recognize objects in images by combining the following two processing streams: the residual stream that carries fine image features at full resolution and the pooling stream that carries image features obtained through a sequence of convolution, pooling, and deconvolution operations.
Network architectures: (a) U-net [6] and (b) the proposed network
Design of full-resolution residual unit (FRRU) [8]
2.2 Training Strategy
We assume that the proposed network is applied to eye structures segmentation based on sparse annotation. Thus, we need to construct a framework to enable the network to effectively learn image features even from less annotated slices for training.
In the case of our application, it is expected that the training and testing subsets of images have no significant differences of geometric and visual characteristics (e.g., location, scale, or contrast) between objects for segmentation because they are derived from the same image volume. Therefore, we here adopt rotation and elastic deformation for data augmentation to efficiently train small geometric variations of eye structures in the images based on less annotated slices for training, although there are many techniques for increasing the amount of training data. Each slice in the training subset is augmented twentyfold by rotating \(-25^\circ \) to \(25^\circ \) at 5 degree intervals and repeating the elastic deformation ten times based on random shifts of \(5 \times 5\) grid points and B-spline interpolation.
Additionally, for more effective network training, we use categorical cross-entropy loss function weighted by the inverse of class frequency to reduce the negative effects of class imbalance (i.e., difference of sizes between different eye structures in the images).
3 Experiments and Results
3.1 Experimental Setup
We validated the segmentation performance of the proposed method on a dataset of eyeball images, which were scanned using a micro-CT scanner (inspeXio SMX-90CT Plus, Shimadzu Co., Japan). The dataset consists of micro-CT volumes of five pig eyeballs, and the size of each volume is 1024 \({\times }\) 1024 \({\times }\) 548 (sagittal \({\times }\) coronal \({\times }\) axial) voxels, with a voxel size of 50 \({\upmu }\)m. Figure 4 shows an example of micro-CT images and label images used for the validation. As a preprocessing step, the original micro-CT images were filtered by using a wavelet-FFT filter [10] and a median filter to remove the ring artifacts and random noises, and subsequently the filtered images were normalized based on the mean and standard deviation on the training subset of images for each micro-CT volume. We defined six labels, including Background, Wall and membrane, Lens, Vitreum, Ciliary body and Zinn’s zonule, and Anterior chamber. The preprocessed images and the corresponding manually annotated images were used for network training and testing.
In this study, for fundamental comparative evaluation, we compared our network with the following two representative networks: FCN-8s [3] and U-net [6]. To evaluate the segmentation performances associated with network architectures, all the networks were trained and tested on the same datasets under the same conditions (i.e., the same learning rate, optimizer, and loss function were assigned to the networks). On the assumption of the semantic segmentation from sparse annotation, 2.5% (i.e., every 40 slices) of all the slices and the remaining slices on the axial plane in each volume were used as training and testing subsets, respectively. The slices of each training subset were augmented by the two data augmentation techniques (i.e., rotation and elastic deformation). Each of the networks was trained from scratch on the augmented training subset of slices for 100 epochs and tested on the testing subset. The segmentation performances were quantitatively and qualitatively evaluated by comparing Dice similarity coefficient (DSC) scores and visualization results between the networks. The networks used for experiments were implemented using KerasFootnote 1 with the Tensorflow backendFootnote 2, and all the experiments were performed on a NVIDIA Quadro P6000 graphic card with 24 GB memory.
3.2 Experimental Results
Table 1 indicates the comparison results of DSC scores of the three networks, including FCN-8s, U-net, and the proposed network. The proposed network could segment eye structures with a mean Dice score of 91.5% and achieve the best segmentation performance of the three networks. In addition, the results showed that the proposed network could segment almost all the labels with higher mean score and lower standard deviation than the other networks. Even on the label of “Ciliary body & Zinn’s zonule” that is hard to segment because of the high variability of shapes, the proposed network provided mean DSC score of more than 85%.
Figure 5 visualizes a part of the segmentation results obtained by the three networks. FCN-8s generalized the segmentation results with jagged edges near the label boundaries, and U-net produced segmentation results including some errors despite the smooth label boundaries. Compared to these conventional networks, we could find that the proposed network generalized more accurate segmentation results with smoother edges for all labels than the other networks.
4 Discussion
As indicated in Table 1, the proposed network achieved high mean DSC scores with low standard deviation for segmenting eye structures from sparse annotation, although only 2.5% of all the slices (i.e., 14 of 548 slices) were used for network training. The proposed network could consistently achieve higher accuracy for segmentation of eye structures with different sizes and shapes, compared to FCN-8s and U-net. This is probably because the proposed network succeeded in learning more robust image features against the change of sizes and shapes in the images. In other words, these results imply a FRRU contributes to obtaining finer features for strong localization.
In addition, Fig. 5 showed that the proposed network could generalize segmentation results with more accurate and smoother class boundaries compared to FCN-8s and U-net, although it produced some false positives. This can be considered to be due to the fact that the loss of fine image features occurred in the training process, especially in the pooling operations. Although both of them had skip connections for obtaining multi-scale features, it is probably difficult to convey image features for precise localization by only the conventional skip connections. Therefore, the network architecture incorporating FRRUs can be very effective to learn multi-scale image features, which conventional architectures have difficulty capturing.
However, even the network with FRRUs failed to provide accurate segmentation results on some slices. Thus, in future work, we will aim to further improve the segmentation accuracy of our network by combining other strategies for obtaining multi-scale image features (e.g., dilated convolutions [11]), and then we will apply our network to segmentation of finer eye structures from higher-resolution images such as X-ray refraction-contrast CT images [12] to create more elaborate eyeball model.
5 Conclusion
In this study, we proposed a FCN architecture and its training scheme for segmenting eye structures from high-resolution images based on sparse annotation. The network architecture consists of a sequence of FRRUs, which enable to effectively combine multi-scale image features for localization and recognition. Experimental results on micro-CT volumes of five pig eyeballs showed that the proposed network outperformed conventional networks and achieved mean segmentation accuracy of more than 90% by training with the weighted loss function on the augmented data, even from very few annotated slices. The proposed segmentation method may have the potential to help create an eyeball model for patient-specific eye surgery simulation.
Notes
References
Joag, M.G., et al.: The bioniko ophthalmic surgery model: an innovative approach for teaching capsulorhexis. Investig. Ophthalmol. Vis. Sci. 55(13), 1295–1295 (2014)
Someya, Y., et al.: Training system using bionic-eye for internal limiting membrane peeling. In: 2016 International Symposium on Micro-NanoMechatronics and Human Science (MHS), pp. 1–3. IEEE (2016)
Long, J., Shelhamer, E., Darrell, T.: Fully convolutional networks for semantic segmentation. In: IEEE Conference on Computer Vision and Pattern Recognition, pp. 3431–3440 (2015)
Badrinarayanan, V., Kendall, A., Cipolla, R.: Segnet: a deep convolutional encoder-decoder architecture for image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 39(12), 2481–2495 (2017)
Lin, G., Milan, A., Shen, C., Reid, I.: Refinenet: multi-path refinement networks for high-resolution semantic segmentation. In: IEEE Conference on Computer Vision and Pattern Recognition, pp. 1925–1934 (2017)
Ronneberger, O., Fischer, P., Brox, T.: U-Net: convolutional networks for biomedical image segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 234–241. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24574-4_28
Çiçek, Ö., Abdulkadir, A., Lienkamp, S.S., Brox, T., Ronneberger, O.: 3D U-Net: learning dense volumetric segmentation from sparse annotation. In: Ourselin, S., Joskowicz, L., Sabuncu, M.R., Unal, G., Wells, W. (eds.) MICCAI 2016. LNCS, vol. 9901, pp. 424–432. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-46723-8_49
Pohlen, T., Hermans, A., Mathias, M., Leibe, B.: Full-resolution residual networks for semantic segmentation in street scenes. In: IEEE Conference on Computer Vision and Pattern Recognition, pp. 4151–4160 (2017)
He, K., Zhang, X., Ren, S., Sun, J.: Deep residual learning for image recognition. In: IEEE Conference on Computer Vision and Pattern Recognition, pp. 770–778 (2016)
Münch, B., Trtik, P., Marone, F., Stampanoni, M.: Stripe and ring artifact removal with combined wavelet-Fourier filtering. Opt. Express 17(10), 8567–8591 (2009)
Yu, F., Koltun, V.: Multi-scale context aggregation by dilated convolutions. In: International Conference on Learning Representations (ICLR) (2016)
Sunaguchi, N., Yuasa, T., Huo, Q., Ichihara, S., Ando, M.: X-ray refraction-contrast computed tomography images using dark-field imaging optics. Appl. Phys. Lett. 97(15), 153701 (2010)
Acknowledgments
Parts of this work were supported by the ImPACT Program of Council for Science, Technology and Innovation (Cabinet Office, Government of Japan), the JSPS KAKENHI (Grant Numbers 26108006, 17K20099, and 17H00867), and the JSPS Bilateral International Collaboration Grants.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Switzerland AG
About this paper
Cite this paper
Sugino, T., Roth, H.R., Oda, M., Mori, K. (2018). Fully Convolutional Network-Based Eyeball Segmentation from Sparse Annotation for Eye Surgery Simulation Model. In: Stoyanov, D., et al. Simulation, Image Processing, and Ultrasound Systems for Assisted Diagnosis and Navigation. POCUS BIVPCS CuRIOUS CPM 2018 2018 2018 2018. Lecture Notes in Computer Science(), vol 11042. Springer, Cham. https://doi.org/10.1007/978-3-030-01045-4_14
Download citation
DOI: https://doi.org/10.1007/978-3-030-01045-4_14
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-01044-7
Online ISBN: 978-3-030-01045-4
eBook Packages: Computer ScienceComputer Science (R0)