Abstract
The composition of the upper respiratory tract microbiome may play an important role in the development of lower respiratory tract infections. Here, we characterised the microbiome of the nasopharynx and oropharynx of hospitalised patients with community-acquired pneumonia (CAP) with unknown aetiology in an attempt to obtain insight into the aetiology of CAP. A random sample of 10 patients hospitalised with CAP previously enrolled in a separate clinical trial (ClinicalTrials.gov registry, Study ID: NCT01248715) in which a complete microbiological workup was not able to define an aetiology were analysed in this pilot study. This larger trial (n = 1,221) enrolled patients from 9 adult hospitals in Louisville, Kentucky, USA. Nasopharyngeal and oropharyngeal swabs were obtained for metagenomic analysis. Polymerase chain reaction (PCR) for Streptococcus pneumoniae was performed in all patients. One patient had a distinct nasophararyngeal microbiome consisting largely of Haemophilus influenzae. This was the only patient with a negative PCR for S. pneumoniae in both nasophararyngeal and oropharyngeal specimens. Overall, substantial differences were found between nasophararyngeal and oropharyngeal microbiomes. The upper respiratory tract microbiome of only one patient suggested H. influenzae as a probable aetiology of CAP. Although this was a pilot study of only 10 patients, the presence of S. pneumoniae in the upper respiratory tract of the other 9 patients warrants further investigation.
Similar content being viewed by others
1. Introduction
Community-acquired pneumonia (CAP) is the leading cause of infectious disease-related death in the United States [1]. The most common bacterial aetiology of CAP in hospitalised patients is Streptococcus pneumoniae, but other important pathogens include Haemophilus influenzae, Staphylococcus aureus, Mycoplasma pneumoniae, Chlamydia pneumoniae, and Legionella pneumophila [2]. One important clinical challenge in the management of hospitalised patients with CAP is that an aetiologic diagnosis cannot be reached in a significant number of patients even when applying state of the art microbiological methods [2]. Several possibilities may explain why an aetiology is not identified in some patients: 1) new unrecognised pathogens may be present; 2) current microbiological methods may not be sensitive enough to detect the pathogens; 3) some pathogens that we normally recognise as non-pathogenic colonisers of the upper respiratory tract may in fact be able to cause CAP.
Colonisation of the upper respiratory tract by pathogenic organisms, followed by microaspiration, is considered the primary pathogenesis of pneumonia [3]. It has been speculated that the normal flora of the upper respiratory tract may play an important role in the colonisation with pathogenic flora. However, this “normal flora” of the upper respiratory tract has not been well defined, particularly for patients with lower respiratory tract infections. With the new capabilities to fully characterise the human microbiota, the possibility now exists to better understand the bacterial communities in the upper airway.
We recently completed a prospective, randomised clinical trial evaluating the role of empiric antiviral therapy in the management of hospitalised patients with lower respiratory tract infections due to influenza virus (ClinicalTrials.gov registry, Study ID: NCT01248715). As part of this trial, hospitalised patients with CAP had a comprehensive microbiological workup including sputum and blood cultures; urinary antigen tests for S. pneumoniae and L. pneumophila; oropharyngeal swabs for polymerase chain reaction (PCR) detection of M. pneumoniae, L. pneumophila, and C. pneumoniae; as well as a nasopharyngeal swab for PCR detection of 12 respiratory viruses. An aetiologic diagnosis was not reached in many patients with CAP in whom this complete microbiological workup was performed.
In an attempt to better characterise the upper respiratory tract microbiome in hospitalised patients with CAP with unknown aetiology, we selected a random sample of 10 patients with CAP from our clinical trial in whom an aetiology was not identified. We speculated that in patients with CAP of unknown aetiology, distinct patterns of the microbiota in the nasopharynx and oropharynx might be identified. Furthermore, these patterns may provide insight into the aetiology of CAP in patients without identification of organisms using current technologies. With this in mind, we performed this pilot study with the objective to characterise the microbiome of the nasopharynx and oropharynx of hospitalised patients with CAP with an unknown aetiology.
2. Methods
2.1 Study design, patients and setting
The pilot study is a retrospective analysis of data collected prospectively from hospitalised adult patients with acute lower respiratory tract infection who were enrolled in a randomised clinical trial (NCT01248715). Enrollment in the clinical trial occurred from October 2010 through April 2013 in 9 hospitals in Louisville, Kentucky, USA. Briefly, all hospitalised patients with acute lower respiratory tract infections who were admitted to any of the 9 adult hospitals were approached for consent and enrollment into the clinical trial. More details on the enrollment and overview of patients are described on https://doi.org/clinicaltrials.gov/ct2/show/NCT01248715, and on the study website, https://doi.org/retostudy.com. Local institutional review board approval was obtained for each study site. In the clinical trial, all specimens with the exception of the nasopharyngeal and oropharyngeal swabs were prospectively collected during the initial days of hospitalisation for clinical care.
In the pilot study, the nasopharyngeal and oropharyngeal swabs were also prospectively collected after obtaining written informed consent from each patient. These swabs were then stored in our biorepository. Clinical data were collected from all patients on a paper case report form by trained study coordinators and/or research associates in our group. Data were then entered into a secure, online data capture system. All data entered into the data capture system were subsequently reviewed by trained clinical research coordinators and/or clinical research associates for data quality issues including out of range values, incorrect data types, and missing values. In the event data inconsistencies were identified, queries were sent to the individuals collecting and/or entering the data for follow-up. Once all queries were resolved, the cases were accepted into the database for analysis. The study protocol and the data collection form for the clinical trial are available online at https://doi.org/retostudy.com.
To validate the CAP diagnosis, the physician principal investigator involved in the pilot study reviewed all cases in the database. We assessed demographic, clinical history, laboratory, radiography, microbiological, therapeutic, and outcome data for all patients. A full list of variables collected for the study can be found on the case report form located on the study website, https://doi.org/retostudy.com/documents/RETOS_CRF.pdf. We included for the current pilot study a random sample of 10 patients satisfying the criteria for CAP and in whom a full microbiological workup failed to identify an aetiology from the 1,221 patients included in the intent-to-treat sample of the clinical trial (NCT01248715). A random number generator was used to select the random sample of patients.
2.2 Study definitions
CAP was defined as the presence of a new pulmonary infiltrate on chest radiograph at the time of hospitalisation that was associated with at least 1 of the following 3 criteria: new or increased cough; an abnormal temperature (<35.6°C or >37°C); or leukocytosis, leukopenia, or left shift pattern on white blood cell count.
A full microbiological workup was defined as having had the following laboratory tests: 1) blood cultures, 2) respiratory cultures (sputum, tracheal aspirate, bronchoalveolar lavage), 3) urinary antigen for S. pneumoniae, 4) urinary antigen for L. pneumophila serogroup 1, 5) nasopharyngeal swab for detection of respiratory viruses (influenza A virus untypable, influenza A virus H3 subtype, influenza A virus H1 subtype, influenza B virus, parainfluenza virus type 1, parainfluenza virus type 2, parainfluenza virus type 3, human metapneumovirus, respiratory syncytial virus [RSV] A, RSV B, adenovirus, and rhinovirus/enterovirus) via the Luminex xTAG Respiratory Viral Panel (Luminex Corporation, USA), 6) oropharyngeal swab for in-house polymerase chain reaction (PCR) detection of atypical pathogens: L. pneumophila, M. pneumoniae, and C. pneumoniae, and 7) rapid test for influenza virus.
Time to clinical stability was defined as the day when the patient met the following criteria: 1) normal or improved symptoms of pneumonia (cough and shortness of breath), 2) lack of fever for at least 8 h (<37.8°C), 3) white blood cell count normal or improving (decrease of >10% from baseline), and 4) oral intake considered adequate by the treating physician.
Length of hospital stay was defined as the number of days between hospital admission and hospital discharge.
2.3 Sample collection and storage
On the day of hospital admission, a nasopharyngeal swab (Copan Flocked Swab: Copan Diagnostics, USA) and an oropharyngeal swab (Copan Flocked Swab: Copan Diagnostics) was collected from hospitalised patients with CAP enrolled in our clinical trial. These samples were placed in 3.0 ml of universal transport media (Quidel, USA), vortexed for 30 s and aliquoted into 3 × 1.0 ml microcentrifuge cryogenic tubes, and stored at −80°C for future microbiome determination.
2.4 PCR
2.4.1 DNA isolation and microbiome sequencing
The total genomic DNA from each swab was isolated with the Maxwell® automated DNA isolation method using the Maxwell® 16 Blood DNA Purification Kit (Promega Corporation, USA). The 16S rRNA genes were amplified using 16S rRNA-specific primers (0.5 µM), 27f (AGAGTTTGATCCTGGCTCAG) and 534r (ATTACCGCGGCTGCTGG) with the FastStart Hifidelity PCR System (Roche Diagnostics Corporation, Switzerland). These primers were anchored with adapters and multiplex identifiers (MIDs) for 454 sequencing to distinguish various samples in a single 454 sequencing reaction. The PCR cycling conditions were 95°C for 5 min, followed by 30 cycles of 94°C for 30 s, 56°C for 30 s, and 72°C for 1 min and 30 s with a final extension period of 8 min at 72°C. Each PCR reaction was performed in triplicate. The PCR amplicon products were pooled and gel purified using QIAGEN gel purification columns (QIAGEN, The Netherlands). The amplicon was quantified using the Quant-iT™ PicoGreen® dsDNA assay kit (Invitrogen: Thermo Fisher Scientific Inc., USA). The pooled, purified amplicons were sequenced using the 454 GS FLX System (Roche Diagnostic Corporation) or the 454 GS Junior System (Roche Diagnostic Corporation) according to manufacturer’s protocols.
2.4.2 Microbiota sequence analysis
The microbial analysis was performed using the QIIME pipeline (QIIME 1.5.0: https://doi.org/www.qiime.org), developed and maintained by Knight and colleagues [4]. Briefly, the quality sequences (200–650 base pair lengths) were de-multiplexed based on their barcodes. The 16S rRNA Operational Taxonomic Units (OTUs) were picked based on 97% sequence identity using UCLUST [5] against the GreenGenes 16S rRNA database (https://doi.org/greengenes.lbl.gov/cgi-bin/nph-citation.cgi) (gg_otus-12_10) [6]. The GreenGenes taxonomies were used to generate the relative abundance of taxa at different levels of phylogeny (phylum, order, class, family, genus, species).
2.4.3 Streptococcus pneumoniae PCR setup
PCR was used on the nasopharyngeal and oropharyngeal swabs to detect S. pneumoniae since the metagenomic analysis did not extend to the species level. The nested PCR protocols were used to detect the presence of S. pneumoniae as described by Murdoch [7]. The Roche FastStart HiFidelity PCR System (Roche Diagnostic Corporation) was used for amplification. Briefly, the first round PCR was targeted to the pneumolysin gene to amplify a 348 base pair region using primers la (5′-ATTTCTGTAACAGCTACCAACGA-3′) and lb (5′-GAATTCCCTGTCTTTTCAAAGTC-3′). The nested PCR (2nd round PCR, inner PCR) was performed using primers lla* (5′-CCCACTCTTCTTGCGGTTGA-3′) and llb (5′-TGAGCCGTTATTTTTTCATACTG-3′) on 1 µl to obtain a 208 base pair inner PCR product. We used clinically identified S. pneumoniae as a positive control and empty/TE buffer (Tris [10 mM]-EDTA [1 mM] pH 7.4)/mock isolation samples were used as a negative control.
2.5 Ethics statement
This study was conducted under the provisions of the Declaration of Helsinki. The prospective randomised clinical trial was registered prior to the enrollment of the first patient in the ClinicalTrials.gov registry (Study ID: NCT01248715, https://doi.org/clinicaltrials.gov/ct2/show/NCT01248715). The randomised clinical trial and the pilot study were also approved by the University of Louisville Human Subjects Program Protection Office (Protocol number 10.0465) prior to approaching patients for enrollment or data collection. Written informed consent was obtained from each patient by trained study coordinators and/or research associates.
3. Results
Baseline patient characteristics as well as clinical outcomes can be found in Table 1. The average age was 63 years (standard deviation [SD] = 16 years), 70% were male, and 50% were admitted directly to the intensive care unit. The average number of days required to reach clinical stability was 3.3 days (SD = 2.6 days), the average length of hospital stay was 5.6 days (SD = 5.6 days), all patients survived to discharge, and 2 patients had died at one-year follow-up.
3.1 Bacterial phyla and genera
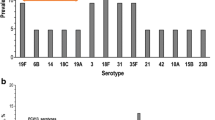
For each patient, the relative percentages of the 4 most important phyla (Firmicutes, Bacteroidetes, Actinobacteria, and Proteobacteria) were plotted (Figure 1). There were clear phylum-level differences between the nasopharyngeal and oropharyngeal microbiota. Figure 2 depicts the most commonly identified genera in the nasopharynx and oropharynx.
3.2 Detection of Streptococcus pneumoniae by nested PCR
We only identified 1 sample (case number 10) that was negative for S. pneumoniae in both the nasopharyngeal and oropharyngeal swabs (Table 2). This sample had a high level of H. influenzae (98%) at the nasopharyngeal locaion.
4. Discussion
We were able to identify 1 patient (case number 10) in whom the analysis of the upper respiratory tract microbiome provided some insight into the aetiology of CAP. This patient was unique with respect to the nasopharyngeal microbiome compared to other patients in our pilot study. The major difference was the substantially more Proteobacteria than Actinobacteria. Furthermore, the phyla Actinobacteria, Firmicutes, and Bacteroidetes were nearly absent in the microbiota in this patient’s nasopharynx. The primary organism within the Proteobacteria family was H. influenzae, which comprised 98% of the sequences. This was also the only patient with negative PCR for S. pneumoniae in both nasopharyngeal and oropharyngeal samples. These data suggest that H. influenzae may have been the aetiologic agent of this patient with CAP of unknown aetiology.
In our sample of patients, we found significant differences in the nasopharyngeal and oropharyngeal microbiomes. These data from patients with CAP is in accordance with recent literature suggesting that the upper respiratory tract microbiome of the nasopharynx is significantly different than the microbiome of the oropharynx in healthy humans [8]. Both microbiomes contain genera of organisms known to be aetiologic agents of CAP. This observation emphasises the need to consider the upper respiratory tract microbiome as two distinct microbial communities: the nasopharyngeal and the oropharyngeal microecosystems. Since microaspiration may originate from either of these two areas of the respiratory tract, future research in this area may need to involve both upper respiratory tract microecosystems.
The Human Microbiome Project indicates that the pharynx of normal individuals is inhabited by four primary phyla: Actinobacteria, Proteobacteria, Firmicutes, and Bacteroidetes. We also identified these four primary phyla in the nasopharynx and oropharynx of hospitalised patients with CAP. The most important pathogens causing CAP are present in the phyla Firmicutes and Proteobacteria. Firmicutes are Gram-positive bacteria that include the Streptococci such as S. pneumoniae, and the Staphylococci. The Proteobacteria include the Enterobacteriaceae, Pseudomonadaceae, and other Gram-negatives such as Haemophilus and Legionella. The Bacteroidetes are Gram-negative bacteria including anaerobes such as Bacteroides, capable of causing anaerobic lung infections. The Actinobacteria are Gram-positive bacteria that include several organisms able to cause sub-acute to chronic pneumonia such as Actinomyces, Streptomyces, and Mycobacterium. In our patients, we also identified several genera that included potential pathogenic organisms able to cause CAP.
Although the nasopharyngeal microbiome of hospitalised patients with pneumonia has not been well described, Chaban and colleagues [9] characterised the bacteria in this body site from patients with influenza infection. These investigators identified the nasopharyngeal microbiome to be relatively evenly distributed between Actinobacteria, Proteobacteria, and Firmicutes, with few Bacteroidetes. Our results showed a similar distribution for Actinobacteria and Firmicutes, but far less Proteobacteria overall. Lemon and colleagues [8] characterised the oropharyngeal microbiome of healthy adults and identified that it was composed largely of Firmicutes, with some cases having more Proteobacteria. This was much more closely aligned with our patients, with the largest phyla overall being Firmicutes.
The need to study both areas of the upper respiratory tract is underscored by our differential detection of S. pneumoniae. From the patients evaluated, the organism was identified in 90% of the patients’ nasopharynx and/or oropharynx. However, only 33% of these 9 patients had positive PCR in both the nasopharynx and oropharynx. It is unclear if the presence of this organism in the upper respiratory tract may indicate lower respiratory tract infection due to S. pneumoniae. Some investigators consider that S. pneumoniae is an under-identified aetiology of CAP due to poor sensitivity of current diagnostic techniques [3]. Since S. pneumoniae may only be colonising the upper respiratory tract of these patients, it is difficult to assess the potential for underdiagnosis of this pathogen as the cause of CAP in these patients. However, microaspiration from the upper respiratory tract plays an important role in the development of pneumonia due to S. pneumoniae. Furthermore, this pathogen is the most common cause of CAP. These two factors suggest that our documentation of S. pneumoniae in the upper respiratory tract of almost all of the patients with CAP of unknown aetiology may provide evidence that the aetiology of CAP in these patients may indeed be S. pneumoniae.
There are several limitations of our pilot study. First, since this was a pilot study, we evaluated only 10 patients, which limits the generalisability of our results. Patients signed the consent form and agreed to participate in our clinical trial on the day of hospital admission, but all patients had received antibiotics in the emergency department before enrollment into the trial. The prior antibiotic use before obtaining the nasopharyngeal and oropharyngeal samples may have biased our results. We were also not able to identify the microbiome down to species-level for all patients due to limitations in our methodology. Further, we looked at the microbiome only from the bacterial perspective. As the field of metagenomics evolves, the complex interactions between bacteria and viruses as aetiologies of CAP such as co-infection with S. pneumoniae and influenza viruses are becoming more relevant. In our study, we did not evaluate the upper respiratory tract microvirome, which may be important in patients with CAP. It is also possible that our high levels of detection of S. pneumoniae were due to detection of pneumolysin presence in some non-pneumococcal Streptococci inhabiting the upper respiratory tract of our patients.
Although our study is descriptive in nature, the data presented is the first step to begin to correlate the upper respiratory tract microbiome with the aetiology of CAP. Future studies of the microbiome in patients with identified aetiologies of lower respiratory tract infections may help to define the probable association of the upper respiratory tract microbiome with particular aetiologic agents of CAP. The upper respiratory tract microecosystems may predispose patients to colonisation with particular pathogenic organisms, influence virulence gene expression of pathogenic organisms already present in the normal microbiome, or impact the immune response to pathogenic organisms. If this were to be the case, reestablishing the upper respiratory tract microbiome homeostasis may have a role in protecting patients from lower respiratory tract infections.
In conclusion, this pilot study suggests that in hospitalised patients with CAP of unknown aetiology, characterising the upper respiratory tract microbiome may be of limited use in defining the aetiology of the patients’ lower respiratory tract infection. However, our results suggest that the nasopharyngeal and oropharyngeal microbiota of hospitalised patients with CAP of unknown aetiology are diverse, and contain several organisms that may be aetiologies of CAP. Furthermore, the presence of S. pneumoniae in the upper respiratory tract microbiome of these patients requires further investigation.
References
World Health Organization. The top 10 causes of death Geneva 2010 [updated 2008 November 29, 2010]. Available from: https://doi.org/www.who.int/mediacentre/factsheets/fs310/en/index.html.
Mandell LA, Wunderink RG, Anzueto A, Bartlett JG, Campbell GD, Dean NC et al.; Infectious Diseases Society of America; American Thoracic Society. Infectious Diseases Society of America/American Thoracic Society consensus guidelines on the management of community-acquired pneumonia in adults. Clin Infect Dis 2007;44 Suppl 2:S27–72. PMID:17278083 https://doi.org/www.dx.doi.org/10.1086/511159
Albrich WC, Madhi SA, Adrian PV, van Niekerk N, Mareletsi T, Cutland C et al. Use of a rapid test of pneumococcal colonization density to diagnose pneumococcal pneumonia. Clin Infect Dis 2012;54:601–9. PMID:22156852 https://doi.org/www.dx.doi.org/10.1093/cid/cir859
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods 2010;7:335–6. PMID:20383131 https://doi.org/www.dx.doi.org/10.1038/nmeth.f.303
Edgar RC. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010;26:2460–1. PMID:20709691 https://doi.org/www.dx.doi.org/10.1093/bioinformatics/btq461
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 2006;72:5069–72. PMID:16820507 https://doi.org/www.dx.doi.org/10.1128/AEM.03006-05
Murdoch DR, Anderson TP, Beynon KA, Chua A, Fleming AM, Laing RT et al. Evaluation of a PCR assay for detection of Streptococcus pneumoniae in respiratory and nonrespiratory samples from adults with community-acquired pneumonia. J Clin Microbiol 2003;41:63–6. PMID:12517826 https://doi.org/www.dx.doi.org/10.1128/JCM.41.1.63-66.2003
Lemon KP, Klepac-Ceraj V, Schiffer HK, Brodie EL, Lynch SV, Kolter R. Comparative analyses of the bacterial microbiota of the human nostril and oropharynx. MBio 2010;1:1. PMID:20802827
Chaban B, Albert A, Links MG, Gardy J, Tang P, Hill JE. Characterization of the upper respiratory tract microbiomes of patients with pandemic H1N1 influenza. PLoS ONE 2013;8:e69559. PMID:23844261 https://doi.org/www.dx.doi.org/10.1371/journal.pone.0069559
Acknowledgements
The authors would like to thank Dr. Ane Uranga, Médico Neumólogo, Hospital of Galdakao-Usansolo, Bilbao, Spain and Ms. Kimberly Buckner, University of Louisville Division of Infectious Diseases Global Health Research Organization, Louisville, Kentucky, USA, for their review of the manuscript and editorial assistance.
Funding Source: The current study was unfunded. The clinical trial from which the random sample of patients were selected was funded by the US Centers for Disease Control and Prevention, ClinicalTrials.gov registry (Study ID: NCT01248715). The funder had no role in the design of this review, compilation and interpretation of material, decision to publish, or preparation of the manuscript.
Competing interests: All authors declare no competing interests.
Author information
Authors and Affiliations
Corresponding author
Additional information
Author contributions: All authors met ICMJE authorship criteria. TW, VJ, RK, PP, FA, RC, BH, JR conceived and designed the research plan. VJ, BH conducted the laboratory research. TW, VJ, PC, RK conducted the data analysis and interpretation. TW, VJ wrote the first draft of the manuscript. All authors critically reviewed the manuscript for important intellectual content. All authors approved the final version of the manuscript.
TW and VJ contributed equally to this work as first authors.
BH and JR contributed equally to this work as senior authors.
Rights and permissions
Copyright: This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
About this article
Cite this article
Wiemken, T.L., Jala, V.R., Kelley, R.R. et al. The upper respiratory tract microbiome of hospitalised patients with community-acquired pneumonia of unknown aetiology: a pilot study. Pneumonia 6, 83–89 (2015). https://doi.org/10.15172/pneu.2015.6/682
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.15172/pneu.2015.6/682