Abstract
Background
Neoadjuvant chemotherapy for breast cancer leads to considerable variability in clinical responses, with only 10 to 20% of cases achieving complete pathologic responses (pCR). Biological and clinical factors that determine the extent of pCR are incompletely understood. Mounting evidence indicates that the patient's immune system contributes to tumor regression and can be modulated by therapies. The cell types most frequently observed with this association are effector tumor infiltrating lymphocytes (TILs), such as cytotoxic T cells, natural killer cells and B cells. We and others have shown that the relative abundance of TILs in breast cancer can be quantified by intratumoral transcript levels of coordinately expressed, immune cell-specific genes. Through expression microarray analysis, we recently discovered three immune gene signatures, or metagenes, that appear to reflect the relative abundance of distinct tumor-infiltrating leukocyte populations. The B/P (B cell/plasma cell), T/NK (T cell/natural killer cell) and M/D (monocyte/dendritic cell) immune metagenes were significantly associated with distant metastasis-free survival of patients with highly proliferative cancer of the basal-like, HER2-enriched and luminal B intrinsic subtypes.
Methods
Given the histopathological evidence that TIL abundance is predictive of neoadjuvant treatment efficacy, we evaluated the therapy-predictive potential of the prognostic immune metagenes. We hypothesized that pre-chemotherapy immune gene signatures would be significantly predictive of tumor response. In a multi-institutional, meta-cohort analysis of 701 breast cancer patients receiving neoadjuvant chemotherapy, gene expression profiles of tumor biopsies were investigated by logistic regression to determine the existence of therapy-predictive interactions between the immune metagenes, tumor proliferative capacity, and intrinsic subtypes.
Results
By univariate analysis, the B/P, T/NK and M/D metagenes were all significantly and positively associated with favorable pathologic responses. In multivariate analyses, proliferative capacity and intrinsic subtype altered the significance of the immune metagenes in different ways, with the M/D and B/P metagenes achieving the greatest overall significance after adjustment for other variables.
Conclusions
Gene expression signatures of infiltrating immune cells carry both prognostic and therapy-predictive value that is impacted by tumor proliferative capacity and intrinsic subtype. Anti-tumor functions of plasma B cells and myeloid-derived antigen-presenting cells may explain more variability in pathologic response to neoadjuvant chemotherapy than previously recognized.
Similar content being viewed by others
Background
Breast cancer is the most common cancer in women worldwide with over 200,000 new cases diagnosed in the US each year [1]. An increasing fraction of these patients are being offered systemic treatment prior to definitive surgery, known as neoadjuvant therapy. While the intent of conventional systemic therapy is to reduce the risk of distant recurrence (that is, for patients with non-metastatic invasive breast cancer), the primary objective of neoadjuvant therapy is to reduce tumor volume, thereby improving surgical outcomes for patients who desire breast conservation or for whom a primary surgical approach is otherwise not medically feasible. Moreover, according to the results of clinical trials in the US and Europe, neoadjuvant chemotherapy is as effective as adjuvant chemotherapy at prolonging patient disease-free survival, distant metastasis-free survival (DMFS) and overall survival [2,3].
Like adjuvant therapy, the current standards of care for neoadjuvant treatment include chemotherapy, endocrine therapy, and biologic therapy (for example, HER2-directed therapy). A corollary benefit of neoadjuvant treatment, however, is that it may serve as an in vivo chemosensitivity test, allowing for early evaluation of the efficacy of systemic therapy and the possible discontinuation of ineffective treatment [4,5]. Neoadjuvant chemotherapy can lead to significant clinical response rates of 60 to 80%, although only 10 to 20% of patients will exhibit a complete pathologic response (pCR) [2,6]. pCR is typically defined as tumor regression marked by the absence of detectable residual disease in the breast and lymph nodes at surgery. Recently, more precise diagnostic models that better quantify the extent of residual disease have been developed [7–9]. For example, measurement of residual cancer burden (RCB) provides a categorical index for tumor responsiveness to neoadjuvant treatment based on size and cellularity of the primary tumor and number and size of involved lymph nodes [9].
The biological mechanisms that influence tumor responsiveness in the neoadjuvant setting are not clearly understood. Routinely administered cytotoxic agents such as anthracyclines and taxanes are known to inhibit replication of rapidly dividing tumor cells by blocking nucleic acid synthesis, or by disrupting microtubule function, respectively. Not surprisingly, markers of tumor cell proliferation, including Ki-67 staining and histologic grade, have been observed to be significantly associated with higher rates of pCR in breast tumors [10,11]. Other therapy-predictive features of breast cancer, such as negative estrogen receptor status and HER2 overexpression, have also been identified [11–13], although not without some degree of controversy [14] and with little indication of clinically applicable predictive value. Mounting evidence now indicates that host-therapy interactions influence tumor responsiveness to neoadjuvant treatment, and that the patient's immune system, in particular, can actively contribute to tumor regression. In mouse models of cancer, where intact tumors were treated with anthracycline-based chemotherapies, tumor regression was observed in immunocompetent mice, while the same tumors transplanted into immunodeficient mice failed to respond [15–18]. Furthermore, anthracycline-based chemotherapies have been observed to induce rapid and prominent tumor infiltration by Th1-oriented effector immune cells in mice and in some human breast cancer patients [19–21]. Indeed, the mere abundance of tumor-infiltrating leukocytes, namely CD3+/CD8+ T lymphocytes, has been robustly correlated with pCR in the neoadjuvant setting [22–25] as well as relapse-free and overall survival of breast cancer patients [25–29]. In more recent years, microarray expression profiling studies in breast and other tumor types have identified immune gene signatures from whole tumor RNA extracts that reflect the abundance of tumor-infiltrating immune cells [30–38]. We and others have found that the biological and phenotypic properties of the genes comprising these signatures implicate distinct immune cell lineages [34–37,39,40], and that combinations of these immune genes correlate with patient outcomes ranging from recurrence-free survival [30,32,36–43] to tumor regression in the neoadjuvant setting [44–47].
Recently, we reported in Nagalla et al. [36] the discovery of three biologically distinct immune gene signatures, or metagenes, in a large microarray dataset comprising 1,954 breast tumor expression profiles. Through gene ontology enrichment analysis and demonstration of immune cell type-specific expression patterns, we provided evidence that these immune metagenes reflect tumor-infiltrating populations of: 1) B cells/plasma B cells (B/P) marked by the high expression of IgG antibody isotype-related genes; 2) a T cell/natural killer cell-specific population (T/NK) likely reflecting a predominantly Th1-type functional orientation; and 3) a monocyte/dendritic cell population (M/D) marked by the expression of myeloid specific markers and a host of major histocompatibility complex (MHC) class II antigen-presenting molecules. Each of these signatures was found to be significantly and positively associated with DMFS of patients. This protective effect, however, was mostly restricted to highly proliferative cancers of the basal-like, HER2-enriched and luminal B (LumB) intrinsic molecular subtypes. By contrast, the same immune signatures exhibited little to no protective effect in tumors of low or intermediate proliferative potential or those classified as luminal A (LumA) or claudin-low (CL) subtypes. In the present work, we sought to evaluate the therapy-predictive potential of these immune metagenes in the context of neoadjuvant chemotherapy for breast cancer, and in the presence of other covariates such as proliferation and intrinsic molecular subtype.
Methods
Microarray data origination and patient characteristics
We assembled a retrospective microarray database (MDACC-701) of breast tumor expression profiles derived from five well-curated, publicly available datasets housed in the NCBI's Gene Expression Omnibus (GEO). The dataset accession numbers are GSE25066, GSE20194, GSE20271, GSE22093 and GSE23988. Specific details of the patient cohorts are described elsewhere [42,48–51] and summarized below. All microarray experiments pertaining to these datasets were conducted at the Department of Pathology, MD Anderson Cancer Center (MDACC), Houston, Texas, as part of several international and multicenter studies conducted between 2000 and 2010. According to previously published reports [48–51] for each study the research protocol was approved by one or more institutional review boards, and all participating patients provided written informed consent consistent with the principles of the Declaration of Helsinki. Expression profiles were generated from RNA samples isolated from fine needle aspirates (FNAs) or needle core biopsies of breast tumors (stage I to III) collected prior to treatment with neoadjuvant chemotherapy. All RNA samples were analyzed on the Affymetrix U133A or U133 PLUS 2.0 GeneChip platforms. In multiple instances, a tumor expression profile was associated with more than one GEO dataset. To create MDACC-701, we downloaded a total of 1,128 tumor profiles from the five datasets then filtered for the unique (nonredundant) profiles using a custom script to measure correlations between all pair-wise combinations. In this manner, redundant profiles (that is, tumor profiles included in more than one dataset) and hybridization repeats could be identified by virtue of high (or perfect) pair-wise correlations. After consolidating the unique profiles, we further excluded a small number of outlier arrays (n = 7) based on low signal intensity distributions. Upon completion of filtering, 701 tumor expression profiles remained. Microarray probe sets were filtered to include only those common to both array platforms (22,277 probe sets). Corresponding patient and clinical characteristics were obtained from supplemental data associated with the original publications or from data associated with the GEO accessions. For redundant profiles, comparison of data entries across the different clinical data sources revealed a small number of discrepancies. In these instances, the discordant clinical data points were re-labeled as 'uncertain' and censored from our analyses. Patient and clinical characteristics of MDACC-701 are summarized in Table 1 and consolidated on a per-sample basis in Additional file 1. Of note, none of the tumor expression profiles of MDACC-701 overlap with the datasets used to discover and characterize the immune metagenes in Nagalla et al. [36].
Microarray data processing
The tumor expression profiles were normalized by the Robust Multi-array Average (RMA) algorithm [52] using R software and the Affy package library file from the Bioconductor project [53]. Normalization was performed within each dataset, and normalized expression values (log2 signal intensities) were corrected for batch effects across datasets using the COMBAT empirical Bayes method [54]. PAM50 subtypes (including the CL classification) were assigned to each sample using previously published methodology [36,55–57].
Metagene construction
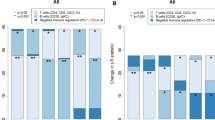
A summary of the content of the individual metagenes is as follows: proliferation (P) metagene (61 probe sets, 54 genes), B/P metagene (65 probe sets, 40 genes), T/NK metagene (52 probe sets, 46 genes) and the M/D metagene (30 probe sets, 19 genes). The proliferation and immune metagenes were constructed as described in Nagalla et al. [36] based on the probe set and gene name assignments listed in Additional File six of that publication. The probe sets 200904_at, 204834_at and 211742_s_at, which overlapped between the T/NK and M/D metagenes, were excluded to enhance distinction between metagenes. Metagene scores were computed for each tumor by averaging the signal intensities of the genes comprising each metagene as described previously [36]. When multiple probe sets corresponded to the same gene designation, these probe sets were averaged first, prior to cross-gene averaging of signal intensities. Tumors were grouped into metagene tertiles by ranking tumors by metagene scores and identifying the 33rd and 66th percentile thresholds. As the cellular composition of a tumor biopsy can differ depending on the tissue sampling method used [58], we sought to compare the metagene score distributions between the surgically acquired tumor specimens that we previously used to derive the metagenes (n = 1,954 tumor samples) [36] and the confirmed FNA biopsy samples that comprise the majority of the current study (n = 482 tumor samples). Both microarray datasets were quantile normalized by the RMA method [52] and similarly corrected for batch effects [54] prior to computing the proliferation and immune metagene scores. The log2 transformed metagene scores were then mean-centered and their distributions examined by boxplot analysis (Additional file 2). While not identical, the major features of the metagene distributions appeared largely conserved between the surgical and FNA specimens, suggesting their general comparability to one another.
Statistical analyses
In the panel of 680 cases having tumor response data, a series of simple logistic regression models were fit examining each of the metagenes (B/P, T/NK, M/D and P) and subtype separately to determine associations with tumor response (odds ratio and 95% confidence interval) (Table 2). Metagenes were entered as continuous variables and subtype was entered as a categorical variable unless otherwise specified. We next examined each immune metagene's association with tumor response while adjusting for proliferation and subtype (Table 2). We then stratified the data into tertiles based on the proliferation metagene and once again examined each immune metagene's association with tumor response (Table 3). Next, we fit a stepwise logistic regression model within each tertile to see whether one (or more) metagenes were independently associated with tumor response. We then examined the association of each immune metagene with tumor response separately by each cancer subtype (basal-like, CL, HER2-enriched (HER2-E), LumB, and LumA; Table 4). In addition, we used a chi-square test to determine whether there was a relationship between treatment type and tumor response. Finally, we fit two stepwise logistic regression models to predict tumor response with 6 or 11 potential predictor variables: estrogen receptor (ER) status, the P metagene, BP, TNK, and MD metagenes and tumor subtype considered as one six-level categorical variable (Table 5) or tumor subtype considered as six individual binary variables (that is, LumA yes/no, LumB yes/no, and so on) (Table 6). Analyses were performed using SAS version 9.3 (SAS Institute Inc., Cary, NC, USA).
Results
We assembled a microarray database of gene expression profiles of breast tumor biopsies from a multicenter meta-cohort of 701 breast tumor patients who received neoadjuvant chemotherapy (Table 1). From this database, we re-constructed the three immune metagenes (B/P, T/NK and M/D) and a proliferation (P) metagene as previously described [36] and as outlined in the Methods section. Briefly, a metagene is defined as a cluster of coordinately expressed gene transcripts whose expression levels, within a tumor, can be averaged to generate a single metagene score that reflects the composite transcriptional activity level of the gene cluster [32,36]. In Nagalla et al. [36] we found that these scores (for each of the three immune metagenes) exhibited prognostic value by Cox regression analysis, with high metagene scores associated with prolonged patient DMFS. The prognostic value, however, was largely restricted to the highly proliferative tumors defined by the upper tertile of the proliferation metagene scores (that is, the most proliferative tumors).
To determine the therapy-predictive value of the immune metagene scores in the neoadjuvant setting, we used simple (that is, single explanatory variable) logistic regression models to examine each metagene, individually, followed by multiple (that is, multiple explanatory variables) logistic regression to measure associations between immune metagenes, the P metagene, tumor subtype and tumor response to chemotherapy. Measures of tumor response were based on previously assigned scores of RCB (0 = complete pathologic response, 1 = minimal residual disease, 2 = moderate residual disease and 3 = extensive residual disease) or determination of the presence or absence of a clinical pCR. Tumors with RCB scores of 0 or 1, or that achieved a pCR (in the absence of assigned RCB scores) were coded as '1' to designate a positive response; all other instances were coded as '0' to designate a negative response. In the group of 680 patients annotated for tumor response, univariate analyses revealed highly significant associations between tumor response and all five covariates (the immune metagenes, the P metagene and intrinsic subtype) with high immune and P metagene scores, and basal-like subtype, being associated positively with tumor response, and LumA subtype being associated with negative tumor response (Table 2).
Next, we investigated the dependence of the therapy-predictive performance of the immune metagenes on tumor proliferative capacity and molecular subtype. First, we examined the association of each metagene with tumor response while adjusting for the proliferation metagene and subtype. As shown in Table 2, the adjusted odds ratios for each immune metagene remained highly significant, indicating that each metagene contributes additive predictive information independent of proliferation and subtype, and is not simply recapitulating information about tumor response already conveyed by those variables. To examine this more closely, we next stratified cases into proliferation (P) tertiles (low (PL), intermediate (PI) and high (PH)) and the association of each immune metagene with tumor response was examined as a function of P tertile (Table 3). Significant positive associations were observed for all immune metagenes within each of the three P tertiles, with the exception of the M/D metagene in the PL tertile. We then fit three stepwise multiple logistic regression models, one for each P tertile, to determine whether or not multiple metagenes would retain significance in a single model. We found that only one immune metagene achieved significance in each P tertile. In the PL tertile, only the B/P metagene remained significant (P = 0.001), while only the M/D metagene remained significant in the PI and PH tertiles (P = 0.001 and P = 0.0002, respectively). This result reflects the degree of collinearity between the three immune metagenes, particularly M/D and T/NK, which have a Spearman correlation of 0.80 (Additional file 3). We next investigated the predictive value of the metagenes in the context of the intrinsic molecular subtypes, as we previously observed the prognostic value of the metagenes to segregate most significantly with the basal, HER2-E and LumB subtypes [36]. As shown in Table 4, the M/D and B/P metagenes achieved statistical significance in certain subtypes, despite potential limitations owing to variability in sample size. While all three metagenes trended towards significant positive associations with tumor response in the basal and LumB subtypes, only the M/D metagene achieved a significant association in these two subtypes. By contrast, the B/P metagene achieved significance in the CL subtype despite small sample size. None of the metagenes displayed a significant association within the HER2-E and LumA subtypes.
Based on reports that indicate an immuno-modulatory role for anthracyclines and taxanes [59–62], we investigated the possible impact of exposure to these drugs in relation to tumor response. Using a chi-square test, we examined a 4 × 2 table (treatment by response) to see whether a relationship existed between treatment type and tumor response in this meta-cohort. No statistically significant association was observed. Furthermore, treatment type did not mediate the observed associations between individual metagenes and tumor response (data not shown).
To better understand the predictive value of the immune metagenes in the presence of other clinical and predictive covariates, we fit multiple logistic regression models on the 662 cases with complete annotation for the variables listed below. Specifically, we fit two stepwise logistic regression models to predict tumor response using either 6 or 11 potential predictor variables: ER status, P, B/P, TN/K, and M/D metagenes, and tumor subtype considered as one six-level categorical variable (Table 5) or tumor subtype considered as six individual binary variables (Table 6). The goal of this analysis was to identify a subset of variables that retained a significant association with tumor response when included together in the logistic regression model. When subtype was considered as one categorical variable, we found that ER status, P, M/D, and B/P metagenes all were retained in the model as statistically significant predictors of tumor response (P <0.05). However, when we re-fit the model with subtypes entered as individual binary variables, we found that the LumA subtype was added as a statistically significant predictor, along with ER status, P and M/D metagenes. In this model, B/P was no longer statistically significant (P >0.05) and was not selected for inclusion by the stepwise procedure. Despite these differences between the two stepwise logistic regression models, both showed a strong and consistent association of ER status, P and M/D metagenes (P <0.007 for all variables) with tumor response, suggesting that these variables each explain different aspects of tumor response to neoadjuvant chemotherapy.
Discussion
Over the past decade, a number of tumor expression profiling studies have identified transcriptomic signatures unique to tumor-infiltrating immune cells. Often revealed by hierarchical clustering techniques or outcome correlation studies, these signatures distinguish multiple different immune cell types [32–37] and recapitulate immunohistochemistry-based observations in breast cancer that link tumor-infiltrating immune cell abundance to disease-free survival and overall survival of patients [30,36–42]. More recently, similar studies involving pre-surgical breast tumor biopsies have begun to demonstrate associations between immunity-related genes and tumor responsiveness to neoadjuvant chemotherapy [22,44,45,63,64]. However, a unified understanding of how immunity-related genes relate to both patient prognosis and therapy prediction has yet to be addressed. In Nagalla et al. [36], we recently reported the discovery of three immune metagenes with highly significant and independent associations with patient DMFS; however, the significance of these associations was found to depend largely on tumor proliferative capacity and intrinsic molecular subtype. In the current study, we hypothesized that the same immune metagenes may exhibit similarly significant associations with tumor response to neoadjuvant chemotherapy. To test this hypothesis, we analyzed a collection of 701 microarray expression profiles of primary breast tumor biopsies and corresponding clinical data, including tumor responsiveness to neoadjuvant chemotherapy. Overall, we found that each of the immune metagenes, B/P, T/NK and M/D, was highly significantly and positively associated with tumor response (Table 2), thereby confirming the duality of their roles as biomarkers of favorable outcome in both patient prognosis and therapy prediction.
However, certain biological variables that influence the strength of these associations were found to vary between the prognostic and therapy-predictive settings. Whereas the prognostic performance of the immune metagenes observed in Nagalla et al. [36] was found to be mostly restricted to highly proliferative tumors (PH), this was not the case for their therapy-predictive performance. With the exception of the M/D metagene, which did not reach significance in the PL tertile, all metagenes were found to be significantly associated with positive tumor response in each of the proliferation tertiles and with similar odds ratios (Table 3). In multivariate analysis, we observed mostly collinear relationships among the metagenes, with only single metagenes retaining significance in each proliferation tertile. Specifically, only the B/P metagene remained significant in the PL tertile, while only the M/D metagene remained significant in the PI and PH tertiles. Taken together, these observations suggest that tumor proliferative capacity may modify the prognostic and therapy-predictive potentials of the immune metagenes in fundamentally different ways. While the prognostic attributes of the immune metagenes (but not their therapy-predictive attributes) exhibit dependency on a high proliferative capacity, the predominating therapy-predictive power of the immune metagenes may vary from one immune compartment to another in a proliferation-dependent manner (for example, B/P versus M/D).
With respect to intrinsic molecular subtypes, the therapy-predictive associations could not be resolved to the same degree as that of the prognostic associations observed in Nagalla et al. due to smaller sample sizes that prevented simultaneous stratification by both subtype and proliferation tertile. However, stratification by subtype alone did reveal several interesting therapy-predictive associations (Table 4). First, the majority of odds ratios trended toward positive and significant tumor responses in three subtypes - basal-like, LumB and CL - while showing few to no associations in the HER2-E and LumA subtypes. Similarly, in Nagalla et al., we found that the immune metagenes were strongly associated with favorable DMFS in the majority of basal-like and LumB tumors, but not LumA tumors. By contrast, however, in Nagalla et al. we observed that the immune metagenes were not associated with prognosis in CL tumors, not as a whole, nor when partitioned into proliferation tertiles. Interestingly, these findings may indicate that CL tumors exemplify a condition wherein the prognostic and therapy-predictive roles of the immune metagenes diverge, rather than parallel one another. However, given the paucity of CL tumors represented in our microarray dataset, this hypothesis warrants further investigation in larger sample populations. Another discordant observation pertained to the HER2-E tumor subtype. While the immune metagenes were prognostic of DMFS in the majority of HER2-E tumors analyzed (that is, the PH population), we did not observe a parallel association with tumor response in HER2-E tumors in the current study. Whether this observation would hold true in the PH subpopulation of HER2-E tumors could not be determined due to limiting sample size.
Taken together, these observations indicate that while the immune metagenes are associated with both patient prognosis and chemotherapy response, the tumor properties that influence these associations (proliferation and subtype) are not consistent in their effects. A plausible explanation may relate to the anti-tumor biology mirrored by the immune metagenes and the impact of chemotherapy on tumor immunogenicity. As surrogate markers of immune cell abundance, and by virtue of their positive associations with both DMFS and chemotherapy response, the immune metagenes appear to reflect the anti-tumor potential of the host immune system. Neoadjuvant chemotherapy is known to impact tumor-specific immune responses in various ways. Central among these is the ability of chemotherapy to enhance tumor-specific immunogenicity. For example, chemotherapy-induced cell death can trigger the release of tumor-associated antigens or cell death-associated molecules leading to a cascade of anti-tumor immune responses that can contribute to tumor regression [16,65,66]. Alternatively, chemotherapeutic agents are known to exert a variety of other immunostimulatory effects, including: 1) induction of MHC class I expression and subsequent presentation of tumor antigens [67]; 2) increased expression of ligands that stimulate tumor-reactive activation of NK and cytotoxic T cells [68,69]; 3) induction of tumor-expressing death receptors responsive to ligands expressed by immune effectors [70]; and 4) the depletion or inactivation of tumor-protective regulatory T cells [71,72]. In the neoadjuvant setting, such mechanisms of chemotherapy-induced immunogenicity may operate independent of tumor proliferation rate and intrinsic molecular subtype. Thus, while it remains uncertain as to why the prognostic power of the immune metagenes is restricted to highly proliferative breast tumors and certain molecular subtypes [36], their therapy-predictive power may be influenced more by the prevalence of chemotherapy-induced immunogenic mechanisms than by tumor phenotypes that dictate immunogenic potential in the absence of neoadjuvant treatment.
As our study involved the comparison of gene expression metrics between prognostic and therapy-predictive settings, an underlying assumption was that microarray expression profiles are comparable between different types of tissue biopsies - namely surgical tumor biopsies (SURGbx) obtained at tumor resection (that is, the basis for our previous prognostic observations) and fine-needle aspirate biopsies (FNAbx) or core biopsies (Cbx) obtained prior to surgery (the basis for our current therapy-predictive observations). Previously, Symmans and colleagues compared the cellular compositions and expression profiles between breast FNAbx and Cbx [58]. They reported similar proportions of immune cell infiltrates (on average, 15% (FNAbx) and 20% (Cbx)) but discordant proportions of malignant epithelial cells (80% (FNAbx) versus 50% (Cbx)) and stromal cells (5% (FNAbx) versus 30% (Cbx)). While unsupervised hierarchical clustering revealed a high degree of gene correlation between patient-matched FNA and core biopsies overall, disproportionate cell compositions among biopsy types were shown to result in skewed distributions for certain cell type-specific gene expression patterns. Whether this is true for SURGbx and FNAbx has, to our knowledge, not been investigated. The proliferation and immune metagenes examined in our current study derived predominantly from expression profiles of FNAbx (confirmed for 69% of samples (n = 482)) but also included a smaller unspecified number of Cbx intermixed with FNAbx (31% of samples). In Nagalla et al. [36] we defined and characterized the metagenes based solely on SURGbx specimens (n = 1,954). Thus, we used this opportunity to compare and contrast metagene score distributions between the SURGbx and FNAbx microarray datasets by box and whisker plot analysis (Additional file 2). In each metagene comparison, both the interquartile ranges (boxes) and the spreads among the lower and upper quartiles (whiskers) showed good concordance between FNAbx and SURGbx datasets, suggesting that the distributions of the proliferation and immune metagenes are fairly comparable among these different biopsy types. Nevertheless, a more rigorous investigation of the impact of biopsy method on gene expression dynamics would be warranted for clinical diagnostic applications.
To date, the published data on immune gene signatures predictive of breast tumor response to neoadjuvant chemotherapy derives from four studies that have focused mainly on genes implicated in the biology of tumor infiltrating lymphocytes [22,44,45,64]. Surprisingly, we observed very little overlap between these four published gene sets and our immune metagenes. The greatest overlap was seen with our T/NK metagene. Of the 46 genes comprising our T/NK metagene, 12 are included in one or more of the published gene sets, with at least two T/NK genes overlapping with each of the four gene sets. By contrast, however, only one of the 40 genes comprising our B/P metagene, and one of the 19 genes comprising our M/D metagene showed overlap with a published gene set. Thus, we conclude that the therapy-predictive attributes of the B/P and MD metagenes are mostly unexplored, representing novel biomarkers of breast tumor response to neoadjuvant chemotherapy.
By stepwise logistic regression, we discerned that the M/D and B/P metagenes gave the most robust therapy-predictive performances among the immune metagenes. In a similar vein, both the B/P and M/D metagenes were strong and independent predictors of DMFS in Nagalla et al., whereby the B/P metagene emerged as the most significant immune covariate by multivariate analysis. In the context of therapy prediction, however, the M/D metagene prevailed as the most significant and additive immunity-related covariate in the final multivariate model.
The genes comprising the M/D metagene are overexpressed in myeloid cell lineages and enriched for functions associated with antigen processing and presentation [36]. Of the 19 genes comprising the M/D metagene, nine are involved in MHC class II-mediated antigen presentation (HLA-DRA, HLA-DRB1, HLA-DMA, HLA-DMB, HLA-DPA1, HLA-DPB1, HLA-DQA1, HLA-DQB1, CD74), suggesting that antigen presentation may constitute the driving biology behind the metagene's therapy-predictive power. While B cells also express MHC class II molecules, the M/D metagene is absent of B cell markers but inclusive of CSF1R, which encodes the classical macrophage colony-stimulating factor receptor that controls growth and differentiation of macrophages and dendritic cells [73]. Thus, the biology underlying the M/D metagene is consistent with a myeloid-driven, anti-tumor immune response engendered by either macrophages (for example, M1-polarized) or dendritic cells functionally oriented towards tumor rejection. Dendritic cells (DCs) are professional antigen presenting cells that coordinate innate and adaptive immune responses to cancer. Dying tumor cells, such as those succumbing to chemotherapy-induced apoptosis, emit danger signals interpreted by DCs as damage-associated molecular patterns (DAMPs) [74]. These signals can induce DC maturation, production of pro-inflammatory cytokines, tumor cell engulfment and subsequent processing and presentation of tumor antigens [15,75,76]. Moreover, anthracyclines and taxanes can stimulate DC-mediated antigen presentation either indirectly, through induction of a DC-responsive immunogenic form of tumor cell death [16], or directly, through DC exposure to chemotherapy (at low or noncytotoxic concentrations) resulting in the up-regulation of MHC and co-stimulatory molecules on the surface of DCs [77]. These observations, and in light of the prognostic and therapy-predictive attributes of the M/D metagene described herein, suggest the possibility that DC-based cancer vaccines [78] could synergize with conventional breast cancer chemotherapeutics, and if administered in the neoadjuvant setting, could prime a durable immunogenic response that not only contributes to primary tumor regression but provides protection against recurrent disease.
Conclusion
Our findings demonstrate the existence of distinct transcriptional footprints of infiltrating effector immune cell subpopulations in breast tumors that are predictive of both chemotherapeutic efficacy and reduced risk of metastatic recurrence. From a biological perspective, these metagenes underscore the important participation of different arms of the immune system in the chemotherapy-induced rejection of established breast tumors as well as the prevention of distant recurrence in the presence or absence of adjuvant treatment [36]. Furthermore, our work indicates that antigen presentation may play a more prominent role in the efficacy of neoadjuvant chemotherapy of breast cancer than previously recognized, and may explain, in part, the variability of pathologic response in the neoadjuvant setting. As reporters of immunogenic potential, the immune metagenes could have functionality as actionable therapeutic markers, particularly in this era of expanding immunotherapies. How the immune metagenes might be harnessed to inform clinical decisions early in the therapeutic sequence warrants further investigation.
Authors' contributions
AA, MAB and RD carried out statistical analyses. JWC and MAB assembled the microarray data and corresponding clinical annotations, controlled for data quality, and performed molecular subtype classifications. SN, RD, JWC and MAB provided input on data interpretation and writing of the manuscript. LDM and AA conceived of the study and wrote the paper. All authors read and approved the final manuscript.
Additional files
Abbreviations
- B/P:
-
B cells/plasma B cells
- Cbx:
-
core biopsy
- CL:
-
claudin-low
- DC:
-
dendritic cell
- DMFS:
-
distant metastasis-free survival
- ER:
-
estrogen receptor
- FNA:
-
fine needle aspirate
- FNAbx:
-
fine-needle aspirate biopsy
- GEO:
-
Gene Expression Omnibus
- HER2-E:
-
HER2-enriched
- LumA:
-
luminal A
- LumB:
-
luminal B
- M/D:
-
monocyte/dendritic cell population
- MDACC:
-
MD Anderson Cancer Center
- MHC:
-
Major histocompatibility complex
- P:
-
Proliferation
- pCR:
-
complete pathologic response
- PH :
-
high proliferation tertile
- PI :
-
intermediate proliferation tertile
- PL :
-
low proliferation tertile
- RCB:
-
Residual cancer burden
- RMA:
-
Robust Multi-array Average
- SURGbx:
-
Surgical tumor biopsy
- T/NK:
-
T cell/natural killer cell-specific population
References
Jemal A, Siegel R, Xu J, Ward E: Cancer statistics, 2010. CA Cancer J Clin. 2010, 60: 277-300. 10.3322/caac.20073. doi:10.3322/caac.20073
Fisher B, Bryant J, Wolmark N, Mamounas E, Brown A, Fisher ER, Wickerham DL, Begovic M, DeCillis A, Robidoux A, Margolese RG, Cruz AB, Hoehn JL, Lees AW, Dimitrov NV, Bear HD: Effect of preoperative chemotherapy on the outcome of women with operable breast cancer. J Clin Oncol. 1998, 16: 2672-2685.
van der Hage JA, van de Velde CJ, Julien JP, Tubiana-Hulin M, Vandervelden C, Duchateau L: Preoperative chemotherapy in primary operable breast cancer: results from the European Organization for Research and Treatment of Cancer trial 10902. J Clin Oncol. 2001, 19: 4224-4237.
Rastogi P, Anderson SJ, Bear HD, Geyer CE, Kahlenberg MS, Robidoux A, Margolese RG, Hoehn JL, Vogel VG, Dakhil SR, Tamkus D, King KM, Pajon ER, Wright MJ, Robert J, Paik S, Mamounas EP, Wolmark N: Preoperative chemotherapy: updates of National Surgical Adjuvant Breast and Bowel Project Protocols B-18 and B-27. J Clin Oncol. 2008, 26: 778-785. 10.1200/JCO.2007.15.0235. doi:10.1200/JCO.2007.15.0235
Schott AF, Hayes DF: Defining the benefits of neoadjuvant chemotherapy for breast cancer. J Clin Oncol. 2012, 30: 1747-1749. 10.1200/JCO.2011.41.3161. doi:10.1200/JCO.2011.41.3161
Smith IC, Heys SD, Hutcheon AW, Miller ID, Payne S, Gilbert FJ, Ah-See AK, Eremin O, Walker LG, Sarkar TK, Eggleton SP, Ogston KN: Neoadjuvant chemotherapy in breast cancer: significantly enhanced response with docetaxel. J Clin Oncol. 2002, 20: 1456-1466. 10.1200/JCO.20.6.1456.
Ellis MJ, Tao Y, Luo J, A’Hern R, Evans DB, Bhatnagar AS, Chaudri Ross HA, von Kameke A, Miller WR, Smith I, Eiermann W, Dowsett M: Outcome prediction for estrogen receptor-positive breast cancer based on postneoadjuvant endocrine therapy tumor characteristics. J Natl Cancer Inst. 2008, 100: 1380-1388. 10.1093/jnci/djn309. doi:10.1093/jnci/djn309
Mathieu MC, Mazouni C, Kesty NC, Zhang Y, Scott V, Passeron J, Arnedos M, Schnabel CA, Delaloge S, Erlander MG, Andre F: Breast Cancer Index predicts pathological complete response and eligibility for breast conserving surgery in breast cancer patients treated with neoadjuvant chemotherapy. Ann Oncol. 2012, 23: 2046-2052. 10.1093/annonc/mdr550. doi:10.1093/annonc/mdr550
Symmans WF, Peintinger F, Hatzis C, Rajan R, Kuerer H, Valero V, Assad L, Poniecka A, Hennessy B, Green M, Buzdar AU, Singletary SE, Hortobagyi GN, Pusztai L: Measurement of residual breast cancer burden to predict survival after neoadjuvant chemotherapy. J Clin Oncol. 2007, 25: 4414-4422. 10.1200/JCO.2007.10.6823. doi:10.1200/JCO.2007.10.6823
Nishimura R, Osako T, Okumura Y, Hayashi M, Arima N: Clinical significance of Ki-67 in neoadjuvant chemotherapy for primary breast cancer as a predictor for chemosensitivity and for prognosis. Breast Cancer. 2010, 17: 269-275. 10.1007/s12282-009-0161-5. doi:10.1007/s12282-009-0161-5
Petit T, Wilt M, Velten M, Millon R, Rodier JF, Borel C, Mors R, Haegele P, Eber M, Ghnassia JP: Comparative value of tumour grade, hormonal receptors, Ki-67, HER-2 and topoisomerase II alpha status as predictive markers in breast cancer patients treated with neoadjuvant anthracycline-based chemotherapy. Eur J Cancer. 2004, 40: 205-211. 10.1016/S0959-8049(03)00675-0.
Andre F, Mazouni C, Liedtke C, Kau SW, Frye D, Green M, Gonzalez-Angulo AM, Symmans WF, Hortobagyi GN, Pusztai L: HER2 expression and efficacy of preoperative paclitaxel/FAC chemotherapy in breast cancer. Breast Cancer Res Treat. 2008, 108: 183-190. 10.1007/s10549-007-9594-8. doi:10.1007/s10549-007-9594-8
Dowsett M, Smith IE, Ebbs SR, Dixon JM, Skene A, Griffith C, Boeddinghaus I, Salter J, Detre S, Hills M, Ashley S, Francis S, Walsh G, A’Hern R: Proliferation and apoptosis as markers of benefit in neoadjuvant endocrine therapy of breast cancer. Clin Cancer Res. 2006, 12: 1024s-1030s. 10.1158/1078-0432.CCR-05-2127. doi:10.1158/1078-0432.CCR-05-2127
Burcombe RJ, Makris A, Richman PI, Daley FM, Noble S, Pittam M, Wright D, Allen SA, Dove J, Wilson GD: Evaluation of ER, PgR, HER-2 and Ki-67 as predictors of response to neoadjuvant anthracycline chemotherapy for operable breast cancer. Br J Cancer. 2005, 92: 147-155. 10.1038/sj.bjc.6602256. doi:10.1038/sj.bjc.6602256
Apetoh L, Ghiringhelli F, Tesniere A, Obeid M, Ortiz C, Criollo A, Mignot G, Maiuri MC, Ullrich E, Saulnier P, Yang H, Amigorena S, Ryffel B, Barrat FJ, Saftig P, Levi F, Lidereau R, Nogues C, Mira JP, Chompret A, Joulin V, Clavel-Chapelon F, Bourhis J, Andre F, Delaloge S, Tursz T, Kroemer G, Zitvogel L: Toll-like receptor 4-dependent contribution of the immune system to anticancer chemotherapy and radiotherapy. Nat Med. 2007, 13: 1050-1059. 10.1038/nm1622. doi:10.1038/nm1622
Casares N, Pequignot MO, Tesniere A, Ghiringhelli F, Roux S, Chaput N, Schmitt E, Hamai A, Hervas-Stubbs S, Obeid M, Coutant F, Metivier D, Pichard E, Aucouturier P, Pierron G, Garrido C, Zitvogel L, Kroemer G: Caspase-dependent immunogenicity of doxorubicin-induced tumor cell death. J Exp Med. 2005, 202: 1691-1701. 10.1084/jem.20050915. doi:10.1084/jem.20050915
Ghiringhelli F, Apetoh L, Tesniere A, Aymeric L, Ma Y, Ortiz C, Vermaelen K, Panaretakis T, Mignot G, Ullrich E, Perfettini JL, Schlemmer F, Tasdemir E, Uhl M, Genin P, Civas A, Ryffel B, Kanellopoulos J, Tschopp J, Andre F, Lidereau R, McLaughlin NM, Haynes NM, Smyth MJ, Kroemer G, Zitvogel L: Activation of the NLRP3 inflammasome in dendritic cells induces IL-1beta-dependent adaptive immunity against tumors. Nat Med. 2009, 15: 1170-1178. 10.1038/nm.2028. doi:10.1038/nm.2028
Michaud M, Martins I, Sukkurwala AQ, Adjemian S, Ma Y, Pellegatti P, Shen S, Kepp O, Scoazec M, Mignot G, Rello-Varona S, Tailler M, Menger L, Vacchelli E, Galluzzi L, Ghiringhelli F, di Virgilio F, Zitvogel L, Kroemer G: Autophagy-dependent anticancer immune responses induced by chemotherapeutic agents in mice. Science. 2011, 334: 1573-1577. 10.1126/science.1208347. doi:10.1126/science.1208347
Demaria S, Volm MD, Shapiro RL, Yee HT, Oratz R, Formenti SC, Muggia F, Symmans WF: Development of tumor-infiltrating lymphocytes in breast cancer after neoadjuvant paclitaxel chemotherapy. Clin Cancer Res. 2001, 7: 3025-3030.
Ladoire S, Arnould L, Apetoh L, Coudert B, Martin F, Chauffert B, Fumoleau P, Ghiringhelli F: Pathologic complete response to neoadjuvant chemotherapy of breast carcinoma is associated with the disappearance of tumor-infiltrating foxp3+ regulatory T cells. Clin Cancer Res. 2008, 14: 2413-2420. 10.1158/1078-0432.CCR-07-4491. doi:10.1158/1078-0432.CCR-07-4491
Ladoire S, Mignot G, Dabakuyo S, Arnould L, Apetoh L, Rebe C, Coudert B, Martin F, Bizollon MH, Vanoli A, Coutant C, Fumoleau P, Bonnetain F, Ghiringhelli F: In situ immune response after neoadjuvant chemotherapy for breast cancer predicts survival. J Pathol. 2011, 224: 389-400. 10.1002/path.2866. doi:10.1002/path.2866
Denkert C, Loibl S, Noske A, Roller M, Muller BM, Komor M, Budczies J, Darb-Esfahani S, Kronenwett R, Hanusch C, von Torne C, Weichert W, Engels K, Solbach C, Schrader I, Dietel M, von Minckwitz G: Tumor-associated lymphocytes as an independent predictor of response to neoadjuvant chemotherapy in breast cancer. J Clin Oncol. 2010, 28: 105-113. 10.1200/JCO.2009.23.7370. doi:10.1200/JCO.2009.23.7370
Hornychova H, Melichar B, Tomsova M, Mergancova J, Urminska H, Ryska A: Tumor-infiltrating lymphocytes predict response to neoadjuvant chemotherapy in patients with breast carcinoma. Cancer Invest. 2008, 26: 1024-1031. 10.1080/07357900802098165. doi:10.1080/07357900802098165
Loi S, Sirtaine N, Piette F, Salgado R, Viale G, Van Eenoo F, Rouas G, Francis P, Crown JP, Hitre E, de Azambuja E, Quinaux E, Di Leo A, Michiels S, Piccart MJ, Sotiriou C: Prognostic and predictive value of tumor-infiltrating lymphocytes in a phase III randomized adjuvant breast cancer trial in node-positive breast cancer comparing the addition of docetaxel to doxorubicin with doxorubicin-based chemotherapy: BIG 02–98. J Clin Oncol. 2013, 31: 860-867. 10.1200/JCO.2011.41.0902. doi:10.1200/JCO.2011.41.0902
Ono M, Tsuda H, Shimizu C, Yamamoto S, Shibata T, Yamamoto H, Hirata T, Yonemori K, Ando M, Tamura K, Katsumata N, Kinoshita T, Takiguchi Y, Tanzawa H, Fujiwara Y: Tumor-infiltrating lymphocytes are correlated with response to neoadjuvant chemotherapy in triple-negative breast cancer. Breast Cancer Res Treat. 2012, 132: 793-805. 10.1007/s10549-011-1554-7. doi:10.1007/s10549-011-1554-7
Aaltomaa S, Lipponen P, Eskelinen M, Kosma VM, Marin S, Alhava E, Syrjanen K: Lymphocyte infiltrates as a prognostic variable in female breast cancer. Eur J Cancer. 1992, 28A: 859-864. 10.1016/0959-8049(92)90134-N.
Andre F, Dieci MV, Dubsky P, Sotiriou C, Curigliano G, Denkert C, Loi S: Molecular pathways: involvement of immune pathways in the therapeutic response and outcome in breast cancer. Clin Cancer Res. 2013, 19: 28-33. 10.1158/1078-0432.CCR-11-2701. doi:10.1158/1078-0432.CCR-11-2701
Lee AH, Gillett CE, Ryder K, Fentiman IS, Miles DW, Millis RR: Different patterns of inflammation and prognosis in invasive carcinoma of the breast. Histopathology. 2006, 48: 692-701. 10.1111/j.1365-2559.2006.02410.x. doi:10.1111/j.1365-2559.2006.02410.x
Mahmoud SM, Paish EC, Powe DG, Macmillan RD, Grainge MJ, Lee AH, Ellis IO, Green AR: Tumor-infiltrating CD8+ lymphocytes predict clinical outcome in breast cancer. J Clin Oncol. 2011, 29: 1949-1955. 10.1200/JCO.2010.30.5037. doi:10.1200/JCO.2010.30.5037
Alexe G, Dalgin GS, Scanfeld D, Tamayo P, Mesirov JP, DeLisi C, Harris L, Barnard N, Martel M, Levine AJ, Ganesan S, Bhanot G: High expression of lymphocyte-associated genes in node-negative HER2+ breast cancers correlates with lower recurrence rates. Cancer Res. 2007, 67: 10669-10676. 10.1158/0008-5472.CAN-07-0539. doi:10.1158/0008-5472.CAN-07-0539
Broet P, Kuznetsov VA, Bergh J, Liu ET, Miller LD: Identifying gene expression changes in breast cancer that distinguish early and late relapse among uncured patients. Bioinformatics. 2006, 22: 1477-1485. 10.1093/bioinformatics/btl110. doi:10.1093/bioinformatics/btl110
Dave SS, Wright G, Tan B, Rosenwald A, Gascoyne RD, Chan WC, Fisher RI, Braziel RM, Rimsza LM, Grogan TM, Miller TP, LeBlanc M, Greiner TC, Weisenburger DD, Lynch JC, Vose J, Armitage JO, Smeland EB, Kvaloy S, Holte H, Delabie J, Connors JM, Lansdorp PM, Ouyang Q, Lister TA, Davies AJ, Norton AJ, Muller-Hermelink HK, Ott G, Campo E, et al: Prediction of survival in follicular lymphoma based on molecular features of tumor-infiltrating immune cells. N Engl J Med. 2004, 351: 2159-2169. 10.1056/NEJMoa041869. doi:10.1056/NEJMoa041869
Donson AM, Birks DK, Barton VN, Wei Q, Kleinschmidt-Demasters BK, Handler MH, Waziri AE, Wang M, Foreman NK: Immune gene and cell enrichment is associated with a good prognosis in ependymoma. J Immunol. 2009, 183: 7428-7440. 10.4049/jimmunol.0902811. doi:10.4049/jimmunol.0902811
Donson AM, Birks DK, Schittone SA, Kleinschmidt-DeMasters BK, Sun DY, Hemenway MF, Handler MH, Waziri AE, Wang M, Foreman NK: Increased immune gene expression and immune cell infiltration in high-grade astrocytoma distinguish long-term from short-term survivors. J Immunol. 2012, 189: 1920-1927. 10.4049/jimmunol.1103373. doi:10.4049/jimmunol.1103373
Griesinger AM, Birks DK, Donson AM, Amani V, Hoffman LM, Waziri A, Wang M, Handler MH, Foreman NK: Characterization of distinct immunophenotypes across pediatric brain tumor types. J Immunol. 2013, 191: 4880-4888. 10.4049/jimmunol.1301966. doi:10.4049/jimmunol.1301966
Nagalla S, Chou JW, Willingham MC, Ruiz J, Vaughn JP, Dubey P, Lash TL, Hamilton-Dutoit SJ, Bergh J, Sotiriou C, Black MA, Miller LD: Interactions between immunity, proliferation and molecular subtype in breast cancer prognosis. Genome Biol. 2013, 14: R34-10.1186/gb-2013-14-4-r34. doi:10.1186/gb-2013-14-4-r34
Rody A, Holtrich U, Pusztai L, Liedtke C, Gaetje R, Ruckhaeberle E, Solbach C, Hanker L, Ahr A, Metzler D, Engels K, Karn T, Kaufmann M: T-cell metagene predicts a favorable prognosis in estrogen receptor-negative and HER2-positive breast cancers. Breast Cancer Res. 2009, 11: R15-10.1186/bcr2234. doi:10.1186/bcr2234
Teschendorff AE, Miremadi A, Pinder SE, Ellis IO, Caldas C: An immune response gene expression module identifies a good prognosis subtype in estrogen receptor negative breast cancer. Genome Biol. 2007, 8: R157-10.1186/gb-2007-8-8-r157. doi:gb-2007-8-8-r157
Ascierto ML, Kmieciak M, Idowu MO, Manjili R, Zhao Y, Grimes M, Dumur C, Wang E, Ramakrishnan V, Wang XY, Bear HD, Marincola FM, Manjili MH: A signature of immune function genes associated with recurrence-free survival in breast cancer patients. Breast Cancer Res Treat. 2012, 131: 871-880. 10.1007/s10549-011-1470-x. doi:10.1007/s10549-011-1470-x
Bianchini G, Qi Y, Alvarez RH, Iwamoto T, Coutant C, Ibrahim NK, Valero V, Cristofanilli M, Green MC, Radvanyi L, Hatzis C, Hortobagyi GN, Andre F, Gianni L, Symmans WF, Pusztai L: Molecular anatomy of breast cancer stroma and its prognostic value in estrogen receptor-positive and -negative cancers. J Clin Oncol. 2010, 28: 4316-4323. 10.1200/JCO.2009.27.2419. doi:10.1200/JCO.2009.27.2419
Desmedt C, Haibe-Kains B, Wirapati P, Buyse M, Larsimont D, Bontempi G, Delorenzi M, Piccart M, Sotiriou C: Biological processes associated with breast cancer clinical outcome depend on the molecular subtypes. Clin Cancer Res. 2008, 14: 5158-5165. 10.1158/1078-0432.CCR-07-4756. doi:10.1158/1078-0432.CCR-07-4756
Iwamoto T, Bianchini G, Booser D, Qi Y, Coutant C, Shiang CY, Santarpia L, Matsuoka J, Hortobagyi GN, Symmans WF, Holmes FA, O'shaughnessy J, Hellerstedt B, Pippen J, Andre F, Simon R, Pusztai L: Gene pathways associated with prognosis and chemotherapy sensitivity in molecular subtypes of breast cancer. J Natl Cancer Inst. 2011, 103: 264-272. 10.1093/jnci/djq524. doi:10.1093/jnci/djq524
Wang E, Miller LD, Ohnmacht GA, Mocellin S, Perez-Diez A, Petersen D, Zhao Y, Simon R, Powell JI, Asaki E, Alexander HR, Duray PH, Herlyn M, Restifo NP, Liu ET, Rosenberg SA, Marincola FM: Prospective molecular profiling of melanoma metastases suggests classifiers of immune responsiveness. Cancer Res. 2002, 62: 3581-3586.
Desmedt C, Di Leo A, de Azambuja E, Larsimont D, Haibe-Kains B, Selleslags J, Delaloge S, Duhem C, Kains JP, Carly B, Maerevoet M, Vindevoghel A, Rouas G, Lallemand F, Durbecq V, Cardoso F, Salgado R, Rovere R, Bontempi G, Michiels S, Buyse M, Nogaret JM, Qi Y, Symmans F, Pusztai L, D’Hondt V, Piccart-Gebhart M, Sotiriou C: Multifactorial approach to predicting resistance to anthracyclines. J Clin Oncol. 2011, 29: 1578-1586. 10.1200/JCO.2010.31.2231. doi:10.1200/JCO.2010.31.2231
Ignatiadis M, Singhal SK, Desmedt C, Haibe-Kains B, Criscitiello C, Andre F, Loi S, Piccart M, Michiels S, Sotiriou C: Gene modules and response to neoadjuvant chemotherapy in breast cancer subtypes: a pooled analysis. J Clin Oncol. 2012, 30: 1996-2004. 10.1200/JCO.2011.39.5624. doi:10.1200/JCO.2011.39.5624
Karn T, Pusztai L, Holtrich U, Iwamoto T, Shiang CY, Schmidt M, Muller V, Solbach C, Gaetje R, Hanker L, Ahr A, Liedtke C, Ruckhaberle E, Kaufmann M, Rody A: Homogeneous datasets of triple negative breast cancers enable the identification of novel prognostic and predictive signatures. PLoS One. 2011, 6: e28403-10.1371/journal.pone.0028403. doi:10.1371/journal.pone.0028403
Mello-Grand M, Singh V, Ghimenti C, Scatolini M, Regolo L, Grosso E, Zambelli A, Da Prada GA, Villani L, Fregoni V, Baiardi P, Marsoni S, Miller WR, Costa A, Chiorino G: Gene expression profiling and prediction of response to hormonal neoadjuvant treatment with anastrozole in surgically resectable breast cancer. Breast Cancer Res Treat. 2010, 121: 399-411. 10.1007/s10549-010-0887-y. doi:10.1007/s10549-010-0887-y
Hatzis C, Pusztai L, Valero V, Booser DJ, Esserman L, Lluch A, Vidaurre T, Holmes F, Souchon E, Wang H, Martin M, Cotrina J, Gomez H, Hubbard R, Chacon JI, Ferrer-Lozano J, Dyer R, Buxton M, Gong Y, Wu Y, Ibrahim N, Andreopoulou E, Ueno NT, Hunt K, Yang W, Nazario A, DeMichele A, O'shaughnessy J, Hortobagyi GN, Symmans WF: A genomic predictor of response and survival following taxane-anthracycline chemotherapy for invasive breast cancer. JAMA. 2011, 305: 1873-1881. 10.1001/jama.2011.593. doi:10.1001/jama.2011.593
Popovici V, Chen W, Gallas BG, Hatzis C, Shi W, Samuelson FW, Nikolsky Y, Tsyganova M, Ishkin A, Nikolskaya T, Hess KR, Valero V, Booser D, Delorenzi M, Hortobagyi GN, Shi L, Symmans WF, Pusztai L: Effect of training-sample size and classification difficulty on the accuracy of genomic predictors. Breast Cancer Res. 2010, 12: R5-10.1186/bcr2468. doi:10.1186/bcr2468
Shi L, Campbell G, Jones WD, Campagne F, Wen Z, Walker SJ, Su Z, Chu TM, Goodsaid FM, Pusztai L, Shaughnessy JD, Oberthuer A, Thomas RS, Paules RS, Fielden M, Barlogie B, Chen W, Du P, Fischer M, Furlanello C, Gallas BD, Ge X, Megherbi DB, Symmans WF, Wang MD, Zhang J, Bitter H, Brors B, Bushel PR, Bylesjo M, et al: The MicroArray Quality Control (MAQC)-II study of common practices for the development and validation of microarray-based predictive models. Nat Biotechnol. 2010, 28: 827-838. 10.1038/nbt.1665. doi:10.1038/nbt.1665
Tabchy A, Valero V, Vidaurre T, Lluch A, Gomez H, Martin M, Qi Y, Barajas-Figueroa LJ, Souchon E, Coutant C, Doimi FD, Ibrahim NK, Gong Y, Hortobagyi GN, Hess KR, Symmans WF, Pusztai L: Evaluation of a 30-gene paclitaxel, fluorouracil, doxorubicin, and cyclophosphamide chemotherapy response predictor in a multicenter randomized trial in breast cancer. Clin Cancer Res. 2010, 16: 5351-5361. 10.1158/1078-0432.CCR-10-1265. doi:10.1158/1078-0432.CCR-10-1265
Irizarry RA, Bolstad BM, Collin F, Cope LM, Hobbs B, Speed TP: Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 2003, 31: e15-10.1093/nar/gng015.
Bioconductor project [], [http://www.bioconductor.org/packages/2.14/bioc/html/affy.html]
Johnson WE, Li C, Rabinovic A: Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics. 2007, 8: 118-127. 10.1093/biostatistics/kxj037. doi:10.1093/biostatistics/kxj037
Lee YF, Miller LD, Chan XB, Black MA, Pang B, Ong CW, Salto-Tellez M, Liu ET, Desai KV: JMJD6 is a driver of cellular proliferation and motility and a marker of poor prognosis in breast cancer. Breast Cancer Res. 2012, 14: R85-10.1186/bcr3200. doi:10.1186/bcr3200
Parker JS, Mullins M, Cheang MC, Leung S, Voduc D, Vickery T, Davies S, Fauron C, He X, Hu Z, Quackenbush JF, Stijleman IJ, Palazzo J, Marron JS, Nobel AB, Mardis E, Nielsen TO, Ellis MJ, Perou CM, Bernard PS: Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol. 2009, 27: 1160-1167. 10.1200/JCO.2008.18.1370. doi:10.1200/JCO.2008.18.1370
Prat A, Parker JS, Karginova O, Fan C, Livasy C, Herschkowitz JI, He X, Perou CM: Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res. 2010, 12: R68-10.1186/bcr2635. doi:10.1186/bcr2635
Symmans WF, Ayers M, Clark EA, Stec J, Hess KR, Sneige N, Buchholz TA, Krishnamurthy S, Ibrahim NK, Buzdar AU, Theriault RL, Rosales MF, Thomas ES, Gwyn KM, Green MC, Syed AR, Hortobagyi GN, Pusztai L: Total RNA yield and microarray gene expression profiles from fine-needle aspiration biopsy and core-needle biopsy samples of breast carcinoma. Cancer. 2003, 97: 2960-2971. 10.1002/cncr.11435. doi:10.1002/cncr.11435
Chan OT, Yang LX: The immunological effects of taxanes. Cancer Immunol Immunother. 2000, 49: 181-185. 10.1007/s002620000122.
Haynes NM, van der Most RG, Lake RA, Smyth MJ: Immunogenic anti-cancer chemotherapy as an emerging concept. Curr Opin Immunol. 2008, 20: 545-557. 10.1016/j.coi.2008.05.008. doi: S0952-7915(08)00082-4
Mattarollo SR, Loi S, Duret H, Ma Y, Zitvogel L, Smyth MJ: Pivotal role of innate and adaptive immunity in anthracycline chemotherapy of established tumors. Cancer Res. 2011, 71: 4809-4820. 10.1158/0008-5472.CAN-11-0753. doi:0008–5472.CAN-11-0753
Tsavaris N, Kosmas C, Vadiaka M, Kanelopoulos P, Boulamatsis D: Immune changes in patients with advanced breast cancer undergoing chemotherapy with taxanes. Br J Cancer. 2002, 87: 21-27. 10.1038/sj.bjc.6600347. doi:10.1038/sj.bjc.6600347
Maj T, S Awek A, Kaleta-Kuratewicz K, Che Mo Ska-Soyta A: 17beta-Estradiol and interferon tau interact in the regulation of the immune response in a model of experimental autoimmune orchitis. J Interferon Cytokine Res. 2011, 31: 825-837. 10.1089/jir.2010.0147. doi:10.1089/jir.2010.0147
West NR, Milne K, Truong PT, Macpherson N, Nelson BH, Watson PH: Tumor-infiltrating lymphocytes predict response to anthracycline-based chemotherapy in estrogen receptor-negative breast cancer. Breast Cancer Res. 2011, 13: R126-10.1186/bcr3072. doi:10.1186/bcr3072
Emens LA, Machiels JP, Reilly RT, Jaffee EM: Chemotherapy: friend or foe to cancer vaccines?. Curr Opin Mol Ther. 2001, 3: 77-84.
Zitvogel L, Kepp O, Kroemer G: Decoding cell death signals in inflammation and immunity. Cell. 2010, 140: 798-804. 10.1016/j.cell.2010.02.015. doi:10.1016/j.cell.2010.02.015
Wan S, Pestka S, Jubin RG, Lyu YL, Tsai YC, Liu LF: Chemotherapeutics and radiation stimulate MHC class I expression through elevated interferon-beta signaling in breast cancer cells. PLoS One. 2012, 7: e32542-10.1371/journal.pone.0032542. doi:10.1371/journal.pone.0032542
Armeanu S, Bitzer M, Lauer UM, Venturelli S, Pathil A, Krusch M, Kaiser S, Jobst J, Smirnow I, Wagner A, Steinle A, Salih HR: Natural killer cell-mediated lysis of hepatoma cells via specific induction of NKG2D ligands by the histone deacetylase inhibitor sodium valproate. Cancer Res. 2005, 65: 6321-6329. 10.1158/0008-5472.CAN-04-4252. doi:10.1158/0008-5472.CAN-04-4252
Gasser S, Orsulic S, Brown EJ, Raulet DH: The DNA damage pathway regulates innate immune system ligands of the NKG2D receptor. Nature. 2005, 436: 1186-1190. 10.1038/nature03884. doi:10.1038/nature03884
Chakraborty M, Abrams SI, Coleman CN, Camphausen K, Schlom J, Hodge JW: External beam radiation of tumors alters phenotype of tumor cells to render them susceptible to vaccine-mediated T-cell killing. Cancer Res. 2004, 64: 4328-4337. 10.1158/0008-5472.CAN-04-0073. doi:10.1158/0008-5472.CAN-04-0073
Ghiringhelli F, Larmonier N, Schmitt E, Parcellier A, Cathelin D, Garrido C, Chauffert B, Solary E, Bonnotte B, Martin F: CD4 + CD25+ regulatory T cells suppress tumor immunity but are sensitive to cyclophosphamide which allows immunotherapy of established tumors to be curative. Eur J Immunol. 2004, 34: 336-344. 10.1002/eji.200324181. doi:10.1002/eji.200324181
Lutsiak ME, Semnani RT, De Pascalis R, Kashmiri SV, Schlom J, Sabzevari H: Inhibition of CD4(+)25+ T regulatory cell function implicated in enhanced immune response by low-dose cyclophosphamide. Blood. 2005, 105: 2862-2868. 10.1182/blood-2004-06-2410. doi:10.1182/blood-2004-06-2410
MacDonald KP, Rowe V, Bofinger HM, Thomas R, Sasmono T, Hume DA, Hill GR: The colony-stimulating factor 1 receptor is expressed on dendritic cells during differentiation and regulates their expansion. J Immunol. 2005, 175: 1399-1405. 10.4049/jimmunol.175.3.1399.
Nace G, Evankovich J, Eid R, Tsung A: Dendritic cells and damage-associated molecular patterns: endogenous danger signals linking innate and adaptive immunity. J Innate Immun. 2012, 4: 6-15. 10.1159/000334245. doi:10.1159/000334245
Obeid M, Tesniere A, Ghiringhelli F, Fimia GM, Apetoh L, Perfettini JL, Castedo M, Mignot G, Panaretakis T, Casares N, Metivier D, Larochette N, van Endert P, Ciccosanti F, Piacentini M, Zitvogel L, Kroemer G: Calreticulin exposure dictates the immunogenicity of cancer cell death. Nat Med. 2007, 13: 54-61. 10.1038/nm1523. doi:10.1038/nm1523
Tesniere A, Apetoh L, Ghiringhelli F, Joza N, Panaretakis T, Kepp O, Schlemmer F, Zitvogel L, Kroemer G: Immunogenic cancer cell death: a key-lock paradigm. Curr Opin Immunol. 2008, 20: 504-511. 10.1016/j.coi.2008.05.007. doi:10.1016/j.coi.2008.05.007
Shurin GV, Tourkova IL, Kaneno R, Shurin MR: Chemotherapeutic agents in noncytotoxic concentrations increase antigen presentation by dendritic cells via an IL-12-dependent mechanism. J Immunol. 2009, 183: 137-144. 10.4049/jimmunol.0900734. doi:10.4049/jimmunol.0900734
Butterfield LH: Dendritic cells in cancer immunotherapy clinical trials: are we making progress?. Frontiers in immunology. 2013, 4: 454-10.3389/fimmu.2013.00454. doi:10.3389/fimmu.2013.00454
Acknowledgements
The Microarray Shared Resource and Biostatistics Core of the Comprehensive Cancer Center of Wake Forest University (supported by NCI CCSG P30CA012197) provided resources for this study. The authors thank Megan J Whelen, MPH (Comprehensive Cancer Center of Wake Forest University) for copyediting, editorial, and production assistance. LDM is supported in part by the Mary Kirkpatrick Professorship for Breast Cancer Research and a research scholar grant from the American Cancer Society (RSG-12-198-01-TBG).
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Electronic supplementary material
13073_2014_80_MOESM2_ESM.docx
Additional file 2: Comparison of metagene distributions by biopsy type. Mean-centered metagene scores derived from microarray expression profiles of surgical samples (SURGbx; n = 1,954) and fine-needle aspirate biopsies (FNAbx; n = 482) were compared by box and whisker plot analysis. The upper and lower bounds of each box delineate the interquartile range (25th to 75th percentiles) and the horizontal black line within the box designates the median of the distribution. Upper and lower whiskers (T-bars) demarcate the 95th and 5th percentiles, respectively. Open circles mark the outliers (below and above the 5th and 95th percentiles, respectively). Red and green dashed lines mark the upper and lower tertile thresholds. (DOCX 61 KB)
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Alistar, A., Chou, J.W., Nagalla, S. et al. Dual roles for immune metagenes in breast cancer prognosis and therapy prediction. Genome Med 6, 80 (2014). https://doi.org/10.1186/s13073-014-0080-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13073-014-0080-8