Abstract
Background
Chronic wounds are frequently colonized or infected with multiple bacterial or fungal species, which can both promote or inhibit each other. Network analyses are helpful to understand the interplay of these species in polymicrobial infections. Our aim was to analyse the network of bacterial and fungal species in chronic wounds.
Methods
Swabs (n = 163) from chronic wound infections (Masanga, Sierra Leone, 2019–2020) were screened for bacterial and fungal species using non-selective agars. Some of these wounds were suspected but not confirmed Buruli ulcer. Species identification was done with MALDI-TOF mass spectrometry. Network analysis was performed to investigate co-occurrence of different species within one patient. All species with n ≥ 10 isolates were taken into account.
Results
Of the 163 patients, 156 had a positive wound culture (median of three different species per patient; range 1–7). Pseudomonas aeruginosa (n = 75) was the dominating species with frequent co-detections of Klebsiella pneumoniae (21 cases; OR = 1.36, 95%CI: 0.63–2.96, p = 0.47), Staphylococcus aureus (14 cases; OR = 1.06, 95%CI: 0.44–2.55, p = 1) and Proteus mirabilis (13 cases; OR = 0.84, 95%CI: 0.35–1.99, p = 0.69).
Conclusion
The culturome of chronic wounds in Sierra Leonean patients is highly diverse and characterized by the co-occurrence of P. aeruginosa, K. pneumoniae and S. aureus.
Similar content being viewed by others
Background
Chronic wounds are common in underserved populations with limited access to healthcare and are frequently colonized or infected with polymicrobial bacterial communities. The composition of the wound microbiome is of clinical relevance as the presence of certain species (e.g. Enterobacter) or a stable microbiota community in diabetic foot ulcera were predictive for delayed healing [1, 2].
The interaction of multiple species within the wound is complex as they affect each other by the secretion of molecules. These molecules can alter the tolerance to antibiotics, virulence or biofilm formation as shown for Pseudomonas aeruginosa and Staphylococcus aureus [3]. The S. aureus protein toxin Panton-Valentine leukocidin (PVL) is of particular interest in resource limited settings as it is associated with severe skin and soft tissue infections (SSTI) and much more common among S. aureus from sub-Saharan Africa (up to 74%) compared to Europe (1.4%) [4,5,6]. One in vitro study suggests that PVL could be a competitive advantage for S. aureus in the interplay with P. aeruginosa, but this finding has not been yet confirmed in vivo [7].
In the past years, network-based analytical approaches were developed to study the structure and interaction of polymicrobial communities [8]. These network analyses make use of mathematical modelling to identify patterns within bacterial communities. While first results on microbial networks are available from chronic wounds in developed countries, this information is absent for resource-limited settings. Results from high-income countries should not be extrapolated to low- and middle-income countries as they differ in host-related factors (e.g. malnutrition, HIV-infection), pathogens (e.g. Mycobacterium ulcerans, Blastomyces, Coccidioides), microbiota, and the environment (e.g. humidity, sanitary system, abundance of flies) [9,10,11]. We therefore performed a network analysis (i) to characterize the bacterial community of chronic wounds in patients from Sierra Leone and (ii) to test if PVL-positive S. aureus is associated with the absence of specific species (e.g. P. aeruginosa).
Methods
Study population
The study made use of an already existing database of bacterial species from chronic wounds in patients (n = 163) from Masanga, Sierra Leone (July 2019–November 2020) [12]. These wounds showed characteristics of Buruli ulcer and could be secondary infections to Mycobacterium ulcerans [12].
In brief, patients with any kind of wounds (e.g. wound originating from trauma, infection, burn or drug reaction) and an informed consent to participate were included. If a person was not of legal age, the guardian gave the informed consent. Patients with closed wounds (e.g. from blunt trauma, haematomas) and surgical site infections were excluded.
Microbiology
After cleaning the wound from bandages, traditional leaves or necrotic skin with a sterile cotton gauze, one swab (Transswab, MWE, Corsham, England) per patient was taken applying slight pressure from both the (undermined) edges and the central areas of the wound (Essen Rotary technique) [13]. Samples were stored in Amies transport medium at 2–7 °C until shipment to Germany (median time between sampling and culture: 3.8 months). Details on the culture conditions are described elsewhere [12]. MALDI-TOF mass spectrometry (Microflex Bruker, Bremen, Germany) and the MBT Compass software (version 4.1.80, Bruker) were used for species identification. All S. aureus isolates were screened for PVL using a commercial test kit (eazyplex® MRSAplus, Amplex, Gars-Bahnhof, Germany).
Network analysis
Network analysis was performed using R 4.2.1 [14]. The R package “igraph” was used for visualization [15]. Interaction networks were generated adapting an example published by Varghese et al., analysing co-occurring and persistent symptoms in COVID-19 [16]. In brief, every species is represented by a node. The size of each node is correlated with the number of observations (precise numbers are additionally reported). Intersects, i.e. co-occurring species within one patient, were visualized by edges. The thickness of an edge corresponds to the number of observations; precise numbers are additionally reported for intersects ≥ 10. A missing edge between two nodes indicates that co-occurrence was never observed for these two species. To reduce the complexity of the network and improve visualization, only species detected in ≥ 10 patients were considered.
Heatmaps, visualizing all species detected and their co-occurrence, were generated using R 4.2.1 and R package “pheatmap” [17]. For S. aureus, we differentiate between PVL-positive (PVL +) and PVL-negative (PVL -) isolates.
Cross-streak assay
To assess the interaction of S. aureus and P. aeruginosa, we performed a cross-streak assay with overnight cultures [7, 18]. One loop tip (ca. 1 µl) of P. aeruginosa was horizontally streaked on a Columbia blood agar plate. An identical amount of one S. aureus colony was streaked vertically crossing the P. aeruginosa streak in the middle of the plate. After incubation (18–24 h, ambient air, 35 ± 1 °C), the growth of S. aureus was scored (0–2 points: 0 = full inhibition by P. aeruginosa, 1 = interruption of the S. aureus streak at the crossing with the P. aeruginosa streak, 2 = no interruption of the S. aureus streak). The scores of the independent tests were added to quantify the S. aureus growth performance (S. aureus growth score).
Statistics
The diversity indices (Shannon, Simpson) were calculated with “R” as implemented in the package “abdiv”. Statistical testing, applying Fisher’s Exact Test, was performed using R and the base function fisher.test (alternative: two.sided). The S. aureus growth scores were compared with the Wilcoxon rank sum test with continuity correction.
Results
Of the 163 eligible patients (11% females, median age of 40 years, range: 0–88), 156 had a positive wound culture and were entered into the final analysis. The majority of wounds were located at the lower limb (85.9%, n = 140) and had a median diameter of 10 cm. Wounds were superficial (84%, n = 137), 53% of them had deep edges (n = 86) [12]. Results from antimicrobial susceptibility testing of these isolates was already reported elsewhere [12].
Three different species per patient (median, range 1–7) were detected, with a total number of 60 different species and 461 isolates in the whole dataset. The most common species were P. aeruginosa (n = 75/461, 16.3%), Klebsiella pneumoniae (n = 42/461, 9.1%), Proteus mirabilis (n = 31/461, 6.7%) and S. aureus (n = 30/461, 6.5%). Of all S. aureus, 43% (n = 13) were methicillin-resistant (mecA positive).
The only fungi were Candida tropicalis (n = 6/461, 1.3%), Candida orthopsilosis (n = 2/461, 0.4%) and Candida krusei (n = 1/461, 0.2%). Anaerobes were rarely detected: Bacteroides fragilis (n = 4/461, 0.9%) and Bacteroides thetaiotaomicron (n = 2/461, 0.4%). All wounds together had a Shannon index of 3.29 and a Simpson index of 0.94.
We performed a network analysis to identify those species that frequently co-occur in individual wounds (Fig. 1). P. aeruginosa was the dominating species with frequent co-detections of K. pneumoniae (21 cases; OR = 1.36, 95%CI: 0.63–2.96, p = 0.47), S. aureus (14 cases; OR = 1.06, 95%CI: 0.44–2.55, p = 1) and P. mirabilis (13 cases; OR = 0.84, 95%CI: 0.35–1.99, p = 0.69). Although the network analysis suggests a triangle pattern of P. aeruginosa, K. pneumoniae and S. aureus (Fig. 1), the analysis of co-occurrence in individual patients reveals that only six patients carried all three species out of 30 being colonized with S. aureus (Supplementary Fig. 1).
Network analyses of bacteria from chronic wound infections, Sierra Leone. Only species with a total number of ≥ 10 isolates were included. Node size and node number correspond to the number of isolates of the respective species. Thickness of grey edges corresponds to the co-occurrence of the displayed species
The PVL protein toxin was detected in 23% (n = 7/30) of all S. aureus isolates.
We tested our hypothesis that the presence of PVL-positive S. aureus is associated with the presence of certain species. We did not find any significant associations for P. aeruginosa (OR = 0.38, 95%CI: 0.03–2.93, p = 0.40), P. mirabilis (OR = 2.57, 95%CI: 0.17–29.79, p = 0.57), S. dysgalactiae (OR = 0, 95%CI: 0–3.65, p = 0.30) or K. pneumoniae (OR = 0.63, 95%CI: 0.05–4.99, p = 1).
Due to the well-described co-detection of P. aeruginosa and S. aureus (irrespective of PVL), we performed a cross-streak assay with a P. aeruginosa isolate from this study (0093, co-isolated from a wound with a PVL-positive S. aureus) and a standard P. aeruginosa strain (ATCC 27,853). These were co-cultured with seven PVL-positive and seven PVL-negative S. aureus isolates from this study. The median S. aureus growth scores were comparable between PVL-positive and PVL-negative isolates (using both 0093 and ATCC 27,853 as competitors, 2.5 vs. 3, p = 0.87, Fig. 2). S. aureus growth strongly depended on the P. aeruginosa competitor: the median S. aureus growth score was significantly higher with the standard strain (ATCC 27,853) compared to the clinical isolate of this study (4 vs. 2, p = 0.0001, Fig. 2).
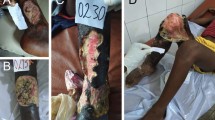
Cross-streak assay with Pseudomonas aeruginosa and Staphylococcus aureus. Two different P. aeruginosa strains (0093, ATCC 27,853, horizontal) were cross-streaked with seven PVL- positive and seven PVL-negative S. aureus from this study (A-C). The growth of S. aureus was scored 0 points (full inhibition by P. aeruginosa, A), 1 point (interruption of the S. aureus streak at the crossing with the P. aeruginosa streak, B) and 2 points (no interruption of the S. aureus streak, C). The scores of three independent tests were added to calculate the S. aureus growth score (min. 0, max. 6). The growth scores were comparable between PVL-positive and PVL-negative S. aureus isolates (D). However, the growth was significantly dependent on the competitor P. aeruginosa strain (E)
Discussion
The main findings of our study were a frequent co-detection of P. aeruginosa and S. aureus and K. pneumoniae in chronic wounds.
The co-detection of P. aeruginosa and S. aureus in chronic infections (e.g. chronic wounds, cystic fibrosis) is well known and reflects a multi-layered interaction in which both species can inhibit (up-regulation of virulence factors) and promote each other (fitness gain, antimicrobial resistance, inhibition of opsonization) [19, 20]. To the best of our knowledge, the impact of PVL on the co-existence has not yet been investigated in detail. In vitro co-culture studies suggest that PVL confers a growth advantage for S. aureus as shown in a cross streak assay with P. aeruginosa wild type and S. aureus lukS-PV mutants [7]. PVL consists of the two subunits lukF-PV and lukS-PV, and the leukocidin is only active if both subunits are present [21]. If a competitive success of PVL-positive S. aureus over P. aeruginosa contributes to the widespread of PVL-positive isolates in Africa is part of current discussions. We did not detect an association of PVL with the absence of P. aeruginosa most likely due to the small sample size of PVL-positive S. aureus. Similarly, in the cross-streak assay (Fig. 2) we did not find any evidence that PVL-positive S. aureus inhibits the growth of P. aeruginosa. In contrast, the growth of S. aureus rather depended on the P. aeruginosa strain with a stronger inhibition of S. aureus by the clinical 0093 isolate compared to the ATCC 27,853 standard strain. This observation is in line with the study by Michelsen et al. who showed that inhibition of S. aureus by P. aeruginosa varies, and is strongest in less human-adapted isolates (e.g. early isolates of chronic infections) [18].
Our study has limitations. First, the long time span between sampling and culture certainly hampered the detection of fastidious bacteria. The low detection of anaerobic bacteria in our collection could suggest that we might underestimated fastidious genera. However, we rate the impact of this bias as low as a similar study that performed culture shortly after sampling in Ghana showed a comparable bacterial spectrum (i.e. predominance of S. aureus, P. aeruginosa or Proteus) in untreated Buruli ulcer cases [22]. In addition, (meta-) genomic profiling would have provided a more detailed picture of the wound microbiome [23]. Second, the aetiology of chronic wound infections in our study is not fully understood. It is possible that they may also be superinfected Buruli ulcers. Third, our findings on the microbial network represent a snapshot of the microbial community. If these communities are stable or if they dynamically change over time should be addressed in longitudinal observations.
Conclusion
The culturome of chronic wounds in Sierra Leonean patients is divers and characterized by the co-detection of P. aeruginosa, K. pneumoniae and S. aureus.
Data availability
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
Change history
25 May 2023
A Correction to this paper has been published: https://doi.org/10.1186/s12879-023-08278-w
References
Verbanic S, Shen Y, Lee J, Deacon JM, Chen IA. Microbial predictors of healing and short-term effect of debridement on the microbiome of chronic wounds. NPJ biofilms and microbiomes. 2020;6(1):21.
Loesche M, Gardner SE, Kalan L, Horwinski J, Zheng Q, Hodkinson BP, Tyldsley AS, Franciscus CL, Hillis SL, Mehta S, et al. Temporal Stability in Chronic Wound Microbiota is Associated with Poor Healing. J Invest Dermatol. 2017;137(1):237–44.
Ibberson CB, Whiteley M. The social life of microbes in chronic infection. Curr Opin Microbiol. 2020;53:44–50.
Young BC, Earle SG, Soeng S, Sar P, Kumar V, Hor S, Sar V, Bousfield R, Sanderson ND, Barker L et al. Panton-Valentine leucocidin is the key determinant of Staphylococcus aureus pyomyositis in a bacterial GWAS. eLife 2019, 8.
Shallcross LJ, Fragaszy E, Johnson AM, Hayward AC. The role of the Panton-Valentine leucocidin toxin in staphylococcal disease: a systematic review and meta-analysis. Lancet Infect Dis. 2013;13(1):43–54.
Schaumburg F, Alabi AS, Peters G, Becker K. New epidemiology of Staphylococcus aureus infection in Africa. Clin Microbiol Infect. 2014;20(7):589–96.
Miller CL, Van Laar TA, Chen T, Karna SLR, Chen P, You T, Leung KP. Global transcriptome responses including small RNAs during mixed-species interactions with methicillin-resistant Staphylococcus aureus and Pseudomonas aeruginosa. MicrobiologyOpen 2017, 6(3):e00427.
Layeghifard M, Hwang DM, Guttman DS. Disentangling interactions in the Microbiome: A Network Perspective. Trends Microbiol. 2017;25(3):217–28.
De Filippo C, Cavalieri D, Di Paola M, Ramazzotti M, Poullet JB, Massart S, Collini S, Pieraccini G, Lionetti P. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc Natl Acad Sci U S A. 2010;107(33):14691–6.
Hammoudi N, Cassagne C, Million M, Ranque S, Kabore O, Drancourt M, Zingue D, Bouam A. Investigation of skin microbiota reveals Mycobacterium ulcerans-aspergillus sp. trans-kingdom communication. Sci Rep. 2021;11(1):3777.
Onwugamba F, Fitzgerald JR, Rochon K, Guardabassi L, Alabi A, Kuhne S, Grobusch MP, Schaumburg F. The role of ‘filth flies’ in the spread of antimicrobial resistance. Travel medicine and infectious disease 2018.
Schaumburg F, Vas Nunes J, Mönnink G, Falama AM, Bangura J, Mathéron H, Conteh A, Sesay M, Sesay A, Grobusch MP. Chronic wounds in Sierra Leone: pathogen spectrum and antimicrobial susceptibility. Infection 2022.
Schwarzkopf A, Dissemond J. Indications and practical implementation of microbiologic diagnostics in patients with chronic wounds. J Dtsch Dermatol Ges. 2015;13(3):203–9.
R. : A language and environment for statistical computing [https://www.R-project.org/]
The igraph. software package for complex network research [https://igraph.org]
Varghese J, Sandmann S, Ochs K, Schrempf IM, Frömmel C, Dugas M, Schmidt HH, Vollenberg R, Tepasse PR. Persistent symptoms and lab abnormalities in patients who recovered from COVID-19. Sci Rep. 2021;11(1):12775.
Pretty, Heatmaps. [https://cran.r-project.org/web/packages/pheatmap/index.html]
Michelsen CF, Christensen AM, Bojer MS, Høiby N, Ingmer H, Jelsbak L. Staphylococcus aureus alters growth activity, autolysis, and antibiotic tolerance in a human host-adapted Pseudomonas aeruginosa lineage. J Bacteriol. 2014;196(22):3903–11.
Biswas L, Götz F. Molecular Mechanisms of Staphylococcus and Pseudomonas interactions in cystic fibrosis. Front Cell Infect Microbiol. 2021;11:824042.
Yung DBY, Sircombe KJ, Pletzer D. Friends or enemies? The complicated relationship between Pseudomonas aeruginosa and Staphylococcus aureus. Mol Microbiol. 2021;116(1):1–15.
Spaan AN, van Strijp JAG, Torres VJ. Leukocidins: staphylococcal bi-component pore-forming toxins find their receptors. Nat Rev Microbiol. 2017;15(7):435–47.
Yeboah-Manu D, Kpeli GS, Ruf M-T, Asan-Ampah K, Quenin-Fosu K, Owusu-Mireku E, Paintsil A, Lamptey I, Anku B, Kwakye-Maclean C, et al. Secondary bacterial infections of Buruli Ulcer Lesions before and after chemotherapy with streptomycin and rifampicin. PLoS Negl Trop Dis. 2013;7(5):e2191.
Uberoi A, Campbell A, Grice EA. Chap. 12 - The wound microbiome. In: Wound Healing, Tissue Repair, and Regeneration in Diabetes edn. Edited by Bagchi D, Das A, Roy S: Academic Press; 2020: 237–258.
Acknowledgements
We thank all laboratory technicians and medical staff at Masanga Hospital, Sierra Leone and the Institute of Medical Microbiology, University of Münster, Germany for their assistance and continuous support.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
SS: Methodology, Software, Formal analysis, Writing - Original Draft, Visualization. JVN: Investigation, Data Curation, Writing - Review & Editing. MPG: Resources, Writing - Review & Editing, Supervision, Funding acquisition. MS: Investigation, Writing - Review & Editing. MAK: Writing - Review & Editing, Supervision. JV: Methodology, Software, Resources, Writing - Review & Editing. FS: Conceptualization, Formal analysis, Investigation, Resources, Data Curation, Writing - Original Draft, Project administration, Funding acquisition. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The National Ethics committee of Sierra Leone, the Sierra Leone Ethics and Scientific Review Committee (approval granted on 25 January 2019) granted ethical approval. All participants signed a written informed consent prior to inclusion. All methods were performed in accordance with the relevant guidelines and regulations.
Consent for publication
Not applicable.
Competing interests
All authors declare that they have no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Sandmann, S., Nunes, J.V., Grobusch, M.P. et al. Network analysis of polymicrobial chronic wound infections in Masanga, Sierra Leone. BMC Infect Dis 23, 250 (2023). https://doi.org/10.1186/s12879-023-08204-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12879-023-08204-0