Abstract
Background
The 2017 World Health Organization (WHO) report has listed extended-spectrum β-lactamase-producing Enterobacterales (ESBL-E) as critical pathogens for public health and requiring urgently new antibiotics. The aim of this study was to characterize phenotypically and genotypically ESBL-E isolated among clinical samples in Dschang, Cameroon.
Methods
A cross-sectional study was conducted during a four-month periods from February to May 2022 in the two biggest hospitals of Dschang. Clinical samples were collected and cultured on Eosin Methylene Blue agar. Suspected growing colonies were biochemically identified using the Enterosystem Kit 18R. Antimicrobial susceptibility testing (AST) was done using the Kirby Bauer disc diffusion method and interpretated according to the CA-SFM recommendations. ESBL phenotypes were double screened using CHROMagar™ ESBL and double disk synergy test (DDST). The detection of resistance genes was performed using conventional and multiplex PCR methods. Results were analyzed with SPSS (version 21) and a p-value < 0.05 was considered statistically significant.
Results
A total of 152 Enterobacterales were isolated among 597 clinical samples including urine, blood, cervico-vaginal, urethral swabs and wound samples. The overall prevalence of ESBL-Enterobacterales was 29.61% (45/152). The most represented ESBL species were Escherichia coli (n = 23; 51.11%), Klebsiella pneumoniae (n = 8; 17.78%) and Citrobacter freundii (n = 6; 13.33%).
Conclusion
This study reveals the high burden of ESBL-E among clinical samples in the regional hospital in Dschang with the most common species being E. coli and K. pneumoniae. It confirmed the high occurrence of blaCTX−M and blaTEM among ESBL-E. The study suggests that implementing antimicrobial stewardship program and real-time surveillance of antimicrobial resistance are needed in the Western region of Cameroon. Moreover, the implementation of infection prevention and control measures (IPC) is essential to curb the dissemination of these bacteria from community to hospital settings. Implementation of national action plan to fight against antimicrobial resistance at the local levels is urgently needed.
Similar content being viewed by others
Background
Extended-spectrum β-lactamase (ESBL) and carbapenemase-producing Enterobacterales have been listed by the World Health Organization (WHO) as critical priority pathogens for which new antimicrobials need to be developed [1]. To escape the activities of the majority of β-lactam antibiotics such as penicillin, first, second and third generations of cephalosporin (C3G), Enterobacterales produced ESBL enzymes leading to increase resistance to all β-lactam except cephamycins, carbapenems and monobactams [2]. Escherichia coli and Klebsiella pneumoniae are among the leading species associated with ESBL production [3]. They are responsible for the increased hospitalization length of stay and deaths particularly in Low-and-Middle Income Countries (LMICs) [3,4,5,6,7,8].
Multi-drug resistance is increasingly being detected in numerous clinical Enterobacterales strains because of the extensive antibiotic used in hospital settings [3]. Despite considerable efforts for their containment, ESBL-producing Enterobacterales (ESBL-E) and multi-drug resistant Enterobacterales (MDR-E) are increasingly implicated in several difficult-to-treat infections especially in sub-Saharan African (sSA) countries [3].
Numerous genes encoding for ESBL enzymes were implicated and described among clinical isolates in Uganda, Nepal, Côte d’Ivoire and Chad with the most important genes being blaCTX−M followed by blaTEM and blaSHV [8,9,10,11]. Moreover, a high prevalence of ESBL-E were also observed from carriage among HIV patients during a COVID-19 pick in 2022 in Cameroon [12].
ESBL enzymes have emerged following chromosomal mutation and acquisition of resistance genes carried on diverse mobile genetic elements (MGEs) including plasmids, integrons, insertion sequences, transposons, genomic islands and bacteriophage [11]. The common transferability of resistance amongst bacteria will likely be associated with increasing rates of MDR infections, although some gaps remain as the burden of MDR-E in community as well as hospitalized patients [11]. This study aims at determining the phenotypic and genotypic characteristics of ESBL-E isolated from clinical samples in two hospitals in Dschang, Cameroon post COVID-19 pandemic.
Methods
Study settings
This study was carried out in the two biggest health care structures of the Dschang District in Western region of Cameroon encoded as H1 and H2 for ethical reasons. The H1 is the biggest health care infrastructure in Dschang while H2 is a private Catholic Medical Center with the same level. Clinical samples were collected and analyzed during a four-month periods, from February 1st to May 20th, 2022. All clinical samples originating from community and hospitalized patients were examined in the respective laboratories of these hospitals.
Samples collection and identification
All collected samples except feces were inoculated onto Eosin Methylene Blue (EMB) agar (CM-EMB100, Rapid Labs). All fecal samples were inoculated into Selenite broth and incubated for 18 to 24 h at 37 °C. After incubation, the inoculum was plated onto and Salmonella-Shigella (SS) agar (CM-SSB244, Rapid Labs) and was then incubated for 18 to 24 h at 37 °C for the isolation of Salmonella spp. as well as Shigella spp. For all growing colonies, oxidase test was performed to identify Gram negative fermenting bacteria. Specie identification was performed using the Enterosystem 18R kits following the manufacturer’s recommendations.
Antimicrobial susceptibility testing (AST) and ESBL screening methods
The AST was performed using the Kirby-Bauer disc diffusion method on Muller Hinton agar following the recommendations of the Antibiogram Committee of the French Society of Microbiology (CA-SFM) guidelines. A panel of 11 antibiotic discs of five different families were tested including: amoxicillin and clavulanic acid (AMC; 20 − 10 µg), ceftazidime (CAZ; 30 µg), cefotaxime (CTX; 30 µg), ceftriaxone (CRO; 30 µg), cefixime (CXM; 30 µg), imipenem (IMP; 10 µg), gentamicin (CN; 30 µg), tobramycin (TOB; 10 µg), ciprofloxacin (CIP; 5 µg), levofloxacin (LEV; 5 µg) and chloramphenicol (CHL; 30 µg). A double ESBL-E screening was done using double-disk synergy method and confirmed using ESBL chromogenic agar CHROMagar ESBL (CHROMagar™ Orientation, Paris - France). The samples were then stored at -20 °C in cryotubes containing trypticase soya supplemented with 20% glycerol.
DNA genomic extraction
Extraction of the genomic DNA of all ESBL-E isolates subjected to molecular characterization was done as previously described [12]. During the procedure, two pure colonies of ESBL-E were introduced into 400 µL of a solution of Tris EDTA (10 mM Tris; 0.1 mM EDTA), then the suspension was boiled at 95 °C for 25 min using a dry bath (MIULab DKT200-1, Lasec International Ltd., Johannesburg, South Africa). This was followed by centrifugation of the suspension at 9500 rpm for 5 min. Finally, 150 µL of the supernatant containing the DNA was transferred to an eppendorf tube and frozen at -20 °C for further analysis.
Detection of ESBL-producing Enterobacterales
Conventional and multiplex PCR methods
A 10 µL PCR master mix solution was used for detection of the blaSHV gene among ESBL-E isolates and consisted of 2.8 µl of nuclease-free water, 0.1 µl of each forward and reverse primer, 5 µl of 2x DreamTaq green PCR master mix (Thermo Fisher Scientific, Lithuania) and 2 µl of DNA. Singleplex PCR was performed using BIO-RAD T100 thermal cycler (Bio-Rad Laboratories, Marnes-la-Coquette, France). The amplification steps were as follows: initial denaturation (94 °C for 30 s), 30 cycles of denaturation at 94 °C for 4 s, elongation at 72 °C for 50 s and final elongation at 72 °C for 5 min.
For the detection of the blaTEM and blaCTX−M genes among the ESBL-E, the reaction occurred in a 10 µl reaction mix with a volume consisting of 5 µL of 2x DreamTaq green PCR Master mix (Thermo Fisher Scientific, Lithuania), 2.6 µL of nuclease-free water, 0.1 µL of each reverse and forward primer and 2 µL of template DNA. The amplification steps for all reactions were as follows: initial denaturation (94 °C for 30 s), 30 cycles of denaturation at 94 °C for 4 s, annealing for 40 s, elongation at 72 °C for 50 s and final elongation at 72 °C for 5 min [13, 14].
PCR products and visualization
Revelation of amplicons was done along with a 100 bp molecular ladder (New England Biolabs, MA, USA) in a 1.5% (w/v) agarose gel electrophoresis. Migration was set at 90 V for 45 min followed by staining with ethidium bromide solution (0.5 µg/mL) for 15 min and rapid destaining in water. The visualization was made under ultraviolet radiation using a G-BOX chemi XL gel documentation system (Syngene, Cambridge, UK).
Data analysis
Socio-demographic and clinical data were entered into Epi Info version 7.2.5.0 and exported to SPSSv21 for analysis. Pearson Chi-square test was used to assess for any differences between the two ESBL phenotype categories with respect to clinical and demographic parameters. The different data, were compared using the independent t-test. A p-value < 0.05 was considered statistically significant.
Results
Socio-demographic characteristics of patients
A total of 597 consecutive clinical samples from patients attended to hospitals was collected from the laboratories of both hospitals H1 and H2 while 124 samples were positive with at least one Enterobacterales with a total of 154 Enterobacterales identified (Fig. 1). Most of the study participants were females, 83.25% (497/597), whereas the age group 20–30 years old (n = 280/597; 46.90%) was most frequent. Overall, 20.77% (n = 124/597) of participants had a culture positive to Enterobacterales, with 29.84% (n = 37/124) of these being infected by an ESBL-E isolates and 76.61% (n = 95/124) by MDR-E. The majority of patients infected by Enterobacterales were living in urban area (n = 101/124; 81.45%) comparing to rural area (18/124; 14.52%) with a significant difference (P < 0.0001).
A total of 75.88% (453/597) and 24.12% (144/597) samples were collected in H1 and H2, respectively. We observed 63.71% (79/124) and 36.29% (45/124) samples positives to Enterobacterales culture in H1 and H2, respectively. The 63.2% (60/95) and 36.8% (35/95) of patients infected by MDR-Enterobacterales were coming from H1 and H2, respectively with a significant difference (P < 0.05). Concerning ESBL-E, 70.3% (26/37) and 29.7% (11/37) of patients infected were coming from H1 and H2, respectively. The age of the participants was between 1 and 95 years as illustrated in Table 1.
Frequency of Enterobacterales isolates
Out of these 597 samples analyzed, 360 (60,30%) were culture positive to bacteria of which 124 (20.77%) were positive to one or more Enterobacterales. Out of the Enterobacterales positive samples, 152 Enterobacterales isolates were identified. Escherichia coli (n = 67; 44.07%), Klebsiella pneumoniae (n = 34; 22.36%), Citrobacter freundii (n = 16; 10.52%) and Serratia liquefaciens (n = 10; 06.57%) were the leading species. The isolates were identified from a various clinical specimens collected: endocervical swabs (n = 75; 49.34%), stool (n = 61; 40.13%) and other samples (n = 16; 10.52%) as illustrated in Table 2. E. coli was mostly detected among endocervical swabs (n = 41/67; 61.19%) and stool (n = 19/67; 28.36%) followed by K. pneumoniae among vaginal swabs (n = 20/34; 58.82%) and stool (n = 8/34; 23.53%) respectively, while C. freundii was only isolated from stool (n = 14/14; 100%). Among all identified Enterobacterales, the majority (122/152; 80.26%) were coming from patients living in urban area. Concerning hospitals, majority of isolates were coming from H1 (93/152; 61.18%) and (59/152; 38.22%) from H2 as illustrated in Table 2.
Occurrence of ESBL-producing Enterobacterales
Out of the 152 non-duplicated Enterobacterales identified, 45 (29.60%) were ESBL producers. The most common ESBL species were E. coli (51.11%; 23/45), K. pneumoniae (17.77; 8/45) and C. freundii (13.33%; 6/45) as described in the Table 3. The proportion of ESBL-E was higher in H1 (71.11%; 32/45) than H2 although without statistical significance (71.11% vs. 28.29%; 13/45) (p = 0.1811). ESBL-E were frequently isolated from vaginal swabs (44.44%; n = 20/45), fecal samples (42.22%; n = 19/45) and urine (6.67%; n = 3/45) (Table 2).
Antibiotic resistant profiles of Enterobacterales and ESBL-E
The antibiotic resistance profiles of Enterobacterales species shows a high level of resistance to amoxicillin-clavulanic acid (139/152; 91.45%), imipenem (95/152; 62.5%), ceftriaxone (81/152; 53.29%) and ceftazidime (76/152; 50%) (Table 4). E. coli showed the highest resistance rates to amoxicillin-clavulanic acid (92.54%), imipenem (61.19%), ceftriaxone (58.21%), ceftazidime (56.72%), levofloxacin (50.75%), ciprofloxacin (47.76%), chloramphenicol (22,39%) and gentamicin (37.31%). K. pneumoniae were mainly resistant to amoxicillin-clavulanic acid (94.18%), imipenem (67.68%) and tobramycin (50%). However, a low resistance level was observed to gentamicin (32.35%) and chloramphenicol (32.35%).
Concerning ESBL-E, a significant level of resistance to non β-lactam antibiotics including ciprofloxacin (55.56%), levofloxacin (46.67%) and gentamicin (24.44%) was detected (Table 3; Fig. 2).
Distribution of multidrug-resistant Enterobacterales
A total of 114 (75%) isolates were MDR with E. coli (n = 52/114; 45.61%) and K. pneumoniae (n = 25/114; 21.93%) being the most prevalent MDR species (Table 5; Fig. 3). In addition, all P. mirabilis (2.63% n = 4/152) were MDR (Table 5). The vaginal samples were more positive to MDR-Enterobacterales (n = 59/114; 51,75%) (Table 4). The majority of MDR-E were resistant to three (n = 39/114; 25.66%) and four (n = 35/114; 23.03%) antibiotic families (Table 5).
Among 15 different resistance profiles of MDR-Enterobacterales, five isolates (n = 5/114; 4.38%) were resistant to nine antibiotics (AMC-CTR-CRX-IMP-CIP-LEV-CN-TOB-CHL) belonging to six different families of antibiotics. Moreover, three isolates (n = 3; 1.97%) were resistant to 11 antibiotics (AMC-CTX-CAZ-CTR-CRX-IMP-CIP-LEV-CN-TOB-CHL) from six different families of antibiotics (Fig. 4).
Characterization of ß-lactamase encoding genes
The results of the molecular characterization shown that blaCTX−M (n = 38/45; 84.44%) was the leading ß-lactamase resistance gene, followed by blaTEM (n = 45; 73.33%) and blaSHV (n = 11; 24.44%) (Fig. 5; Table 6). Interestingly 64.44% (29/45) of ESBL-E isolates carried more than one gene, some isolates carried concomitantly the three genes (n = 10/45; 22.22%). It was reported that 100% (8/8) of ESBL-producing K. pneumoniae carried the gene blaCTX−M (n = 8/8; 100%) and 75% (6/8) of them carried the three genes (Table 6).
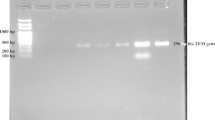
Agarose gel electrophoresis of PCR products (blaCTX-M, blaTEM and blaSHV) of some ESBL-producing Enterobacterales isolated from clinical samples
Ladder: 100 bp DNA ladder; PC: positive control; NC: negative control; 1 & 2: CTX-M positive, TEM positive; 3: CTX-M positive, TEM negative; 4 : CTX-M negative, TEM positive ; 5 & 6 : SHV positive ; 7 : SHV negative
Discussion
Extended-spectrum β-lactamase-producing Enterobacterales remain a public health threat with important societal and economic repercussions as demonstrated [15]. In this study, the prevalence of ESBL-E among clinical samples was 29.61% (n = 45/152) in the two highest hospital settings in Dschang district. These results could be explained by the fact that the H1 hospital is in the process of urbanization with the multiplication of street medicine and access of population to self-medication. In addition, this result is higher than those obtained in Central African Republic where the evaluation of increasing prevalence of antimicrobial resistance among ESBL-Enterobacteriaceae uropathogens in Bangui shows a prevalence of 19.3% in 2006 [16]. This could also be explained by the fact that they have only worked on urine samples compared to this study where all clinical samples were included. This result is lower than a report observed in Nepal reporting a prevalence (39.13%) of CTX-M β-lactamases producing multi-drug resistant Escherichia coli and Klebsiella pneumoniae among patients attending Bir Hospital [10]. This difference could be explained by the fact that they have worked on the clinical samples and have selected only MDR-Enterobacterales and processed for further ESBL confirmation. Our result is also in contrast with the study conducted in Ghana where the prevalence was (49.3%) and this very high prevalence could be justified by the fact that ESBL-E was an issue already identified and the lack of awareness on misuse and overuse of antibiotics have been previously described [17].
The female (79.84%) was more frequently positive to ESBL-E than male (20.16%). This finding could be explained because endocervical swabs were the most samples collected and the majority of Enterobacterales were isolated from this specimen. In addition, these bacteria are commonly observed in the intestinal tract and can colonize the vaginal tract among women given the proximity of the vagina and rectum in women as described by a report conducted by Bercion et al. in Central African Republic were 60% of Enterobacteriaceae were isolated from woman [16].
Similarly, endocervical swabs (20/45; 44.44%) were the clinical samples most positive to ESBL-E. This finding is in contrast with a study conducted in Sudan in 2020, where most of the ESBL-E isolates were detected from urine samples (44%, n = 75) and wound swabs (44%, n = 75) respectively [18]. This could be explained by the fact that female was most represented in the study and observed in this region. Moreover, the lack of water, poor hygienic conditions combined with consumption of counterfeit antibiotics, poor education level could contribute to the self-contamination [9].
This study reveals the predominance of E. coli (n = 67; 44.08%), K. pneumoniae (n = 34; 24.34%) and C. freundii (n = 14; 9.21%). These results are similar to those reported in Uganda, during a study carried out in Mulango hospital concerning clinical samples in which they identify E. coli as the most organism (53.9%), followed by K. pneumoniae (28.7%) [8].
This is in agreement with a study conducted in Ethiopia by Teklu et al. in 2017, where Enterobacterales especially E. coli (n = 228; 53.5%) and K. pneumoniae (n = 103; 24.1%) were the most bacteria isolated among clinical samples [6]. In addition, E. coli (n = 23; 34.33%) and K. pneumoniae (n = 8; 23.53%) were the leading ESBL producers. Our findings are similar to several studies investigated worldwide especially in Africa countries like Burkina Faso, where a study shown that ESBL-E were mostly represented among clinical samples with E. coli (67.5%) and K. pneumoniae (26%) [19], in Ethiopia where E. coli (52.2%) and K. pneumoniae (78.6%) were the most detected ESBL-E in clinical samples collected from hospitals in Addis Ababa [6], and in study reported in Uganda where K. pneumonia had the highest rate of ESBL producers (72.7%) among clinical samples in Mulago hospital after a cross-sectional study in 2014 [8].
ESBL-E showed the high resistance rates of more than 60% to all ß-lactams family including penicillin, cephalosporins and carbapenems. This result is similar to those obtained in Nigeria in 2016, where high resistance rate was observed among ESBL-E isolates with the high resistance profile with ceftriaxone (92.3%), aztreonam (96.8%), cefpodoxime (96.3%), cefotaxime (98.8%) and trimethoprim/sulfamethoxazole (90%) [20]. This high resistance level to ß-lactams family of ESBL-E has been already described by Teklu et al. in Ethiopia with ceftazidime (100%) and ceftriaxone (100%) [6].This could be explained by the extensive and unregulated use of β-lactam antibiotics as a first-line medication to treat infectious diseases in Africa [9, 10]. Numerous reports have already demonstrated that in African countries, hospitalized patients received numerous antibiotics for the therapeutic purpose without antibiotic susceptibility testing, which contribute to the selective pressure on the microbiome and increase antimicrobial resistance [21]. High level of MDR-E. coli and MDR-K. pneumoniae have been observed and this could be explained by the excessive and inappropriate use of antibiotics in Dschang, where antibiotics are easily accessible over the counter without a prescription [22].
ESBL-E. coli and ESBL-K. pneumoniae are commonly responsible of hospital-and community-acquired infections [18]. The blaCTX−M and blaTEM were the most frequent ß-lactamase genes detected among ESBL-E. coli and ESBL-K. pneumoniae with prevalence ranging from 74 to 100%. These findings are in agreement with those obtained in Nepal where the prevalence of blaCTX−M among clinical ESBL-E. coli and ESBL-K. pneumoniae were 93.81% and 78.94% isolated from clinical samples respectively [10]. This finding also agreed with a study conducted in Egypt where blaCTX−M was the most common ESBL genes among ESBL-producing E. coli with a prevalence of 83.3% [21]. These results could be explained by the powerful ability of this gene to hydrolyze numerous β-lactam antibiotics including ceſtazidime, cefotaxime and aztreonam, which probably offers a selective advantage when they are overuse or misuse concomitantly. Moreover, it is plausible because the third generation of cephalosporins including ceftriaxone and cefixime are extensively used as a first line treatment among hospitalized and community patients having vaginal, urinary tract and bloodstream infections in the H1 and H2 hospitals, this could contribute to the expansion of blaCTX−M in the region [23]. However, the relatively low prevalence of blaSHV has already been reported in Africa countries especially in Chad where there had no blaSHV gene among clinical samples [9]. This result agrees with the results obtained in numerous studies conducted during the last ten years in LMIC countries and confirm that blaSHV is relatively infrequent in Africa [2].
Limits
The limitations of our study are the unable detect the genes responsible for carbapenem resistance, or to detect other resistance gene coding for β-lactam resistance. In addition, this study was limited to two health facilities in the Dschang district, it would be interesting to extend the study to the whole of Cameroon to obtain more relevant results.
Conclusion
This study reveals the high burden of MDR-E and ESBL-E among clinical samples in the regional hospital in Dschang with the most common species being E. coli and K. pneumoniae. It confirmed the high occurrence of blaCTX−M and blaTEM among ESBL-E. The study suggests that implementing antimicrobial stewardship program and real-time surveillance of antimicrobial resistance are needed in the Western region of Cameroon. Moreover, the implementation of infection prevention and control measures (IPC) is essential to curb the dissemination of these bacteria from community to hospital settings. It will be also important to reduce the burden and subsequent morbidity and mortality rates associated with antimicrobial resistance in the country by implementing national action plan to fight against antimicrobial resistance at the local levels.
Data Availability
The data are available upon request in accordance with confidentiality and privacy regulations from the corresponding author.
Abbreviations
- MDR:
-
multidrug resistance
- ESBL:
-
Extended Spectrum β-lactamase
- ESBL-E:
-
Extended Spectrum β-lactamase producing-Enterobacterales
References
OMS. L’OMS publie une liste de bactéries contre lesquelles il est urgent d’avoir de nouveaux antibiotiques. 2017. p. 1–3.
Ouchar MO, Kempf M, Lounnas M, Tidjani A, Hide M, Benavides JA, et al. Epidemiology and prevalence of extended-spectrum β-lactamase- and carbapenemase-producing Enterobacteriaceae in humans, animals and the environment in West and Central Africa. Int J Antimicrob Agents. 2021;57(1):106203.
Murray CJ, Ikuta KS, Sharara F, Swetschinski L, Robles Aguilar G, Gray A, et al. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet. 2022;399(10325):629–55.
Vora S, Auckenthaler R. Que signifie «bêtalactamases à Spectre élargi» en pratique? Maladies infectieuses. Rev Med Suisse. 2009;5(220):1991–4.
Founou LL, Founou RC, Allam M, Ismail A, Djoko CF, Essack SY. Genome sequencing of extended-spectrum β-lactamase (ESBL)-producing Klebsiella pneumoniae isolated from pigs and abattoir workers in Cameroon. Front Microbiol. 2018;9:1–12.
Teklu DS, Negeri AA, Legese MH, Bedada TL, Woldemariam HK, Tullu KD. Extended-spectrum beta-lactamase production and multi-drug resistance among Enterobacteriaceae isolated in Addis Ababa, Ethiopia. Antimicrob Resist Infect Control. 2019;8(1):1–12.
Pandit R, Awal B, Shrestha SS, Joshi G, Rijal BP, Parajuli NP. Extended-Spectrum β -Lactamase (ESBL) Genotypes among Multidrug-Resistant Uropathogenic Escherichia coli Clinical Isolates from a Teaching Hospital of Nepal. Interdiscip Perspect Infect Dis. 2020;2020.
Kateregga JN, Kantume R, Atuhaire C, Lubowa MN, Ndukui JG. Phenotypic expression and prevalence of ESBL-producing Enterobacteriaceae in samples collected from patients in various wards of Mulago Hospital, Uganda. BMC Pharmacol Toxicol. 2015;16:14.
Ouchar MO, Lounnas M, Hide M, Dumont Y, Tidjani A, Kamougam K, et al. High prevalence and characterization of extended-spectrum ß-lactamase producing Enterobacteriaceae in Chadian hospitals. BMC Infect Dis. 2019;19(1):205.
Koirala S, Khadka S, Sapkota S, Sharma S, Khanal S, Thapa A et al. Prevalence of CTX-M β -Lactamases Producing Multidrug Resistant Escherichia coli and Klebsiella pneumoniae among Patients Attending Bir Hospital, Nepal. Biomed Res Int. 2021;2021.
Onduru OG, Mkakosya RS, Aboud S, Rumisha SF. Genetic determinants of resistance among ESBL-Producing Enterobacteriaceae in Community and Hospital settings in East, Central, and Southern Africa: a systematic review and Meta-analysis of prevalence. Can J Infect Dis Med Microbiol. 2021;2021:5153237.
Zemtsa RJ, Noubom M, Founou LL, Dimani BD, Koudoum PL, Mbossi AD et al. ESBL - Producing Enterobacterales Isolated from Carriage Samples among HIV Infected Women in Yaoundé, Cameroon. 2022;1–13.
Strauß LM, Dahms C, Becker K, Kramer A, Kaase M, Mellmann A. Development and evaluation of a novel universal β-lactamase gene subtyping assay for blaSHV, blaTEM and blaCTX-M using clinical and livestock-associated Escherichia coli. J Antimicrob Chemother. 2015;70(3):710–5.
Batchelor M, Hopkins K, Threlfall EJ, Clifton-Hadley FA, Stallwood AD, Davies RH, et al. Bla(CTX-M) genes in clinical Salmonella isolates recovered from humans in England and Wales from 1992 to 2003. Antimicrob Agents Chemother. 2005;49(4):1319–22.
Founou RC, Founou LL, Essack SY. Clinical and economic impact of antibiotic resistance in developing countries: a systematic review and meta-analysis. PLoS ONE. 2017;12:e0189621.
Bercion R, Mossoro-Kpinde D, Manirakiza A, Le Faou A. Increasing prevalence of antimicrobial resistance among Enterobacteriaceae uropathogens in Bangui, Central African Republic. J Infect Dev Ctries. 2009;3(3):187–90.
Obeng-Nkrumah N, Twum-Danso K, Krogfelt KA, Newman MJ. High levels of extended-spectrum beta-lactamases in a major teaching hospital in Ghana: the need for regular monitoring and evaluation of antibiotic resistance. Am J Trop Med Hyg. 2013;89(5):960–4.
Dirar M, Bilal N, Ibrahim ME, Hamid M. Resistance patterns and phenotypic detection of β-lactamase enzymes among Enterobacteriaceae isolates from Referral hospitals in Khartoum State, Sudan. Cureus. 2020;12(3):7260.
Ouedraogo A-S, Sanou M, Kissou A, Sanou S, Solaré H, Kaboré F, et al. High prevalence of extended-spectrum ß-lactamase producing enterobacteriaceae among clinical isolates in Burkina Faso. BMC Infect Dis. 2016;16:326.
Duru C, Olanipekun G, Odili V, Kocmich N, Rezac A, Ajose TO, et al. Molecular characterization of invasive Enterobacteriaceae from pediatric patients in Central and Northwestern Nigeria. PLoS ONE. 2020;15(10):e0230037.
Ramadan H, Soliman AM, Hiott LM, Elbediwi M, Woodley TA, Chattaway MA, et al. Emergence of Multidrug-Resistant Escherichia coli Producing CTX-M, MCR-1, and FosA in Retail Food from Egypt. Front Cell Infect Microbiol. 2021;11:681588.
Mba B, Ars F, Cl KT, Ang N. Appréhention Du Risque Et Perception Par Les consmmateurs: Cas Des Médicaments Dans La Ville De Dschang-Cameroun. Glob J Manag Bus Res B Econ Commer. 2014;14(7).
Müller-Schulte E, Tuo MN, Akoua-Koffi C, Schaumburg F, Becker SL. High prevalence of ESBL-producing Klebsiella pneumoniae in clinical samples from central Côte d’Ivoire. Int J Infect Dis IJID off Publ Int Soc Infect Dis. 2020;91:207–9.
Acknowledgements
We thank all staff members of two healthcare structures and all participants who agreed to participate to this study.
Funding
Raspail Carrel Founou received funding from the Mérieux Institute, Lyon France for the CAREFOOD project. CAREFOOD project had supported all the molecular aspect of this study. This work was also supported by the Research Institute of Centre of Expertise and Biological Diagnostic of Cameroon (CEDBCAM-RI). The funders had no role in the study design, nor the decision to submit the work for publication. Any opinions, findings and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the organizations or agencies that provided support for the project.
Author information
Authors and Affiliations
Contributions
Conceptualization: OAN, RCF and MN; Methodology: RCF, LLF; Software: OAN; Validation: RCF, LLF and MN; Formal analysis: OAN, LLF, RCF; Investigation: OAN, BDD, PLK, AM, JRZ, CSM, LTT; Resources: RCF, LLF and MN; Writing original draft: OAN and RCF; Review and editing: OAN, RCF and LLF; Visualization: OAN, RCF; Supervision: RCF, LLF and MN; Project administration: RCF and LLF.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This research has been approved by the Regional Ethics Committee for Health Research for the Western region (N° 2023/01/009/CE/CRERSH-OU/VP).The study was approved by the Centre of Expertise and Biological Diagnostic of Cameroon (CEDBCAM-RI) under the number (N° 001/02/22/LA/CEDBCAM-RI/DG). The study was conducted in accordance with the declaration of Helsinki. In addition, the research authorizations of the various healthcare structures have been granted. All methods and protocols used were approved by the CEDBCAM-RI in accordance with the relevant international guidelines and regulations for research laboratory ethics. Informed written consent was provided by the caregivers or the child’s legal guardian in their preferred language (English or/and French) for the child to participate in this study.
Consent for publication
Not applicable.
Competing of interest
The authors declare no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Nkengkana, O.A., Founou, R.C., Founou, L.L. et al. Phenotypic and genotypic characterization of multidrug resistant and extended-spectrum β-lactamase-producing Enterobacterales isolated from clinical samples in the western region in Cameroon. BMC Infect Dis 23, 819 (2023). https://doi.org/10.1186/s12879-023-08742-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12879-023-08742-7