Abstract
Background
Gene flow between species, across ploidal levels, and even between evolutionary lineages is a common phenomenon in the genus Arabidopsis. However, apart from two genetically fully stabilized allotetraploid species that have been investigated in detail, the extent and temporal dynamics of hybridization are not well understood. An introgression zone, with tetraploid A. arenosa introgressing into A. lyrata subsp. petraea in the Eastern Austrian Forealps and subsequent expansion towards pannonical lowlands, was described previously based on morphological observations as well as molecular data using microsatellite and plastid DNA markers. Here we investigate the spatio-temporal context of this suture zone, making use of the potential of next-generation sequencing and whole-genome data. By utilizing a combination of nuclear and plastid genomic data, the extent, direction and temporal dynamics of gene flow are elucidated in detail and Late Pleistocene evolutionary processes are resolved.
Results
Analysis of nuclear genomic data significantly recognizes the clinal structure of the introgression zone, but also reveals that hybridization and introgression is more common and substantial than previously thought. Also tetraploid A. lyrata and A. arenosa subsp. borbasii from outside the previously defined suture zone show genomic signals of past introgression. A. lyrata is shown to serve usually as the maternal parent in these hybridizations, but one exception is identified from plastome-based phylogenetic reconstruction. Using plastid phylogenomics with secondary time calibration, the origin of A. lyrata and A. arenosa lineages is pre-dating the last three glaciation complexes (approx. 550,000 years ago). Hybridization and introgression followed during the last two glacial-interglacial periods (since approx. 300,000 years ago) with later secondary contact at the northern and southern border of the introgression zone during the Holocene.
Conclusions
Footprints of adaptive introgression in the Northeastern Forealps are older than expected and predate the Last Glaciation Maximum. This correlates well with high genetic diversity found within areas that served as refuge area multiple times. Our data also provide some first hints that early introgressed and presumably preadapted populations account for successful and rapid postglacial re-colonization and range expansion.
Similar content being viewed by others
Background
Hybridization is an important driver for speciation [1], particularly in plants [2]. Frequently, hybridization is associated with polyploidization (allopolyploidization); and polyploid speciation (neopolyploidization; see [3]) events have been estimated to make up 24% of all speciation events in present-day angiosperms, with almost half of them allopolyploidizations [4]. This high frequency is thought to be linked to the advantages of (allo-) polyploids compared to their diploid ancestors. Through the loss of genetic constraints on duplicated alleles, new gene functions and gene families can evolve [5]. Many allopolyploid species are able to colonize new ecological niches, and these are not necessarily restricted to niches intermediate of their ancestors [6].
A special case of hybridization is introgression, where as a consequence of back-crossing with one of the parental species the genomic contribution of one species is increasingly larger than the other parent’s. This can result in clinal genetic and ecological differentiation [7,8,9]. With the genomic tools now available, introgression has become a popular study area, particularly where genes of adaptive value introgress preferentially. This so called ‘adaptive introgression’ has been described from a number of species in both animals [10] (e.g. Heliconious butterflies [11]; sticklebacks [12]) and plants (e.g. sunflower [13], Arabidopsis [14]; Silene [15]; Populus [16]).
The model genus Arabidopsis provides great opportunities to study hybridization and introgression, as several cases of hybridization have been documented between Arabidopsis species and even between major evolutionary lineages. Two stabilized allopolyploid species, A. suecica [17,18,19,20] and A. kamchatica [21,22,23,24] have been described. Also gene flow between different Arabidopsis species is more common [25] than previously assumed, and is found even across ploidal levels [26]. Our introgression study system was first characterized by Schmickl and Koch [7], who reported a suture zone stretching from the eastern Austrian Forealps in the south to the Danube valley in the Wachau region and further to the north-eastern border of Austria. In this area A. arenosa (hereafter called ‘Arenosa’) introgressed into A. lyrata (hereafter called ‘Lyrata’). This region is also a major center of genetic diversity for both Lyrata and Arenosa [27]. A first indication of the hybridization was given by the morphological intermediacy of many Lyrata populations in the area, and subsequent analyses showed that the respective populations are indeed exclusively tetraploid [7].
Taxonomically, the Arenosa populations in Austria can be assigned to the tetraploid mountain taxon A. arenosa subsp. borbasii, with only few populations at rural sites (railway tracks, e.g.) belonging to the tetraploid lowland taxon A. arenosa subsp. arenosa [28], both of which most likely originated through autopolyploidization [29,30,31]. In contrast, Lyrata populations in the area belong to the European subspecies A. lyrata subsp. petraea, which is generally diploid in Central Europe with few local exceptions [24, 32]. However, the introgressed tetraploid Lyrata populations represent fully stabilized polyploids. Evidence from the maternally inherited plastid genome suggests that gene flow is unidirectional and Lyrata acts as the maternal parent, while Arenosa is the pollen donor [7, 33]. Asymmetrical gene flow is frequently found among natural Arabidopsis hybrids and is often favoring Lyrata as maternal parent [32]. In the offspring of artificial crosses between diploid Lyrata and diploid representatives of the Arenosa group from the Carpathian Mountains, asymmetrical fitness has been detected and has been ascribed to cytonuclear incompatibilities [33]. However, the fitness of Arenosa maternal hybrids was found to be higher in these crosses, while in our study area Lyrata type plastids are found indicating higher fitness in Lyrata maternal hybrids [7].
It has been hypothesized that the first hybrids between Lyrata and Arenosa in the Wachau occurred within the Late Pleistocene, and subsequent secondary contact at the northern edge of the introgression zone was thought to be postglacial [7]. The allotetraploid species in Arabidopsis, A. suecica and A. kamchatica, also both likely originated during the Late Pleistocene [17, 21]; and A. suecica is most likely of postglacial origin less than 20 kya (thousand years ago) [19]. On a larger geographic scale gene flow between the two study species and even across ploidal levels has been described extensively before [7, 26], however data available to date are not appropriate to estimate temporal evolutionary scenarios of hybridization and gene flow due to limited marker resolution and evolutionary models to be applied (e.g., in case of microsatellites). Additionally, extensive gene flow between all lineages of Arabidopsis, including A. thaliana [25], suggests that reproductive isolation in Arabidopsis is rather caused by geographic isolation than genetic incompatibilities and that gene flow may have occurred even long after lineage diversification [25].
Present-day species distribution patterns and vegetation composition in Central Europe is heavily influenced by glaciation and deglaciation cycles during the Pleistocene and in particular by the last three major glaciation/deglaciation cycles with three glaciations or glaciation complexes [named Elster (480–410 kya), Saalian (380–130 kya), Weichselian (115–11.7 kya) in Northern Europe; or Mindel, Riss, Würm in the European Alps, respectively] [34]. Large parts of the Alps were covered with ice and snow during these cold periods, and plants had to retreat to refugia for periglacial survival, from where they recolonized suitable habitats in warming interglacial periods. These refugia were identified mostly at the southern, but also the eastern border of the European Alps [35]. Maximum extent of the ice sheet in the Austrian Alps during the last two glaciation complexes were reached about 20 kya in the Würm and 135 kya in the Riss glaciation [36]. It has been hypothesized that some populations of Lyrata survived the last glacial maximum (LGM, Würm glaciation) north of the Alps [37, 38], and large effective population sizes for the population in Plech, Germany as well as for the ancestral population of Lyrata have been estimated [39, 40]. The diploid Lyrata populations in the Eastern Austrian Forealps that might have served as the maternal parent for introgressed tetraploid Lyrata were also hypothesized to have survived the LGM in the cryptic refugia dominated by open rock habitats and cold-adapted vegetation they still inhabit today [7, 41].
The Arabidopsis introgression zone in the Wachau is of particular interest because of its potential to study adaptation to substrate type, increased temperature and drought stress. Diploid Lyrata are usually restricted to limestone and dolomite, whereas tetraploid Lyrata are found on siliceous bedrock, suggesting the acquisition of genes adapted to this bedrock type through introgressive hybridization with Arenosa [7].
The genetic and genomic tools available for A. thaliana are easily applicable in the remainder of the genus, thus making Arabidopsis an ideal model system for comparative evolutionary biology [42, 43]. Next-generation sequencing (NGS) techniques and high quality genomes sequences from two species of the genus, A. thaliana (TAIR: https://www.arabidopsis.org) and A. lyrata [44], allow for open questions from previous research now to be investigated in more detail.
With this study, we aim to first test the idea of a unidirectional introgression zone using genome-wide DNA analysis, we further aim to quantify the genetic levels of introgression along the presumed south-north gradient, and finally we elaborate on a spatio-temporal setting for this complex scenario of repetitive Pleistocene genetic isolation and subsequent gene flow.
Results
We analyzed whole-genome re-sequencing data of Arenosa and Lyrata from an introgression zone in northeastern Austria [7]. We included all closely related taxa from adjacent areas to set our findings into a broader evolutionary context. All of these related taxa belong to the Arenosa group and were sampled in the Carpathian Mountains, and for simplicity those will be addressed here as ‘Carpathian Arenosa’.
Spatial distribution of genetic variation and genetic admixture
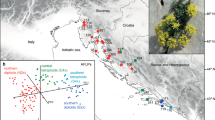
Genetic clustering was used to detect population structure and affinity to genetic clusters. The STRUCTURE analysis [45] based on nuclear genome-wide SNP calling characterizes the introgression zone. The most likely number of genetic clusters [46] was estimated to K = 2, and similarity among runs with 10 independent, randomly chosen subsets of synonymous SNPs was very high (> 0.97). The two detected genetic clusters generally represent the Arenosa and Lyrata gene pools (Fig. 1). While Carpathian diploid/tetraploid Arenosa and diploid Lyrata show no traces of admixture, tetraploid Arenosa from Austria as well as tetraploid Lyrata were partially assigned to one or the other gene pool. Only two samples of Carpathian Arenosa had a partial, although minute, genetic assignment to the Lyrata cluster (0.2% in SK08-h, A. petrogena subsp. exoleta and 2% in 915,139-g, A. petrogena subsp. petrogena). Within tetraploid Lyrata we detected a gradient of introgression from the eastern Austrian Forealps northwards towards the Danube valley in the Wachau region (Fig. 1 b, c), as had been shown previously using a small set of nuclear microsatellites [7]. The mean proportion of Arenosa genome in tetraploid Lyrata was highest in the south of this Wachau suture zone, with a population mean of 57% for pop. 137, and lowest in the northern population with a mean of 9% for pop. 37. Two tetraploid Lyrata populations from outside the introgression zone were also investigated, and both of them also showed partial assignment to the Arenosa cluster: Among them, pop. Moed exhibits an even higher proportion of Arenosa genome (12%) than the northernmost population from the introgression zone. With a genetic assignment of 7% to the Arenosa cluster, pop. 95 (Czech Republic) had an even lower affinity to the Lyrata cluster.
Genetic clustering analysis from STRUCTURE [45]. Results for mean values over 10 independent runs at K = 2 are shown. Runs are based on 10 randomly sampled subsets of the complete dataset of 15,454 genes, using a fraction of 0.05 of synonymous sites per subset. a bar plot showing the genetic assignment of each sample, sorted by taxon and population. b Mean values over all individuals as pie charts in each population in the complete sampling area. c Mean values over all individuals in each population in enlarged suture zone in northeastern Austria and adjacent regions. The Wachau region is indicated with a dashed line. Tetraploid populations are marked by black circles around the pie charts. Bedrock type was categorized based on the geological map of Austria (KM500 Austria from http://www.geologie.ac.at) under the Open Data licence Creative Commons 3.0 Österreich (CC BY 3.0 AT)
All (tetraploid) Arenosa from the study area also showed signatures of admixture with Lyrata. The southernmost populations carry the smallest amount of Lyrata in their genomes, with only 4%, while the population adjacent to the southern border of the introgression zone showed a much larger proportion of Lyrata genome with 17% (pop. 402,140), thereby also demonstrating higher admixture than the northernmost population of Lyrata in the introgression zone. However, the population adjacent to the introgression zone in the north had a much lower amount of admixture (pop. 118, with 6%). Pop. One hundred seventy six from Czech Republic, well outside the reported introgression zone, also showed a relatively high amount of admixture (12%), as did the highly endemic and high alpine taxon A. arenosa subsp. arenosa var. intermedia (pop. 210, 4%).
Results from phylogenetic network reconstruction using SplitsTree [47] (Additional file 1) were consistent with genetic assignment in STRUCTURE. Both Arenosa and Lyrata were recognized as distinct groups, with diploids at the respective tips of the network and all tetraploids in between. The network also recognized the intermediate position of both the tetraploid introgressed Lyrata and tetraploid Arenosa. Their position followed the level of admixture suggested by STRUCTURE analysis, with pops. 37, 95 and Moed close to diploid Lyrata, and more admixed populations towards the Arenosa group, and with pop. One hundred thirty seven from the southernmost border of the suture zone closest to Arenosa. Also within tetraploid Arenosa, the position towards the Lyrata group depends on the level of admixture. Pops. 402,140 and 176, that showed the highest degree of admixture in STRUCTURE analysis, were closest to each other and closest to the Lyrata group. All other samples from this taxon, as well as endemic high alpine A. arenosa subsp. arenosa var. intermedia, clustered together (Additional file 1).
Polymorphisms are shared within and between species, cytotypes and populations
Considering a scenario of an introgression zone, we expect the introgressed populations to share their polymorphism with either of the parental species. We also expect an increasing amount of polymorphism shared with one of the parental species along a clinal gradient following the suture zone from the South to the North; while the amount of polymorphism shared with the other parental source is decreasing. We assessed shared polymorphism between groups by counting the number of shared SNPs between the investigated groups, namely diploid and tetraploid Lyrata and Arenosa. The results are displayed in Venn diagrams, considering all samples (Fig. 2a) as well as tetraploid Lyrata populations along the introgression zone separately (Fig. 2b-e).
Percentage of shared polymorphism between Lyrata and Arenosa. Only biallelic SNPs were considered, and loci with missing data were ignored. Four groups were evaluated in each analysis: Diploid Arenosa, tetraploid Arenosa, diploid Lyrata and tetraploid Lyrata (A), or the respective population from the introgression zone from south to north only (B-E). a: all tetraploid Lyrata were included, b: only population 137, c: only population 85, d: only population 55, e: only population 37. f: map highlighting the populations in the introgression zone (bold). g: Trends across the introgression zone (south to north) for selected examples are shown and were taken from diagrams B to E. The map was created under the Open Data licence Creative Commons 3.0 Österreich (CC BY 3.0 AT)
In the dataset comprising all populations of tetraploid Lyrata from the area of interest (pops. 37, 55, 85 and 137), a total number of 2,505,371 biallelic SNPs or 5,010,742 alleles was analyzed. The highest total number of alleles was found within tetraploid Arenosa (78.72%), followed by diploid Arenosa (73.35%), tetraploid Lyrata (72.72%) and finally diploid Lyrata (60.42%). With approximately 52%, more than half of the alleles were shared between all four groups. The amount of private alleles was highest in tetraploid Arenosa (10.16%), followed by diploid Arenosa (8.38%). A high amount of private alleles was also detected in tetraploid Lyrata (6.37%), while diploid Lyrata had the lowest amount of private alleles (3.63%). In contrast, when considering only one of the tetraploid Lyrata populations, the amount of private alleles in tetraploid Lyrata was lower (1.47% - 2.08%).
As expected, the amount of polymorphism shared between tetraploid Arenosa and tetraploid Lyrata was decreasing (from 61.49 to 55.17%; Fig. 2b-e) following the introgression zone from south to north. However, the amount of polymorphism shared between diploid and tetraploid Lyrata did not differ significantly (varying between 54.46 and 54.79%, Fig. 2b-e). A more detailed perspective is provided with Fig. 2g, and the most severe decrease along the cline from south to north is found with uniquely shared polymorphisms between Arenosa and tetraploid Lyrata (from 6.29 to 1.67%).
Phylogenetic reconstruction based on the plastid genome provides the maternal perspective
To follow the maternal origin of tetraploid Lyrata, we reconstructed a phylogenetic tree based on the complete sequence of the maternally inherited plastid genome using maximum likelihood (ML). Capsella bursa-pastoris, Capsella rubella and Camelina sativa were used as outgroup with sister relationship to Arabidopsis [48,49,50], and from within the genus Arabidopsis A. thaliana and A. cebennensis were included as outgroup species to Lyrata and Arenosa [25]. Bootstrap values were generally high and above 95%, however the backbone of Lyrata and Arenosa clades had only low support (Fig. 3).
Maximum likelihood phylogenetic reconstruction of complete plastid genome sequences from Lyrata and Arenosa in Central Europe. Bootstrap values from 1000 bootstrap replicates are indicated (** for bootstrap support of 100% and * for support of 95–99%). Capsella bursa-pastoris, Capsella rubella and Camelina sativa were set as outgroup. Taxa are indicated with color codes at sample names, tetraploids are indicated in bold. Four clades of Lyrata are categorized with a separate color code: yellow, orange and red bars and marked a, b, c/d, respectively
The occurrence of polyploids (indicated in bold) is not restricted to one single clade, but is a recurring pattern across the entire tree (Fig. 3). This applies to both species groups, but may be more astonishing in Lyrata, as this group represents only one taxon in Europe and not several different taxa with distinct ecological niches such as Austrian and Carpathian Arenosa. The two species groups fall into three major clades (indicated in grey in Fig. 3), with one clade shared between them (clade 2). Moreover, also tetraploid Lyrata are found in two major clades (clades 2 and 3), and can furthermore be split into four deeply divergent haplotype groups (A-D in Fig. 3).
One individual from the southernmost population, 137-c, has a plastid type closely related to Arenosa pop. 201. These two populations are also geographically close, indicating recent and ongoing gene flow at the southern edge of the introgression zone. Maternal gene flow in the other direction is evident north of the introgression zone, where Arenosa from the southern Czech Republic (pop. 176) carries a Lyrata plastid type B. Also diploid populations of Lyrata can be found in two major clades, with all samples from pop. Seventy four found exclusively in clade 2. Generally, diploid Lyrata investigated here can be divided into four clades (Fig. 3). Three different populations were sampled from Lower Austria, and several individuals from Germany and the Czech Republic. While geographically close groups are not necessarily closely related, individuals within the groups form well supported monophyletic clades with no exceptions. In contrast, no such pattern could be detected in Arenosa, where neither populations nor taxa group together. Apart from the aforementioned 137-c with an Arenosa plastid, tetraploid Lyrata contains plastid types closely related to two of the four groups found in diploid Lyrata. The two populations in the northern part of the introgression zone (pops. 37 and 55) as well as the sample from Czech Republic (pop. 95) are in the clade of Czech and German diploids, and the more southern populations group with Austrian pop. 115 (with the exception of pop. 137, which has plastid types from both groups), thus showing a clear separation into northern and southern clade (indicated by the dashed line in Fig. 4b).
Divergence time estimates based on complete plastid genome sequences from Lyrata and Arenosa in Central Europe. a: Map showing plastome clades (defined for Lyrata) and Arenosa plastid types from phylogenetic reconstruction. Color codes correspond to plastome clades from Figs. 3, 4c. Dashed grey line indicates the separation of northern and southern plastome types. b: Detailed zoom of map shown in A with information on snowline and ice sheet extensions during the Last Glacial Maximum (20 kya), and refuge area for plant species indicated according to different vegetation zones [35]. Dashed grey line indicates the major biogeographical separation of northern and southern plastome types. The map was created under the Open Data licence Creative Commons 3.0 Österreich (CC BY 3.0 AT). c: Combined BEAST results from two independent MCMC runs of 1*108 generations each. Secondary calibration was performed using the results from Hohmann et al. (2015) [3] for the split of the outgroup genera, Capsella and Camelina, as well as for the Arabidopsis crown age and root height. Divergence time estimates are shown with their 95% HPD intervals, only node ages ≥0.05 mya are given. Taxa are indicated with color codes at sample names, tetraploids in bold. Four clades of Lyrata are indicated with yellow, orange and red bars (a, b, c, and d)
Divergence time estimations indicate late Pleistocene differentiation
Divergence time estimation was based on the complete plastid genome and conducted in BEAST [51]. The topology of the resulting tree (Fig. 4) is consistent with the results from ML phylogenetic reconstruction, with differences only at splits with low bootstrap support in ML analysis. The crown age of outcrossing Arabidopsis, represented here by the split between A. cebennensis and the clade of Arenosa and Lyrata, was estimated to 0.66 million years ago (mya), and therefore is predating the last three glaciations [Mindel, Riss, Würm; corresponding to Elster (480–410 kya), Saalian (380–130 kya), Weichselian (115–11.7 kya) in Northern Europe]. The onset of lineage diversification follows shortly thereafter, at 0.55 mya, and within 100 ky the three major plastid lineages split (Fig. 4). Few taxonomically or geographically meaningful deep splits of about 300 kya exist; but most splits are more recent (<140 kya) and most likely peri- and postglacial. Divergence between the four clades in Lyrata is deep, with pop. 74 (clade A) having split from the other 0.55 mya (million years ago) before the Mindel glaciation. However, here chloroplast capture of an Arenosa-plastid type or incomplete lineage sorting might have occurred to explain this finding. Pop. 88 (clade D) has a split time placed in the Early Riss glacial period 0.31 mya. The two clades consistently containing tetraploids (clades B and C) split 0.16 mya during Late Riss glaciation, but before the maximum extent of the ice sheets 135 kya [36]. The deepest split within the group north of the Alps (clade B) is only 0.09 mya, while the crown age of the southern group (clade C) is 0.14 mya, also set during the cold period of Riss glaciation, shortly after divergence from clade B. Very recent divergence between plastid genomes found in Arenosa and Lyrata can be observed in two cases. The split of tetraploid Lyrata sample 137-c from all individuals of pop. Two hundred One from Arenosa is younger than 50,000 years. And the split of the Czech Arenosa sample 176R-03 from its next relative, a plastid type found in diploid Lyrata from Germany is of about 60,000 years of age. Both of these cases are indicative of recent (possibly postglacial) gene flow and plastid capture between the Arenosa and Lyrata lineage. It is worth to mention that also the plastome split between Capsella rubella and Capsella bursa-pastoris (1.57 mya, 95% confidence interval 0.98–2.31 mya) correlates well with a species split-time estimates (minimum age) of about 1 mya based on whole nuclear genome data and using coalescent models [52].
Discussion
Genetic clustering indicates an enlarged suture zone
Genetic cluster analysis significantly shows the introgression of Arenosa into tetraploid Lyrata in our study area (Fig. 1). The amount of genetic contribution of Arenosa is highest in the Eastern Austrian Forealps in the south of the suture zone described earlier from the lowlands [7] and is represented by pop. 137. This is also indicated by SplitsTree analysis, where this population is closest to the Arenosa samples (Additional file 1). Following the sampled populations northwards, the amount of genomic contribution of Arenosa is decreasing, and SplitsTree places the northernmost pop. Thirty seven closest to diploid Lyrata. This is in line with patterns previously reported and described using few microsatellite markers [7]. However, STRUCTURE results show that the extent of introgression is much larger than previously known and expected. Gene flow between Arenosa and Lyrata is possible on the tetraploid level, while none are found in diploids [26, 53]. A congruent result was found here. In STRUCTURE analysis, diploids showed no signs of admixture, with the exception of one Carpathian Arenosa; however, the amount of Lyrata genome in this individual is very low with only 2%. In Lyrata, two tetraploids from outside the introgression zone were investigated, and both of them show traces of the Arenosa gene pool. The population close to Vienna (pop. Moed) was known to be tetraploid before, and had been investigated in several studies [26, 54]. The other population in the Czech Republic however was never studied in detail, and was shown recently to be tetraploid [28]. The amount of Arenosa genome in both these populations is surprisingly high since it was assumed that these populations are typical A. lyrata (e.g. based on morphology, [7]), and with 12%, the population from Mödling even has a higher amount than that found in the northernmost population of the introgression zone. Interestingly, the morphological intermediacy between Arenosa and Lyrata that has been reported for the introgression zone [7] cannot be found in Mödling, and the ecology of the population, growing on rocky calcareous outcrops and old limestone walls is very different from that of the introgression zone and is closest to diploid Lyrata; therefore and because of similar ecological niches of diploid Lyrata and tetraploid populations from Mödling an autotetraploid origin had been assumed previously. However, both STRUCTURE results and SplitsTree network reconstruction strongly suggest gene flow from Arenosa into this population as well, thus indicating an enlarged suture zone (including regions around pops. 95 (Czech Republic) and Mödling) and providing first evidence for a more complex spatio-temporal evolutionary scenario.
Divergence time estimates indicates inter- and periglacial diversification
Some geographically close populations of diploid Lyrata in the Eastern Austrian Forealps show deep divergence in plastid genome phylogenetic reconstruction. The two south-easternmost populations, pops. Eighty eight and 74, have plastid types divergent from all other Lyrata plastids, and divergence time estimates of 0.31 mya and 0.55 mya, respectively. SplitsTree network analysis based on the nuclear genome data also places these two populations the furthest from the introgression zone, although data from the nuclear genome place them closely together (Additional file 1). However, the occurrence of diploid Lyrata in this region is often restricted to cryptic refugia with open rocks and cold-adapted relic vegetation types, and survival during the last glaciation cycles in these refuge areas has been previously hypothesized [7]. The isolated positions of pops. 88 and 74 in the chloroplast phylogenetic reconstruction as well as in SplitsTree network confirm this hypothesis, and coincides with potential refuge areas for mountain plants on calcareous bedrock in the area in question with sufficient large effective population sizes to maintain high genetic diversity [55] (Figs. 1, 4). Split times among plastid genomes and its correlation with the last four glaciations and respective maximum extent of the ice sheet (if known) is shown with Fig. 5 (detailed blow-up of Fig. 4). It is remarkable that the vast majority of splits in the plastome phylogeny is placed into glaciation periods maybe indicating populations of related species forced into ‘melting suture zones’. The crown group age of tetraploid Lyrata is 0.16 mya (with the exception of sample 137c that carries an Arenosa type plastid) (Figs. 1, 5). This coincides with the ending of the Riss glacial or beginning of the last interglacial period, indicating a first occurrence of tetraploid Lyrata around this time. Ice extent of the glaciers in Eastern Austria at that time reached far into the Foreland [36], and we can hypothesize that species were forced into genetic contact at lower elevations. Signatures of past diversification and contact of gene pools are also evident from the distribution patterns of plastid genetic diversity and are highest in the montane refuge areas of the northeastern Alps (Fig. 4; south of the indicated dashed line). This montane refuge consists of different bedrock types, often changing over short distances and thereby providing the chance for limestone adapted Lyrata and siliceous bedrock adapted Arenosa to come into close genetic contact.
Divergence time estimates based on complete plastid genome sequences from Lyrata and Arenosa in Central Europe. Detail from Fig. 4 with focus on the last ~700 kya. Divergence time estimates are shown with their 95% HPD intervals. Taxa are indicated with color codes at sample names, tetraploids in bold. Four clades of Lyrata are indicated with yellow, orange and red bars (a, b, c, and d). Glaciation complexes are indicated in lighter blue, maximum extent of the ice sheets during Würm and Riss glaciation in dark blue lines
The majority of splits in the plastome tree (Fig. 5) is found during the Würm glaciation indicating an increased net diversification rate. Presumably, effective population sizes were large and allowed to maintain significant proportions of the genetic variation. This has been described earlier for Lyrata populations from northern Bavaria in Germany (exemplified here with pops. 112 and 133) with a split time from Bohemian populations calculated with 39,000 generations (minimum of 78,000 years) and also placed completely into Riss glaciation with effective population sizes of approximately 175,000 individuals [40, 56] estimated an origin of autotetraploid Arenosa populations in the Eastern Alps also to 11,000–30,000 generations ago. Divergence between A. halleri and A. lyrata, representing the timing of rapid diversification of the major lineages within Arabidopsis, was estimated to have occurred 337,000 years ago [57]. Such studies give valuable insights into coalescent-based estimates relevant for the introgression zone, as the establishment of hybrids must postdate the origin of their diploid ancestors. However, compared to the plastid perspective, these estimates are younger (see also Table 1 for a comparison of divergence time estimates in Arabidopsis). This could be due to the non-recombining nature of plastid genomes that might lead to overestimated divergence times, in particular when the number of samples is low and coalescent analyses could underestimate divergence in the presence of gene flow, which was reported in high amounts in Arabidopsis [25]. Considering the limitations from both approaches however, the first establishment of tetraploid (and introgressed) Lyrata is still likely to have occurred either during the last alpine interglacial period (Riss-Würm interglacial: around 120 kya) or maybe during or around the maximum glaciation of the Riss glaciation (135 kya) with glaciers reaching into the Forelands and thereby also forcing plant populations of related species into ‘melting suture zones’. Two cases of secondary contact with intrapopulational plastid type variation are apparent from phylogenetic reconstruction and BEAST analysis at the extended borders of the introgression zone: (1) tetraploid Arenosa pop. 176R-03 from southern Czech Republic carries a plastid type closely related to diploid Lyrata from Germany (Lyrata type B), and (2) tetraploid Lyrata sample 137-c from the southernmost population of the introgression zone carries a putative Arenosa plastid type, while other representatives of this population carry a typical Lyrata plastid type. This suggests that sample 137-c might have acquired a plastid from Arenosa through chloroplast capture. Both of these instances are relatively recent (peri- or postglacial), and indicate that gene flow between major evolutionary lineages in Arabidopsis is an ongoing process rather than a concluded event (or series of events) from the past.
Directional gene flow and the origin of the hybrids
Directional gene flow between Arabidopsis species seems to be not unusual and has been described in hybrid A. suecica (maternal A. thaliana X paternal A. arenosa) and A. kamchatica (A. lyrata X A. halleri), both of hybrid origin, and introgressed A. lyrata (maternal A. lyrata X paternal A. arenosa) [7, 23, 24, 27, 32]. Arabidopsis kamchatica evolved several times independently (polytopic origins) and maternal contribution depends on the regions considered (e.g., A. lyrata as maternal parent in China, A. halleri as maternal parent in Japan). Combining these few examples there is some evidence that A. lyrata more often serves as maternal parent in introgression and hybridization scenarios, whereas A. arenosa plays a prominent role as paternal donor. Exactly this situation has been introduced in a suture zone along the Danube valley in the Wachau region in Lower Austria [7] and has been significantly demonstrated herein (Figs. 1, 4). Predictions from elevated mutational rates of plastid evolution in Lyrata when compared to Arenosa suggest that maternal Lyrata hybrids should have a lower fitness because of potential cytonuclear incompatibilities [33], but the opposite seems to be true in this natural introgression zone. Almost all tetraploid introgressed Lyrata have a plastid type belonging to Lyrata, indicating Arenosa usually acts as a pollen donor in the wild. This indicates that strong postzygotic and exogenous factors (e.g., climate and soil composition) act on first generation hybrids, thereby counteracting endogenously caused fitness constraints. One exception to this observation however could be in pop. One hundred thirty seven in the south of the introgression zone, where sample 137-c has a plastid type closely related to the neighboring pop. Two hundred one from Arenosa. The recent divergence suggests secondary contact of these taxa after the last glacial maximum in the Holocene. Although cytonuclear incompatibilities have been hypothesized to be the cause for the asymmetry of fitness in Arenosa and Lyrata crosses [33], maternal effects on gene expression might also play a role in the fitness asymmetry observed. In intraspecific A. lyrata hybrids, a strong bias towards the expression of maternal alleles was detected [61]. These effects could be mitigated by the high amount of introgression in the genome in pop. One hundred thirty seven (~57% of Arenosa, as indicated by STRUCTURE analysis), thus allowing for gene flow also in the reverse direction.
On a smaller scale gene flow from Lyrata to Arenosa can be observed in several populations, although putatively followed by backcrossing to Arenosa, not Lyrata, as indicated by plastid types. Results from STRUCTURE show that not only tetraploid Lyrata, but also the nearby populations of Arenosa show traces of introgression. This has not been shown before based on microsatellite loci used by Schmickl and Koch [7], and the extent observed in this study is somewhat surprisingly high. All populations of Arenosa from the study area investigated here show traces of Lyrata genome. While partial genetic assignment to the Lyrata cluster had been detected for populations directly adjacent to the introgression zone before, more specifically in the area around Mariazell in the eastern Austrian Forealps in the south and in the Wachau region at the north shore of the Danube river in the north [7], the results in this study indicates also partial assignment of an Arenosa population in the southern Czech Republic to Lyrata. Moreover, admixture was detected south of the introgression zone as far as to the distribution area of small range endemic and high alpine taxon A. arenosa subsp. arenosa var. intermedia. (pop. 210; Fig. 1). Tetraploid Carpathian Arenosa show no assignment to the Lyrata cluster at all, thus making polyploidy as the only explanation rather unlikely. Nevertheless, the traces of Lyrata genome found in high-alpine A. arenosa subsp. arenosa var. intermedia are surprising, considering the highly endemic and isolated distribution of this taxon in formerly alpine refugia.
Our findings with remarkable high amounts of Lyrata genome reported here from all populations of Arenosa within our total study area in Austria stand in sharp contrast to a recent study of Arenosa in Austria also based on whole genome sequence data, which found introgression from Lyrata into Arenosa only in one single serpentine adapted population [62]. Furthermore, analysis of shared polymorphism indicates a high amount of unique variants in tetraploid Lyrata (Fig. 2), and analyses of reduced datasets including only one population of this taxon show that only part of this variation is shared between introgressed populations. The number of private alleles in tetraploid Lyrata is surprisingly high, considering that they allegedly arose through hybridization of diploid A. lyrata subsp. petraea with A. arenosa subsp. borbasii, both of which are well represented in our analyses. A recent hybridization event should result in a low number of private alleles in hybrids, as they recruit all their genetic variation from the parental species. The high amount might also suggest that the hybridization occurred long ago in the “southern” montane refugia and diversity hotspot (Fig. 4), and therefore new variants could accumulate in the populations and afterwards migrated north towards the Danube river system. Alternatively, we might argue that the parental species were not fully recovered in our sampling, e.g. that populations now extinct served as parents or that the other species of Arabidopsis that is present in Austria, A. halleri, also played a role in the origin of tetraploid Lyrata. Considering the high levels of gene flow between all lineages of Arabidopsis [25], this does not seem unlikely. A comprehensive survey of Novikova et al. 2016 [25] did not show extensive gene flow between A. lyrata and A. halleri, but a preliminary population-based microsatellite study (Koch et al., unpublished) exemplified various A. halleri populations that are introgressed with A. arenosa, and we might assume that via this “bridge” A. halleri alleles might be able to enter even the genome of tetraploid A. lyrata.
Introgression and its potential for adaptive evolution – A perspective
We have shown herein that there was a primary introgression from Arenosa into a maternal Lyrata genomic background in the northeastern Forealps around the maximum glaciation of the Riss glaciation (135 kya) with glaciers forcing populations of Arenosa and Lyrata into ‘melting suture zones’ at lower elevation. These regions are also of intermediate and/or mixed bedrock types and might have provided the environmental heterogeneity to enable introgressed populations to adapt to the new environment. These populations have served as genetic source for the subsequent colonization of lowland regions towards the Danube river (Wachau). This finding also correlates with the fact that tetraploid Lyrata populations in the Wachau are found on siliceous and intermediate bedrock types (Fig. 1c). Consequently one might have to consider adaptive evolution acting on substrate specific genes as shown earlier for bedrock type adaptation in North American Arabidopsis lyrata [63]. Evidence from the maternally inherited plastid genome suggests that gene flow was effectively unidirectional with Lyrata as the maternal parent. This was explained partially with cytonuclear incompatibilities [33]. However, it is also likely that we have to consider trade-offs in ecological traits in introgressed populations. Arenosa is a short-lived biennial and mostly single rosette-forming plant, which is adapted to produce many seeds with high immediate germination rates enabling the species to colonize rapidly disturbed habitats. In contrast, European Lyrata is a long-lived perennial with creeping stolons. Its flowering season is prolonged and seed production is limited with seeds of delayed germination. Consequently, introgressed and colonizing Lyrata populations might combine different parental traits, which are under respective selective regimes. However, the reason for the preference of Lyrata as maternal background in tetraploid populations is unclear, and maybe this is related to intrinsic processes such as stabilizing the polyploid genome and enabling regular meiosis [29,30,31]. We have set-up reciprocal crossing experiments with most of the populations and are analyzing respective ecological key traits in the offsprings, that can be discussed within a reliable genomic context and an evolutionary spatio-temporal scenario.
Conclusions
Here we present a first detailed analysis of an Arabidopsis introgression zone in Austria based on genome-wide polymorphism data. The complex and highly dynamic system of past and ongoing gene flow between A. lyrata subsp. petraea and A. arenosa subsp. borbasii provides a great opportunity to study a number of evolutionary processes, such as adaptive introgression, asymmetrical fitness and periglacial isolation of populations in detail. The focus of our study lies on describing the population structure of the study system. Results from previous studies were significantly confirmed, and a surprisingly high amount of introgression, including from outside the originally described introgression zone, was detected. This adds to recent evidence suggesting that in Arabidopsis restrictions to gene flow are caused by geographic barriers rather than by genetic incompatibilities and past inter-lineage gene flow occurred frequently. Using chloroplast phylogenomics and corresponding divergence time estimations, our study strongly suggests a late Pleistocene origin of tetraploid introgressed Lyrata in the Eastern Austrian Forealps, most likely during the last inter- and periglacial period. The hybrids subsequently expanded northwards and colonized the entire introgression zone in the lowland Danube river system, with secondary contact to Arenosa possibly occurring after the LGM in the North close to the Bohemian Massif..
Methods
Plant material and sampling strategy
Focusing on an inter-species introgression zone in Eastern Austria stretching from the Northeastern Forealps to the Danube river we investigated 71 accessions in total representing samples of the Arenosa and Lyrata lineage [25]. Based on previous results [7, 27] transects were sampled representing clinal variation but also potential centers of genetic diversity and glacial refugia. The sampling also included individuals from Southeastern Germany, Czech Republic and Slovakia to set the study system into a broader phylogeographic and evolutionary context. The final dataset included diploid and tetraploid Lyrata, introgressed tetraploid Lyrata, various tetraploid Arenosa as putative parental source and diploid species from the Arenosa lineage representing the ancestral diploid Arenosa genepool. Sampling is displayed in Fig. 6 and further details are given with Additional file 2. Vouchers have been prepared and are deposited with the herbaria in Heidelberg (HEID) and Zurich (UZH). Details are provided with Additional file 2.
Phylogenetic analyses included Arabidopsis cebennensis as closest relative to Lyrata and Arenosa as outgroup taxon [28]. For plastome analyses sequences of Arabidopsis thaliana and Capsella bursa-pastoris were retrieved fromNCBI/Genbank and from a recent study focusing on the Brassicaceae family (Hohmann et al., 2015 [3]; Capsella rubella and Camelina sativa).
DNA isolation
Total genomic DNA was extracted from herbarium vouchers, silica dried or fresh leaf material either following the CTAB protocol [64] or using the Invisorb Spin Plant Mini kit (STRATEC Biomedical AG, Birkenfeld, Germany). Fresh material was collected in the Botanical Garden Heidelberg and immediately frozen in liquid nitrogen until further use. Dried leaf material was immediately silica dried or taken from herbarium vouchers. Depending on leaf size, one to three leaves were used per extraction. Material was homogenized either by grinding with a pistil on liquid nitrogen, or using a Precellys®24 homogenizer (Bertin Technologies, Montigny-le-Brettonneux, France) at 5000 rounds per minute for two times 10 s with five 2.5 mm glass beads. The Invisorb Spin Plant Mini kit was used following the manufacturer’s instructions, including the optional step of RNA digestion. The following modifications were applied to the CTAB protocol: DNA pellets were washed twice with 70% ethanol and then dissolved in 100 μl TE-buffer (10 mM Tris-HCl, 1 mM EDTA, pH 7.5) supplemented with 2 units of RNase A, RNA digestion was performed at 37 °C for 1 h. Quality (high molecular weight) and quantity (> 5 ng/μl) of DNA was checked rigorously prior to library preparation. DNA quality and fragment length was checked on 1% agarose gels, and concentration was assessed via fluorescence spectroscopy using a high-sensitivity, double-stranded DNA specific dye with the Qubit® dsDNA HS Assay (Thermo Fisher Scientific, Waltham, Massachusetts, USA).
Library preparation and next-generation sequencing
Libraries were sequenced at the Deep Sequencing Core Facility, University of Heidelberg, and were prepared from total genomic DNA with the Illumina TruSeq Kit (Illumina, Inc., San Diego, California, U.S.) in paired-end mode with insert size 200–400 base pairs (bp). Read length was 100 bp. Samples were barcoded for multiplexing, and on average 4 tetraploid or 8 diploid individuals were sequenced on one lane to obtain intermediate sequencing depth. Sequencing was conducted on Illumina HiSeq2000 (Illumina, Inc.).
For details regarding library preparation method, see Additional file 2.
Assembly and annotation of plastid genomes
Assembly of plastid genomes generally followed the procedure described in Hohmann et al. (2015) [3] and Novikova et al. (2016) [25]. To recover the complete plastome sequence the reads were assembled de novo in CLC Genomics workbench version 6.0.4 (CLC bio, Aarhus, Denmark). Failed reads were discarded upon import, and imported reads were trimmed for sequencing quality as well as for remaining adapter sequences. A minimum per-base quality of 0.001 (corresponding to a phred-based score of 30) was applied, with a minimum remaining length of 50 bp. Only paired reads were used further.
The legacy version of the CLC Genomics de novo algorithm was used with distance settings matching the insert size of sequencing libraries. Sequences were required to have a sequence similarity of 0.9 over a length fraction of 0.9 for assembly, and the default parameters were used for mismatch, insertion and deletion cost (2, 3, and 3, respectively). Random assignment was used for non-specific matches, and conflicts were resolved by voting. An automatic word size was applied, and minimum contig length was generally 2000 bp, but adjusted depending on read number to ensure sufficient computational resources. Reads were mapped back to contigs to achieve an accurate assembly and be able to detect inconsistencies. Contigs belonging to the plastid genome are easily identified by their high coverage, as plastids have a high copy number in each cell, comparable to that of nuclear encoded ribosomal DNA and approximately 10 times higher than that of the mitochondrial genome [65]. The browser version of NCBIs nucleotide blast was used to confirm the identification as plastid sequence with default settings and detect the sequence direction. These were manually aligned to a reference in PhyDE version 0.9971 [66]. Already assembled sequences were used as a reference, and since here only additional individuals from already known populations were sequenced, the already assembled genomes from respective populations were used. Contigs often overlapped, but where they did not the sequence of the reference was inserted into a pseudo genome. This in turn was used as a reference for mapping in a round of quality control. The legacy version of the CLC Genomics mapping algorithm was used with the same settings as for de novo assembly to map all reads back to the hybrid sequence of reference and newly assembled contigs. Subsequently, Variant Detection was conducted in CLC Genomics to identify differences between reads and reference, ignoring non-specific matches and broken pairs. To find all differences such as SNPs and short indels, the minimum coverage was set to 1 and variant probability to 0.1. As the plastid genome is haploid, the maximum expected number of variants was 1. Long indels, which will not be identified by Variant Detection, were located manually. If variants were found, the newly assembled genome sequence was adjusted and the mapping as well as variant detection steps were repeated until no further differences could be detected.
Sequences were annotated in Geneious version 7.1.7 (Biomatters Ltd., Auckland, New Zealand). This was achieved by aligning the newly generated plastid genome sequences to the A. thaliana reference (Genbank accession AP000423) using the automatic alignment parameters, and then transferring the annotations. Annotations were checked manually for the correct assignment of start- and stop codons, and annotation features were adjusted if needed.
Alignments of plastomes
Complete plastid genomes were aligned using the MAFFT version 7.017 [67–69] plugin in Geneious v7.1.7 (Biomatters Ltd., Auckland, New Zealand) with the FFT-NS-ix1000 algorithm. The default parameters 200 PAM/k = 2 scoring matrix, gap open penalty of 1.53 and offset value of 0.123 were used. C. rubella, C. bursa-pastoris and C. sativa were included as outgroup to the genus Arabidopsis, and A. thaliana as well as A. cebennensis were included as relatives to the Arenosa and Lyrata. Subsequently, the alignment was cut into segments representing single exons, introns and intergenic spacers. Regions with two overlapping annotations were used only once. The segments on the Inverted Repeat were included only once, resulting in a dataset of 265 alignments. All alignments were realigned using the stand-alone version of MAFFT version 7.123b [69], with the iterative refinement method with local pairwise alignment information (command line arguments: --localpair --retree 6 --maxiterate 16), and gapped positions were excluded using Gblocks version 0.91b [70] with default block settings, except for minimum block length, which was set to 2 bp. Seven alignments were excluded because of their block length. These were generally the short sequences sharing two annotations. The remaining 257 alignments were used for further analysis.
PartitionFinder version 1.1.1 [71, 72] was used to find an optimal partitioning scheme and evolutionary model for the alignments. Only models implemented in BEAST were tested, with branch lengths unlinked and BIC for model selection in a greedy search. The best partitioning scheme consisted of 2 subsets. One subset contained mostly coding regions as well as all segments from the Inverted Repeat region, with best model GTR + Γ, possibly caused by the slower substitution rate of the Inverted Repeat region [73] and coding regions. The second subset contained mostly intergenic regions and introns, with best evolutionary model GTR + Γ + I. For details on partitioning scheme see Additional file 3. Alignments are available with Additional file 4.
Phylogenetic reconstructions
Maximum likelihood (ML) trees of plastome sequence data were reconstructed using RAxML version 8.1.16 [74]. The partitioned dataset was used as input, with individual per partition branch length optimization, for a rapid bootstrap analysis with 1000 bootstrap replicates followed by a thorough ML search. RAxML does not allow for individual per partition evolutionary models, therefore GTRGAMMA was used for both partitions, also following the recommendation of the RAxML user manual. The clade of C. sativa, C. rubella and C. bursa-pastoris was set as outgroup. Output was visualized in FigTree version 1.4.1 [51].
Divergence time estimations
Divergence times of plastome lineages were estimated using BEAST version 1.7.5 [51]. The partitioned alignments according to the best partitioning scheme as described above were used, with unlinked substitution and clock models and a combined tree. GTR + G and GTR + G + I were used as substitution models for the two subsets, respectively, with 4 gamma categories. The clock model was uncorrelated log normal relaxed clock [75] with starting rate 1.0 × 10−4 for both subsets, and estimated rates. Tree prior for the combined tree was Speciation: Birth-Death Incomplete Sampling [76]. A user-defined starting tree was generated by converting the ML tree into a chronogram and adjusting node ages to fit the priors in R version 3.2.3 [77] using the function ‘chronos’ included in the R package APE version 2.13 [78].
Secondary calibration was applied following Novikova et al. (2016) [25] and using the estimates from Hohmann et al. (2015) [3]. The split between A. thaliana and the rest of Arabidopsis, i.e. the Arabidopsis crown age, was set to 5.96 mya, and the split of the two outgroup genera, Camelina and Capsella, was set to 7.35 mya [3]. A normal distribution with standard deviation 1.0 was used for both of these calibration points, as this was a good fit with the 95% HPD intervals from Hohmann et al. 2015 [3]. The tree root height, i.e. the Arabidopsis stem age, was constrained using a normal distribution with a mean value of 8.16 mya and standard deviation of 1.0.
Two independent MCMC chains were run for 1 × 108 generations, sampling parameters every 1 × 104 generations. Subsequently, the two analyses were combined in LogCombiner version 1.7.5 [51], discarding the first 1 × 107 generations of each run as burn-in, and the resulting 18,000 trees were combined to a maximum clade credibility tree with median node heights in TreeAnnotator version 1.7.5 [51]. The tree including the 95% HPD intervals of divergence time estimates was visualized in FigTree version 1.4.1 [51].
The combined tree file including 95% HPD intervals for all nodes are available in Additional file 5.
Mapping of the reads form the nuclear genome
Reads of Arenosa and Lyrata samples were mapped against the A. thaliana reference genome (TAIR: https://www.arabidopsis.org). The also available genome sequence of A. lyrata was not used in order to prevent a bias of better mapping of one given species to an ingroup reference. Mapping was performed using the BWA-MEM [79] algorithm from BWA version 0.7.8 [80] with an increased penalty for unpaired read pairs to 15 and default settings for all other parameters. Duplicate mapped reads were then removed with SAMtools version 0.1.19 [81, 82]. Indel realignment was performed with IndelRealigner from GATK version 3.3 [83], and the alignment was filtered for primary and unique reads with SAMtools (flags –F 256 –q 5). Since the sampling included diploid as well as tetraploid individuals, variant calling could not be performed for all samples simultaneously. Therefore, variants were first called for individual samples in GATKs Unified Genotyper for both SNPs and indels for all confident sites [84], and subsequently combined using GATKs CombineVariants with filtered sites treated as uncalled. In addition to quality filters applied by default by GATK we used only sites of intermediate coverage, thereby excluding potential duplicated regions and pseudo-genes. Coverage distribution was calculated with GATKs Pileup, and coverage density was calculated. GATKs CallableLoci tool was used to find intervals with a minimum coverage corresponding to the ploidy level of the sample and a maximum coverage of 0.95 of the coverage density, and variant calling was performed only in intervals that fulfilled the criteria for coverage.
After combining the variants, a set of 15,454 genes was extracted with the intersect function of the BEDTools program collection [85]. These genes were already used in Novikova et al. (2016) [25] as they were well covered in a data set of genomic sequence from all taxa within the genus Arabidopsis, and they were proven to be single-copy using the same coverage criteria as described above. This resulted in a subsampling of the complete genome comprising 39,046,724 bp including 3′- and 5′-UTR as well as introns, of which 7,581,148 had SNPs in the dataset. Only SNPs were used for downstream analyses, and only variant sites were selected. An additional filtering step for sites with 50% missing data or more, or with frequency 1 for the derived allele, i.e. invariant sites, resulted in 5,473,967 sites, out of these 1,414,525 are synonymous SNPs. Annotations were added using SnpEff version 4.2 [86].
Genetic clustering analysis
Clustering analysis based on nuclear genome data was conducted only on a subset of data. This was necessary to rescale to a computationally feasible dataset, and was done by randomly choosing a fraction of 0.05 of synonymous variants from the original file using GATKs SelectVariants. Ten independent datasets were generated, and used as input for STRUCTURE analyses. Since the function ‘fraction’ uses a probabilistic approach, the number of SNPs varies between input files (between 70,383 and 70,475). To include diploid and tetraploid individuals into the same analysis, diploid data was duplicated similar to the procedure described in Hohmann et al. 2014 [28].
STRUCTURE version 2.3.4 [45, 87] was run with allele frequencies correlated in the admixture algorithm. Ten thousand MCMC steps were discarded as burn-in, and followed by 20,000 iterations for K = 1–6 (number of clusters). Instead of 10 independent runs for each K, as is commonly done when using STRUCTURE, the 10 subsets of data were used as input for different runs.
To demonstrate that the genetic assignment of individuals is robust to the random subsampling performed here, CLUMPP version 1.1.2 [88] was used to find the matching permutation for each run. Input files for CLUMPP were generated with the stand-alone version of structureHarvester version 0.6.94 [89]. The R script structure-sum [90] was used to calculate similarity between runs as well as determine the best number of clusters according to the method described by Evanno et al. (2005) [46]. Results from structure-sum are shown in Additional file 6. Distruct version 1.1 [91] was used to plot mean STRUCTURE results for the optimal number of clusters.
Shared polymorphism among taxa and cytotypes
To determine the number of shared polymorphisms within the nuclear data, four groups were compared, namely diploids and tetraploids of Lyrata and Arenosa, respectively. Tetraploid Lyrata from outside the introgression zone (pops. 95 and Moed) were not considered in this analysis. In addition, the same analyses were conducted using subsets of tetraploid Lyrata comprising only samples of one of the four populations from the introgression zone. Only biallelic SNPs were selected, and sites with missing data were discarded. The amount of alleles shared between groups was counted and Venn diagrams were drawn in R version 3.2.3 [76] using the library VennDiagram version 1.6.17 [92], and further details are shown with Addiitonal 7.
SplitsTree analysis of nuclear data
The full set of variants in the 15,454 genes, over all samples, with filtering for more than 50% missing data as described above, i.e. 5,473,967 SNPs, was used for network reconstruction. Here also multiallelic SNPs could be used. Fasta files were generated in GATK using FastaAlternateReferenceMaker [82]. IUPAC ambiguity codes were inserted for heterozygous SNPs. A phylogenetic network was reconstructed in SplitsTree version 4.14.2 [47] using the equal angle NeighborNet with 1000 bootstrap replicates.
Change history
11 April 2018
Upon publication of the original article [1], the authors had flagged that there was an error in Fig. 1c, as the key in this figure was displaying incorrectly. The colours had not displayed in the key in the final published article, and instead appear as plain white.
Abbreviations
- Kya:
-
Thousand years ago
- ML:
-
Maximum likelihood
- Mya:
-
Million years ago
References
Mallet J. Hybrid speciation. Nature. 2007;446:279–83.
Rieseberg LH, Willis JH. Plant speciation. Science. 2007;317:910–4.
Hohmann N, Wolf EM, Lysak MA, Koch MA. A time-calibrated road map of Brassicaceae species radiation and evolutionary history. Plant Cell. 2015;10:2770–84.
Barker MS, Arrigo N, Baniaga AE, Li Z, Levin DA. On the relative abundance of autopolyploids and allopolyploids. New Phytol. 2016;210:391–8.
van den Bergh E, Hofberger JA, Schranz, ME. Flower power and the mustard bomb: comparative analysis of gene and genome duplications in glucosinolate biosynthetic pathway evolution in Cleomaceae and Brassicaceae. Am J Bot 2016, 7:1212–1222.
Blaine Marchant D, Soltis DE, Soltis PS. Patterns of abiotic niche shifts in allopolyploids relative to their progenitors. New Phytol. 2016;212:708–18.
Schmickl R, Koch MA. Arabidopsis hybrid speciation processes. Proc Natl Acad Sci U S A. 2011;108:14192–7.
Canestrelli D, Bisconti R, Nascetti G. Extensive unidirectional introgression between two salamander lineages of ancient divergence and its evolutionary implications. Sci Rep. 2014;4:6516.
Harrison RG, Larson EL. Heterogeneous genome divergence, differential introgression, and the origin and structure of hybrid zones. Mol Ecol. 2016;25:2454–66.
Hedrick PW. Adaptive introgression in animals: examples and comparison to new mutation and standing variation as sources of adaptive variation. Mol Ecol. 2013;22:4606–18.
Pardo-Diaz C, Salazar C, Baxter SW, Merot C, Figueiredo-Ready W, Joron M, McMillan WO, Jiggins CD. Adaptive introgression across species boundaries in Heliconius butterflies. PLoS Genet. 2012;8:e1002752.
Lucek K, Lemoine M, Seehausen O. Contemporary ecotypic divergence during a recent range expansion was facilitated by adaptive introgression. J Evol Biol. 2014;27:2233–48.
Whitney KD, Randell RA, Rieseberg LH. Adaptive introgression of herbivore resistance traits in the weedy sunflower Helianthus annuus. Am Nat. 2006;167:794–807.
Castric V, Bechsgaard J, Schierup MH, Vekemans X. Repeated adaptive introgression at a gene under multiallelic balancing selection. PLoS Genet. 2008;4:e1000168.
Minder AM, Widmer A. A population genomic analysis of species boundaries: neutral processes, adaptive divergence and introgression between two hybridizing plant species. Mol Ecol. 2008;17:1552–63.
Suarez-Gonzalez A, Hefer CA, Christe C, Corea O, Lexer C, Cronk QCB, Douglas CJ. Genomic and functional approaches reveal a case of adaptive introgression from Populus balsamifera (balsam poplar) in P. trichocarpa (black cottonwood). Mol Ecol. 2016;25:2427–42.
Jakobsson M, Hagenblad J, Tavare S, Sall T, Halldén C, Lind-Halldén C, Nordborg M. A unique recent origin of the allotetraploid species Arabidopsis suecica: evidence from nuclear DNA markers. Mol Biol Evol. 2006;23:1217–31.
Mummenhoff K, Hurka H. Subunit polypeptide composition of rubisco and the origin of allopolyploid Arabidopsis suecica (Brassicaceae). Biochem Sys Ecol. 1994;22:807–11.
Novikova PY, Tsuchimatsu T, Simon S, Nizhynska V, Voronin V, Burns R, Fedorenko OM, Holm S, Säll T, Prat E, Marande W, Castric V, Nordborg M. Genome sequencing reveals the origin of the allotetraploid Arabidopsis suecica. Mol Biol Evol. 2017;34:957–68.
Säll T, Jakobsson M, Lind-Halldén C, Halldén C. Chloroplast DNA indicates a single origin of the allotetraploid Arabidopsis suecica. J Evol Biol. 2003;16:1019–29.
Shimizu-Inatsugi R, Lihova J, Iwanaga H, Kudoh H, Marhold K, Savolainen O, Watanabe K, Yakubov VV, Shimizu KK. The allopolyploid Arabidopsis kamchatica originated from multiple individuals of Arabidopsis lyrata and Arabidopsis halleri. Mol Ecol. 2009;18:4024–48.
Shimizu KK, Fujii S, Marhold K, Watanabe K, Kudoh H. Arabidopsis kamchatica (Fisch. Ex DC.) K. Shimizu & Kudoh and A. kamchatica subsp. kawasakiana (Makino) K. Shimizu & Kudoh, new combinations. Acta Phytotax Geobot. 2005;56:163–72.
Schmickl R, Jørgensen MH, Brysting A, Koch MA. Phylogeographic implications for north American arctic Arabidopsis lyrata. Plant Ecol Divers. 2008;1:245–54.
Schmickl R, Jørgensen MH, Brysting AK, Koch MA. The evolutionary history of the Arabidopsis lyrata Complex: a hybrid in the amphi-Beringian area closes a large distribution gap and builds up a genetic barrier. BMC Evol Biol. 2010;10:98.
Novikova PY, Hohmann N, Nizhynska V, Tsuchimatsu T, Ali J, Muir G, Guggisberg A, Paape T, Schmid K, Fedorenko OM, et al. Sequencing of the genus Arabidopsis identifies a complex history of nonbifurcating speciation and abundant trans-specific polymorphism. Nat Genet. 2016;48:1077–82.
Jørgensen MH, Ehrich D, Schmickl R, Koch MA, Brysting AK. Interspecific and interploidal gene flow in central European Arabidopsis (Brassicaceae). BMC Evol Biol. 2011;11:346.
Koch MA, Matschinger M. Evolution and genetic differentiation among relatives of Arabidopsis Thaliana. Proc Natl Acad Sci U S A. 2007;104:6272–7.
Hohmann N, Schmickl R, Chiang TY, Lučanová M, Kolář F, Marhold K, Koch MA. Taming the wild: resolving the gene pools of non-model Arabidopsis lineages. BMC Evol Biol. 2014;14:e224.
Hollister JD, Arnold BJ, Svedin E, Xue KS, Dilkes BP, Bomblies K. Genetic adaptation associated with genome-doubling in autotetraploid Arabidopsis arenosa. PLoS Genet. 2012;8:e1003093.
Yant L, Hollister JD, Wright KM, Arnold BJ, Higins JD, Franklin FC, Bomblies K. Meiotic adaptation to genome duplication in Arabidopsis arenosa. Curr Biol. 2013;23:2151–6.
Wright KM, Arnold B, Xue K, Surinova M, O'Connell J, Bomblies K. Selection on meiosis genes in diploid and tetraploid Arabidopsis arenosa. Mol Biol Evol. 2015;32:944–55.
Koch MA, Wernisch M, Schmickl R. Arabidopsis thaliana’s wild relatives: an updated overview on systematics, taxonomy and evolution. Taxon. 2008;57:933–43.
Muir G, Ruiz-Duarte P, Hohmann N, Mable BK, Novikova P, Schmickl R, Guggisberg A, Koch MA. Exogenous selection rather than cytonuclear incompatibilities shapes asymmetrical fitness of reciprocal Arabidopsis hybrids. Ecol Evol. 2015;5:1734–45.
Cohen KM, Gibbard PL. Global chronostratigraphical correlation table for the last 2.7 million years v. 2016a. Cambridge: Subcommission on quaternary Stratigraphy, International Commission on Stratigraphy; 2016. https://www.quaternary.stratigraphy.org.uk/charts. Accessed 11 Apr 2017
Tribsch A, Schönswetter P. Patterns of endemism and comparative phylogeography confirm palaeo-environmental evidence for Pleistocene refugia in the eastern alps. Taxon. 2003;52:477–97.
van Husen D. Geological processes during the Quarternary. Mitt Österr Geol Ges 2000, 92: 135–156.
Clauss MJ, Mitchell-Olds T. Population genetic structure of Arabidopsis lyrata in Europe. Mol Ecol. 2006;15:2753–66.
Muller MH, Leppala J, Savolainen O. Genome-wide effects of postglacial colonization in Arabidopsis lyrata. Heredity. 2008;100:47–58.
Ross-Ibarra J, Wright SI, Foxe JP, Kawabe A, DeRose-Wilson L, Gos G, Charlesworth D, Gaut BS. Patterns of polymorphism and demographic history in natural populations of Arabidopsis lyrata. PLoS One. 2008;3:e2411.
Pyhäjärvi T, Aalto E, Savolainen O. Time scales of divergence and speciation among natural populations and subspecies of Arabidopsis lyrata (Brassicaceae). Am J Bot. 2012;99:1314–22.
Ansell SW, Stenoien HK, Grundmann M, Schneider H, Hemp A, Bauer N, Russell SJ, Vogel JC. Population structure and historical biogeography of European Arabidopsis lyrata. Heredity. 2010;105:543–53.
Bomblies K, Weigel D. Arabidopsis - a model genus for speciation. Curr Opin Genet Dev. 2007;17:500–4.
Claus M, Koch MA. Arabidopsis and its poorly known relatives. Trends Plant Sci. 2006;11:449–59.
Hu TT, Pattyn P, Bakker EG, Cao J, Cheng JF, Clark RM, Fahlgren N, Fawcett JA, Grimwood J, Gundlach H, et al. The Arabidopsis lyrata genome sequence and the basis of rapid genome size change. Nature Genet. 2011;43:476–81.
Pritchard JK, Stephens M, Donnelly P. Inference of population structure using multilocus genotype data. Genetics. 2000;155:945–59.
Evanno G, Regnaut S, Goudet J. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol. 2005;14:2611–20.
Huson DH, Bryant D. Application of phylogenetic networks in evolutionary studies. Mol Biol Evol. 2006;23:254–67.
Koch MA, Kiefer M. Genome evolution among cruciferous plants - a lecture from the comparison of the genetic maps of three diploid species: Capsella rubella, Arabidopsis lyrata ssp. petraea and Arabidopsis thaliana. Am J Bot. 2005;92:761–7.
Couvreur TLP, Franzke A, Al-Shehbaz IA, Bakker FT, Koch MA, Mummenhoff K. Molecular phylogenetics, temporal diversification and principles of evolution in the mustard family (Brassicaceae). Mol Biol Evol. 2010;27:55–71.
Huang CH, Sun R, Zeng YH, Zhang N, Cai L, Zhang Q, Koch MA, Al-Shehbaz IA, Edger PP, Pires JC, Tan DY, Zhong Y, Ma H. Resolution of Brassicaceae phylogeny using nuclear genes uncovers nested radiations and supports convergent morphological evolution. Mol Biol Evol. 2016;33:349–412.
Drummond AJ, Suchard MA, Xie D, Rambaut A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol. 2012;29:1969–73.
Douglas GM, Gos G, Steige KA, Salcedo A, Holm K, Josephs EB, Arunkumar R, Ågren JA, Hazzouri KM, Wang W, Platts AE, Williamson RJ, Neuffer B, Lascoux M, Slotte T, Wright SI. Hybrid origins and the earliest stages of diploidization in the highly successful recent polyploid Capsella bursa-pastoris. Proc Natl Acad Sci U S A. 2015;112:2806–11.
Lafon-Placette C, Johannessen IM, Hornslien KS, Ali MF, Bjerkan KN, Bramsiepe J, Glöckle BM, Rebernig CA, Brysting AK, Grini PE, Köhler C. Endosperm-based hybridization barriers explain the pattern of gene flow between Arabidopsis lyrata and Arabidopsis arenosa in Central Europe. Proc Natl Acad Sci U S A. 2017;114:E1027–35.
Dart S, Kron P, Mable BK. Characterizing polyploidy in Arabidopsis lyrata using chromosome counts and flow cytometry. Can J Bot. 2004;82:185–97.
Schönswetter P, Stehlik I, Holderegger R, Tribsch A. Molecular evidence for glacial refugia of mountain plants in the European alps. Mol Ecol. 2005;14:3547–55.
Arnold B, Kim ST, Bomblies K. Single geographic origin of a widespread autotetraploid Arabidopsis arenosa lineage followed by interploidy admixture. Mol Biol Evol. 2015;32:1382–95.
Roux C, Castric V, Pauwels M, Wright SI, Saumitou-Laprade P, Vekemans X. Does speciation between Arabidopsis halleri and Arabidopsis lyrata coincide with major changes in a molecular target of adaptation? PLoS One. 2011;6:e26872.
Beilstein MA, Nagalingum NS, Clements MD, Manchester SR, Mathews S. Dated molecular phylogenies indicate Miocene origin for Arabidopsis thaliana. Proc Natl Acad Sci U S A. 2010;107:18724–28.
Koch MA, Haubold B, Mitchell-Olds T. Comparative evolutionary analysis of chalcone synthase and alcohol dehydrogenase loci in Arabidopsis, Arabis and related genera. Mol Biol Evol. 2000;17:1483–98.
Ossowski S, Schneeberger K, Lucas-Lledo JI, Warthmann N, Clark RM, Shaw RG, Weigel D, Lynch M. The rate and molecular spectrum of spontaneous mutations in Arabidopsis thaliana. Science. 2010;327:92–4.
Videvall E, Sletvold N, Hagenblad J, Agren J, Hansson B. Strong maternal effects on gene expression in Arabidopsis lyrata hybrids. Mol Biol Evol. 2016;33:984–94.
Arnold BJ, Lahner B, DaCosta JM, Weisman CM, Hollister JD, Salt DE, Bomblies K, Yant L. Borrowed alleles and convergence in serpentine adaptation. Proc Natl Acad Sci U S A. 2016;113:8320–5.
Turner TL, Bourne EC, Von Wettberg EJ, Hu TT, Nuzhdin SV. Population resequencing reveals local adaptation of Arabidopsis lyrata to serpentine soils. Nat Genetics 2010, 42: 260–263.
Doyle JJ, Doyle JL. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull. 1987;19:11–5.
Straub SC, Parks M, Weitemier K, Fishbein M, Cronn RC, Liston A. Navigating the tip of the genomic iceberg: next-generation sequencing for plant systematics. Am J Bot. 2012;99:1–16.
Müller K, Quandt D, Müller J, Neinhuis C. PhyDE, version 0.9971: Phylogenetic data editor. 2010. Available at http://www.phyde.de.
Katoh K, Misawa K, Kuma K, Miyata T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transformation. Nucleic Acids Res. 2002;30:3059–66.
Katoh K, Kuma K, Toh H, Miyata T. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res. 2005;33:511–8.
Katoh K, Standley DM. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 2013;30:772–80.
Castresana J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol. 2000;17:540–52.
Lanfear R, Calcott B, Ho SY, Guindon S. Partitionfinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol. 2012;29:1695–701.
Lanfear R. Selecting optimal partitioning schemes for phylogenomic datasets. BMC Evol Biol. 2014;14:82.
Wolfe KH, Li W-H, Sharp PM. Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAs. Proc Natl Acad Sci U S A. 1987;84:9054–8.
Stamatakis A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 2014;30:1312–3.
Drummond AJ, Ho SYW, Phillips MJ, Rambaut A. Relaxed phylogenetics and dating with confidence. PLoS Biol. 2006;4:e88.
Stadler T. On incomplete sampling under birth-death models and connections to the sampling-based coalescent. J Theor Biol. 2009;261:58–66.
R Core Team. R: a language and environment for statistical computing. Vienna: R Foundation for Statistical Computing; 2014. URL https://www.R-project.org
Paradis E, Claude J, Strimmer K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics. 2004;20:289–90.
Li H. 2013. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv:1303.3997v2 [q-bio.GN].
Li H, Durbin R. Fast and accurate short read alignment with burrows-wheeler transform. Bioinformatics. 2009;25:1754–60.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R. 1000 genome project data processing subgroup. The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25:2078–9.
Li H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics. 2011;27:2987–93.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–303.
DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, Philippakis AA, del Angel G, Rivas MA, Hanna M, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nature Genet 2011, 43: 491–498.
Quinlan AR, Hall IM. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics. 2010;26:841–2.
Cingolani P, Platts A, Wang le L, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila Melanogaster strain w1118; iso-2; iso-3. Fly. 2012;6:80–92.
Falush D, Stephens M, Pritchard JK. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics. 2003;164:1567–87.
Jakobsson M, Rosenberg NA. CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics. 2007;23:1801–6.
Earl DA, vonHoldt BM. STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour. 2011;4:359–61.
Ehrich D, Gaudeul M, Assefa A, Koch MA, Mummenhoff K, Nemomissa S, Intrabiodiv C, Brochmann C. Genetic consequences of Pleistocene range shifts: contrast between the Arctic, the alps and the east African mountains. Mol Ecol. 2007;16:2542–59.
Rosenberg NA. Distruct: a program for the graphical display of population structure. Mol Ecol Notes. 2003;4:137–8.
Chen H. 2016. VennDiagram: generate high-resolution Venn and Euler plots. R package version 1.6.17. https://CRAN.R-project.org/package=VennDiagram.
Acknowledgments
We are grateful to Polina Novikova (Ghent University) developing the data pipeline to further analyze genomic data and all colleagues contributing with additional field samples (Alex Widmer, Alessia Guggisberg from Zurich, Graham Muir, Long Li from Heidelberg).
Funding
Funding from DFG (Deutsche Forschungsgemeinschaft) to MAK is appreciated within the priority research program SPP1529 (Adaptomics) and under the project code KO2302/14–1. We acknowledge financial support by Deutsche Forschungsgemeinschaft and Ruprecht-Karls-Universität Heidelberg within the funding programme Open Access Publishing. The funding body played no role in the design of the study and collection, analysis, and interpretation of data and in writing the manuscript.
Availability of data and materials
The datasets generated and analyzed herein are available in Genbank/ENA and provided with Additional file 2 and ERZ437650 -ERZ437677 (plastid genomes).
Author information
Authors and Affiliations
Contributions
MAK designed the study and set up the experiments. NH processed the data. NH and MAK analyzed and interpreted the data. MAK and NH wrote the manuscript. This manuscript is close to the final chapter of the PhD thesis of NH. Both authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
All plant material was collected in the wild, and no permissions are needed to obtain the material. Our study fully complies with regulations of CBD (convention biological diversity) and the Nagoya protocol. Voucher specimens have been deposited with the Heidelberg herbarium (HEID) and according information is provided with the accession list.
Consent for publication
Not applicable
Competing interests
Both authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Additional files
Additional file 1:
Result of SplitsTree network reconstruction. (PDF 770 kb)
Additional file 2:
Accession list and population details. (XLSX 28 kb)
Additional file 3:
Partitioning scheme of plastid DNA sequence data for phylogenetic analysis. (TXT 14 kb)
Additional file 4:
Alignment of plastid DNA sequence data for phylogenetic analysis. (TXT 9059 kb)
Additional file 5:
Combined tree file (plastome data) with BEAST output. (TXT 147 kb)
Additional file 6:
Structure-sum output, analysis of deltaK and similarity values between STRUCTURE runs. (PDF 165 kb)
Additional file 7:
VENN diagrams with absolute numbers of SNPs. (PDF 2549 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Hohmann, N., Koch, M.A. An Arabidopsis introgression zone studied at high spatio-temporal resolution: interglacial and multiple genetic contact exemplified using whole nuclear and plastid genomes. BMC Genomics 18, 810 (2017). https://doi.org/10.1186/s12864-017-4220-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12864-017-4220-6