Abstract
Background
Bacterial endocarditis is a recognised disease in humans and animals. In humans, infection with Coxiella burnetii can cause endocarditis, but this has not been investigated thoroughly in animals. Endocarditis in cattle is a common post-mortem finding in abattoirs and studies have identified Trueperella pyogenes as a major cause. Despite exposure of cattle to C. burnetii, the significance of this particular bacterium for development and progression of endocarditis has not been studied in detail. Cardiac valves of cattle affected with endocarditis (n = 100) were examined by histology, fluorescence in situ hybridization (FISH) and real time quantitative polymerase chain reaction (PCR). Serum was examined for anti-C. burnetii antibodies by enzyme-linked immunosorbent assay (ELISA).
Results
Serology revealed that 70% of the cattle were positive for antibodies to C. burnetii, while PCR analysis identified 25% of endocarditis valve samples as being positive. C. burnetii was not detected by FISH, probably due to the low infection levels. Most cattle had chronic valvular vegetative endocarditis with lesions being characterised by a core of fibrous tissue covered by significant amounts of fibrin, sometimes with areas of liquefaction, and with a coagulum covering the surface. In a few cases, including the case with the highest infection level, lesions were characterized by extensive fibrosis and calcification. Histologically, bacteria other than C. burnetii were observed in most cases.
Conclusions
The presence of C. burnetii DNA is relatively common in cattle affected with valvular endocarditis. The role of C. burnetii remains however unknown as lesions did not differ between C. burnetii infected and non-infected cattle and because T. pyogenes–like bacteria were present in the inflamed valves; a bacterium able to induce the observed lesions. Heart valves of normal cattle should be investigated to assess if C. burnetii may be present without preexisting lesions.
Similar content being viewed by others
Background
Coxiella burnetii is a Gram-negative obligate intracellular bacterium that infects a wide range of mammalian species, and causes the disease syndrome Q fever. Human cases of Q fever are generally regarded as being associated with exposure to domestic ruminants although a significant proportion of cases do not report a direct contact to animals [1].
In humans, infection is either subclinical or results in a self-limiting febrile illness. The infection may become chronic and lead to development of endocarditis in those individuals with predisposing conditions, such as valvulopathy, prosthetic valve implants, vascular abnormalities or immunosuppression [2]. In addition, infection during pregnancy carries an increased risk of miscarriage [3–5].
The diagnosis of Q fever in animals is typically associated with abortion or delivery of weak or stillborn offspring. Such reproductive consequences occur sporadically in cattle, while flock outbreaks have been reported in goats and sheep [6, 7]. There is little in the way of published research into non-reproductive clinical manifestations of Q fever in ruminant species, despite the documented high seroprevalence against C. burnetii reported in livestock [8]. However, circulating C. burnetii DNA has been detected sporadically in blood of cattle thus indicating that some animals occasionally develop coxiellaemia [9]. Based on the comparative aspects in humans, where C. burnetii is a well-known cause of endocarditis, it could be suspected that C. burnetii may also be implicated in the development or progression of endocarditis in cattle under certain circumstances.
Valvular endocarditis is a well-recognised condition in cattle, with an estimated prevalence of 1–2% observed during post-mortem inspection at abattoirs [10, 11]. The aetiology has been investigated using routine microbiological techniques in several studies and the cultureable bacterial flora of bovine endocarditis is well-known. Trueperella pyogenes, a major opportunistic pathogen of cattle, has been identified in the majority of cases [12–16], although streptococci were reported in an older study [17]. However, some bacteria, such as C. burnetii, cannot be cultured by standard bacteriological methods as the bacterium requires cell cultures for propagation due to its intracellular nature. Studies specifically targeting C. burnetii in bovine endocarditis cases have not been done, but the hypothesis of C. burnetii being associated with endocarditis in animals has been tested in northern sea otters, which inhabit an environment where marine mammals are exposed to C. burnetii; however C. burnetii was not found in cases of endocarditis [18].
Danish dairy cattle are frequently seropositive for C. burnetii thus showing a widespread exposure to this bacterium [19]. As endocarditis is a common finding in Danish slaughter cattle as well [11] and as cattle is expected to experience episodes of coxiellaemia [9], we performed a study to investigate if C. burnetii could be detected in inflamed cardiac valves of Danish cattle.
Methods
Study population and samples
Cardiac valves and blood samples were obtained from cattle (n = 100), where there was evidence of endocarditis at post mortem inspection at three Danish abattoirs (Danish Crown Tonder, Holsted and Aalborg), during a 12 month period. The heart was incised during routine post-mortem examination and the endocardium inspected for lesions. If present, a representative tissue sample was taken from the inflamed valve and blood was obtained from the heart or large veins. The samples and information on the location of cardiac lesions and presence of other lesions were submitted to the University of Copenhagen.
The tissue specimens were inspected upon arrival and transverse sections of 5 mm were made, aiming to preserve both the base and the surface of the lesion. Two neighbouring biopsy specimens were placed in either 10% neutral buffered formalin or in tubes without fixative, the latter stored at −80 °C until DNA extraction. Blood samples were centrifuged at 2000 g for 10 min and the serum stored at −80 °C until analysis. Data for each animal were obtained from the Danish Central Cattle Database and included breed, gender, herd of origin, and dates of birth and of slaughter.
Histopathology
The formalin fixed samples were routinely processed for histopathology, embedded in paraffin, sectioned at 3 μm, and stained with haematoxylin and eosin (H&E). Light microscopy was performed non-blinded for cases 1–50, while cases 51–100 were examined blinded to results of laboratory analysis (PCR and ELISA) by one researcher (JSA). Sections of a single case (Case #43) was additionally stained using periodic acid–Schiff (PAS), phosphotungstic acid haematoxylin (PTAH) and by the Masson’s trichrome method for connective tissue.
Serology
Serum samples were tested for antibodies to C. burnetii using an indirect enzyme-linked immunosorbent assay (ELISA) (LSIVet Ruminant Q Fever Serum/Milk ELISA Kit, Laboratoire Service International) according to the manufacturer’s instructions. Briefly, serum was diluted 1:400 in dilution buffer and transferred to wells of ELISA plates coated with antigen (total volume 100 μL). The plates were incubated for 1 h at 37 °C followed by washing three times and incubation with 100 μL anti-ruminant IgG peroxidase conjugate for 1 h at 37 °C. After washing three times, wells were incubated with 100 μL tetramethylbenzidine substrate for 10 min at room temperature (around 22 °C) in the dark. Colour development was stopped by adding 100 μL 0.5 M H2SO4. Absorbance values were measured at 450 nm (OD450). Antibody reactivity was calculated using the sample to positive ratio (S/P) calculated as (Sample OD – Negative OD) / (Positive OD – Negative OD) × 100. The S/P values were categorised as negative (S/P ratio ≤40) or positive (S/P ratio > 40).
Real-time PCR
DNA was extracted using the DNeasy Blood & Tissue Kit (Qiagen), according to the manufacturer’s instructions. Samples of approximately 4 mm3 in size were processed from the surface of vegetative changes. DNA samples were tested by a real-time PCR designed to detect the C. burnetii-specific IS1111a element [20] using specific primers and probe (Sense: 5′-CATCACATTGCCGCGTTTAC-3′, antisense: 5′-GGTTGGTCCCTCGACAACAT-3′, probe: FAM-AATCCCCAACAACACCTCCTTATTCCCAC-BHQ1). An inhibition control was used which utilizes the primers for the IS1111a element and a dedicated probe (YakimaYellow-ACATAATCTCTCCGACCCCACACTTCCATAC-BHQ1).
Reactions were performed using a 7500 Fast Real Time PCR system (Applied Biosystems) and consisted of 5 μL DNA template, 400 nM of each primer and 200 nM of each probe in 10 μL PerfeCTa Multiplex qPCR Supermix, uracil-N-glycosylase (2×) (New England Biolabs), with 0.4 μL Low Rox dye (Quanta BioSciences), 1 μL of inhibition control, and distilled water up to a total volume of 15 μL. An initial uracil DNA glycosylase (UDG) incubation for 5 min at 45 °C and denaturation/activation for 60 s at 95 °C was followed by 50 cycles of denaturation for 10 s at 95 °C, annealing/elongation for 30 s at 60 °C. Results were analysed with 7500 Fast System Software (Applied Biosystems) and scored negative when no amplicon was generated, or when the cycle threshold (Ct) value was >40; positive for Ct values ≤36, and inconclusive for Ct values between 36 and 40. When inhibition of the internal positive control was observed, samples were retested at a dilution 1 in 10.
Fluorescence in situ hybridization (FISH)
Cases were selected (n = 10) that demonstrated the highest infection load (i.e. lowest Ct values on real-time PCR) for C. burnetii, and controls (n = 10) were randomly selected from those with negative serology and PCR results. Serial sections of 3 μm thickness were prepared and subjected to FISH examination, blinded to the test results. Two oligonucleotide probes targeting the 16S rRNA were used: Eub-338; 5′-GCTGCCTCCCGTAGGAGT-3′ for the entire domain Bacteria and S-S-C.burnetti-443; 5′-CTTGAGAATTTCTTCCCC-3′ for C. burnetii [21]. The oligonucleotide probes were labelled at the 5′ end with fluorescein isothiocyanate (FITC) (Eub-338 and S-S-C.burnetti-443), or Cy3 (Eub-338 and S-S-C.burnetti-443) (Eurofins MWG Operon). The FITC-labelled Eub-338 and the Cy3-labelled S-S-C.burnetti-443 probes were used in a double hybridization, whereas the two other probes were used individually.
Hybridization was performed at 45 °C with 40 μL of hybridization buffer (100 mM Tris [pH 7.2], 0.9 M NaCl, 0.1% sodium dodecyl sulfate) and 200 ng of each probe for 16 h in a Sequenza slide rack (Thermo Shandon). The sections were washed three times at 45 °C in hybridization buffer for 15 min and subsequently three times in washing solution (100 mM Tris [pH 7.2], 0.9 M NaCl). The sections were rinsed in water, air dried, and mounted in Vectashield (Vector Laboratories) for fluorescence microscopy. An Axioimager M1 fluorescence microscope, equipped with a 100-W HBO lamp and filter sets 43 and 38 were used to visualize Cy3 and FITC, respectively.
Statistical analysis
Data were analysed in SAS version 9.4 (SAS Institute). ELISA and PCR results were considered to be ordinal variables and testing of associations between age and ELISA or PCR values were evaluated by Spearman’s correlation, linear regression and analysis of variance. Associations between gender and breed versus ELISA and PCR values were evaluated in contingency tables using the chi-square test. Associations between ELISA and PCR results were tested by Spearman’s correlation analysis and associations between ELISA and PCR results, categorized as test positive and test negative, were evaluated by Kappa estimation and McNemar’s test. Associations between type and number of heart valves affected was analysed in contingency tables using Fisher’s exact test.
Results
The samples originated from dairy cattle, mainly Holsteins (83%), and most cases were female (96%). Mean ages of males and females were 20.5 months (SD 12.2 months) and 53.0 months (SD 23.0 months), respectively. The cases originated from 92 herds (two animals from each of eight herds and one animal from each of the remaining 84 herds). The right atrioventricular (tricuspid) valve was most frequently affected (58%), followed by the left atrioventricular (mitral) (31%), pulmonary (24%) and aortic valve (16%), as recorded by the meat inspection personnel at the abattoirs. A likely source of systemic infection was identified in 17 cases (peritonitis/traumatic reticuloperitonitis, n = 11; liver abscesses, n = 6). Other lesions consistent with bacteraemia, such as embolic pneumonia, nephritis or osteomyelitis, were reported in around 50% of the endocarditis cases.
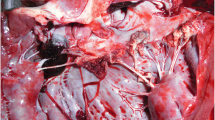
Most cases (n = 95) demonstrated chronic valvular vegetative endocarditis, affecting one or more valves, sometimes with adjacent mural lesions. Lesions were characterised by a core of pale fibrous tissue covered by significant amounts of fibrin and with a coagulum covering the surface (Fig. 1). Intralesional liquefaction was present in 27 cases. The lesions varied in size from <1 cm to several cm. One case had only acute/subacute lesions, which were characterized by haemorrhagic less extensive lesions without grossly visible granulation tissue. In other cases (n = 4), including Case #43, there was evidence of more chronic, organised lesions, characterized by extensive fibrosis and calcification (Fig. 2).
Chronic bovine fibrinous endocarditis. a gross pathology showing two areas of endocardial inflammation. The largest lesion, which is located in the valve (V), is cut through thus exposing a central core of fibrous tissue (F) that is externally covered by fibrin and clots (arrow). A minor mural lesion is similarly covered by fibrin and a clot; b + c Photomicrographs of endocardial lesions showing a core of granulation tissue (G) that is externally bordered by a zone of suppurative inflammation (I). The surface of the lesions is covered by fibrin (F) in which multiple bacterial colonies are embedded (arrows). c A focus of calcification (C) is present on the surface, which is also covered by a layer of neutrophils. Such calcified foci are consecutively embedded first in the fibrin and later in the granulation tissue as the lesion continues to expand by superficial opposition of fibrin and organisation into fibrous tissue from the base. (H&E)
Chronic bovine fibrous calcified endocarditis. a Gross pathology showing extensive fibrosis and thickening of the stroma (*) and widespread superficial calcification (arrows). Formalin fixed specimen. b Histopathology of the lesion, showing extensive fibrosis (*) and areas of calcification (arrows). (H&E)
In terms of the histopathology findings, in general there was a core consisting of fibrous tissue, with inflamed granulation tissue at the periphery. This was covered by a zone of suppurative inflammation with overlying layers of fibrin containing basophilic bacterial colonies and neutrophils, either widely dispersed or as accumulations of bacteria and debris (Fig. 1). An acute/subacute case was characterized by intra-valvular haemorrhage, slight suppurative inflammation and endothelial ulceration covered by fibrin in some locations, while more established lesions were characterised by valvular fibrosis and intact endothelium. In some of the chronic cases (n = 21), calcification was present within the layers of fibrin, in the zone of intense inflammation and in the fibrous core. The most widespread calcification was observed in chronic fibrous cases with intact/healed endothelium. Osseous metaplasia was present in one case.
None of the cases examined for C. burnetii by FISH were found to be positive, by use of the FITC nor Cy3 labelled probes. Similar to the H&E stained sections bacterial colonies of varying seize were observed in the fibrin layers of the suppurative inflammation when hybridized with the probe for the domain Bacteria (Fig. 3) with a bacterial morphology varying from coccoid to small rod-like.
Serology revealed that 70% of the endocarditis cases were positive for antibodies against C. burnetii. PCR analysis of heart valve lesions identified 25% as being positive, 52% negative and 23% were inconclusive. Ct values for the positive samples ranged from 33 to 36 (i.e. low copy number), except for one animal, a 7-year-old Jersey cow (Case #43) that demonstrated a Ct value of 13.7 in the aortic valve, and an ELISA S/P value of 82%.
Figure 4 is a scatter plot of the relations between ELISA and PCR values. Regression analysis showed there was no statistical significant association between ELISA and PCR values (P = 0.28), and when these were converted to categorical data (i.e. test positive vs. negative), estimation of Kappa (0.12) with associated McNemar’s test also failed to show any significant agreement. However, Spearman’s correlation analysis showed a weak association (R S = −0.225, R S 2 = 0.05, P = 0.025). Spearman’s correlation, regression analysis and analysis of variance showed no significant associations between age and ELISA or PCR results. There was no significant relationship between gender or breed versus ELISA or PCR results.
Scatter plot of Coxiella burnetii DNA in bovine endocarditis lesions and corresponding serology results. ELISA S/P values >40 are considered to be positive, those ≤40 are considered to be negative. PCR Ct values ≤36 are considered to be positive, those between 36 and 40 are inconclusive and those >40 are considered negative. Samples with no amplicon present (i.e. below the detection limit) were assigned a nominal Ct value of 41. Cut-off values are indicated by red lines and inconclusive by a dotted line
Discussion
In humans, detection of C. burnetii by culture, PCR or immunohistochemistry of a cardiac valve is required for making a definitive diagnosis of Q fever endocarditis [22]. However, in a number of PCR- or culture-positive cases, there are no signs of inflammation [23], highlighting the challenges associated with making this diagnosis. Fulminant Q fever valvular endocarditis in humans is characterized by inflammation of either acute or chronic active nature, dominated by focal or diffuse infiltrations by lymphocytes, histiocytes and occasional plasma cells. Microabscesses and necrosis are present in some cases and the surface is covered by fibrinous vegetations and leukocytic infiltrations, while the valvular stroma shows extensive fibrosis and calcification. Macrophages containing intracellular bacteria may be demonstrated in the vegetations by use of special stains or immunohistochemistry [24–26].
The bovine endocarditis lesions examined in the present study were similar to those described in human Q fever endocarditis, although they were probably more advanced in nature as bovine endocarditis may remain undiagnosed for a long period of time due to lack obvious clinical signs until severely compromised valvular function. Despite the similarities in histopathology, the association between the development and progression of endocarditis and C. burnetii infection is not clear. Huge numbers of bacteria with morphology and staining abilities consistent with T. pyogenes were commonly visualised within the endocarditis lesions thus showing that development and progression of the lesion may be associated with another primary bacterial infection rather than with C. burnetii. Also around 50% of the cattle had lesions consistent with pyaemia, a condition likely to be the main reason for many animals to develop endocarditis. Bacterial culturing was not performed as the present study focused on detection of C. burnetii only and because the culturable bacterial flora of bovine endocarditis has been examined thoroughly in several studies [10, 12, 15]. The PCR results indicated that even in positive samples, the copy number (indicative of infection ‘load’) was low in all but one animal (Case #43). It is therefore tempting to speculate that C. burnetii contributed to the endocarditis in most cases as a co-inhabitant of a primary T. pyogenes pyaemia-associated endocarditis.
A secondary role of C. burnetii in the development of bovine endocarditis is also supported by the negative FISH results. The FISH technique only detects bacteria in active growth, so in addition to a low infection level, negative FISH results may also indicate presence of only inactive bacteria. Furthermore, C. burnetii is difficult to visualize due its small size. So unless present in bacteria-loaded macrophages, which indicates intracellular multiplication, detection of C. burnetii by FISH is challenging. As macrophages loaded with C. burnetii were absent, a major role of C. burnetii in the development and progression of bovine endocarditis seems less likely.
In the specimen with the most conclusively positive PCR result (Case #43), the pathology was dominated by extensive fibrosis and calcification. Such lesions have been reported in cases of human Q fever endocarditis and more commonly occur in association with C. burnetii than in lesions caused by another aetiology [26]. Whether extensive fibrosis and calcification is associated with C. burnetii in cattle remains to be studied. This type of lesion may represent the end-stage of healing of a previous vegetative endocarditis. Calcification within the fibrin layers, at the borders between fibrin and granulation tissue and within the granulation tissue was found in many of the examined cases and suggests that calcification develops within the fibrin layer and gradually becomes embedded in the connective tissue core, during re-organisation of the fibrinous mass from its base. Such calcification could persist for prolonged periods in the fibrotic core, either as inert calcification or within granulomas, if associated with a foreign body reaction. Examination of other cases is needed to determine whether extensive fibrosis and calcification are associated with C. burnetii endocarditis, or if the presence of C. burnetii in such cases simply reflects that the animal was at risk of C. burnetii colonisation due to a pre-existing long lasting valvulopathy.
Most of the cattle examined (70%) were seropositive for antibody reactivity to C. burnetii by ELISA. In Denmark, recent studies have shown that 59–79% of bulk milk samples of dairy cattle herds are antibody positive [27, 28] thus demonstrating that Danish dairy cattle are commonly exposed to this organism. The presence of C. burnetii DNA in 25% of the endocarditis valves shows that cattle may experience coxiellaemia following exposure, which potentially might lead to colonisation of the cardiac valves when other predisposing conditions are present. This case series focused on detection of C. burnetii in cases of endocarditis and did not include normal heart valves, so a possible presence of C. burnetii in normal heart valves remains uncertain. It is therefore not possible to determine the role of C. burnetii in the initiation of endocardial inflammation as only grossly affected valves were included in the study. A case-control study should be conducted to obtain more information on the role of C. burnetii in the development of bovine valvular endocarditis. However, as valvulopathy, including pre-existing valvular endocarditis, is a recognized risk factor for developing C. burnetii associated endocarditis in humans, cattle with such lesions most likely have a higher risk for C. burnetii valvular colonisation than normal cattle. In cases of vegetative endocarditis, the continued apposition of fibrin probably gives C. burnetii bacteria passing the valves in case of coxiellaemia a greater chance of being trapped and subsequently infecting the valve than in animals without pre-existing valvulopathy.
Because the statistical analysis showed a weak correlation between the ELISA and PCR values (P = 0.025), and regression analysis and the estimation of Kappa with the associated McNemar’s test did not show any significant agreement between PCR and ELISA results, we conclude that the relationship between the serology and PCR data is not conclusive. Given the high seroprevalence in the cattle population and the relatively low prevalence of endocarditis, it is not likely that serology can be used as part of the diagnostic approach for Q fever endocarditis. The cow with high infection load (Case #43) illustrates this point as it had a relatively low ELISA S/P value. Serology and PCR have been used to study cows excreting C. burnetii in milk or vaginal discharge, but the association is usually weak [29–31], although many cows that persistently excrete C. burnetii organisms in the milk often show high antibody titres [32].
Conclusions
The findings indicate the presence of C. burnetii DNA in a proportion of heart valve lesions in cattle. The exact role of C. burnetii in the development and progression of lesions remains unknown as most cases were also infected by huge numbers of T. pyogenes; a bacterium known to be associated with severe endocarditis in cattle. C. burnetii PCR positive and negative cases could not be distinguished pathologically, but it is noteworthy that the cow with the highest infection level had extensive valvular fibrosis, widespread calcification and an intact endothelium thus being morphologically unique in this case series. Cattle affected with vegetative endocarditis are usually condemned as unfit for human consumption and therefore are unlikely to pose a zoonotic risk to humans. There is a need for more research into the clinical consequences of Q fever in ruminants, beyond those focusing on reproductive disease. The spectrum of clinical signs seen in the human infection should be taken into consideration, when assessing the possible health implications in livestock. In particular, the role of C. burnetii in bovine endocarditis merits further investigation, including examination for its possible presence in normal heart valves.
Abbreviations
- C. burnetii :
-
Coxiella burnetii
- Ct:
-
Cycle threshold
- ELISA:
-
Enzyme-linked immunosorbent assay
- H&E:
-
Haematoxylin and eosin
- PCR:
-
Polymerase chain reaction
- S/P:
-
Sample to positive ratio
- SD:
-
Standard deviation
- T. pyogenes :
-
Trueperella pyogenes
References
Arricau-Bouvery N, Rodolakis A. Is Q fever an emerging or re-emerging zoonosis? Vet Res. 2005;36:327–49.
Brouqui P, Raoult D. Endocarditis due to rare and fastidious bacteria. Clin Microbiol Rev. 2001;14:177–207.
Angelakis E, Raoult D. Q fever. Vet Microbiol. 2010;40:297–309.
Million M, Roblot F, Carles D, D’Amato F, Protopopescu C, Carrieri MP, Raoult D. Reevaluation of the risk of fetal death and malformation after Q fever. Clin Infect Dis. 2014;59:256–60.
Nielsen SY, Molbak K, Henriksen TB, Krogfelt KA, Larsen CS, Villumsen S. Adverse pregnancy outcomes and Coxiella burnetii antibodies in pregnant women, Denmark. Emerg Infect Dis. 2014;20:925–31.
Agerholm JS. Coxiella burnetii associated reproductive disorders in domestic animals - a critical review. Acta Vet Scand. 2013;55:13.
Garcia-Ispierto I, Tutusaus J, López-Gatius F. Does Coxiella burnetii affect reproduction in cattle? A clinical update. Reprod Dom Anim. 2014;49:529–35.
Guatteo R, Seegers H, Taurel AF, Joly A, Beaudeau F. Prevalence of Coxiella burnetii infection in domestic ruminants: a critical review. Vet Microbiol. 2011;149:1–16.
Das DP, Malik SV, Rawool DB, Das S, Shoukat S, Gandham RK, et al. Isolation of Coxiella burnetii from bovines with history of reproductive disorders in India and phylogenetic inference based on the partial sequencing of IS1111 element. Infect Genet Evol. 2014;22:67–71.
Strehle U, Welbers N. Die Pyogenes-Endokarditis des Rindes. Tierärztl Umsch. 1987;42:869–78.
Denwood M, Houe H, Forkman B, Nielsen SS. Random effect selection in generalised linear models: a practical application to slaughterhouse surveillance data in Denmark. In: Proc Ann Meeting Soc Vet Epidemiol Prev Med, Ghent, Belgium. 2015. p. 135–45.
Power HT, Rebhun WC. Bacterial endocarditis in adult dairy cattle. J Amer Vet Med Assoc. 1983;182:806–8.
Narucka U, van den Berg J, Nouws JF, Okma BD, Peelen JP, Soethout AE. Afwijingen bij slachtdieren. V. Endocarditis bij slachtvarkens, zeugen en runderen. Tijdschr Diergeneeskd. 1985;110:776–9.
Healy AM. Endocarditis in cattle. A review of 22 cases. Irish Vet J. 1996;49:43–8.
Müller M, Platz S, Ehrlein J, Ewringmann T, Mölle G, Weber A. Bakteriell bedingte Thromboembolie bei Milchkühen – eine retrospektive Auswertung von 31 Sektionsfällen unter besonderer Berücksichtigung des Ursachenkomplexes. Berl Munch Tierarztl Wochenschr. 2005;118:121–7.
Buczinski S, Francoz D, Fecteau G, DiFruscia R. Heart disease in cattle with clinical signs of heart failure: 59 cases. Can Vet J. 2010;51:1123–9.
Evans ETR. Bacterial endocarditis of cattle. Vet Rec. 1957;69:1190–206.
Duncan C, Gill VA, Worman K, Burek-Huntington K, Pabilonia KL, Johnson S, et al. Coxiella burnetii exposure in northern sea otters (Enhydra lutris kenyoni). Dis Aquat Org. 2015;114:83–7.
Paul S, Agger JF, Agerholm JS, Markussen B. Prevalence and risk factors of Coxiella burnetii seropositivity in Danish beef and dairy cattle at slaughter adjusted for test uncertainty. Prev Vet Med. 2014;113:504–11.
Roest HJ, Ruuls RC, Tilburg JJ, Nabuurs-Franssen MH, Klaassen CH, Vellema P, et al. Molecular epidemiology of Coxiella burnetii from ruminants in Q fever outbreak, the Netherlands. Emerg Infect Dis. 2011;17:668–75.
Jensen TK, Montgomery DL, Jaeger PT, Lindhardt T, Agerholm JS, Bille-Hansen V, Boye M. Application of fluorescent in situ hybridisation for demonstration of Coxiella burnetii in placentas from ruminant abortions. Acta Pathol Microbiol Immunol Scand. 2007;115:347–53.
Raoult D. Chronic Q, fever: expert opinion versus literature analysis and consensus. J Infect. 2012;65:102–8.
Grisoli D, Million M, Edouard S, Thuny F, Lepidi H, Collart F, et al. Latent Q fever endocarditis in patients undergoing routine valve surgery. J Heart Valve Dis. 2014;23:735–43.
Brouqui P, Dumler JS, Raoult D. Immunohistologic demonstration of Coxiella burnetii in the valves of patients with Q fever endocarditis. Amer J Med. 1994;97:451–8.
Bruneval P, Choucair J, Paraf F, Casalta JP, Raoult D, Scherchen F, Mainardi JL. Detection of fastidious bacteria in cardiac valves in cases of blood culture negative endocarditis. J Clin Pathol. 2001;54:238–40.
Lepidi H, Houpikian P, Liang Z, Raoult D. Cardiac valves in patients with Q fever endocarditis: microbiological, molecular, and histologic studies. J Infect Dis. 2003;187:1097–106.
Agger JF, Christoffersen AB, Rattenborg E, Nielsen J, Agerholm JS. Prevalence of Coxiella burnetii antibodies in Danish dairy herds. Acta Vet Scand. 2010;52:5.
Agger JF, Paul S. Increasing prevalence of Coxiella burnetii seropositive Danish dairy cattle herds. Acta Vet Scand. 2014;56:46.
Guatteo R, Joly A, Beaudeau F. Shedding and serological patterns of dairy cows following abortions associated with Coxiella burnetii DNA detection. Vet Microbiol. 2012;155:430–3.
Natale A, Bucci G, Capello K, Barberio A, Tavella A, Nardelli S, et al. Old and new diagnostic approaches for Q fever diagnosis: correlation among serological (CFT, ELISA) and molecular analyses. Comp Immunol Microbiol Infect Dis. 2012;35:375–9.
Niemczuk K, Szymańska-Czerwińska M, Śmietanka K, Bocian Ł. Comparison of diagnostic potential of serological, molecular and cell culture methods for detection of Q fever in ruminants. Vet Microbiol. 2014;171:147–52.
Guatteo R, Beaudeau F, Joly A, Seegers H. Coxiella burnetii shedding by dairy cows. Vet Res. 2007;38:849–60.
Acknowledgements
The meat inspection staff at the slaughter houses is greatly acknowledged for submission of samples. Michiel Kroese (CVI) is greatly acknowledged for technical assistance with the PCR analysis and the C. burnetii ELISA while Heidi Holm, Anne Dorte Roed and Annie Ravn Pedersen are greatly acknowledged for technical assistance with histology, sample handling and FISH, respectively.
Funding
This study was funded by the authors’ institutions.
Availability of data and materials
The dataset is available upon request from the corresponding author.
Authors’ contributions
JSA designed the study, organized the submission of samples, performed gross and microscopic examination and drafted the manuscript. TKJ performed the FISH examinations. JFA did the statistics. MYE and HIJR supervised the PCR and ELISA analyses. All authors participated in the interpretation of data and writing the manuscript. All authors have read and approved the final version of the manuscript.
Competing interests
The authors declare that they have no competing interests.
Consent for publication
Not applicable.
Ethics approval and consent to participate
This study did not require any institutional or official approval.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Agerholm, J.S., Jensen, T.K., Agger, J.F. et al. Presence of Coxiella burnetii DNA in inflamed bovine cardiac valves. BMC Vet Res 13, 69 (2016). https://doi.org/10.1186/s12917-017-0988-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12917-017-0988-5