Abstract
Background
Porcine tonsils are lympho-epithelial tissues, colonized by numerous bacteria and viruses, that act as a reservoir for both host-specific pathogens and zoonotic pathogens with a high potential of transmission to humans. There are no existing studies describing the development of the tonsillar microbiome. We sequenced 16S rRNA genes from tonsillar samples of pigs to follow the development of the microbial communities from birth through weaning. Samples derived from sows were also analyzed to determine potential sources for the tonsil microbiome in piglets.
Results
The composition of the newborn piglet tonsil microbiome could be differentiated by litter and had strong similarity to the sow teat skin as well as sow vaginal microbiome. The tonsil microbiome in these young piglets was mainly dominated by members of the Pasteurellaceae, Moraxellaceae, and Streptococcaceae families, while there were some transient members of the microbiome that were abundant at specific times, such as Staphylococcaceae in newborns and Fusobacteriaceae and Leptotrichiaceae in weeks 2 and 3. The microbiome initially differed between litters but over the following 3 weeks the communities of different litters converged in composition and then diverged in week 4 due to a combination of changes and stresses associated with weaning, including a shift from milk to a solid diet, in-feed Carbadox® and room change.
Conclusions
A significant portion of the tonsil microbiome was acquired either at birth from the sow vaginal tract or within a few hours post-birth from the sow teat skin. Our data demonstrate a temporal succession in the development of the pig tonsillar microbiome through the first weeks of life, with a convergence in the composition of the microbiome in all piglets by 3 weeks of age. The combination of management practices associated with weaning coincided with dramatic shifts in the tonsillar microbiome.
Similar content being viewed by others
Background
Tonsils are lympho-epithelial tissues located at the junction of the oropharynx and nasopharynx that play a key role in surveillance of inhaled or ingested pathogens [1]. In pigs, the tonsils are colonized by numerous bacteria and viruses, and act as a reservoir for both host-specific pathogens and zoonotic pathogens with a high potential of transmission to humans [1,2,3,4]. Bacterial pathogens such as Actinobacillus pleuropneumoniae, Streptococcus suis, and Salmonella enterica are frequently found in tonsils of asymptomatic animals. Under conditions of stress, such as animal transport, these pathogens can spread to the lower respiratory and gastrointestinal tracts and be transmitted to other animals, including humans [5]. It has been suggested that the tonsillar microbiome plays a preventive role in host colonization by pathogenic microorganisms [6,7,8] and also exerts regulatory roles in maintaining immune homeostasis, providing resistance to infection [9, 10].
In contrast to the rising number of studies characterizing the intestinal microbiome of mammals, there are few available studies characterizing the tonsillar microbiome and its development in those species. We have previously characterized the core microbiome of the tonsils in healthy 18–20 week old pigs [11]. In that study, members of the families Pasteurellaceae, Moraxellaceae, Streptococcaceae, Fusobacteriaceae, Veillonellaceae, Enterobacteriaceae, Neisseriaceae, and Peptostreptococcaceae, as well as the order Clostridiales, constituted the core tonsil microbiome in these grower-finisher pigs. Tonsils remain an under-explored habitat of the mammalian microbiome, and how the tonsil microbiome is established, how the structure of this community affects acquisition and carriage of pathogens, and how it contributes to health and disease, in animals or in humans, are not well understood.
Currently, there are no studies following the development of tonsillar microbiomes in humans or in pigs, while other microbiomes, such as the intestinal microbiome in mammals, have been more broadly studied in the last decade. Studies conducted in pigs and humans have suggested that the development of the intestinal microbiome is a gradual and successional process [12,13,14,15] with a significant fluctuation after weaning [16]. The cessation of milk feeding as well as supplementation of other diets and feeding sources generate changes in the intestinal microbiota [17,18,19,20,21]. It is unknown if a similar successional process occurs in tonsils.
It has been shown for humans and pigs that in the development of intestinal microbial communities there is, initially, substantial individual variation in community composition that tends to shift over time, eventually becoming similar in the major phyla across individuals [14]. Environmental factors such as the maternal microbiota can play a relevant role in the initial development of the intestinal microbiota [12, 14, 22]. Nevertheless, the developing microbiome is composed of interacting bacteria and not simply randomly assembled microorganisms [13].
The goal of the current study was to utilize culture-independent, high-throughput sequencing of 16S rRNA genes to follow the development of the tonsillar microbial communities in pigs from birth through weaning.
Methods
Animals
The Michigan State University Institutional Animal Care and Use Committee approved this study and the animal procedures. Ten crossbred sows (Yorkshire x Hampshire) from the high health status herd at the Michigan State University Swine Teaching and Research Center were used in this study, with full approval from the facility coordinators and the Department of Animal Sciences. This herd has no history of recent respiratory disease and is considered free of Actinobacillus pleuropneumoniae, Mycoplasma hyopneumoniae, and Porcine Respiratory and Reproductive Syndrome virus by medical history. This herd experienced a recent outbreak of porcine epidemic diarrhea virus (PEDV), which was under control at the time this study was initiated. The herd has a history of vaccination against porcine circovirus type 2 (PCV2), erysipelas and atrophic rhinitis.
This herd was managed as a farrow to finishing facility with ~ 200 sows. Newborn piglets received a single intramuscular injection of Iron-Dextran. Piglets were weaned at 3 weeks of age (21 to 24 days – average weight 18 pounds) and moved to a nursery room, with litters maintained as pen mates. The weaned piglets were introduced to a solid diet based on a pellet ration (Pig 1300®, Akey Nutrition, Brookville, OH) supplemented with Carbadox® at a dose of 50 g/ton.
Individually identified sows of different parity, from primiparous (pregnant for the first time) to multiparous (pregnant multiple times), were purposely selected for this study and included two first parity sows (ear tags 1700 and 1707, respectively), 1 s parity sow (1631), one fifth parity sow (1445), one sixth parity sow (1402) and one tenth parity sow (1711). Four piglets from each of these six sows, selected randomly, were sampled within a period no longer than 8 h post-birth (PB) (PB were piglets which might have interacted with other piglets or the sow before sampling) and then at 1, 2, 3 and 4 weeks of age. An additional four freshly delivered piglets were randomly selected (N = 16 piglets) from each of 4 crossbred Yorkshire x Hampshire sows (1 s parity sow (1785), two third parity sows (1604, 1760) and one fourth parity sow (1704)), and sampled immediately at birth (AB) (AB piglets were sampled before they had any contact with external sources other than vaginal sources), avoiding any contact of the piglet with either the sow teats or the pen environment prior to sampling, with the purpose of determining the status of the microbiome right at birth. The litter from sow 1604 was sampled in 2014; the remaining three litters were sampled in 2015.
Collection of microbiome samples
Samples of the tonsil microbiome were collected from sows and piglets using either cytology brushes (Cytosoft™, Medical Packaging Corporation, Camarillo, CA) for very small piglets or tonsil brushes developed by our group and validated in previous studies [11]. Collection of samples was as previously described [11]. Briefly, samples were collected at approximately the same time of day for each sampling time. Sow tonsillar samples were collected before they were fed. The pigs’ movement was restricted either by holding them firmly wrapped with a towel, or by using a snare on larger animals. The mouth was held open by using a mouth speculum while the tonsils were brushed. Right and left tonsils were brushed for approximately ten times each, rotating the brush in a clockwise fashion. Brushes were removed from the pig’s mouth and placed into a 50 ml sterile test tube containing 20 ml of 80% ice-cold ethanol. Samples were stored at − 20 °C until processed.
The sow vaginal microbiome was sampled by introducing a sterile cotton swab approximately 8 cm into the vaginal tract and rubbing the vaginal walls with the swab while turning the swab in a clockwise fashion. The teat microbiome was also collected by using a cotton swab and rubbing the teat surface of at least 10 teats per sow. Vaginal and teat swabs were placed individually into 50 ml sterile test tubes containing 20 ml of 80% ice-cold ethanol. Samples were stored at − 20 °C until processed.
The sow fecal microbiome was sampled by collecting approximately 5 g of feces directly from the rectum. Samples were placed individually into 50 ml sterile test tubes containing 20 ml of 80% ice-cold ethanol. Samples were stored at − 20 °C until processed.
Isolation of community DNA
Sample extraction was performed as previously described [11]. Briefly, the 20 ml of 80% ice-cold ethanol containing the brushes, swabs or feces with the microbiome samples were thoroughly vortexed for 1 min, divided into equal volumes and transferred to two sterile acid washed Corex® tubes, and centrifuged in a refrigerated Sorvall SS-34 rotor at 16,000 x g for 30 min. After centrifugation, the supernatant was removed and discarded. The pellet of one tube was suspended in 5 ml of ice-cold 80% ethanol and archived at − 20 °C. The second pellet was suspended in 0.25 ml of phosphate buffered saline, pH 7, and transferred to PowerBead tubes (MoBio Laboratories, Carlsbad, CA) and vigorously shaken for approximately 2 min at room temperature using a MiniBeadBeater-16 (BioSpec Products, Inc., Bartlesville, OK). An exception to this protocol was followed with fecal samples, where both pellets were suspended in 0.5 ml of phosphate buffered saline, pH 7. Community DNA was then extracted using a PowerSoil DNA Isolation Kit (MoBio Laboratories, Carlsbad, CA) following the manufacturer’s instructions. The concentration of extracted community DNA was determined by spectrophotometry, using a Nanodrop (Thermo Scientific, Wilmington, DE). Each sample was then split in two vials; one was archived at − 80 °C and the other was processed for sequencing.
Illumina sequencing and sequence analysis
For Illumina sequencing, samples were processed at the Michigan State University Research Technology Support Facility (RTSF) using an Illumina MiSeq platform. Negative controls consisting of either DNA-free water or MoBio C6 solution (“blank library controls”, [23]) and positive controls consisting of either Escherichia coli DH5α genomic DNA or a well-characterized activated sludge polymicrobial community [24] were included in the sequencing runs. Briefly, the V4 region of the 16S rRNA gene of the community DNA was amplified using uniquely indexed primers for each sample, as described by Caporaso [25]. After PCR, amplification products were normalized using an Invitrogen SequalPrep normalization plate. The normalized samples were pooled and PCR reaction cleanup was done with AMPure XP beads. After quality control and quantitation, the pool was loaded on an Illumina MiSeq v2 flow cell and sequenced with a 500-cycle v2 reagent kit (PE250 reads). Base calling was performed by Illumina Real Time Analysis Software (RTA) v1.18.54 and output of RTA demultiplexed and converted to FastQ files with Illumina Bcl2fastq v1.8.4.
Amplicon analysis was performed using the open-source, platform-independent, community-supported software program mothur v.1.38.0 (http://www.mothur.org) [26]. Processing of the raw sequencing data was done according to the mothur standard operating procedure (http://www.mothur.org/wiki/MiSeq_SOP) [27]. Alignment was accomplished using the mothur-formatted version 123 of Silva 16S rRNA gene database [28]. Sequences were classified and any sequences classified as Chloroplast, Mitochondria, unknown, Archaea, or Eukaryota were removed from the data set. Subsampling at 5907 sequences per sample was performed, followed by a preclustering of the sequences and removal of chimeric sequences using a mothur-formatted version of the Ribosomal Database Project (RDP) training set version 14 and uchime, based on the mothur protocol. Sequences were classified into Operational Taxonomic Units (OTUs) of ≥97% sequence identity. Singleton and doubleton reads were removed, followed by subsampling at 3776 sequences per sample. Because the negative controls consistently showed high levels of contaminants that skewed the results seen with some of the low biomass samples such as those from the newborn piglets, especially those from Litter 1700, 4 OTUs (Ralstonia, Bacillaceae 1, Burkholderia, and Brevundimonas) were also removed from the data set prior to final analysis. A SIMPER comparison of Litter 1700 piglet and sow samples to the relevant negative controls showed 66.1% or greater dissimilarity between the negative controls and the pig samples, rejecting the hypothesis that the sequence data for Litter and sow 1700 samples is due to contamination only [23]. Therefore these samples were included in the final data set. For the final analysis of the data, samples were subsampled to 1979 reads per sample. The full data set analyzed is given in Additional file 1: Table S1.
Diversity and statistical analysis
The statistical analysis was performed using a clustering cutoff of 3% for the processed sequences. Mothur output files were used to estimate diversity indexes and core microbiomes. PAST3 was used for generation of the UPGMA dendogram file, SIMPER, coordinate analysis of the samples, two dimensional scatter plot and 95% concentration ellipses [29]. ImageJ was used to measure the area of the ellipses for the two dimensional scatter plot [30]. Dendogram construction was performed using FigTree v.1.4.2. (http://tree.bio.ed.ac.uk/software/figtree/). RStudio (Version 0.99.446; https://www.rstudio.com/) and libraries: gplots (https://CRAN.R-project.org/package=gplots), plot3D (https://CRAN.R-project.org/package=plot3D), and rgl (https://CRAN.R-project.org/package=rgl) were used to generate heatmaps and tridimensional scatter plots, respectively. Inkscape 0.91 (https://inkscape.org/en/download/mac-os/), was used to process images and edit labels.
Availability of supporting data
Raw sequence data is available at the NCBI database (SRA accession number: SRP110992) and the code for the mothur analysis is available at (https://figshare.com/s/2c98593a953cc9bb1366).
Results
A total of 171 samples derived from the tonsillar microbiome of piglets and sow tonsils, teat skin, vagina and feces as well as control samples were processed. One hundred forty-four pig samples were collected in Spring 2014, and an additional 12 pig samples were collected in Fall 2015. A total of 26 samples with less than 3776 final reads were not included in the final analysis. One of the approaches that we used was comparative analysis of core microbiomes. We defined the core microbiome for a specific litter as the OTU members of the microbiome that were present in at least 75% of piglets of a litter, when litters had at least 4 samples; otherwise core OTUs were defined as OTUs present in all the samples. For the sow samples, where 6 samples were analyzed, we considered OTUs present in 66.6% of the samples to be core OTUs. Further, core OTUs were defined as present in a minimum relative abundance equal or higher than 1% and/or 0.1% of total reads for a selected litter and/or period (Additional file 2: Table S2).
The tonsillar microbiome found in PB samples clusters by litter
Among the 156 samples collected, 40 samples contain the tonsillar microbiome of newborn piglets (Table 1). Twenty four of these 40 samples were collected from PB piglets and the remaining 16 samples were collected from AB piglets. Six samples were discarded from the analysis due to low number of reads.
A dendrogram clustering of PB piglet samples and sow derived samples, based on the Bray-Curtis dissimilarity index (Fig. 1) showed that samples from each litter clustered together with littermates. Microbiomes derived from piglets from primiparous sows (Litter 1700 and 1707) clustered in separated clades, as did the piglets from sow 1402. In general, PB piglet samples were more closely associated with samples derived from the sow teat microbiome as opposed to other sow samples. However, piglets from litter 1700 also clustered with the sample from the sow vaginal microbiome.
Samples from the sow tonsils and sow feces clustered together by source. Samples from the vaginal tract of multiparous sows clustered together, but were distinct from those from the two primiparous sows.
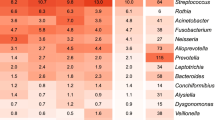
A heat-map representation of the major OTUs reveals the differences driving the clustering (Fig. 2). In the PB piglets from sows 1445, 1631, and 1711, all multiparous sows, the most abundant OTUs were Pasteurellaceae, Streptococcus, Moraxella, Rothia, and Staphylococcus (OTUs 001, 002, 003, 007, and 009, respectively). PB piglets from sow 1402 (parity 6) contained significant numbers of Pasteurellaceae, Streptococcus, and Moraxella (OTUs 001, 002, and 003) but also contained large numbers of Enterobacteriaceae (OTU005) and Clostridium sensu strictu (OTU010). In contrast, piglets derived from primiparous sows 1700 and 1707 had a very low abundance of Pasteurellaceae, Streptococcus, and Moraxella (OTUs 001, 002 and 003), but a high abundance of Staphylococcus (OTU009) as well as several anaerobic organisms.
Thirty most abundant Operational Taxonomic Units (OTUs) for piglets and sow samples. Heat-map showing the relative abundance of the top 20 OTUs identified for all samples. The figure shows the relative abundance of the OTUs for each PB piglet and sow samples, as well as the average for the litters and sow samples
A significant proportion of the microbiome of the piglets came from maternal sources
We processed samples derived from teat skin, vagina and tonsil from six sows belonging to the PB group (Table 1), and fecal samples from two of these sows. We compared and traced which OTUs were identified as core microbiome for each litter and if they were identified as core for microbiomes derived from the different sow sources (Additional file 2: Table S2).
The OTUs identified as core for PB litters varied between litters. As was seen in Fig. 2, Pasteurellaceae, Streptococcus and Moraxella (OTUs 001, 002 and 003) were identified as core with a minimum relative abundance equal or higher than 1% and were found in high proportions in most piglets from multiparous sows but not piglets from primiparous sows. When sow sources for these three OTUs were examined, OTU001 (Pasteurellaceae) was identified as a core organism in the sow vaginal tract, representing on average 20.1% of the vaginal microbiome. However, OTU001 was present in only low amounts in the vaginal tracts from primiparous sows. In contrast, OTU002 (Streptococcus) was found in both vaginal and teat skin samples, while OTU003 (Moraxella) was found mainly in teat skin samples; both were more prevalent in samples from multiparous sows. In addition, Rothia (OTU007) and Staphylococcus (OTU009) were identified in high proportions and as core for most PB litters regardless of sow parity, as well as core for the sow teat microbiome, but not as part of the core microbiome for other sow samples.
These results suggested that the PB piglets had acquired new organisms from teat skin or other sources within the first few hours of life. To test this hypothesis, we collected samples from piglets immediately at birth (AB), prior to any contact with sources other than the uterus and the sow vaginal tract. These samples were very sparse in content, and unfortunately the resulting sequence data was dominated by organisms known to be frequent contaminants in DNA extraction kits and library preparation kits, including Comamonadaceae, Sphingomonadaceae and Xanthomonadaceae [23]. However, when these likely contaminants were deleted from the data, it was clear that the organisms most frequently seen were organisms associated mainly with the sow vaginal tract, including OTU002 Streptococcus, Corynebacterium (OTU034 and 051, and several anaerobic organisms including multiple OTUs of Clostridiaceae and Peptostreptococcaceae. OTU001 Pasteurellaceae was also found although in low numbers.
An analysis of the microbiome derived from sow sources as well as from PB piglets at the family level showing the 20 most abundant taxa is presented in Fig. 3. These taxa generally include multiple OTUs. Streptococcaceae (20 OTUs), Staphylococcaceae (4 OTUs), Micrococcaceae (10 OTUs), Pasteurellaceae (21 OTUs) Moraxellaceae (29 OTUs) and families belonging to the Clostridiales (746 OTUs) were present in high abundance in the PB piglets, and were also found in the sow vagina samples and/or the sow teat skin samples.
Tonsil communities of newborn piglets differed initially between litters but by 3 weeks of age clustered together reflecting similar composition
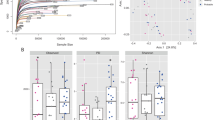
We inquired if, over time, the tonsillar communities of piglets reached a common microbiome. A principal coordinate analysis (PCoA) based on Bray-Curtis distances for the PB newborn through week 4 samples (Fig. 4a and b) showed a dynamic pattern in the samples. The PB newborns were widely spread although two distinct clusters were detectable. Over the next 3 weeks the microbiomes formed increasingly tighter clusters reflecting greater similarity as the animals aged. However, in the fourth week there was a dramatic shift, where, instead of continuing to cluster together more tightly as observed in previous weeks, samples were scattered.
Principal Coordinate Analysis (PCoA) characterizing the tonsillar microbiome from PB piglets through the different sampling times. Two dimensional plotting illustrating the distribution of microbiome in the first two axis, the 95% concentration ellipses for newborn piglets through 4 weeks and the relative area for the ellipses (a). Three-dimensional plot illustrating the main three axes for the distribution of the microbiome of PB piglets since newborn through 4 weeks (b)
Further analysis using a SIMPER approach based on Bray-Curtis distances (Additional file 3: Table S3) reinforced what we observed in the PCoA plots. Microbiome samples from PB piglets were dissimilar when compared with others and the value of dissimilarity fluctuated, with the lowest dissimilarity being 45.8% for litter 1445 vs 1711 and the highest being 94.04% for litter 1631 vs 1707. Although there was a substantial variability in the overall dissimilarity between litters, as time advanced the dissimilarity value between litters decreased and reached the lowest values in the third week, when values were as low as 23.5% for litters 1402 vs 1707 and the highest was 37.3% for litters 1402 vs 1445.
To visualize the temporal patterns of distribution of different bacterial families, we charted the mean value for the 20 most commonly identified families per sampling time (Fig. 5). While there were litter to litter variations in the microbiome of PB newborn piglets, as shown in Fig. 3, the differences between litters disappeared through the successional development of the microbiome in the following weeks.
Members of the families Micrococcaceae and Staphylococcaceae were abundant in PB newborns but decreased drastically and almost disappeared in the following weeks. By 1 week of age, the tonsil microbiome was dominated by members of the Pasteurellaceae, Moraxellaceae, and Streptococcaceae families, a dominance that remained throughout the lives of these pigs (unpublished data). In contrast, some members of the microbiome appeared and disappeared over time, such as Porphyromonadaceae and Flavobacteriaceae which appeared at weeks 1 to 3, as well as Fusobacteriaceae and Leptotrichiaceae that were present during weeks 2 and 3.
The transition between third and fourth week represents a critical period for the development of the microbiome
According to MSU farm management practices, piglets were weaned between the third and fourth week. We investigated if the management practices experienced by the piglets (weaning, shift to a nursery room and introduction to a pelleted ration supplemented with Carbadox®), correlated with the timing of shifts in community structures.
An analysis of the members of core microbiome (Additional files 4 and 5: Table S4 and Table S5) demonstrated that for the third week, OTU001 (Pasteurellaceae), OTU002 (Streptococcus), OTU003 (Moraxella), OTU006 (Porphyromonas), OTU017 (Fusobacterium) and OTU026 (Flavobacteriaceae), identified as core with a minimum relative abundance equal or higher than 1%, were shared among all the litters, and represented on average 70.1% of the identified OTUs. In addition, OTU012 (Streptococcus), OTU15 (Prevotellaceae), OTU018 (Peptostreptococcaceae) and OTU024 (Bacillales Incertae Sedis XI) were core members at a minimum relative abundance of 0.1% or higher. OTU001 (Pasteurellaceae), OTU002 (Streptococcus) and OTU003 (Moraxella) dominated the microbiome, together representing on average 59.5% of the identified OTUs. An analysis at the family level (Fig. 5) for the same period shows the families Moraxellaceae and Pasteurellaceae as the most abundant families with an abundance of almost 55.6%, followed by members of the families Streptococcaceae, Porphyromonadaceae, Fusobacteriaceae, Leptotrichiaceae and Flavobacteriaceae, together with an abundance of 24.5%.
The fourth week showed a different panorama, where only two OTUs, OTU001 (Pasteurellaceae) and OTU002 (Streptococcus), were identified as core at 1% or higher for all the litters. However, in contrast to the third week, these two OTUs represented on average 67.6% of the identified OTUs. In the fourth week, members of the Streptococcaceae family increased dramatically 5–6 fold compared to week 3, Pasteurellaceae remained at the same levels, and members of Moraxellaceae decreased 4 fold. Members of Porphyromonadaceae, Flavobacteriaceae, Fusobacteriaceae, and Leptotrichiaceae decreased to negligible numbers in week 4.
Furthermore, a SIMPER analysis (Additional file 3: Table S3) indicated that the dissimilarity in percentages between litters showed changes; instead of continuing the downward trend as observed in previous weeks, the dissimilarity values generally increased by 10–20%, with the lowest being 32.04% between litters 1631 and 1402 and the highest 47.98% between 1700 and 1445.
Discussion
In this study, we followed the development of the tonsillar microbial communities in pigs from birth through weaning. We focused on identification of the source of bacteria found in the tonsils, the successional development, and the apparent effect of weaning on the tonsillar microbiome.
Analysis of the microbiome in the PB piglets showed that the most abundant organisms were Streptococcus, Staphylococcus, Moraxella, Rothia, and Pasteurellaceae (OTUs 002, 009, 003, 007, and 001 respectively). Based on comparative analysis of putative source microbiomes and piglet tonsil microbiomes, as well as cultivation studies, we concluded that while the Pasteurellaceae and Streptococcus were most likely acquired from the sow vaginal tract during the birth process, Moraxella, Staphylococcus and Rothia were likely acquired from the sow teat skin (or milk, which we did not sample) with a few hours after birth.
To determine what organisms were actually acquired during birth, we collected samples from piglets immediately after they were born (AB piglets), prior to contact with teat skin or other sources except the sow vaginal tract. As described in results, these samples were very sparse and the microbiome identified in them was heavily biased by likely contaminants from either the DNA extraction kit or library construction kit used. This has recently become a recognized problem in microbiome studies on low microbial biomass samples [23, 31, 32]. While we ran appropriate positive controls and zero template library construction controls, we did not run DNA extraction controls with the same kit used in this study. We have done this with a different lot number of the same kit, and used the information from that experiment as well as other published studies [23] to help analyze the data on the microbiome in AB piglets.
When we removed what we considered to be contaminants from the data set for AB piglets, the remaining organisms included OTU002 Streptococcus and OTU001 Pasteurellaceae, both of which colonized, multiplied, and persisted for the life of these pigs. It should be noted that we have cultured Streptococcus suis from at least half of these AB piglet samples, as well as from sow vaginal samples, providing additional support for the conclusion that this organism is acquired during the birth process. We also identified organisms such as OTU016 Clostridium sensu strictu and two OTUs (034 and 051) identified as Corynebacterium that were found in the sow vaginal tract and in AB and PB piglets, but not older piglets, suggesting that some of the organisms acquired during birth do not persist in the tonsils.
If these organisms failed to adhere to the tonsil epithelium, they would be washed out of the oropharynx, and other organisms acquired from the vaginal tract, such as Streptococcus, that do adhere to the tonsil epithelium would be able to colonize and multiply. Further, once the piglets had contact with other sow sources such as the teat skin or milk, new colonizers such as Staphylococcus could be acquired, leading to a sequential development of the tonsil microbiome starting within the first few hours of life. The clustering pattern observed in Fig. 1, where samples from three of the four multiparous litters clustered together as a group with sow teat skin samples and litters born from primiparous sows clustered with the cognate teat and vaginal or just teat samples, reinforce that teat and/or vaginal samples from sow are the initial source of the microbiome for PB newborns. Our findings are supported by the results of other studies, such as the study of Mandar and Mikelsaar [33], which characterized the initial colonization of the external ear canal in newborn humans and compared the microorganisms found with the vaginal flora of their mothers and concluded that there is a significant influence of the vaginal microflora in the initial microbial population found in the newborns. Further, Bokulich et al. [12], studying the effect of antibiotics, birthmode and diet in the development of fecal microbiota in children during early life, suggested that early colonizers are transmitted to children from maternal microbiota. They found that children delivered vaginally shared more OTUs derived from the vagina than children delivered by ceasarean. Additionally, they identified that the infant fecal microbiota was initially associated with vaginal and rectal maternal microbiota, but later was more associated with maternal fecal microbiota. Further, a study of the development of the human intestinal microbiome suggested that the bacterial population detected in human infants in early stages of life might be determined by specific bacteria to which infant was previously exposed, based on similarity patterns observed between infant samples and maternal sources as breast milk and vaginal swabs [14]. These authors identified Streptococcus and Staphylococcus as well as other aerobes as first colonizers [14]. Similarly, we observed that piglets sampled within short period after being born had high proportions of Streptococcus, Moraxella and Staphylococcus in their tonsils.
Our results demonstrated a strong litter effect in the tonsillar microbiome in PB piglets (Fig. 3). However, over the following 3 weeks there was a gradual successional development in the tonsil microbiomes of all piglets and by the third week the microbiomes of all piglets from all litters were highly similar (Fig. 4). This was true even for a litter that did not share the same farrowing room, as was the case for piglets from litter 1445 (data not shown). This is in contrast to the reported development of the pig intestinal microbiome over the first few weeks of life, where no obvious effect of litter was seen [34].
In this successional development, some organisms such as Staphylococcus and Micrococcaceae that were found in high proportions in the PB piglets decreased dramatically within the next 2 weeks, suggesting a role only as initial weak colonizers. These organisms are commonly isolated from multiple skin locations [35, 36] and vagina [37] and are likely to be vertically transmitted from sow to offspring. Over the same period there was a concomitant increase in stronger colonizers, particularly members of the families Pasteurellaceae and Moraxellaceae, as well as maintenance of the levels of Streptococcaceae. These three families, which contain both commensals and pathogens that are residents of mucosal surfaces of animal and humans, comprise a large proportion of the tonsil microbiome throughout the lives of pigs. Members of the order Clostridiales were also identified as a small but consistent part of the microbiome throughout the first 3 weeks, which is not surprising since they were identified as part of the core tonsillar microbiome of pigs [11] as well as members of intestinal microbiome [15, 18, 34, 38].
Over this period we also found several transitory OTUs or families that appeared and disappeared at specific time points. For example, Porphyromonadaceae, Prevotellaceae, and Flavobacteriaceae appeared at week 1 and increased slightly over the next 2 weeks. Similarly, Fusobacteriaceae and Leptotrichiaceae appeared at weeks 2 and 3. By week three, this successional development of the tonsil microbiome of all of the piglets, regardless of litter, led to a distinct common consortium of bacterial species.
Similarly, Palmer et al. [14], studied the microbiome profiles from human infant stool samples and suggested that although initially the microbiome was very distinct between individuals, over time it converged towards a common profile. The authors followed the development of the intestinal microbiome in 14 human infants, and showed that there was a considerable variation in the colonization process among individuals. Each infant had a distinct arrangement of bacterial species that it acquired and maintained. The acquired microorganisms had a temporal pattern in which they appeared and disappeared; however, they reached a stable population over time, with some taxonomic groups persisting while the presence of other taxa was only transient [14]. The authors reported the occurrence of significant shifts in the population assembly which seemed to stabilize over time.
At 3 weeks of age, the piglets were weaned onto a solid ration containing the growth promoter Carbadox® and were moved from the farrowing room to the nursery, while kept in groups with their littermates. Comparison of the microbiome composition for the third and fourth weeks showed a major shift associated with the significant stress event of weaning, with its environmental, social and feed changes. Our results are in concordance with other studies following the development of the intestinal microbiome in humans and pigs, which reported significant changes in microbiome composition associated with life events [13, 14]. In pigs, it has been demonstrated that the transition from nursery to weaning is associated with a significant change in the intestinal microbiota [18, 39]. However, the effect of this transition on the tonsillar microbiome has not previously been studied in pigs or humans. In this study, we have demonstrated that the transition from farrowing to a nursery room in parallel with weaning and supplementation of the diet with Carbadox® coincided with a major shift in the tonsil microbiota.
The most obvious effect of weaning was the 5–6 fold increase in members of the Streptococcaceae, from an average of 8% before weaning to ~ 43% of the total identified families after weaning. These organisms were primarily members of the genus Streptococcus. There was a concomitant decrease for members of the Moraxellaceae family, which decreased approximately 4-fold, from ~ 32% before weaning to ~ 8% after weaning. In addition, Fusobacteriaceae, Leptotrichiaceae, Porphyromonadaceae and Flavobacteriaceae decreased dramatically. However, at the same time, the proportion represented by Pasteurellaceae remained constant.
It is not clear whether a single stress, such as change in food or change in environment or application of antibiotic, or a combination of stresses was responsible for the major disruption in the tonsil microbiome at this time. It has been demonstrated in multiple studies that the intestinal microbiota composition was deeply perturbed when the host was treated with antibiotics [8, 12, 14, 40,41,42,43]. In our study, the piglets were supplemented with Carbadox® in food at the time of weaning. It was reported by Looft et al. [40] that the structure and composition of the intestinal community of pigs supplemented with Carbadox® changed significantly, where the relative abundance of Prevotella increased associated with Carbadox® administration as a result of decreased abundance of other bacteria. Another study correlating changes in microbiota with changes of diet during nursing and weaning found that the fecal population of Prevotellaceae increased ~ 50-fold in weaned pigs compared with nursing animals [18]. In our study, members of Prevotellaceae, a minor population in the tonsils, decreased slightly from 2.7% for week three to 1.9% for week four. We can only speculate about the opposite results identified in our studies, since we are comparing different niches (feces vs tonsils) and we did not have controls that were not fed Carbadox® as this was not a goal of our study.
It has been suggested that the environment also plays a relevant role in the initial acquisition of the microbiome. Mulder et al. [10], followed the development of gut microbiota and the potential impacts of early environmental changes, and demonstrated a major impact in the microbial diversity related with those changes and that the impact of those changes are preserved through adulthood. One of the major changes experienced by the weaned piglets is the separation from the sow and the introduction to new food. A deep impact in the intestinal microbial composition has been seen associated with cessation of breast feeding and introduction to a different diet [19].
We observed members of the microbiota that were present all the time, some in high relative abundance while others were found in low abundance. Similarly, there were also transient members whose exact role in the development of the tonsil microbiome is unclear but worth investigating in future studies. It is possible that these transient organisms appeared as secondary colonizers but were displaced by other members of the microbiota, or that these microorganisms were adversely affected by the stressful event of weaning and/or the Carbadox® supplementation and thus disappeared. We have an imprecise idea about the true role of Carbadox® in the development of the tonsillar microbiome, since our study was not specifically intended to answer this question. We unfortunately did not collect sow milk samples, or samples from the pen/cage floor which might have given us a better idea of the possible sources of the PB and subsequent microbiome.
There are many questions that arise from our work that will be the subject of future research. However, this study lays the foundation of our knowledge of how the tonsillar microbiome develops in pigs in the first hours and weeks of age and how weaning affects this microbiome. To the best of our knowledge, this is the first published study that follows the development of the tonsillar microbiome in any mammal during the first weeks of life.
Conclusions
Our data demonstrate a temporal succession in the development of the pig tonsillar microbiome through the first weeks of life. Many of the organisms found in the piglet oropharynx and tonsils immediately after birth appear to wash out rapidly, within the first few hours of life, while other organisms acquired from the vaginal tract, such as Streptococcaceae and Pasteurellaceae, colonize and multiply. Additional organisms are also acquired rapidly from the sow teat skin, and possibly from milk, and eventually also from feces. The composition of the PB newborn piglet tonsil microbiome initially can be differentiated by litter and clusters mainly with the sow teat skin microbiome. Nevertheless, over the next 3 weeks, the composition and structure of the tonsil microbiome reaches a common point of development, showing a high degree of similarity among all piglets, regardless of litter and just prior to weaning. However, there was a dramatic change in the post weaning tonsillar microbiome, which was likely engendered by a combination of change in diet, change in environment, and addition of in- feed antibiotic, demonstrating the effect that weaning management practices exert in shaping tonsillar microbial communities. This research demonstrates the need for further studies to elucidate the role of antibiotic supplementation of feed in the development of tonsillar microbial communities, specifically when administered during the highly susceptible time of weaning.
Abbreviations
- AB:
-
Neonatal piglets sampled at birth
- NCBI:
-
National Center for Biotechnology Information
- OTU:
-
Operational Taxonomic Unit
- PB:
-
Neonatal piglets sampled within 8 h post-birth
- PCoA:
-
Principal Coordinate Analysis
- RDP:
-
Ribosomal Database Project
- SIMPER:
-
Similarity percentages
- UPGMA:
-
Unweighted pair group method with arithmetic mean
References
Horter DC, Yoon KJ, Zimmerman JJ. A review of porcine tonsils in immunity and disease. Anim Health Res Rev. 2003;4:143–55.
Smith TC, Harper AL, Nair R, Wardyn SE, Hanson BM, Ferguson DD, Dressler AE. Emerging Swine Zoonoses. Vector Borne Zoonotic Dis. 2011;11:1225–34.
MacInnes JI, Gottschalk M, Lone AG, Metcalf DS, Ojha S, Rosendal T, Watson SB, Friendship RM. Prevalence of Actinobacillus pleuropneumoniae, Actinobacillus suis, Haemophilus parasuis, Pasteurella multocida, and Streptococcus suis in representative Ontario swine herds. Can J Vet Res. 2008;72:242–8.
Kernaghan S, Bujold AR, MacInnes JI. The microbiome of the soft palate of swine. Anim Health Res Rev. 2012;13:110–20.
Williams LP, Newell KW. Salmonella excretion in joy-riding pigsSalmonella excretion in joy-riding pigs. Am J Public Health Nations Health. 1970;60:926–9.
Weyrich LS, Feaga HA, Park J, Muse SJ, Safi CY, Rolin OY, Young SE, Harvill ET. Resident microbiota affect bordetella pertussis infectious dose and host specificity. J Infect Dis. 2014;209:913–21.
Ghartey JP, Smith BC, Chen ZG, Buckley N, Lo YT, Ratner AJ, Herold BC, Burk RD. Lactobacillus crispatus dominant vaginal microbiome is associated with inhibitory activity of female genital tract secretions against Escherichia coli. PLoS One. 2014;9:8.
Croswell A, Amir E, Teggatz P, Barman M, Salzman NH. Prolonged impact of antibiotics on intestinal microbial ecology and susceptibility to enteric Salmonella infection. Infect Immun. 2009;77:2741–53.
Willing BP, Vacharaksa A, Croxen M, Thanachayanont T, Finlay BB. Altering host resistance to infections through microbial transplantation. PLoS One. 2011;6:9.
Mulder IE, Schmidt B, Stokes CR, Lewis M, Bailey M, Aminov RI, Prosser JI, Gill BP, Pluske JR, Mayer CD, et al. Environmentally-acquired bacteria influence microbial diversity and natural innate immune responses at gut surfaces. BMC Biol. 2009;7:20.
Lowe BA, Marsh TL, Isaacs-Cosgrove N, Kirkwood RN, Kiupel M, Mulks MH. Defining the “core microbiome” of the microbial communities in the tonsils of healthy pigs. BMC Microbiol. 2012;12:14.
Bokulich NA, Chung J, Battaglia T, Henderson N, Jay M, Li H, Lieber AD, Wu F, Perez-Perez GI, Chen Y, et al. Antibiotics, birth mode, and diet shape microbiome maturation during early life. Sci Transl Med. 2016;8:13.
Koenig JE, Spor A, Scalfone N, Fricker AD, Stombaugh J, Knight R, Angenent LT, Ley RE. Succession of microbial consortia in the developing infant gut microbiome. Proc Natl Acad Sci U S A. 2011;108:4578–85.
Palmer C, Bik EM, DiGiulio DB, Relman DA, Brown PO. Development of the human infant intestinal microbiota. PLoS Biol. 2007;5:1556–73.
Inoue R, Tsukahara T, Nakanishi N, Ushida K. Development of the intestinal microbiota in the piglet. J Gen Appl Microbiol. 2005;51:257–65.
Konstantinov SR, Awati AA, Williams BA, Miller BG, Jones P, Stokes CR, Akkermans ADL, Smidt H, De Vos WM. Post-natal development of the porcine microbiota composition and activities. Environ Microbiol. 2006;8:1191–9.
Stephens WZ, Burns AR, Stagaman K, Wong S, Rawls JF, Guillemin K, Bohannan BJM. The composition of the zebrafish intestinal microbial community varies across development. ISME J. 2016;10:644–54.
Frese SA, Parker K, Calvert CC, Mills DA. Diet shapes the gut microbiome of pigs during nursing and weaning. Microbiome. 2015;3:10.
Bergstrom A, Skov TH, Bahl MI, Roager HM, Christensen LB, Ejlerskov KT, Molgaard C, Michaelsen KF, Licht TR. Establishment of intestinal microbiota during early life: a longitudinal, explorative study of a large cohort of Danish infants. Appl Environ Microbiol. 2014;80:2889–900.
Konstantinov SR, Favier CF, Zhu WY, Williams BA, Kluss J, Souffrant WB, de Vos WM, Akkermans ADL, Smidt H. Microbial diversity studies of the porcine gastrointestinal ecosystem during weaning transition. Anim Res. 2004;53:317–24.
Konstantinov SR, Zhu WY, Williams BA, Tamminga S, de Vos WM, Akkermans ADL. Effect of fermentable carbohydrates on piglet faecal bacterial communities as revealed by denaturing gradient gel electrophoresis analysis of 16S ribosomal DNA. FEMS Microbiol Ecol. 2003;43:225–35.
Thompson CL, Wang B, Holmes AJ. The immediate environment during postnatal development has long-term impact on gut community structure in pigs. ISME J. 2008;2:739–48.
Kim D, Hofstaedter CE, Zhao C, Mattei L, Tanes C, Clarke E, Lauder A, Sherrill-Mix S, Chehoud C, Kelsen J, et al. Optimizing methods and dodging pitfalls in microbiome research. Microbiome. 2017;5:14.
Liu WT, Marsh TL, Cheng H, Forney LJ. Characterization of microbial diversity by determining terminal restriction fragment length polymorphisms of genes encoding 16S rRNA. Appl Environ Microbiol. 1997;63:4516–22.
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci U S A. 2011;108:4516–22.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol. 2009;75:7537–41.
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol. 2013;79:5112–20.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Gloeckner FO. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013;41:D590–6.
Hammer O, Harper DAT, Ryan PD. PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron. 2001;4:1.
Schneider CA, Rasband WS, Eliceiri KW. NIH image to ImageJ: 25 years of image analysis. Nat Methods. 2012;9:671–5.
Glassing A, Dowd SE, Galandiuk S, Davis B, Chiodini RJ. Inherent bacterial DNA contamination of extraction and sequencing reagents may affect interpretation of microbiota in low bacterial biomass samples. Gut Pathogens. 2016;8:12.
Lauder AP, Roche AM, Sherrill-Mix S, Bailey A, Laughlin AL, Bittinger K, Leite R, Elovitz MA, Parry S, Bushman FD. Comparison of placenta samples with contamination controls does not provide evidence for a distinct placenta microbiota. Microbiome. 2016;4:29.
Mandar R, Mikelsaar M. Transmission of mother's microflora to the newborn at birth. Biol Neonate. 1996;69:30–5.
Park SJ, Kim J, Lee TS, Rhee SK, Kim H. Characterization of the fecal microbiome in different swine groups by high-throughput sequencing. Anaerobe. 2014;28:157–62.
Braem G, De Vliegher S, Verbist B, Heyndrickx M, Leroy F, De Vuyst L. Culture-independent exploration of the teat apex microbiota of dairy cows reveals a wide bacterial species diversity. Vet Microbiol. 2012;157:383–90.
Grice EA, Kong HH, Conlan S, Deming CB, Davis J, Young AC, Bouffard GG, Blakesley RW, Murray PR, Green ED, et al. Topographical and temporal diversity of the human skin microbiome. Science. 2009;324:1190–2.
Wegener HC, Skovjensen EW. A longitudinal study of Staphylococcus hyicus colonization of vagina of gilts and transmission to piglets. Epidemiol Infect. 1992;109:433–44.
Mach N, Berri M, Estelle J, Levenez F, Lemonnier G, Denis C, Leplat JJ, Chevaleyre C, Billon Y, Dore J, et al. Early-life establishment of the swine gut microbiome and impact on host phenotypes. Environ Microbiol Rep. 2015;7:554–69.
Alain B, Pajarillo E, Chae J-P, Balolong MP, Bum Kim H, Kang D-K. Assessment of fecal bacterial diversity among healthy piglets during the weaning transition. J Gen Appl Microbiol. 2014;60:140–6.
Looft T, Allen HK, Casey TA, Alt DP, Stanton TB. Carbadox has both temporary and lasting effects on the swine gut microbiota. Front Microbiol. 2014;5:9.
Looft T, Johnson TA, Allen HK, Bayles DO, Alt DP, Stedtfeld RD, Sul WJ, Stedtfeld TM, Chai BL, Cole JR, et al. In-feed antibiotic effects on the swine intestinal microbiome. Proc Natl Acad Sci U S A. 2012;109:1691–6.
Rettedal E, Vilain S, Lindblom S, Lehnert K, Scofield C, George S, Clay S, Kaushik RS, Rosa AJM, Francis D, Brozel VS. Alteration of the ileal microbiota of weanling piglets by the growth-promoting antibiotic chlortetracycline. Appl Environ Microbiol. 2009;75:5489–95.
Sekirov I, Tam NM, Jogova M, Robertson ML, Li YL, Lupp C, Finlay BB. Antibiotic-induced perturbations of the intestinal microbiota alter host susceptibility to enteric infection. Infect Immun. 2008;76:4726–36.
Acknowledgements
We would like to thank Kevin Turner, Chris Rozeboom, Scott Kramer and Puja Basu for their instrumental assistance with the sample collection during the study.
Funding
This research was supported by Endowed Research Funds from the College of Veterinary Medicine, Michigan State University, and by the Discretionary Funding Initiative through the Office of the Vice-President for Research and Graduate Studies, Michigan State University. Universidad de Pamplona, Pamplona – Colombia and Colciencias – Colombia, Programa de Formación Doctoral “Francisco José de Caldas” - Credito Educativo - Convocatoria 529 supported Luis Carlos Peña-Cortes. These funding sources did not contribute to the design of the study, to collection, analysis, and interpretation of data, or to writing the manuscript.
Availability of data and materials
Raw sequence data is available at the NCBI database (SRA accession number: SRP110992) and the code for the mothur analysis is available at (https://figshare.com/s/2c98593a953cc9bb1366).
Author information
Authors and Affiliations
Contributions
LCPC, MHM, and JF conceived and designed the study. LCPC, RML, JF, and MHM collected the samples. LCPC and RML processed the samples. LCPC, TM, RML and MHM analyzed the data. LCPC, TM and MHM drafted the manuscript. All authors revised and edited the manuscript, gave final approval for the version to be published, and agreed to be accountable for all aspects of the work.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The Michigan State University Institutional Animal Care and Use Committee approved this study and the animal procedures.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Additional files
Additional file 1:
Table S1. OTU table for the full dataset. (XLSX 1547 kb)
Additional file 2:
Table S2. Core microbiome at OTU level for litters and sow samples. Excel sheet containing values for Core OTU identified for litters and sow samples at 1 and 0.1% of minimum relative abundance. (XLSX 20 kb)
Additional file 3:
Table S3. SIMPER analysis between litters through different sampling times. Excel sheet containing values from SIMPER analysis between different litters through the sampling time (Birth through fourth week). (XLSX 10 kb)
Additional file 4:
Table S4. Core microbiome at OTU level for third week. Excel sheet containing values for Core OTU identified for 3 weeks old piglets at 1 and 0.1% of minimum relative abundance. (XLSX 16 kb)
Additional file 5:
Table S5. Core microbiome at OTU level for fourth week. Excel sheet containing values for Core OTU identified for 4 weeks old piglets at 1 and 0.1% of minimum relative abundance. (XLSX 15 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Pena Cortes, L.C., LeVeque, R.M., Funk, J. et al. Development of the tonsillar microbiome in pigs from newborn through weaning. BMC Microbiol 18, 35 (2018). https://doi.org/10.1186/s12866-018-1176-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12866-018-1176-x