Abstract
Background
Presenilin-1 (PSEN1) is one of the causative genes for early onset Alzheimer’s disease (EOAD). Recently, emerging studies reported several novel PSEN1 mutations among Asian. We describe a male with EOAD had a pathogenic PSEN1 mutation.
Case presentation
A 53-year-old male presented with memory decline, followed by difficulty in finding ways. Patient had positive family history, since his mother and one of his brother was also affected with dementia. Brain magnetic resonance imaging (MRI) scan showed mild degree of atrophy of bilateral hippocampus and parietal lobe. 18F-Florbetaben-PET (FBB-PET) revealed increased amyloid deposition in bilateral frontal, parietal, temporal lobe and precuneus. Whole exome analysis revealed a heterozygous, probably pathogenic PSEN1 (c.695G > T, p.W165C) mutation. Interestingly, Trp165Cys mutation is located in trans membrane (TM)-III region, which is conserved between PSEN1/PSEN2. In vitro studies revealed that PSEN1 Trp165Cys could result in disturbances in amyloid metabolism. This prediction was confirmed by structure predictions and previous in vitro studies that the p.Trp165Cys could result in decreased Aβ42/Aβ40 ratios.
Conclusion
We report a case of EOAD having a pathogenic PSEN1 (Trp165Cys) confirmed with in silico and in vitro predictions.
Similar content being viewed by others
Background
Alzheimer’s disease (AD) (MIM #104300) is a neurodegenerative disease among elderlies, which is resulted by abnormal protein assembly inside the brain. Extracellular and intracellular amyloid beta (Aβ) and Tau protein, respectively, were associated as the main hallmarks of the disease. Early onset AD (EOAD) and late onset AD (LOAD) are the two main forms of the disease. Three genes were verified as causative factor for EOAD: amyloid precursor protein (APP) (MIM #104760) [1], presenilin 1 (PSEN1) (MIM #104311) [2], and presenilin 2 (PSEN2) (MIM #600759) [3]. Approximately 300 mutations of PSEN1, PSEN2, and APP in 635 affected individuals or families have been reported in the Dementia Mutation Database [4] (https://www.alzforum.org/mutations). Majority of mutations were observed in PSEN1 [5,6,7] (n = 219, 76.6%) with over 230 mutations reported as pathogenic in the Alzforum database (https://www.alzforum.org/mutations/psen-1), as compared to APP (n = 51, 17.8%), and PSEN2 (n = 16, 5.6%) [8,9,10,11].
PSEN1 protein contains nine transmembrane (TM) domains, connected with hydrophilic loop regions. As the member of γ- secretase complex, PSEN1 could function as a catalytic subunit of aspartyl protease, involved in the cleavage of C99 residue in APP protein into β-amyloid (Aβ) peptide. PSEN1 mutations may impair the γ-secretase processing, resulting in altered of Aβ production. Gain-of-function mutations could increase the amyoid processing and the ratio Aβ42/Aβ40 [12, 13]. Loss- of- function mutations may reduce protective mechanisms, such as α-secretase cleavage [14]. In addition, accumulation of amyloid peptides may also associated with the reduced Aβ42 clearance [15] and neuronal loss [15,16,17]. In majority of patients, disease occurred at 40–50 years of age [18,19,20,21]. Several cases of young onset AD have been reported, where patients were less than 30–40 years of age [6, 7, 22,23,24,25,26,27,28,29,30,31]. Guerrio et al. (2010) designed an algorithm for variants in EOAD causative genes [32] on prediction of the pathogenic nature of novel mutations. Investigating patients carrying novel as well as previously known mutations along with the associated phenotypes will aid in classification of these variants and may eventually support genetic counseling [33]. In this study, we reported a pathogenic PSEN1 W165C mutation as determined by genetic testing in a Korean patient with EOAD.
Case presentation
A 53-year-old man with 13 years of education presented with progressive memory decline. At aged 50, he complained forgetfulness of meetings or details of story, and repeating the same questions. He had difficulty in orienting to date and in finding way to a new place. His past medical history revealed myocardial infarction with proper medical treatment. His Korean version of Mini-Mental Status Examination (K-MMSE) score was 21/30 and clinical dementia rating scale (CDR) score was 0.5 at three years after symptom onset. And follow-up K-MMSE score at six years after symptom onset was 19/30 and CDR score was 1. His brain MRI at three years after symptom onset revealed mild atrophy of bilateral hippocampus and parietal lobe (Fig. 1a). FBB-PET at five years after symptom onset showed increased amyloid deposition in bilateral parietal, frontal, temporal lobe and precuneus (Fig. 1b). The patient had an APOE ε2 /ε3 polymorphism.
Brain functional and structural neuroimaging data of the proband at diagnosis. a. Axial FLAIR, coronal, and sagittal T1 images of brain MRI, arrows pointing at left-dominant bilateral temporal lobe atrophy. b. FDG-PET, arrows pointing at hypometabolism in left temporal cortex, right anterior temporal cortex and bilateral frontal cortex
A 53-year-old man (III-1, Fig. 2) visited the Seoul National University Bundang Hospital with gradually impaired cognitive function over the previous years. The proband’s family history had a strong family history of dementia, and presented several family members affected by EOAD (Fig. 2). His mother (II-2) suffered from AD with onset in her fifties and deceased. The patient was one of the 4 siblings, comprising 2 brothers and 2 sisters. His first younger brother (III-2) was also diagnosed AD deceased in his forties, and had 2 children. His second younger brother (III-5) and his two younger sisters (III-3, III-4) displayed normal cognitive function. The health condition of the rest of his family members remained unknown, since all living family members and relatives declined to provide any additional information regarding their health.
Genetic analysis of PSEN1 and structural prediction of mutant PSEN 1 protein
Methods
An in depth genetic screen was performed using a specifically expanded panel of 50 causative and risk factor genes for various neurodegenerative disorders [34]. Whole exome sequencing (WES) was performed in Novogene. Standard Sanger sequencing was carried out by BioNeer Inc. (Dajeon, Republic of Korea) [34]. Big Dye Terminator Cyclic sequencing was performed using the ABI 3730XL DNA Analyzer (Bioneer Inc., Dajeon, Republic of Korea). Sequencing data was analyzed using NCBI Blast (http://blast.ncbi.nlm.nih.gov/Blast.cgi) and the chromatograms were screened using the DNA BASER (http://www.dnabaser.com) tool. Possible novel mutations were checked in the Korean Reference Genome Database (KRGDB; http://nih.go.kr/menu.es?mid=a50303020300), which was obtained by whole genome sequencing of 622 healthy Korean individuals. The mutations were also screened against Broad Institute’s Genome Association Database (genome AD, http://gnomad.broadinstitute.org) and 1000 Genomes (http://www.1000genomes.org/) databases.
The possible pathogenic nature of missense variants was predicted using simple online tools, such as PolyPhen-2 (http://genetics.bwh.harvard.edu/pph2), Sorting Intolerant from Tolerant (SIFT; http://sift.jcvi.org/), and PROVEAN (http://provean.jcvi.org) algorithms. ExPasy analysis was also performed (https://www.expasy.org/) using different parameters, such as Kyte and Doolittle hydrophobicity index, bulkiness, and polarity. Mutant and normal prion protein structures were compared by 3D modelling. Protein structures were built using the Raptor X web server (http://raptorx.uchicago.edu/), while Discovery Studio 3.5 Visualizer (Accelrys, San Diego, USA) was used to display the 3D images [35].
Results
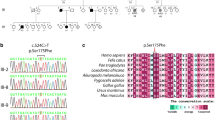
A heterozygous G > T substitution (chr14; g.73653575: G > T) was discovered and confirmed to occur in the PSEN1 coding region using both WES and standard sequencing. This mutation caused the change from tryptophan to cysteine (c.495G > C; p.Trp165Cys) substitution, located at exon 6 of PSEN1 gene, and in transmembrane (TM) helix domain-III of the PSEN1 protein (Fig. 3).
The mutation is associated with EOAD patient with memory decline, followed by difficulty in finding ways and had a strong family history of AD; however, the specific mechanism is not functionally uncovered. The mutation was found in subject a Korean patient with EOAD and not observed in KRGDB, ExAC, and 1000genome control data sets. All in silico pathogenicity programs predicted the mutation to be deleterious. Figure 4 predicted that abnormal conformation inside the helix results in the mutated form due to an abnormal intra-or intermolecular disulfide bridge associated with the potential re-activities with metals or other compounds with thiol groups. Additionally, cysteine is not common recorded in the helix, further suggesting that this mutation might lead to abnormal conformation within the TM region. The intramolecular interactions may also change with the mutation: Trp165 has strong interaction with Ser169 (two hydrogen bonds), and forms another hydrogen bond with Val161. Cys165 changes the hydrogen bonds significantly. The interaction with Ser169 and Val 169 remained, but with Ser169, only one hydrogen bond was visible. Two new interactions could be seen with Ile162 and Ile168 (Fig. 5) In addition, the mutation is localized to trans membrane-III region conserved between PSEN1/PSEN2 and expected to affect Ab42 levels. This hypothesis was previously demonstrated that PSEN1 W165C led to increase Aβ42 and decreased Aβ40, resulting in elevated Aβ42/Aβ40 ratio in gaining loss of function in presenilin [36,37,38,39].
Discussions
Since four different populations have been previously described the PSEN1 Trp165 from four familial AD cases (Table 1), the 165 codon seems to be a very vulnerable site. Initially, PSEN1 Trp165Cys mutation was found in a French family, with a codon combination of TGG > TGC. Mutation was associated with strongly was positive family history, since several affected family members were identified in three generations. Age of onset ranged between 37 and 47 years in the relatives with disease. No details were available on clinical symptoms of affected patients [36]. Second case of Trp165Cys was discovered with alternative codon exchange of TGG > TGT in an Indian family. Affected patients developed disease in their 40s, and disease phenotypes were rapid progressive disease progression and cerebral/cerebellar atrophies [37]. Our case was associated with probable EOAD case in a male patient, and his family members presented AD in their 40s. It may be difficult to find out, whether there could be a common founder between the Indian and Korean families. Since the Korea and India may be geographically isolated from each other, we suggest that PSEN1 Trp165Cys occur independently in these two families.
PSEN1 Trp165Cys is located in the TM-III region of PSEN1 protein. An exchange from native amino acid to Cys may increase the risk of an abnormal intramolecular disulfide bond formation with another Cys. These new S-S bonds may create novel inter- or intramolecular structures, involving in pathogenic mechanisms. The potential mechanism was displayed and revealed the Fig. 5. In addition, at the residue, another mutation to glycine (Gly, G) was previously documented in a Japanese family with young onset AD [39], suggesting that this residue may be critical for PSEN1 function. Interestingly, the mutation is located to TM-III region conserved between PSEN1/PSEN2 and expected to affect Ab42 levels. This prediction was previously demonstrated that PSEN1 Trp165Cys resulted in increased Aβ42 and decreased Aβ40, respectively. It could lead to elevated Aβ42/Aβ40 ratio in gaining loss of function in presenilin [36,37,38,39], verifying as a pathogenic mutation, involved in EOAD. This mutation was associated with rapid progression of disease, since the duration from the first clinical symptoms to the death ranged 4–10 years. Earlier onset of disease (37–47 years) was observed in a French family, but there was no information on the clinical symptoms in this family (Table 1).
Several pathogenic PSEN genes are found to cluster within the predicted α-helical TMs. Among other TMs, TM3 has been identified as one of the critical site in PSEN1, where several familiar AD (FAD) mutations were found. Recently, more than twenty mutations associated with FAD have been reported in TM-III of PSEN1 [36, 40,41,42,43,44,45,46,47,48,49,50] (Fig. 6), and several of them were associated with EOAD (Table 2). Among these mutations, the PSEN1 Trp165Cys mutation is of particular interest because it may be associated with disease early onset. Moreover, the Trp165Cys mutation could cause increase in the Aβ42/Aβ total ratio [36,37,38,39], similarly to other FAD-associated PSEN1 H163R [51], H163Y [52], L166P [53], I167del [54], S170F [31] S170P [40], L174del [44], L173 W [36] and L174 M [45] mutations.
In our case, the PSEN1 p.Trp165Cys variant has been identified in a patient with early onset of age (50s years at diagnosis), suggesting that disease phenotype may be the result of amino acid substitution in this conservative residue. Furthermore, the amino acid position 165, located in the TM-III of PSEN1 indicated a significant phylogenetic conservation among vertebrates, and in homologous proteins such as PSEN2, suggesting that the position is of functional significance. Importantly, the patient’s mother and his brother were also affected by AD that is likely to involve autosomal dominant AD.
Guerrio et al. (2010) designed an algorithm on mutations on PSENs. which may be helpful in prediction on their pathogenic nature [32]. PSEN1 Trp165Cys may be a definitely pathogenic mutation. The Korean case of PSEN1 Trp165Cys may be associated with positive family history of disease, since the mother and one of the brother of patient was affected with AD. This is the third case of Trp165Cys, described all around the world, and EOAD was observed in all cases of AD. All of these findings suggested that Trp165 may be an important in PSEN1, since it bound two pathogenic mutations, Trp165Cys and Trp165Gly [36,37,38]. Functional studies, performed by Sun et al. (2016) revealed that mutation may impair the gamma secretase activity, resulting in elevated amyloid beta 42 production [55]. Our findings confirmed the significance of PSEN1 Trp165Cys in EOAD.
Conclusions
We confirm that PSEN1 p.Trp165Cys may be commonly associated with EOAD. Our findings were consistent with the previously reported cases of this mutation, and supported the hypothesis that PSs contribute the identification of at risk relatives who may be potential candidates for clinical trials.
Availability of data and materials
Not applicable.
Abbreviations
- AD:
-
Alzheimer’s disease
- APP:
-
Amyloid precursor protein
- Aβ:
-
β-amyloid
- CDR:
-
clinical dementia rating scale
- EOAD:
-
Early-onset Alzheimer’s disease
- FDG-PET:
-
Fludeoxyglucose- positron emission tomography
- KRGDB:
-
Korean Reference Genome Database
- MMSE:
-
Minimental state examination
- MRI:
-
Magnetic resonance imaging
- PolyPhen2:
-
Polymorphism phenotyping v2
- PS1:
-
Presenilin-1
- PSEN1:
-
Presenilin 1
- PSEN2:
-
Presenilin 2
- SIFT:
-
Sorting intolerant from tolerant
- TM-III:
-
Transmembrane segment III
- WES:
-
Whole exome sequencing
References
Goate A, Chartier-Harlin MC, Mullan M, Brown J, Crawford F, Fidani L, Giuffra L, Haynes A, Irving N, James L, et al. Segregation of a missense mutation in the amyloid precursor protein gene with familial Alzheimer's disease. Nature. 1991;349(6311):704–6.
Sherrington R, Rogaev EI, Liang Y, Rogaeva EA, Levesque G, Ikeda M, Chi H, Lin C, Li G, Holman K, et al. Cloning of a gene bearing missense mutations in early-onset familial Alzheimer's disease. Nature. 1995;375(6534):754–60.
Levy-Lahad E, Wasco W, Poorkaj P, Romano DM, Oshima J, Pettingell WH, Yu CE, Jondro PD, Schmidt SD, Wang K, et al. Candidate gene for the chromosome 1 familial Alzheimer's disease locus. Science. 1995;269(5226):973–7.
Cruts M, Theuns J, Van Broeckhoven C. Locus-specific mutation databases for neurodegenerative brain diseases. Hum Mutat. 2012;33(9):1340–4.
Loy CT, Schofield PR, Turner AM, Kwok JB. Genetics of dementia. Lancet. 2014;383(9919):828–40.
Giau VV, Bagyinszky E, Yang YS, Youn YC, An SSA, Kim SY. Genetic analyses of early-onset Alzheimer’s disease using next generation sequencing. Sci Rep. 2019;9(1):8368.
Giau VV, Senanarong V, Bagyinszky E, An SSA, Kim S. Analysis of 50 neurodegenerative genes in clinically diagnosed early-onset Alzheimer's disease. Int J Mol Sci. 2019;20(6).
Raux G, Guyant-Marechal L, Martin C, Bou J, Penet C, Brice A, Hannequin D, Frebourg T, Campion D. Molecular diagnosis of autosomal dominant early onset Alzheimer's disease: an update. J Med Genet. 2005;42(10):793–5.
Janssen JC, Beck JA, Campbell TA, Dickinson A, Fox NC, Harvey RJ, Houlden H, Rossor MN, Collinge J. Early onset familial Alzheimer's disease: mutation frequency in 31 families. Neurology. 2003;60(2):235–9.
Giau VV, Bagyinszky E, An SSA, Kim S. Clinical genetic strategies for early onset neurodegenerative diseases. Molecular & Cellular Toxicology. 2018;14(2):123–42.
Giau VV, Pyun JM, Bagyinszky E, An SSA, Kim S. A pathogenic PSEN2 p.His169Asn mutation associated with early-onset Alzheimer's disease. Clin Interv Aging. 2018;13:1321–9.
Tolia A, De Strooper B. Structure and function of gamma-secretase. Semin Cell Dev Biol. 2009;20(2):211–8.
De Strooper B. Aph-1, Pen-2, and Nicastrin with Presenilin generate an active gamma-secretase complex. Neuron. 2003;38(1):9–12.
Giau VV, Lee H, Shim KH, Bagyinszky E, An SSA. Genome-editing applications of CRISPR-Cas9 to promote in vitro studies of Alzheimer's disease. Clin Interv Aging. 2018;13:221–33.
Fernandez MA, Klutkowski JA, Freret T, Wolfe MS. Alzheimer presenilin-1 mutations dramatically reduce trimming of long amyloid beta-peptides (Abeta) by gamma-secretase to increase 42-to-40-residue Abeta. J Biol Chem. 2014;289(45):31043–52.
Xia D, Watanabe H, Wu B, Lee SH, Li Y, Tsvetkov E, Bolshakov VY, Shen J, Kelleher RJ 3rd. Presenilin-1 knockin mice reveal loss-of-function mechanism for familial Alzheimer's disease. Neuron. 2015;85(5):967–81.
Haass C, Selkoe DJ. Soluble protein oligomers in neurodegeneration: lessons from the Alzheimer's amyloid beta-peptide. Nat Rev Mol Cell Biol. 2007;8(2):101–12.
Kim J, Bagyinszky E, Chang YH, Choe G, Choi BO, An SS, Kim S. A novel PSEN1 H163P mutation in a patient with early-onset Alzheimer's disease: clinical, neuroimaging, and neuropathological findings. Neurosci Lett. 2012;530(2):109–14.
Roeber S, Muller-Sarnowski F, Kress J, Edbauer D, Kuhlmann T, Tuttelmann F, Schindler C, Winter P, Arzberger T, Muller U, et al. Three novel presenilin 1 mutations marking the wide spectrum of age at onset and clinical patterns in familial Alzheimer's disease. J Neural Transm (Vienna). 2015;122(12):1715–9.
Sassi C, Guerreiro R, Gibbs R, Ding J, Lupton MK, Troakes C, Lunnon K, Al-Sarraj S, Brown KS, Medway C, et al. Exome sequencing identifies 2 novel presenilin 1 mutations (p.L166V and p.S230R) in British early-onset Alzheimer's disease. Neurobiol Aging. 2014;35(10):2422.e2413–26.
Knight WD, Kennedy J, Mead S, Rossor MN, Beck J, Collinge J, Mummery C. A novel presenilin 1 deletion (p.L166del) associated with early onset familial Alzheimer's disease. Eur J Neurol. 2007;14(7):829–31.
Larner AJ, Doran M. Clinical phenotypic heterogeneity of Alzheimer's disease associated with mutations of the presenilin-1 gene. J Neurol. 2006;253(2):139–58.
Giau VV, Wang MJ, Bagyinszky E, Youn YC, An SSA, Kim S. Novel PSEN1 p.Gly417Ala mutation in a Korean patient with early-onset Alzheimer's disease with parkinsonism. Neurobiol Aging. 2018;72:188.e113–7.
Bagyinszky E, Lee HM, Van Giau V, Koh SB, Jeong JH, An SSA, Kim S. PSEN1 p.Thr116Ile variant in two Korean families with young onset Alzheimer's disease. Int J Mol Sci. 2018;19(9):2604.
Park J, An SSA, Giau VV, Shim K, Youn YC, Bagyinszky E, Kim S. Identification of a novel PSEN1 mutation (Leu232Pro) in a Korean patient with early-onset Alzheimer's disease and a family history of dementia. Neurobiol Aging. 2017;56:212.e211–7.
Pantieri R, Pardini M, Cecconi M, Dagna-Bricarelli F, Vitali A, Piccini A, Russo R, Borghi R, Tabaton M. A novel presenilin 1 L166H mutation in a pseudo-sporadic case of early-onset Alzheimer's disease. Neurol Sci. 2005;26(5):349–50.
Ezquerra M, Carnero C, Blesa R, Oliva R. A novel presenilin 1 mutation (Leu166Arg) associated with early-onset Alzheimer disease. Arch Neurol. 2000;57(4):485–8.
Guo J, Wei J, Liao S, Wang L, Jiang H, Tang B. A novel presenilin 1 mutation (Ser169del) in a Chinese family with early-onset Alzheimer's disease. Neurosci Lett. 2010;468(1):34–7.
Taddei K, Kwok JB, Kril JJ, Halliday GM, Creasey H, Hallupp M, Fisher C, Brooks WS, Chung C, Andrews C, et al. Two novel presenilin-1 mutations (Ser169Leu and Pro436Gln) associated with very early onset Alzheimer's disease. Neuroreport. 1998;9(14):3335–9.
Ezquerra M, Carnero C, Blesa R, Gelpi JL, Ballesta F, Oliva R. A presenilin 1 mutation (Ser169Pro) associated with early-onset AD and myoclonic seizures. Neurology. 1999;52(3):566–70.
Snider BJ, Norton J, Coats MA, Chakraverty S, Hou CE, Jervis R, Lendon CL, Goate AM, McKeel DW Jr, Morris JC. Novel presenilin 1 mutation (S170F) causing Alzheimer disease with Lewy bodies in the third decade of life. Arch Neurol. 2005;62(12):1821–30.
Guerreiro RJ, Baquero M, Blesa R, Boada M, Bras JM, Bullido MJ, Calado A, Crook R, Ferreira C, Frank A, et al. Genetic screening of Alzheimer's disease genes in Iberian and African samples yields novel mutations in presenilins and APP. Neurobiol Aging. 2010;31(5):725–31.
Moulder KL, Snider BJ, Mills SL, Buckles VD, Santacruz AM, Bateman RJ, Morris JC. Dominantly inherited Alzheimer network: facilitating research and clinical trials. Alzheimers Res Ther. 2013;5(5):48.
Van Giau V, An SSA, Bagyinszky E, Kim S. Gene panels and primers for next generation sequencing studies on neurodegenerative disorders. Molecular & Cellular Toxicology. 2015;11(2):89–143.
Källberg M, Wang H, Wang S, Peng J, Wang Z, Lu H, Xu J. Template-based protein structure modeling using the RaptorX web server. Nat Protoc. 2012;7(8):1511–22.
Campion D, Dumanchin C, Hannequin D, Dubois B, Belliard S, Puel M, Thomas-Anterion C, Michon A, Martin C, Charbonnier F, et al. Early-onset autosomal dominant Alzheimer disease: prevalence, genetic heterogeneity, and mutation spectrum. Am J Hum Genet. 1999;65(3):664–70.
Syama A, Sen S, Kota LN, Viswanath B, Purushottam M, Varghese M, Jain S, Panicker MM, Mukherjee O. Mutation burden profile in familial Alzheimer's disease cases from India. Neurobiol Aging. 2018;64:158.e157–13.
Wallon D, Rousseau S, Rovelet-Lecrux A, Quillard-Muraine M, Guyant-Marechal L, Martinaud O, Pariente J, Puel M, Rollin-Sillaire A, Pasquier F, et al. The French series of autosomal dominant early onset Alzheimer's disease cases: mutation spectrum and cerebrospinal fluid biomarkers. J Alzheimers Dis. 2012;30(4):847–56.
Higuchi S, Yoshino A, Matsui T. A novel PS1 mutation (W165G) in a Japanese family with early-onset Alzheimer’s disease. Alzheimers Rep. 2000;3:227–31.
Carecchio M, Picillo M, Valletta L, Elia AE, Haack TB, Cozzolino A, Vitale A, Garavaglia B, Iuso A, Bagella CF, et al. Rare causes of early-onset dystonia-parkinsonism with cognitive impairment: a de novo PSEN-1 mutation. neurogenetics. 2017;18(3):175–8.
Ramirez-Duenas MG, Rogaeva EA, Leal CA, Lin C, Ramirez-Casillas GA, Hernandez-Romo JA, St George-Hyslop PH, Cantu JM. A novel Leu171Pro mutation in presenilin-1 gene in a Mexican family with early onset Alzheimer disease. Ann Genet. 1998;41(3):149–53.
Jin SC, Pastor P, Cooper B, Cervantes S, Benitez BA, Razquin C, Goate A, Cruchaga C. Pooled-DNA sequencing identifies novel causative variants in PSEN1, GRN and MAPT in a clinical early-onset and familial Alzheimer's disease Ibero-American cohort. Alzheimers Res Ther. 2012;4(4):34.
Kasuga K, Ohno T, Ishihara T, Miyashita A, Kuwano R, Onodera O, Nishizawa M, Ikeuchi T. Depression and psychiatric symptoms preceding onset of dementia in a family with early-onset Alzheimer disease with a novel PSEN1 mutation. J Neurol. 2009;256(8):1351–3.
Tiedt HO, Lueschow A, Winter P, Muller U. Previously not recognized deletion in presenilin-1 (p.Leu174del.) in a patient with early-onset familial Alzheimer's disease. Neurosci Lett. 2013;544:115–8.
Tedde A, Nacmias B, Ciantelli M, Forleo P, Cellini E, Bagnoli S, Piccini C, Caffarra P, Ghidoni E, Paganini M, et al. Identification of new Presenilin gene mutations in early-onset familial Alzheimer disease. Arch Neurol. 2003;60(11):1541–4.
Klunemann HH, Rogaeva E, Neumann M, Kretzschmar HA, Kandel M, Toulina A, Sato C, Salehi-Rad S, Pfister K, Klein HE, et al. Novel PS1 mutation in a Bavarian kindred with familial Alzheimer disease. Alzheimer Dis Assoc Disord. 2004;18(4):256–8.
Colacicco AM, Panza F, Basile AM, Solfrizzi V, Capurso C, D’Introno A, Torres F, Capurso S, Cozza S, Flora R, et al. F175S change and a novel polymorphism in Presenilin-1 gene in late-onset familial Alzheimer’s disease. Eur Neurol. 2002;47(4):209–13.
Muller U, Winter P, Graeber MB. A presenilin 1 mutation in the first case of Alzheimer's disease. Lancet Neurol. 2013;12(2):129–30.
Rogaeva EA, Fafel KC, Song YQ, Medeiros H, Sato C, Liang Y, Richard E, Rogaev EI, Frommelt P, Sadovnick AD, et al. Screening for PS1 mutations in a referral-based series of AD cases: 21 novel mutations. Neurology. 2001;57(4):621–5.
Sassi C, Guerreiro R, Gibbs R, Ding J, Lupton MK, Troakes C, Al-Sarraj S, Niblock M, Gallo JM, Adnan J, et al. Investigating the role of rare coding variability in Mendelian dementia genes (APP, PSEN1, PSEN2, GRN, MAPT, and PRNP) in late-onset Alzheimer's disease. Neurobiol Aging. 2014;35(12):2881.e2881–6.
Martin C, Charbonnier F, Flaman J-M, Frebourg T, Moreau V, Campion D, Calenda A, Bellis M, Heilig R, Mallet J, et al. Mutations of the presenilin I gene in families with early-onset Alzheimer's disease. Hum Mol Genet. 1995;4(12):2373–7.
Clark RF, Hutton M, Fuldner M, Froelich S, Karran E, Talbot C, Crook R, Lendon C, Prihar G, He C, et al. The structure of the presenilin 1 (S182) gene and identification of six novel mutations in early onset AD families. Nat Genet. 1995;11(2):219–22.
Moehlmann T, Winkler E, Xia X, Edbauer D, Murrell J, Capell A, Kaether C, Zheng H, Ghetti B, Haass C, et al. Presenilin-1 mutations of leucine 166 equally affect the generation of the notch and APP intracellular domains independent of their effect on Abeta 42 production. Proc Natl Acad Sci U S A. 2002;99(12):8025–30.
Jiao B, Tang B, Liu X, Xu J, Wang Y, Zhou L, Zhang F, Yan X, Zhou Y, Shen L. Mutational analysis in early-onset familial Alzheimer's disease in mainland China. Neurobiol Aging. 2014;35(8):1957.e1951–6.
Sun L, Zhou R, Yang G, Shi Y. Analysis of 138 pathogenic mutations in presenilin-1 on the in vitro production of Abeta42 and Abeta40 peptides by gamma-secretase. Proc Natl Acad Sci U S A. 2017;114(4):E476–e485.
Acknowledgements
Not applicable.
Funding
This work was supported by a National Research Foundation of Korea (NRF) Grants, awarded by the Korean government (MEST, No. 2017R1A2B4012636 & 2017R1C1B5017807).
Dr. An SS receives the National Research Foundation of Korea (NRF) grant funded by the Korea government (MEST, No. 2017R1A2B4012636). Dr. Eva Bagyinszky receives the National Research Foundation of Korea (NRF) grant funded by the Korea government (MEST, No. 2017R1C1B5017807). Dr. Giau VV reports no disclosure. Dr. Pyun JM reports no disclosure. Dr. Suh J reports no disclosure. Dr. Kim SY reports no disclosure.
Author information
Authors and Affiliations
Contributions
VVG and EB: drafting the manuscript for content and doing genetic analysis. EV: preparing the samples and revising the manuscript. JMP and JS: verifying the MRI/PET results. JMP: interpreting the clinical data and revising the manuscript. VVG and EV: predicting presenilin 1 protein structure. SYK: analysis or interpretation of data, doing study supervision, obtaining funding. VVG and SSAA: drafting/revising the manuscript and interpretation of clinical data. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This study was conducted with approval from the Institutional Review Board of Seoul National University College of Medicine & Neurocognitive Behavior Center, Seoul National University Bundang Hospital (B-1302/192–006).
Consent for Publication
Written informed consent was obtained from the patient for publication of this Case report and any accompanying images. A copy of the written consent is available for review by the Editor of this journal.
Competing interests
The authors declare that they have no competing of interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Van Giau, V., Pyun, JM., Suh, J. et al. A pathogenic PSEN1 Trp165Cys mutation associated with early-onset Alzheimer’s disease. BMC Neurol 19, 188 (2019). https://doi.org/10.1186/s12883-019-1419-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12883-019-1419-y