Abstract
Background
Neurogenesis control and the prevention of premature differentiation in the vertebrate embryo are crucial processes, allowing the formation of late-born cell types and ensuring the correct shape and cytoarchitecture of the brain. Members of the Hairy/Enhancer of Split (Hairy/E(spl)) family of bHLH-Orange transcription factors, such as zebrafish Her3, 5, 9 and 11, are implicated in the local inhibition of neurogenesis to maintain progenitor pools within the early neural plate. To better understand how these factors exert their inhibitory function, we aimed to isolate some of their functional interactors.
Results
We used a yeast two-hybrid screen with Her5 as bait and recovered a novel zebrafish Hairy/E(spl) factor - Her8a. Using phylogenetic and synteny analyses, we demonstrate that her8a evolved from an ancient duplicate of Hes6 that was recently lost in the mammalian lineage. We show that her8a is expressed across the mid- and anterior hindbrain from the start of segmentation. Through knockdown and misexpression experiments, we demonstrate that Her8a is a negative regulator of neurogenesis and plays an essential role in generating progenitor pools within rhombomeres 2 and 4 - a role resembling that of Her3. Her8a co-purifies with Her3, suggesting that Her8a-Her3 heterodimers may be relevant in this domain of the neural plate, where both proteins are co-expressed. Finally, we demonstrate that her8a expression is independent of Notch signaling at the early neural plate stage but that SoxB factors play a role in its expression, linking patterning information to neurogenesis control. Overall, the regulation and function of Her8a differ strikingly from those of its closest relative in other vertebrates - the Hes6-like proteins.
Conclusions
Our results characterize the phylogeny, expression and functional interactions involving a new Her factor, Her8a, and highlight the complex interplay of E(spl) proteins that generates the neurogenesis pattern of the zebrafish early neural plate.
Similar content being viewed by others
Background
Neurogenesis in the early vertebrate neural plate begins at stereotyped loci - termed proneural clusters -, which prefigure the localization of the earliest neuronal groups and the architecture of the primary embryonic neuronal scaffold. These proneural clusters consist of spatially defined progenitor groups engaged in active neurogenesis, within which committed precursors expressing higher levels of proneural genes (such as neurogenin or achaete-scute-like genes, respectively neurog1 and ascl1 in zebrafish) are singled out to differentiate first. An identical scaffold is found in all vertebrate embryos, highlighting the robustness and functional relevance of this organization [1–4]. Dissecting the regulatory cascades involved in this process is therefore of universal importance.
The control of neurogenesis progression within proneural clusters relies on Hairy/Enhancer of split (E(spl)) factors (Hes in mouse; Her in zebrafish). These transcription factors belong to the basic-helix-loop-helix (bHLH) family, characterized by a DNA-binding basic domain and an HLH domain composed of two alpha helices intervened by a loop of a few amino acids [5]. In addition, Hairy/Enhancer of split (E(spl)) factors contain an Orange domain, which is most probably involved in protein-protein interactions, and a WRPW C-terminal tetrapeptide, which mediates transcriptional repression (reviewed in [6, 7]). During the so-called process of lateral inhibition, the expression of Notch ligands in committed precursors activates Notch signaling in neighbouring progenitors, which in turn induces expression of Hes/Her factors. The latter down-regulate proneural genes, hence maintaining Notch-receiving cells in a progenitor state. Reflecting the intermingled distribution of committed and transiently inhibited progenitors, the proneural and E(spl) genes are expressed in a salt-and-pepper fashion within proneural clusters. E(Spl) factors expressed in proneural clusters in zebrafish include her4.1 [8–10], hes5/her15, her2 and her12 [11]. In agreement with the lateral inhibition model, her4.1 expression is positively regulated by Notch, and inhibits expression of neurog1 [10].
Recent work has demonstrated that proneural clusters are delimited negatively, through a process of active neurogenesis suppression taking place in surrounding areas (reviewed in [12]). These "inhibited" areas, so-called "progenitor pools", are transiently maintained in an refractory state to be recruited in later events of neuronal production, and are organized as tight groups of adjacent cells at stereotyped positions within the neural plate. Major progenitor pools can be found at the presumptive midbrain-hindbrain boundary (MHB) [3, 13] and in longitudinal stripes separating the columns of presumptive moto- and lateral neurons in the hindbrain, or moto-, inter- and sensory neurons in the spinal cord [11, 14, 15]. At the least, the MHB pool is maintained until adulthood in zebrafish, where it participates in the generation of adult-born neurons and oligodendrocytes [16]. Embryonic progenitor pools are characterised by the expression of a specific set of transcription factors, including Zic, BF1/Anf and Rx family members [17] as well as Hes/Her proteins. In zebrafish, the combinatorial expression of a distinct set of her genes - which to date includes her3, her5, her9 and her11 - characterises all progenitor pools [12], while in mouse the genes Hes1, Hes3 and Hes5 share sustained expression in adjacent cells of the MHB pool, for example [18–20]. These her/Hes genes exhibit functional similarities and have been implicated in progenitor pool maintenance: their misexpression inhibits neurogenesis, whereas loss-of-function causes premature expression of proneural genes in at least part of their expression domains [11, 13, 15, 19–22]. In addition, her3/5/9/11 as well as Hes1 at the mouse MHB all demonstrate an irregular association with Notch: while Hes/her genes in proneural clusters are activated by Notch signaling, the expression of her3/5/9/11 and Hes1 at the MHB is controlled in a Notch-independent manner. The mechanisms accounting for these specific features remain unknown.
The HLH region of Hes/Her factors functions as a dimerisation domain, and the formation of hetero- and homodimers as well as further possible interactions through the Orange domain are key components of the specificity of the actions of these proteins. Heterodimerisation can involve closely related members of the Her/Hes family, or several different transcription factors or transcriptional cofactors [7]. In order to better understand the mechanism of action of Her factors expressed in progenitor pools, and the pathways regulating their activity, we performed a yeast two-hybrid screen using the HLH and Orange domains of Her5 as bait. This led to the recovery of Her8a, a novel Her factor of the Hes6 subfamily expressed in a broad manner at the presumptive midbrain-hindbrain domain of the early zebrafish neural plate. Morpholino-mediated knockdown and misexpression studies establish Her8a as a negative-regulator of neurogenesis playing an essential role in maintaining progenitor pools of rhombomeres (r) 2 and 4. her8a knockdown produces a similar phenotype to that of her3 knockdown, and co-purification demonstrates that Her8a dimerises with Her3. At the MHB however, we show that the predominant activity is exerted by the combination of Her3, 5, 9 and 11. Together, our results identify a new player in progenitor pool formation and highlight the region-specific combinatorial activity of E(spl) factors in this process.
Results
Identification of Her8a as a potential binding partner for Her proteins
To recover binding partners for Her proteins, we used a yeast two-hybrid screen where a 181-amino acid fragment of Her5 (excluding the basic domain and the WRPW motif) was screened against an 18-20 hpf embryo zebrafish library. This screen returned 280 positive clones, from a total of 76.1 million tested interactions. These 280 positive clones represented 75 unique protein-protein interactions. The quality of these interactions was graded using a PBS scoring system - where A is the highest score of confidence, B is very good, C is good, D is low and N/A is used when no score could be assigned (see Materials and Methods). Our screen returned 6 As, 9 Bs, 2 Cs, 49 Ds and 9 N/As (see Additional file 1, Table S1 for a detailed description of all recovered candidates). Gene ontology enrichment analysis of the recovered binding partners revealed an enrichment of proteins involved in protein transport and also heterodimerisation (see Additional file 2, Table S2). Among these candidates, we note the presence of 7 distinct Her factors, in agreement with the postulated capacity for protein heterodimerisation within this family [7]. As further indication of the validity of the assay, Her5 was found to bind with Her11 with a score of B, an interaction that had been shown previously in our laboratory [22]. Her8a, scored A and corresponding to a new E(spl) family member, proved very strongly expressed in the midbrain-hindbrain (MH) area (see below), and we consequently focused on this factor.
Her8a is a new Hes6-like protein, the ortholog of which was lost in the mammalian lineage
The predicted Her8a protein displays a bHLH and an orange domain, and harbors a WRPW motif at its C-terminus - characteristics common to the Hairy/E(spl) family. Sequence comparisons of the bHLH domain of this family classify Her8a, together with zebrafish Her13 (previously Her13.1), Hes6 (previously Her13.2) and Her8.2, within the subfamily showing highest homology to mouse Hes6 [23]. Proteins of this subfamily exhibit a shortened loop when compared to other Hairy-E(spl) members [24], such as mouse Hes1 and Hes5 (Additional file 3, Figure S1). Hes6-like proteins also share substantial similarity outside the bHLH domains (Additional file 3, Figure S1), allowing their phylogeny to be studied using extended protein sequences. This confirmed the existence of two subfamilies of Hes6-like proteins, Hes6.1 and Hes6.2, comprising respectively Her13/Hes6 and Her8a/Her8.2 (Figure 1A) (and see [23]), encoded by gene pairs (Figure 1B). Importantly, it also resolved for the first time their relationship with the single mammalian Hes6 protein, as sequence alignments directly assign mammalian Hes6 to the Hes6.1 subfamily (Figure 1A). hes6.2-like genes are found neither in eutherian mammals nor in marsupials, but exist in all other phyla, suggesting a late secondary loss of this gene shortly after the divergence of eutherians and marsupials from the monotreme lineage. Finally, and as expected from the whole genome duplication undergone in the teleost fish lineage subsequent to its divergence from other vertebrates [25], followed by secondary gene loss, teleost species exhibit three or four Hes6-like genes. Synteny analyses (Figure 1C) indeed identify a conserved orthologous gene pair (locus 1) as well as a conserved duplicate (locus 2). This duplicate only contains the hes6.1-like member and, based on the situation in other vertebrates, most likely lost the hes6.2-like gene. Strikingly, in zebrafish, the latter gene (her8a) was kept and transferred onto a different genomic location (locus 3). Together, these results indicate that her8a is the ortholog of her8.2 and that it is located in a unique genomic setting compared to other related genes. It also assigns her8a to a subfamily of Hes6-like genes that is closely related to but not directly orthologous to mammalian Hes6.
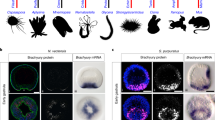
her8a encodes a Hes6-like E(spl) protein closely related to, but not directly orthologous to, mammalian Hes6. A. Phylogenetic tree depicting protein relationship within the Hes6 subfamily, based on the bHLH and Orange domain sequences. Note the absence of Hes6.2 proteins in marsupials and eutherian mammals. In zebrafish, the closest relative to Her8a is Her8.2, and the closest relative to Her13 is Hes6. B. Genomic organization of Hes6-like genes through evolution, confirming that the generation of the two Hes6.1 and Hes6.2 genes is ancestral and that Hes6.2 was secondarily lost within the mammalian taxa subsequent to the divergence of marsupials and eutherian mammals from monotremes. "?" indicates cases where genomic linkage cannot be resolved at present, in the absence of a genome sequence for the corresponding species. C. Genomic organization of the areas surrounding Hes6-like genes in teleost species compared to mouse. With the exception of zebrafish, for all of the teleosts studied, synteny analyses suggest a secondary loss of the hes6.2 duplicate (missing from locus 2) after the duplication of locus 1. In zebrafish, this gene (her8a) has been relocated to a third locus on a distinct chromosome. 1-3: genomic loci.
her8ais expressed across progenitor pools and proneural clusters in the anterior neural plate
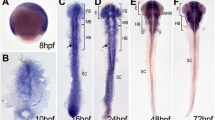
Among hes6-like genes, we found that her8a was the only one with expression in the anterior neural plate at the 3-somite stage. At this stage, her13 was restricted to a portion of the presumptive lateral proneural clusters in the anterior spinal cord (Additional file 4, Figure S2). As shown previously, hes6 was found to be expressed in the tail bud and posterior paraxial mesoderm (Thisse and Thisse, 2004), and her8.2 was very weakly or not expressed (not shown). In contrast, following weak expression during gastrulation [26], her8a displayed strong expression in the neural plate from the tailbud stage onwards. At tailbud, her8a is expressed throughout the neural plate, with the exception of the eye field and the midline, with strongest intensity at presumptive midbrain and hindbrain levels (Figure 2A, bracket). At early segmentation stages, the domain of strong her8a expression overlaps with that of genes expressed in progenitor pools within the presumptive mid- and hindbrain, such as her3 (Figure 2B) and her5 (Figure 2C) (purple arrows). It also encompasses the proneural clusters located in the mid- and anterior hindbrain area (e.g. compare with neurog1 expression, Figure 3A). At 10 somites her8a is expressed in stripes in the hindbrain, with denser expression in rhombomeres (r) 1 and 3-5 (Figure 2D,E). Expression in the midbrain persists. From 24hpf onwards, her8a acquires a distinct expression profile, highlighting neurogenic domains throughout the central nervous system, with weaker staining at the midbrain-hindbrain boundary (mhb) (arrow) and at the zona limitans intrathalamica (Figure 2F) (asterisk). Cross sections of the brain at this stage reveal that her8a expression is confined to the progenitor, ventricular domain, largely complementary to the expression of the post-mitotic neuronal marker HuC/D (Figure 2G). From 48hpf through to adult, her8a remains expressed in proliferation/ventricular zones throughout the brain (Figure 2H,I, and data not shown).
Embryonic expression of her8a. Expression is revealed by whole-mount in situ hybridization at the stages indicated (som: somites, hpf: hours post-fertilization) together with other positional marker genes (B,C,E, color-coded) or proteins (G, immunocytochemistry for the neuronal marker HuC/D in green). A-E and H are dorsal views of flat-mounted embryos, F,I are lateral views, all embryos are viewed anterior left. G is a cross section of a 24hpf embryo at midbrain levels (as indicated in F). At early neural plate stages, the domain of strongest her8a expression covers the mid- and anterior hindbrain (bracket in A-C). It overlaps the presumptive midbrain-hindbrain boundary (her5-positive, purple arrow in C) and the progenitor pools separating medial and lateral hindbrain neurons (her3-positive, purple arrow in B). It extends into rhombomere 2 (white arrowhead in C), more posterior rhombomeres being more weakly labeled. At 10 somites, expression in the midbrain is maintained. It resolves in stripes in the rhombencephalon (D,E). It avoids the midbrain-hindbrain boundary and zona limitans intrathalamica (asterisk). From 24 hpf onwards, her8a characterizes the ventricular zone and progenitor domains of the neural tube (G-I, arrows in I point to the ventricular zone and the arrowhead points to the progenitor domain of the optic tectum). Abbreviations: e: eye field, mid: presumptive midbrain, mhb: midbrain-hindbrain boundary, r: rhombomere. Scale bars: 100 μm.
Gain of Her8a function inhibits neurogenesis
Mouse and Xenopus Hes6 proteins are known as positive regulators of differentiation [24, 27]. In this context, the expression of her8a across both pro-neural and non-neurogenic domains was puzzling and prompted us to explore Her8a function.
In a first gain-of-function approach, embryos were injected with her8a capped mRNA encoding the full-length protein at the one cell stage. They were subsequently fixed and analyzed at 3 somites. We observed that her8a misexpression caused a complete loss of neurog1 expression throughout the embryo (Figure 3A,B) (77% of cases, n = 22). Co-labeling with tp63 (previously ΔNp63), which highlights the border of the epidermal ectoderm juxtaposed to the neural plate [28] (Figure 3A, white arrowheads), showed that the size or morphology of the neural plate were not affected (100% of cases, n = 22), suggesting that overexpressing Her8a specifically blocks neurogenesis without an effect on neural plate formation. This was confirmed by expression analyses for patterning markers such as barhl2, her5 and her9, which highlight distinct domains along the entire antero-posterior axis of the neural plate [11, 29, 30] (Additional file 5, Figure S3). Together, these results indicate that her8a is capable of inhibiting neurogenesis, at least at non-physiological concentrations, and this even across domains normally co-expressing neurog1 and her8a such as the proneural clusters of the mid- and anterior hindbrain (vcc, r2MN, r2l, R4MN, r4l).
Like Her3, Her8a activity maintains the non-neurogenic areas of rhombomeres 2 and 4. A,B. neurog1 expression highlights the proneural clusters at the 3-somite stage (black arrows in A) and is eliminated upon her8a overexpression (B, embryo injected with her8a capped mRNA). Expression of tp63, which highlights the neural plate border (white arrowheads in A), is unchanged. C-F. Compared expression of neurog1 in control embryos (C) and embryos injected with her8aMO (D), her3MO (E) or both MOs (F) shows ectopic neurogenesis between the medial and lateral proneural clusters of r2 and r4 (blue arrows in D-F, compare with white arrowheads in C) when Her8a and/or Her3 activities are blocked. Few "ectopic" neurog1-positive can sometimes be found between the vcc and mnr2; this is however highly variable between individuals and observed in both control and morphant embryos. A-F are dorsal views of flat-mounted embryos, anterior left. Abbreviations: black arrows indicate proneural clusters: IN: presumptive interneurons, MN: presumptive motoneurons, r2: rhombomere 2, r4: rhombomere 4, r2l: lateral neurons of rhombomere 2, r4l: lateral neurons of rhombomere 4, SN: presumptive sensory neurons, vcc: ventro-caudal cluster. G. RT-PCR analysis of her8a expression (left and middle panels) in embryos injected with her8aMO1 ("mo1") and her8aMO2 ("mo2") versus control embryos ("ctr"). Low levels of full length, normally spliced her8a transcripts are detectable in morphants (left and middle panels, arrows) while abnormally spliced transcripts including all or part of intron 1 become produced (stars). Expression of βactin2, used as RT-PCR control, is indentical in all samples (right panel). The scheme at the top indicates the genomic structure of her8a, the position of exons (E, purple) and introns (black bars), the binding sites of her8aMO1 and MO2 (red), the position of RT-PCR primers (blue arrows) and the length of amplified wild-type products (excluding introns) (blue bars).
Her8a is required to maintain the proper neurogenesis pattern in rhombomeres 2 and 4 and acts as binding partner for Her3
To better appreciate the endogenous requirements for Her8a, we next turned to a loss-of-function approach. Embryos at the one-cell stage were injected with morpholinos (MO) directed against the donor splice site of her8a exon 1 (MO1), the acceptor splice site of her8a exon 2 (MO2), or the her8a ATG (MO3), and were analyzed at 3 somites. Reverse transcription PCR was used to reveal strong down-regulation of expression and abnormal splicing of her8a transcripts with both MO1 and MO2, whereas other genes, such as βactin2, remained unaffected (Figure 3G). These observations substantiate that her8aMOs lead to knock-down of her8a expression.
Blocking Her8a resulted in an ectopic expression of neurog1 within the normally non-neurogenic area separating motor- and lateral proneural clusters of r2 and 4 (Figure 3C, arrowheads, Figure 3D, arrows) (80% of cases, n = 60). Although the location of neurog1-positive cells can be slightly variable from embryo to embryo, this phenotype was robust and never observed in wild-type animals. The results obtained with the three MOs were strictly identical (Additional file 6, Figure S4 for a comparison of MO1 and MO2, and data not shown), confirming their specificity. We will compile these data below. This knockdown phenotype strikingly resembles the published effect of her3 knockdown [15], which we further confirmed (Figure 3E) (56% of cases, n = 50). Given that the r2/r4 area of ectopic neurogenesis is where the intense expression of her8a overlaps with that of her3 (Figure 1B, Figure 4B), these identical phenotypes suggest a direct or indirect functional interaction of Her3 and Her8a in controlling common progenitors. As a first obvious possibility, we tested whether her3 and her8a influence each other's expression. Ruling out this scenario, her8aMO-injected embryos displayed a normal expression of her3, and her3MO embryos a normal expression of her8a (not shown) (100% of cases, n = 37 and 24, respectively). A following hypothesis is that her3 and her8a act in a dose-dependent manner to compensate for each other outside r2 and r4, with r2 and r4 showing highest sensitivity to the amount of "Her3+Her8a" proteins. A similar situation was previously demonstrated for Her5+Her11 at the MHB [22]. In this case, we expect that the knockdown of both genes would produce a phenotype of a greater magnitude than the ectopic expression of proneural markers in r2 and r4. To test this possibility, we simultaneously blocked Her3 and Her8a by the co-injection of her3MO and her8aMO (at the same concentrations shown to produce the individual phenotypes). This produced no additional effect on neurog1 expression (Figure 3F) (80% of cases, n = 20). Likewise, the co-injection of her3MO and her8aMO in amounts just below their effective doses (0.75 mM for her8a and 0.375 mM for her3) induced no phenotype in r2 and r4 (not shown). Together, these observations indicate that these two factors alone are not compensating for each other to repress neurog1 in other areas of the embryo.
The heterodimerisation potential of full-length Her8a and Her3 proteins may account for their identical loss-of-function phenotypes. A. Co-affinity purification of full-length Her3 and Her8a proteins in HEK293T cells. When lysates from cells expressing both N-Strep/Flag-Her3 (N-SF-Her3) and Myc-tagged Her8a (myc-Her8a) proteins are eluted from a STREP-Tactin resin, myc-Her8a is observed to co-purify with N-SF-Her3 (lane 4). B-E. Schematized summary of the expression patterns (B) and loss-of-function phenotypes (C-E) of her8a and her3. Genes expression are color-coded and her5/her11 expression is indicated as a landmark (see also Figure 5H). Abbreviations: as in Figure 3; tg: trigeminal ganglion.
As a final alternative hypothesis, and based on the recovery of Her8a as binding to a Her bHLH domain in yeast cells, we tested whether Her8a and Her3 could act as necessary heterodimerisation partners. As no commercial antibodies are available for Her3 and Her8a, and attempts by our laboratory to have them manufactured failed, we chose a co-purification approach using tagged versions of the full-length zebrafish proteins recombinantly expressed in HEK293T cells. By purifying Strep/Flag-tagged Her3 via its Strep-tag II moiety, Myc-tagged Her8a was successfully co-purified (Figure 4A), demonstrating that both proteins interact with each other. This interaction may be relevant to the maintenance of the progenitor pools within r2 and r4, where both her3 and her8a are strongly expressed, and hereby account for the identical phenotypes of Her3 and Her8a loss of function (summarized in Figure 4B-E).
In the absence of Her3, 5, 9 and 11 activity, endogenous Her8a alone is insufficient to preserve neurog1-free progenitor pools in the midbrain-hindbrain domain
The results above indicate that, although her8a is expressed across the entire MH domain, it is only strictly required to block neurogenesis in r2 and r4. This raises the question of which her genes combination encodes the endogenous pattern of neurogenesis inhibition in the MH domain, and whether this combination involves her8a expression. To address this issue, we used double in situ hybridization to re-analyze expression of the progenitor pools genes, comparing her3, 5, 9 and 11. Our data confirmed the full overlap of her5 and her11 (Figure 5A,D) as well as the extension of her3 longitudinal stripes into the presumptive MHB domain [15] (Figure 5B, arrows), and revealed a previously unreported expression of her9 coinciding with the antero-lateral aspects of the her5/11 territory (Figure 5C, arrows) (summarized in Figure 5K,L). Previous loss-of-function experiments of combinations of these genes never achieved a full neurogenic phenotype: concomitantly blocking her5 and her11 induced neurog1 medially but only to a lesser extent in mediolateral and lateral MHB domains [22] (Figure 5M,N), and the co-inhibition of her3 and her9 largely recapitulated her3 loss-of-function in the MH area, with a restricted induction of neurog1 within r2 and r4 [11] (Figure 5O,P). We found that the down-regulation of all four factors together, through the coinjection of the relevant gripNA antisense oligonucleotides, was required to generate a large neurog1-positive domain across the presumptive MHB and r2 (Figure 5E,F) (74% of cases, n = 19) - although the most lateral aspects of the neural plate remained neurog1-free -. In these conditions however, her8a expression remained unperturbed (Figure 5G,H) (100% of cases, n = 39) (schematized in Figure 5Q). Together, these results demonstrate that the endogenous activity of Her8a, in the absence of other progenitor pools Her factors, is insufficient to inhibit neurog1 expression in the MH area. her8a expression also appears insensitive to the combined expression levels of Her3/5/9/11 and to the neurogenic status of this neural plate domain. When following the fate of ectopic neurog1-expressing progenitors in the absence of Her3/5/9 and 11 activities, we found however that only a subset were maintained until 24hpf. These were located ventrally across the midbrain-hindbrain boundary, immediately posterior to vcc-derived neurons (Figure 5I,J).
The activities of Her3, Her5, Her9 and Her11 together account for the progenitor pool pattern of the midbrain-hindbrain area and do not influence her8a expression. A-D. A comparison of the expression patterns of her3, her5, her9, her11 and neurog1 at 3 somites in the midbrain-hindbrain (MH) area using double in situ hybridization (color-coded), on dorsal views of flat-mounted embryos. Arrows point to the her5/her11 domains co-expressing her3 or her9. E-J. Compared neurogenesis in control embryos (E,G,I) and embryos co-injected with the four gripNA antisense oligonucleotides directed against her3, 5, 9 and 11 transcripts (see Methods) (F,H,J). E,F. Expression of neurog1 (revealed by in situ hybridization against gfp in -8.4neurog1:gfp transgenic embryos) [62]: the majority of the MHB/r2 area is induced to express neurog1. G,H. The expression her8a is unaltered upon injection of the four gripNAs. I,J. Detection of GFP in -8.4neurog1:gfp embryos at 24 hpf (sagittal view, confocal projection of a 20 μm section of the neural tube). Ectopic neurons are formed ventrally across the midbrain-hindbrain boundary (position of the boundary indicated by the white bar), in a location normally devoid of GFP-positive cells (arrowheads). K,L. Summarized compared expression of her3, 5, 9 and 11 (K), also together with her8a (L). M-Q. Summary of the combined loss-of-function results for MH-expressed her genes, from Geling et al. [13] (M), Ninkovic et al. [22] (N), Bae et al. [11] (O,P) and the present paper (Q). Abbreviations: mid: midbrain, MN: presumptive motoneurons, r: rhombomere, r2l: lateral neurons of rhombomere 2, vcc: ventro-caudal cluster.
Endogenous her8aexpression in the early neural plate is independent of Notch signaling but requires the expression of SoxB factors
Although many E(spl) transcription factors are downstream effectors of Notch signalling, previous work has shown that zebrafish her genes expressed in progenitor pools, such as her3, 5, 9 and 11 [11, 13, 31] exhibit a non-canonical regulation by Notch: they do not require Notch for their expression, and are insensitive to or transcriptionally inhibited upon ectopic Notch activation. This is in contrast to other family members such as her4.1 that are expressed in neurogenic zones and are activated by Notch signaling [32].
Unusually, her8a is expressed across both progenitor pools and proneural clusters in the early neural plate. To analyze the effect of ectopic Notch activation, we overexpressed the intracellular domain of zebrafish Notch1a (NICD) [10] through capped mRNA injection at the one-cell stage. We could replicate previously published results [10] showing that NICD misexpression completely downregulates neurogenesis throughout the early neural plate (Figure 6A,B) (88% of cases, n = 16). We found that overexpression of NICD causes ectopic or enhanced expression of her8a throughout the embryo (Figure 6C,D) (100% of cases, n = 17), although this induction was weaker at the caudal end of the neural plate (Figure 6D, asterisk). The latter observation was repeated when studying the her4.1 target gene (Figure 6E,F) (100% of cases, n = 13), suggesting a generally lower ability of this neural plate area to respond to NICD overexpression, rather than a her8a-specific feature in Notch response. Next, to determine whether her8a expression depends on endogenous Notch signaling, we incubated embryos between the 50% epiboly and 3-somite stages into the gamma-secretase inhibitor DAPT, which blocks Notch by preventing the cleavage of NICD and has a strong neurogenic effect [33, 34]. As anticipated from previous studies [33], DAPT treatment increased the amount of neurog1-positive cells within each proneural cluster (Figure 6G-H>') (79% of cases, n = 19). However, it failed to reproducibly affect the expression of her8a (Figure 6C,I-J') (100% of cases, n = 15). Together, these results indicate that endogenous Notch signaling is not required for her8a expression in the early neural plate.
her8a expression is independent of endogenous Notch signaling in the early neural plate. Dorsal views of flat-mounted embryos analyzed at the 3-somite stage for the expression of the genes indicated. A-F. Ectopic Notch activation (injection of capped NICD mRNA) abolished neurog1 expression (B) and activates her8a (D) and her4.1 (F) compared to non-injected embryos (ctrl). G-J'. Notch blockade (incubation in the gamma-secretase inhibitor DAPT) increases neurog1 expression within proneural clusters (H) but leaves her8a expression intact (J) at early neural plate stages compared to embryos treated with a vehicle only (ctrl). G'-J' are high magnification views of the areas boxed in G-J.
In order to gain further insight into the endogenous mechanisms controlling her8a regulation at these stages, we scanned the her8a promoter (100 bp downstream and 1000 bp upstream of the ATG start site) with the ModelInspector program (Genomatix) [35]. This revealed a potential Sox (Sry-related HMG box) binding site (SORY_OCT1_01) at position 873-897(+). ModelInspector uses Genomatix's in-house Promoter Module Library, which includes experimentally verified models for functional promoter subunits. In this case the promoter sequence was derived from a publication describing the activation of the mouse Fgf4 enhancer by Sox2 and Oct-3 [36]. This led us to investigate the possibility that a member of the sox gene family is controlling her8a expression. Mouse Sox2 is a member of Group B1, a subdivision of Sox genes involved in neural development [37]. Within this subgroup, we focused on the zebrafish genes sox2 [37], sox3 [37], sox19a and sox19b [38] (see [37] for phylogenetic description), excluding sox1a and sox1b, which are not expressed at the MHB at early embryonic stages [26, 37]. In addition, we also investigated sox21a (previously sox21 or sox30), a member of the related subgroup B2 with specific MHB expression at early embryonic stages [39]. We found that expression of these different sox genes overlapped all or part of the her8a-positive domain at the 5-somite stage: sox2 and sox3 displayed strongest overlap with her8a expression in the anterior hindbrain (mostly r3) (Figure 7A,B), sox21a at the MHB (Figure 7E), while sox19a and b were intensely expressed throughout the MH (Figure 7C,D). In addition, sox2, 3, 19a and 19b all displayed an expression identical to her8a in the presumptive telencephalon and ventral diencephalon, excluding the eye field (Figure 7A-D, compare with F). To analyze the role of these genes in controlling her8a expression, we used MOs targeting their ATG start site [40–43]. A single MO was used to inhibit sox2 and 3, which share the sequence surrounding their ATG. When injected individually at the one-cell stage, none of these MOs produced a phenotype on her8a expression at the 3-somite stage (not shown) (n = 25). However, the combined knock-down of all five sox genes at once caused reduced her8a staining in the MH area (Figure 7G-I) (100% of cases, n = 18), indicating that these factors cooperate, possibly in a dose-dependent manner, to enhance her8a expression within the early neural plate. In a reverse step, we analyzed whether these genes were linked by a positive regulatory loop. We found however that blocking Her8a function upon her8aMO injection had no effect on sox genes expression at the 3-somite stage (not shown) (100% of cases, n = 10 for each sox gene tested).
her8a expression in the early neural plate is partially dependent on the expression of SoxB factors. Dorsal views of flat-mounted embryos analyzed at the 5-somite stage (A-F) and 3-somite stage (G-H) for the expression of the genes indicated (color-coded). A-F. her8a expression overlaps with that of the Sox family members sox2, sox3, sox19a, sox19b and sox21a, including the mid- and anterior hindbrain (red brackets). G-I. the simultaneous knockdown of sox2/3/19a/19b and 21a causes a reduction of her8a expression in the MH area of the early neural plate (white arrows).
Discussion
Her8a is a neurogenesis repressor in the early zebrafish neural plate
Two lines of evidence demonstrate that Her8a can act as a repressor of neurogenesis: firstly, the overexpression of full-length her8a causes a complete loss of neurog1 expression in the early embryo; secondly, we show that morpholino-mediated knockdown of her8a causes ectopic neurog1 expression in rhombomeres 2 and 4. These results are surprising, since the Hes6-like factors studied to date tend to exhibit neurogenesis-promoting activity. When ectopically expressed, Hes6 promotes neurogenesis in the Xenopus embryo [27], the differentiation of cortical neurons at the expense of astrocytes in the mouse [44, 45], and the differentiation of retinal precursor cells into photoreceptors in mouse retinal explants [24]. These activities at least in part involve functionally antagonizing Hes1, since it was shown that Hes6 alone cannot bind the canonical E(spl) binding site (N box) [24, 45]. Rather, Hes6 dimerises with Hes1 and modifies its DNA binding properties [24], its capacity for recruiting the co-repressor Groucho or its stability [44]. Interestingly at least some of these properties appear controlled by the loop domain of Hes6, which is five amino acids shorter than that of other E(spl) proteins (see Additional file 3, Figure S1). Indeed, the addition of five amino acid residues into the loop of Hes6 confers Hes1-like repressor activity on the N box, while conversely, the removal of five amino acid residues from the loop of Hes1 completely ceases repression activity and confers Hes6-like activity [24]. We observed that Her8a has an intermediate loop-length compared to Hes6 and Hes1 (Additional file 3, Figure S1). Although the functional significance of this feature remains a matter for investigation, it is possible that it confers specific mechanistic properties to Her8a that distinguish it from Hes6 and bring it closer to the mode of action of Hes1-like proteins. In addition, our phylogenetic and synteny analyses revealed the complex evolution of Hes6-like genes and, contrary to previous belief, that Her8a is not a direct ortholog of Hes6. Rather, Hes6 orthologs comprise zebrafish Her13 and Hes6, both of which have the same loop length as mouse Hes6. We show here that zebrafish her13 is specifically expressed in a pattern coincident with neurogenesis (Additional file 4, Figure S2) reminiscent of Hes6 expression in the developing nervous system of both mouse and Xenopus, highlighting committed progenitors or early neurons [24, 27]. Thus we would predict that zebrafish Her13, rather than Her8a, shares functional properties with mammalian Hes6. The function of Hes6.2 subfamily proteins, to which Her8a belongs, has not been thoroughly tested, largely due to their absence in mammals. In the chicken neural tube however, Hes6.2 exerts a neurogenesis promoting activity [46], suggesting that Hes6.2 proteins may differ in their activities. We further propose that the splitting of her8a from locus 2 to a distinct genomic location (Figure 1C) permitted the acquisition of a unique expression profile for this gene in zebrafish.
Combined Her activities generate the midbrain-hindbrain progenitor pool pattern through different modalities
Our loss-of-function studies demonstrate that Her8a activity is necessary to prevent neurogenesis within the mediolateral territory of r2 and 4, hence keeping the proneural clusters for moto- and lateral neurons spatially separated within the anterior hindbrain. Combined with the fact that endogenous her8a expression does not depend on Notch at this developmental stage (Figure 6), this function is typical of a "pre-patterning" activity, comparable to that exhibited by the other E(spl) factors Her3, 5, 9 and 11 that delimit the territories competent for neurog1 expression within the neural plate [11, 13, 15]. Despite this functional relevance, the phenotype of her8a morphants appears very restricted compared to the broad expression of her8a, which encompasses the entire mid- and anterior hindbrain (Figure 4B). Functional redundancy and dosage effects have been described for other members of the E(spl) family in the mouse neural tube [20] and the zebrafish early neural plate [11, 22]. For example, Her5 and Her11 act in an equivalent and dose-dependent manner to block neurog1 expression in the medial and lateral aspects of the presumptive MHB [22], and Her3 and Her9 also cooperate to inhibit neurogenesis within the longitudinal stripe separating the presumptive moto- and interneuron clusters of the spinal cord [11]. We found that all her genes analyzed here (her3, 5, 9, 11 and 8a) were at least partially co-expressed within the presumptive MH (Figure 5L), strongly suggesting that redundancy may account for the normal development of this area in her8a morphants. The situation in the hindbrain, however, appears different. The only two her genes highlighting progenitor pools in r2-4 are her3 and her8a. While morphant embryos for each of these genes have an identical phenotype, our results argue against a dose-dependent mechanism involving Her3 and Her8a. Indeed, we found that the co-injection of her8aMO and her3MO at active doses did not produce an additional phenotype (Figure 3F) and that, if both morpholinos were injected together in amounts just below their effective concentration, ectopic neurog1 expression was not observed. Given that the two factors do not regulate each other's expression, these experiments suggest that the presence of each factor individually, rather than their overall dose, is relevant to maintain neurogenesis inhibition within r2 and r4. Although Her8a was isolated as a binding partner for Her5 in yeast cells, co-purification shows that the full length Her8a and Her3 proteins heterodimerise (Figure 4A), and the overlapping expression of her8a and her3 makes it possible that this interaction occurs in vivo. Her proteins can dimerise with a variety of partners, as also supported by our yeast two-hybrid results (Additional file 1, Table S1), and Her-Her heterodimers display enhanced stability over homodimers [7, 22]. A parsimonious interpretation of our results is therefore that the heterodimerisation of Her8a and Her3 is required for sufficient activity of these factors in r2 and r4. Alternatively, the individual activities of Her3 and Her8a may control complementary properties necessary to maintain the progenitor pool cell state.
In spite of the high level of her8a expression across the MHB progenitor pool, the results of the present paper also identify that the decisive inhibition of neurog1 expression in this location is played by other factors, namely Her3, 5, 9 and 11. Her5 and 11 were known for their dose-dependent redundant functions, accounting for neurogenesis inhibition across part of this domain [22]. Through knocking-down all four genes, we could achieve for the first time the transformation of most of the MHB into a neurogenic domain (Figure 5), while leaving her8a expression intact. Collectively, our findings show that the progenitor pool pattern of the midbrain and anterior hindbrain is established by the joined activities of five prepatterning E(spl) factors which act in different combination in the MHB and rhombomere domains. They also suggest that distinct mechanisms of action of these factors may be involved in these two domains.
Importantly however, we observed that the massive neurogenic phenotype induced upon blocking Her3/5/9/11 E(spl) activities is only partially followed by neuronal differentiation (Figure 5). In fact, ectopic neurons are restricted to the ventrolateral aspects of the midbrain-hindbrain boundary, like upon blocking Her5 function alone [21]. The corresponding progenitor population may be particularly prone to neuronal differentiation. For all other progenitors, our observations suggest the need for a further commitment event, independent of Her3/5/9/11 activities, to achieve neuronal differentiation following neurog1 induction. Blocking Notch signaling concomitantly to Her3/5/9/11 did not allow further neurogenesis progression (C. Stigloher, unpub.). Persistent her8a expression in this context may contribute to neurogenesis reversion, although it was not possible to evaluate this possibility as embryos blocked for the activities of all five E(spl) factors developed abnormally.
her8aexpression in the early neural plate is controlled by Sox transcription factors but not Notch signaling
Two types of her genes have been recently distinguished based on their Notch response profile: those acting as Notch mediators, depending on Notch signaling for their expression and overexpressed upon Notch activation, and "non-canonical" her genes endogenously independent of Notch and repressed when Notch is experimentally activated (reviewed in [12]). The former class comprises zebrafish her4.1 and her15, expressed in active neurogenic domains such as proneural clusters of the early neural plate [9–11]; the latter class is composed of her3, 5, 9 and 11, expressed in progenitor pools [11, 13, 15, 21, 22]. Our results illustrate that her8a is unusual in its expression pattern, which overlaps both proneural clusters and progenitor pools. This property is shared with its ortholog Hes6.2 in chicken [46]. her8a also shows a distinctive response to Notch signaling among hairy/E(spl) genes within the early neural plate, since it endogenously does not depend on Notch signaling but responds positively to the experimental activation of Notch (Figure 6). In agreement with the latter observation, we could identify Su(H) binding sites in the upstream regulatory sequence of her8a. However, this potential appears not to be used within proneural clusters of the early neural plate, demonstrating that Her8a is not a mediator of lateral inhibition. This is also in agreement with its uniform rather than salt-and-pepper expression profile. We found nevertheless that overexpressing her8a abolishes neurog1 expression even in proneural clusters where the two genes are normally co-expressed. Although we cannot ascertain that high overexpression levels mimic endogenous Her8a activity, one hypothesis reconciling this different information is that Her8a function within proneural clusters may generally dampen neurog1 expression, contributing to the function of other Her factors in Notch-inhibited precursors, and ensuring a proper differentiation schedule in committed progenitors. Although not further analyzed in this paper, we noted also that her8a expression becomes dependent on Notch signaling at later developmental stages (K. Webb, unpublished).
Our analyses of her8a expression in morphant contexts for other Her factors did not highlight cross-regulations, although we found several consensus N and E boxes within the 600 bp upstream of the her8a start site. Previous work demonstrated the positive regulation of Xenopus Hes6 by proneural bHLH proteins, in particular Neurogenin [27]. Given the presence of E boxes on the her8a promoter, and the co-expression of her8a and neurog1 in proneural clusters, it will be interesting to test whether her8a expression is also positively controlled by proneural factors in these locations. The Ngn1/Hes6 cascade is positively reinforcing proneural activity in Xenopus [27], but our functional data would predict an opposite outcome for a Neurog1/Her8a regulation in zebrafish.
Finally, our results show that the levels of her8a expression are under control of a combination of SoxB1 and B2 factors (Sox2/3/19a/19b and Sox21a, respectively) that display intense and partially overlapping expression within the anterior neural plate (Figure 7). In a recent study, Okuda et al. [41] demonstrated that SoxB1 factors function redundantly to control several successive aspects of zebrafish nervous system development, including neural plate patterning and primary neurogenesis. Although single morphants do not harbor a visible phenotype, her3 expression fails to be induced in quadruple SoxB1 morphants, strongly suggesting that the four factors act redundantly to activate her3 transcription [41]. In a comparable manner, we found that individual SoxB1/B2 morphants display no phenotype, while her8a expression is reduced in the MH domain when all five SoxB1/B2 proteins are abolished. Although we have not tested all possible knock-down combinations, and in particular did not assess the individual relevance of Sox21a in the context of the quadruple knock-out for SoxB1 proteins, these results demonstrate that her8a expression levels are under control of the activity of at least partially redundant SoxB proteins. Expression of these factors is an integral part of the mechanisms patterning the early embryo [40, 41], linking her8a expression with neural plate regionalization. The identification of a Sox2 binding site within the her8a enhancer, and the fact that all SoxB proteins recognize a similar binding motif in vitro, further suggests that part of this control may be direct. In support of this hypothesis, direct binding of SoxB1 factors onto the her3 enhancer has been demonstrated [41]. Like several other SoxB2 proteins, Sox21b was shown to act as a transcriptional inhibitor during dorsoventral patterning of the zebrafish gastrula [40] and generally promotes neurogenesis [47]. It can however act as an activator in other contexts [48], and its specific effect on MH neurogenesis and her genes needs to be directly evaluated. It was recently proposed that SoxB1 transcription factors and Notch cooperate through distinct mechanisms in their control of neurogenesis inhibition, including the inhibition of proneural protein activity and the transcriptional upregulation of Hes/her genes, respectively [49]. Our results and those of Okuda et al. [41] suggest yet another level of regulation, where SoxB proteins directly control the level of expression of some her genes. Whether this is limited to Notch-independent contexts, such as with the regulation of her3 and her8a, remains to be addressed.
Conclusions
In this work, we identify the Hairy/E(spl) transcription factor Her8a as a local inhibitor of neurogenesis in the developing hindbrain. Specifically, we show that Her8a function, like Her3 [11], is required to generate the non-neurogenic progenitor pools normally separating the presumptive moto- and lateral neurons of r2 and 4. We demonstrate that Her8a is a binding partner for Her3 and we propose that this interaction may be functionally relevant in r2 and r4. We further show that Her8a alone is not sufficient to inhibit neurog1 expression in the presumptive MHB area; this event depends, in contrast, on the combined activities of four other E(spl) factors, Her3, 5, 9 and 11. Unlike canonical E(spl) genes, we demonstrate that her8a does not depend on Notch signaling for its expression at early neural plate stages, and we identify a combination of SoxB factors that together enhance her8a expression. Finally, using phylogenetic analyses, we show that Her8a belongs to a Hes6-like subfamily that was recently lost in the mammalian lineage. This observation provides a context for the strikingly divergent functions of Her8a from Hes6; Hes6, which was previously believed to be the mammalian ortholog of Her8a, displays proneural activity. Together, our results characterize the phylogeny, expression and functional cascades involving a new Her factor, and highlight the complex interplay of E(spl) proteins that generates the neurogenesis pattern of the zebrafish midbrain-hindbrain area.
Methods
Yeast Two-Hybrid Analysis
Yeast two-hybrid screening was performed by Hybrigenics, S.A., Paris, France (http://www.hybrigenics.com). The coding sequence for amino acids 20 to 201 of the Danio rerio Her5 protein (GenBank proteic accession number gi: 18858797)
(amino acid sequence DRINQSLETLRMLLLENTNNEKLKNPKVEKAEILESVVHFLRAEQASETDPFQITRVKRARTEES
DEDVESPCKRQSYHDGMRTCLLRVSNFITGKSHEFGQELEKACENIHKSHSRQVQLLSTPSLIEPQVHLYEDPSQQHLAHVQLSNS
CTPSGCSKLAQRTVPAMTSSPKQPVMLCDPV)
was PCR-amplified and cloned into pB29 as an N-terminal fusion to LexA (N-Her5-LexA-C). The construct was checked by sequencing the entire insert and used as a bait to screen a random-primed Danio rerio embryo (stages 18-20 hpf) cDNA library constructed into pP6. pB29 and pP6 derive from the original pBTM116 [50] and pGADGH [51] plasmids, respectively. 76 million clones (7.6 -fold the complexity of the library) were screened using a mating approach with Y187 (mata) and L40DGal4 (mata) yeast strains as previously described [52]. 280 His+ colonies were selected on a medium lacking tryptophan, leucine and histidine, and supplemented with 2 mM 3-aminotriazole to handle bait autoactivation. The prey fragments of the positive clones were amplified by PCR and sequenced at their 5' and 3' junctions. The resulting sequences were used to identify the corresponding interacting proteins in the GenBank database (NCBI) using a fully automated procedure. For each interaction, a Predicted Biological Score (PBS) was computed to assess interaction reliability. This score represents the probability of an interaction being nonspecific. PBS relies on two different levels of analysis; the algorithm and methods used in the calculation are described in detail in Formstecher et al. [53]. Briefly, at first a local score takes into account the redundancy and independency of prey fragments (i.e. the times the interaction was detected with different independent clones and whether it was detected with different or the same fragments), as well as the distribution of reading frames and stop codons in overlapping fragments. Thus, interactions detected with several and different fragments are ranked with a very high confidence score and interactions detected with a single independent fragment are ranked with a moderate confidence score. Secondly, a global score takes into account the interactions found in all the screens performed at Hybrigenics using the same library. This global score represents the probability of an interaction being nonspecific. For practical use, the scores were divided into four categories, from A (highest confidence) to D (lowest confidence). A fifth category (E) specifically flags interactions involving highly connected prey domains previously found several times in screens performed on libraries derived from the same organism. Finally, several of these highly connected domains have been confirmed as false-positives of the technique and are now tagged as F. The PBS scores have been shown to positively correlate with the biological significance of interactions [54, 55].
Gene ontology analysis
Gene ontology enrichment analysis was performed on the recovered yeast-2-hybrid candidates from the categories A, B and C using the AmiGO "Term Enrichment tool" [56] (available at http://amigo.geneontology.org/cgi-bin/amigo/term_enrichment), using the following settings: ZFIN database as a background set, the maximum p-value set at 0.05 and a minimum number of gene products of two.
Sequence alignment, protein domain identification, phylogenetic and synteny analyses
Protein sequences were retrieved by using a tblastn search [57] against the non redundant database on NCBI or on Ensembl genomic data (current release of genomes, July 2010). For non-annotated sequences and to further support the expression of the predicted gene, a search for expressed sequence tags was also performed by tblastn on the EST database of the NCBI server. A list of all sequences used for the molecular phylogeny and their genomic locations is provided in Additional file 7, Table S3. Protein sequences were aligned using the ClustalX software. Only unambiguously aligned residues were retained to build the phylogenetic tree: namely the basic, HLH and Orange domains (see Additional file 3, Figure S1). Maximum likelihood phylogeny was constructed using PhyML (substitution model: JTT, number of substitution rate categories: 4; gamma distribution parameter estimated; proportion of invariable sites estimated) [58]. Branch support was estimated by approximate likelihood-ratio test (aLRT, SH-like) [59]. For the synteny analysis, the position and orientation of annotated genes surrounding hes6 family members was retrieved using the Ensembl genome browser.
Zebrafish strains
Adult zebrafish were maintained using standard fish-keeping protocols and in accordance with institute guidelines for animal welfare (defined by the Regierung von Oberbayern and the Services Vétérinaires de l'Essonne). Wildtype (AB) embryos were obtained through natural matings and were staged according to Kimmel et al. [60].
In situ hybridization
In situ hybridization on embryos was performed as previously described [22, 61] using the following probes: gfp [62], neurog1 [63], deltaNP63 [64], her4.1 [10], her5 [29], her9 [65], her3 [11] and barhl2 [30]. For the her8a and her13 probes, 648bp and 785bp fragments (respectively) were cloned into pCRII-TOPO (Invitrogen) from cDNA from 24hpf AB embryos using the following primers: for her8a forward 5'CACTGCTTGGAAGCAAATGA34, reverse 5'GACTTGGCGTGTGATTGATG3' and for her13 forward 5'TTTCTGTCCAACCCCTTCTG3', reverse 5'GATCCAATCCGATGTTGCTT3' (PCR conditions available on request). The successful clones were verified by sequencing. RNA probes were synthesized following published protocols [66]. For immunohistochemisty the primary antibody was mouse Anti-Hu (diluted 1:1500) (A21271; Molecular Probes) revealed using Cy-2.
Whole-mount immunohistochemistry
GFP of the -8.4neurog1:gfp line [67] was detected using a chicken primary antibody (GFP-1020, Aves Labs, Inc) followed by a secondary antibody anti-chicken coupled to Alexa-488 (Molecular Probes, Invitrogen).
RNA and morpholino injections
Capped RNAs were synthesized using Ambion mMessage mMachine Kit and embryos were injected at the one-cell stage. For her8a overexpression, full-length her8a was cloned by PCR (forward primer: 5'AATAATGACGGCCTCCAACA3'; reverse primer: 5'GGCTGCATTCATTCACCAG3') and cloned into pXT7. her8a capped mRNA was injected at a concentration of 62.5 ng/μl. NICD overexpression was achieved by injecting capped RNA for nic, which encodes the NICD fragment of zebrafish Notch1 [9, 10]. Morpholinos were purchased from Gene-Tools (Philomath, USA) and gripNAs were purchased from Active Motif (Carlsbad, USA). Two her8a splice morpholinos were used, targeted against the donor and acceptor sites of her8a exons 1 and 2 (respectively her8aMO1 and MO2, see Figure 3G). Both her8aMO1 (ATGTGACATTACCTTTCGCTCCTCT) and MO2 (CGCAGCTAAAATGATAGAAAGCATG) were injected at 1mM. Their efficiency was evaluated by RT-PCR following standard protocols on pools of 25 embryos at the 3-somite stage, in comparison to βactin2 expression, using the following PCR primers (FL: primer designed to amplify the full-length her8 transcript; E2: primer designed to amplify the E1-E2 domain; FW: forward primer; REV: reverse primer): her8a-FL-FW: AATAATGACGGCCTCCAACA; her8a-FL-REV: GGCTGCATTCATTCACCAG; her8a-E2-REV: TCCTCTCTCTGCGTTTCTTCTC; βactin2-FWD: AAGGCCAACAGGGAAAAGAT; βactin2-REV: GTGGTACGACCAGAGGCATAC (expected size 109bp). A third her8a MO, targeted against the her8a ATG (MO3: CATTGCCCATGTTGGAGGCCGTCAT) was also used and injected at 1.5 mM. The her3 morpholino [15] was injected at 0.5mM. For the combined knockdown of her3, her5, her9 and her11 the following gripNAs were used, each at 0.4mM: her3 gripNA (AGCCATTGTCCTTAAATG) (overlapping the MO sequence published in [15]), her5 gripNA (GGTTCGCTCATTTTGTGT) [22], her9 gripNA (TGATTTTTACCTTTCTAT) (overlapping the MO sequence published in [11]) and her11 gripNA (AGTCGGTGTGCTCTTCAT) [22]. For the combined knock-down of sox genes, we used: sox2/sox3 morpholino (CTCGGTTTCCATACATGTTATACATT) [42, 43] at 1mM (this MO was initially reported to target sox2 only, but we found that its target site is shared between sox2 and sox3), sox19a morpholino (TGCTGTACATGGCTGCCAACAGAAG) [41] at 1 mM, sox19b morpholino (TAGCCCTTTTCTCAAAACAAACCTG) at 0.25mM, sox21a morpholino (CATGGGCTTTGCCATTTCTTGATAC) (overlapping with the sox21aMO used in [40] at 1mM.
DAPT treatment
DAPT treatment was carried out according to Geling et al. [33]. Embryos were placed in 100 μm DAPT (Alexis Biochemicals), 1% DMSO in embryo medium from 50% epiboly to 3 somites. Control embryos received a corresponding treatment with 1% DMSO. After treatment, the embryos were fixed in 4% PFA overnight at 4°C before being processed for in situ hybridisation.
Co-affinity purification
The full length coding sequences of her3 and her8a were subcloned into the N-Strep/Flag (SF) TAP [68] and myc pCS3+ vectors, respectively. SF-TAP-tagged Her3 and Myc-tagged-Her8a were co-expressed in HEK293T cells. For this purpose 2 μg of pCS3-myc-her8a was co-transfected with either 6 μg pcDNA3.0-SF-TAP-her3 or control vector per 14 cm culture dish of HEK293T cells. After 48 h, cells were lysed in lysis buffer (TBS supplemented with Complete protease inhibitor (Roche), phosphatase inhibitor cocktails I and II (Sigma) and 0.5% NP-40 (Roche)). Beside the solubilization of cytoplasmic proteins, this condition allows the extraction of nuclear proteins which are not tightly bond to DNA. After incubation for 30 min at 4°C cell debris including nuclei were pelleted by centrifugation at 10,000 xg for 10 min, 4°C. The supernatants were filtered through 22-μm syringe filtration units (Millipore). Cleared lysates from two 14 cm culture dishes containing 5-6 mg total protein were incubated with 25 μl Strep-Tactin resin (IBA) for 2 h. After incubation the resin was washed 3 times with lysis buffer. The SF-TAP-tagged Her3 protein was eluted with 100 μl desthiobiotin elution buffer (IBA) and subjected to SDS PAGE. For detection, proteins were blotted onto PVDF membranes (GE Healthcare). The blots were incubated for 1 h with blocking solution (5% dry milk powder in TBST). The SF-TAP-tagged Her3 was detected by incubation of the blots with rabbit polyclonal anti FLAG antibodies (Sigma, 1:2000 in blocking reagent) overnight and secondary anti-rabbit antibodies (Jackson Immuno Research, 1:15,000 in blocking reagent) for 2 h. After each antibody incubation step, the blots were washed 4 times 5 min with TBST. For detection of Myc-Her8a, the blots were stripped using standard protocols and incubated with mouse anti-Myc (Cell signalling, 1:2000) overnight and secondary anti-mouse antibodies (Jackson Immuno Research, 1:15,000 in blocking reagent) for 2 h. Antibody-antigen complexes were visualized using the ECL+ chemiluminescence detection system (GE Healthcare) on Hyperfilms (GE Healthcare).
References
Easter SS, Burrill J, Marcus RC, Ross L, Taylor JSH, Wilson SW: Initial tract formation in the vertebrate brain. Progress in Brain Research. 1994, 102: 79-93.
Wilson SW, Ross LS, Parrett T, Easter SSJ: The development of a simple scaffold of axon tracts in the brain o the embryonic zebrafish Brachydanio rerio. Development. 1990, 108: 121-145.
Bally-Cuif L, Goridis C, Santoni MJ: The mouse NCAM gene displays a biphasic expression pattern during neural tube development. Development. 1993, 117: 543-552.
Chédotal A, Pourquié O, Sotelo C: Initial tract formation in the brain of the chick embryo: selective expression of the BEN/SC1/DM-GRASP cell adhesion molecule. Eur J Neurosci. 1995, 7: 198-212. 10.1111/j.1460-9568.1995.tb01056.x.
Massari ME, Murre C: Helix-loop-helix proteins: regulators of transcription in eucaryotic organisms. Mol Cell Biol. 2000, 20: 429-440. 10.1128/MCB.20.2.429-440.2000.
Fisher A, Caudy M: The function of hairy-related bHLH repressor proteins in cell fate decisions. Bioessays. 1998, 20: 298-306. 10.1002/(SICI)1521-1878(199804)20:4<298::AID-BIES6>3.0.CO;2-M.
Fischer A, Gessler M: Delta-Notch--and then? Protein interactions and proposed modes of repression by Hes and Hey bHLH factors. Nucleic Acids Res. 2007, 35: 4583-4596. 10.1093/nar/gkm477.
Appel B, Givan LA, Eisen JS: Delta-Notch signaling and lateral inhibition in zebrafish spinal cord development. BMC Dev Biol. 2001, 1: 13-10.1186/1471-213X-1-13.
Haddon C, Smithers L, Schneider-Maunoury S, Coche T, Henrique D, Lewis J: Multiple delta genes and lateral inhibition in zebrafish primary neurogenesis. Development. 1998, 125: 359-370.
Takke C, Dornseifer P, v Weizsäcker E, Campos-Ortega JA: her4, a zebrafish homologue of the Drosophila neurogenic gene E(spl), is a target of NOTCH signalling. Development. 1999, 126: 1811-1821.
Bae YK, Shimizu T, Hibi M: Patterning of proneuronal and inter-proneuronal domains by hairy- and enhancer of split-related genes in zebrafish neuroectoderm. Development. 2005, 132: 1375-1385. 10.1242/dev.01710.
Stigloher C, Chapouton P, Adolf B, Bally-Cuif L: Identification of neural progenitor pools by E(Spl) factors in the embryonic and adult brain. Brain Res Bull. 2008, 75: 266-273. 10.1016/j.brainresbull.2007.10.032.
Geling A, Itoh M, Tallafuss A, Chapouton P, Tannhäuser B, Kuwada JY, Chitnis AB, Bally-Cuif L: bHLH transcription factor Her5 links patterning to regional inhibition of neurogenesis at the midbrain-hindbrain boundary. Development. 2003, 130: 1591-1604. 10.1242/dev.00375.
Brewster R, Lee J, Ruiz i Altaba A: Gli/Zic factors pattern the neural plate by defining domains of cell differentiation. Nature. 1998, 393: 579-583. 10.1038/31242.
Hans S, Scheer N, Riedl I, v Weizsacker E, Blader P, Campos-Ortega JA: her3, a zebrafish member of the hairy-E(spl) family, is repressed by Notch signalling. Development. 2004, 131: 2957-2969. 10.1242/dev.01167.
Chapouton P, Adolf B, Leucht C, Ryu S, Driever W, Bally-Cuif L: her5 expression reveals a pool of neural stem cells in the adult zebrafish midbrain. Development. 2006, 133: 4293-4303. 10.1242/dev.02573.
Bally-Cuif L, Hammerschmidt M: Induction and patterning of neuronal development, and its connection to cell cycle control. Curr Opin Neurobiol. 2003, 13: 16-25. 10.1016/S0959-4388(03)00015-1.
Baek JH, Hatakeyama J, Sakamoto S, Ohtsuka T, Kageyama R: Persistent and high levels of Hes1 expression regulate boundary formation in the developing central nervous system. Development. 2006, 133: 2467-2476. 10.1242/dev.02403.
Hirata H, Tomita K, Bessho Y, Kageyama R: Hes1 and Hes3 regulate maintenance of the isthmic organizer and development of the mid/hindbrain. Embo J. 2001, 20: 4454-4466. 10.1093/emboj/20.16.4454.
Hatakeyama J, Bessho Y, Katoh K, Ookawara S, Fujioka M, Guillemot F, Kageyama R: Hes genes regulate size, shape and histogenesis of the nervous system by control of the timing of neural stem cell differentiation. Development. 2004, 131: 5539-5550. 10.1242/dev.01436.
Geling A, Plessy C, Rastegar S, Strähle U, Bally-Cuif L: Her5 acts as a prepattern factor that blocks neurogenin1 and coe2 expression upstream of Notch to inhibit neurogenesis at the midbrain-hindbrain boundary. Development. 2004, 131: 1993-2006. 10.1242/dev.01093.
Ninkovic J, Tallafuss A, Leucht C, Topczewski J, Tannhauser B, Solnica-Krezel L, Bally-Cuif L: Inhibition of neurogenesis at the zebrafish midbrain-hindbrain boundary by the combined and dose-dependent activity of a new hairy/E(spl) gene pair. Development. 2005, 132: 75-88.
Gajewski M, Elmasri H, Girschick M, Sieger D, Winkler C: Comparative analysis of her genes during fish somitogenesis suggests a mouse/chick-like mode of oscillation in medaka. Dev Genes Evol. 2006, 216: 315-332. 10.1007/s00427-006-0059-6.
Bae S, Bessho Y, Hojo M, Kageyama R: The bHLH gene Hes6, an inhibitor of Hes1, promotes neuronal differentiation. Development. 2000, 127: 2933-2943.
Jaillon O, Aury JM, Brunet F, Petit JL, Stange-Thomann N, Mauceli E, Bouneau L, Fischer C, Ozouf-Costaz C, Bernot A, Nicaud S, Jaffe D, Fisher S, Lutfalla G, Dossat C, Segurens B, Dasilva C, Salanoubat M, Levy M, Boudet N, Castellano S, Anthouard V, Jubin C, Castelli V, Katinka M, Vacherie B, Biemont C, Skalli Z, Cattolico L, Poulain J, De Berardinis V, Cruaud C, Duprat S, Brottier P, Coutanceau JP, Gouzy J, Parra G, Lardier G, Chapple C, McKernan KJ, McEwan P, Bosak S, Kellis M, Volff JN, Guigo R, Zody MC, Mesirov J, Lindblad-Toh K, Birren B, Nusbaum C, Kahn D, Robinson-Rechavi M, Laudet V, Schachter V, Quetier F, Saurin W, Scarpelli C, Wincker P, Lander ES, Weissenbach J, Roest Crollius H: Genome duplication in the teleost fish Tetraodon nigroviridis reveals the early vertebrate proto-karyotype. Nature. 2004, 431: 946-957. 10.1038/nature03025.
Thisse B, Thisse C: Fast release clones: a high throughput expression analysis. ZFIN Direct Data Submission. 2004
Koyano-Nakagawa N, Kim J, Anderson D, Kintner C: Hes6 acts in a positive feedback loop with the neurogenins to promote neuronal differentiation. Development. 2000, 127: 4203-4216.
Bakkers J, Hild M, Kramer C, Furutani-Seiki M, Hammerschmidt M: Zebrafish DeltaNp63 is a direct target of Bmp signaling and encodes a transcriptional repressor blocking neural specification in the ventral ectoderm. Dev Cell. 2002, 2: 617-627. 10.1016/S1534-5807(02)00163-6.
Müller M, v. Weizsäcker E, Campos-Ortega JA: Transcription of a zebrafish gene of the hairy-Enhancer of split family delineates the midbrain anlage in the neural plate. Dev Genes Evol. 1996, 206: 153-160. 10.1007/s004270050041.
Colombo A, Reig G, Mione M, Concha ML: Zebrafish BarH-like genes define discrete neural domains in the early embryo. Gene Expr Patterns. 2006, 6: 347-352. 10.1016/j.modgep.2005.09.011.
Latimer AJ, Shin J, Appel B: her9 promotes floor plate development in zebrafish. Dev Dyn. 2005, 232: 1098-1104. 10.1002/dvdy.20264.
Yeo SY, Kim M, Kim HS, Huh TL, Chitnis AB: Fluorescent protein expression driven by her4 regulatory elements reveals the spatiotemporal pattern of Notch signaling in the nervous system of zebrafish embryos. Dev Biol. 2007, 301: 555-567. 10.1016/j.ydbio.2006.10.020.
Geling A, Steiner H, Willem M, Bally-Cuif L, Haass C: A gamma-secretase inhibitor blocks Notch signaling in vivo and causes a severe neurogenic phenotype in zebrafish. EMBO Rep. 2002, 3: 688-694. 10.1093/embo-reports/kvf124.
Crawford TQ, Roelink H: The notch response inhibitor DAPT enhances neuronal differentiation in embryonic stem cell-derived embryoid bodies independently of sonic hedgehog signaling. Dev Dyn. 2007, 236: 886-892. 10.1002/dvdy.21083.
Frech K, Danescu-Mayer J, Werner T: A novel method to develop highly specific models for regulatory units detects a new LTR in GenBank which contains a functional promoter. J Mol Biol. 1997, 270: 674-687. 10.1006/jmbi.1997.1140.
Ambrosetti DC, Basilico C, Dailey L: Synergistic activation of the fibroblast growth factor 4 enhancer by Sox2 and Oct-3 depends on protein-protein interactions facilitated by a specific spatial arrangement of factor binding sites. Mol Cell Biol. 1997, 17: 6321-6329.
Okuda Y, Yoda H, Uchikawa M, Furutani-Seiki M, Takeda H, Kondoh H, Kamachi Y: Comparative genomic and expression analysis of group B1 sox genes in zebrafish indicates their diversification during vertebrate evolution. Dev Dyn. 2006, 235: 811-825. 10.1002/dvdy.20678.
Thisse B, Pflumio S, Fürthauer M, Loppin B, Heyer V, Degrave A, Woehl R, Lux A, Steffan T, Charbonnier XQ, Thisse C: Expression of the zebrafish genome during embryogenesis. ZFIN Direct Data Submission. 2001
Rimini R, Beltrame M, Argenton F, Szymczak D, Cotelli F, Bianchi ME: Expression patterns of zebrafish sox11A, sox11B and sox21. Mech Dev. 1999, 89: 167-171. 10.1016/S0925-4773(99)00199-9.
Argenton F, Giudici S, Deflorian G, Cimbro S, Cotelli F, Beltrame M: Ectopic expression and knockdown of a zebrafish sox21 reveal its role as a transcriptional repressor in early development. Mech Dev. 2004, 121: 131-142. 10.1016/j.mod.2004.01.001.
Okuda Y, Ogura E, Kondoh H, Kamachi Y: B1 SOX coordinate cell specification with patterning and morphogenesis in the early zebrafish embryo. PLoS Genet. 2010, 6: e1000936-10.1371/journal.pgen.1000936.
Christen B, Robles V, Raya M, Paramonov I, Belmonte JC: Regeneration and reprogramming compared. BMC Biol. 2010, 8: 5-10.1186/1741-7007-8-5.
Pujic Z, Omori Y, Tsujikawa M, Thisse B, Thisse C, Malicki J: Reverse genetic analysis of neurogenesis in the zebrafish retina. Dev Biol. 2006, 293: 330-347. 10.1016/j.ydbio.2005.12.056.
Gratton MO, Torban E, Jasmin SB, Theriault FM, German MS, Stifani S: Hes6 promotes cortical neurogenesis and inhibits Hes1 transcription repression activity by multiple mechanisms. Mol Cell Biol. 2003, 23: 6922-6935. 10.1128/MCB.23.19.6922-6935.2003.
Jhas S, Ciura S, Belanger-Jasmin S, Dong Z, Llamosas E, Theriault FM, Joachim K, Tang Y, Liu L, Liu J, Stifani S: Hes6 inhibits astrocyte differentiation and promotes neurogenesis through different mechanisms. J Neurosci. 2006, 26: 11061-11071. 10.1523/JNEUROSCI.1358-06.2006.
Fior R, Henrique D: A novel hes5/hes6 circuitry of negative regulation controls Notch activity during neurogenesis. Dev Biol. 2005, 281: 318-333. 10.1016/j.ydbio.2005.03.017.
Sandberg M, Kallstrom M, Muhr J: Sox21 promotes the progression of vertebrate neurogenesis. Nat Neurosci. 2005, 8: 995-1001. 10.1038/nn1493.
Hwang CK, Wu X, Wang G, Kim CS, Loh HH: Mouse mu opioid receptor distal promoter transcriptional regulation by SOX proteins. J Biol Chem. 2003, 278: 3742-3750. 10.1074/jbc.M208780200.
Holmberg J, Hansson E, Malewicz M, Sandberg M, Perlmann T, Lendahl U, Muhr J: SoxB1 transcription factors and Notch signaling use distinct mechanisms to regulate proneural gene function and neural progenitor differentiation. Development. 2008, 135: 1843-1851. 10.1242/dev.020180.
Vojtek AB, Hollenberg SM: Ras-Raf interaction: two-hybrid analysis. Methods Enzymol. 1995, 255: 331-342.
Bartel PL: Unsing the two-hybrid system to detect protein-protein interactions. In Cellular Interactions in Development: A Practical Approach. 1993, 153-179.
Fromont-Racine M, Rain JC, Legrain P: Toward a functional analysis of the yeast genome through exhaustive two-hybrid screens. Nat Genet. 1997, 16: 277-282. 10.1038/ng0797-277.
Formstecher E, Aresta S, Collura V, Hamburger A, Meil A, Trehin A, Reverdy C, Betin V, Maire S, Brun C, Jacq B, Arpin M, Bellaiche Y, Bellusci S, Benaroch P, Bornens M, Chanet R, Chavrier P, Delattre O, Doye V, Fehon R, Faye G, Galli T, Girault JA, Goud B, de Gunzburg J, Johannes L, Junier MP, Mirouse V, Mukherjee A, Papadopoulo D, Perez F, Plessis A, Rosse C, Saule S, Stoppa-Lyonnet D, Vincent A, White M, Legrain P, Wojcik J, Camonis J, Daviet L: Protein interaction mapping: a Drosophila case study. Genome Res. 2005, 15: 376-384. 10.1101/gr.2659105.
Rain JC, Selig L, De Reuse H, Battaglia V, Reverdy C, Simon S, Lenzen G, Petel F, Wojcik J, Schachter V, Chemama Y, Labigne A, Legrain P: The protein-protein interaction map of Helicobacter pylori. Nature. 2001, 409: 211-215. 10.1038/35051615.
Wojcik J, Boneca IG, Legrain P: Prediction, assessment and validation of protein interaction maps in bacteria. J Mol Biol. 2002, 323: 763-770. 10.1016/S0022-2836(02)01009-4.
Carbon S, Ireland A, Mungall CJ, Shu S, Marshall B, Lewis S: AmiGO: online access to ontology and annotation data. Bioinformatics. 2009, 25: 288-289. 10.1093/bioinformatics/btn615.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ: Basic local alignment search tool. J Mol Biol. 1990, 215: 403-410.
Guindon S, Gascuel O: A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol. 2003, 52: 696-704. 10.1080/10635150390235520.
Anisimova M, Gascuel O: Approximate likelihood-ratio test for branches: A fast, accurate, and powerful alternative. Syst Biol. 2006, 55: 539-552. 10.1080/10635150600755453.
Kimmel CB, Ballard WW, Kimmel SR, Ullmann B, Schilling TF: Stages of embryonic development of the zebrafish. Dev Dyn. 1995, 203: 253-310. 10.1002/aja.1002030302.
Hammerschmidt M, Bitgood MJ, McMahon AP: Protein kinase A is a common negative regulator of Hedgehog signaling in the vertebrate embryo. Genes Dev. 1996, 10: 647-658. 10.1101/gad.10.6.647.
Blader P, Plessy C, Strahle U: Multiple regulatory elements with spatially and temporally distinct activities control neurogenin1 expression in primary neurons of the zebrafish embryo. Mech Dev. 2003, 120: 211-218. 10.1016/S0925-4773(02)00413-6.
Korzh V, Sleptsova I, Liao J, He J, Gong Z: Expression of zebrafish bHLH genes ngn1 and nrd defines distinct stages of Neural differentiation. Dev Dynam. 1998, 213: 92-104. 10.1002/(SICI)1097-0177(199809)213:1<92::AID-AJA9>3.0.CO;2-T.
Kudoh T, Tsang M, Hukriede NA, Chen X, Dedekian M, Clarke CJ, Kiang A, Schultz S, Epstein JA, Toyama R, Dawid IB: A gene expression screen in zebrafish embryogenesis. Genome Res. 2001, 11: 1979-1987. 10.1101/gr.209601.
Leve C, Gajewski M, Rohr KB, Tautz D: Homologues of c-hairy1 (her9) and lunatic fringe in zebrafish are expressed in the developing central nervous system, but not in the presomitic mesoderm. Dev Genes Evol. 2001, 211: 493-500. 10.1007/s00427-001-0181-4.
Hauptmann G, Gerster T: Two-colour whole-mount in situ hybridization to vertebrate and Drosophila embryos. Trends Genet. 1994, 10: 266-
Blader P, Lam CS, Rastegar S, Scardigli R, Nicod JC, Simplicio N, Plessy C, Fischer N, Schuurmans C, Guillemot F, Strahle U: Conserved and acquired features of neurogenin1 regulation. Development. 2004, 131: 5627-5637. 10.1242/dev.01455.
Gloeckner CJ, Boldt K, Schumacher A, Roepman R, Ueffing M: A novel tandem affinity purification strategy for the efficient isolation and characterisation of native protein complexes. Proteomics. 2007, 7: 4228-4234. 10.1002/pmic.200700038.
Acknowledgements and Funding
We thank colleagues from the past and present Bally-Cuif lab for their valuable input into this project, and the fish house teams at the Helmholtz Zentrum München and CNRS. Work in the LBC laboratory at the Helmholtz Zentrum München was funded by the Helmholtz Association, the EU ZF-Models integrated project (contract No. LSHG-CT-2003-503466) and the Center for Protein Science-Munich (CIPSM). Work in the LBC lab at the CNRS is supported by funds from the EU 7th framework integrated projects NeuroXsys and ZF-Health, the Agence Nationale pour la Recherche, the Ecole des Neurosciences de Paris, the Fondation pour la Recherche Médicale, the PIME program and the Schlumberger Association. MC has been recipient of a Marie Curie Intra-European Fellowship and an EMBO Long Term postdoctoral fellowship. Work of MU and CJG is supported by the Helmholtz Alliance for Mental Health in an Aging Society (HelMA, FKZ: HA-215) and the BMBF (neurogenesis from brain and skin cells, FKZ: 01 GN 1009 C), respectively.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Authors' contributions
KJW designed the two-hybrid assay, characterized the recovered proteins, identified her8a and carried out its initial functional characterization. She participated in the design of the study and helped to draft the manuscript. MC conducted the genomic and evolutionary analyses of Her8a, validated the efficiency of MO1 and MO2, conducted the functional characterization of Her8a using MO2 (including late fate analyses), and participated in the analyses of her8a regulation by Sox proteins. She drafted the relevant parts of the manuscript and provided critical input on the study as a whole. CJG developed the STREP/Flag assay in the laboratory of MU and conducted the co-purification experiments under his supervision and advice. He wrote the relevant sections of the manuscript. CS characterized the combined activities of her3, 5, 9 and 11 and provided critical input during the study. BB and ST carried out the expression analyses in situ. LBC conceived the study, participated in its design and coordination, and drafted the manuscript together with KJW. All authors read and approved the manuscript.
Katharine J Webb, Marion Coolen contributed equally to this work.
Electronic supplementary material
12861_2010_631_MOESM1_ESM.DOC
Additional file 1: Table S1. Summary of yeast two-hybrid results. The 75 unique protein-protein interactions recovered are listed, ordered according to their interaction scores (PBS: Protein Binding Score). The recovered her genes are underlined. (DOC 140 KB)
12861_2010_631_MOESM2_ESM.DOC
Additional file 2: Table S2. Gene Ontology enrichment analysis. This analysis was conducted on the recovered yeast-2-hybrid candidates from the categories A, B and C. (DOC 44 KB)
12861_2010_631_MOESM3_ESM.PDF
Additional file 3: Figure S1. Alignment of mouse (Mm) Hes1, Hes5 and Hes6 with Hes6-like proteins from zebrafish (Dr) (Her8a, Her8.2, Her13 and Hes6) and other species (Fr: Fugu rubripes, Ol: Oryzias latipes, Gg: Gallus gallus, Hs: Homo sapiens, Xt: Xenopus tropicalis). Protein domains are color-coded: green: basic domain, purple: HLH domain, orange: orange domain. Note the signature of the Hes6-like subfamily: the loops of Her13 and Hes6 contain 5 less amino acids when compared with other members, and the loops of Her8a and Her8.2 have 3 less amino acids (domain overlayed with the black bar). In addition, the Orange domains of Hes6-like proteins are 65-86% identical to each other, while they show only 23-37% identity with the Orange domains of other Her/Hes proteins. Finally, similarities extending beyond the domains above also identify the Hes6.1 and Hes6.2 subfamilies within Hes6-like proteins (hatched red and blue boxes). (PDF 3 MB)
12861_2010_631_MOESM4_ESM.JPEG
Additional file 4: Figure S2. her13 expression highlights early neurogenesis domains during zebrafish embryonic development. Whole-mount in situ hybridization for her13 (blue staining) at the stages indicated (A-C and E are lateral views, D and F are dorsal views, all with anterior to the left). Note her13 expression in proneural clusters encompassing presumptive spinal interneurons (arrows) and sensory neurons (arrowheads), trigeminal ganglion neurons (asterisk), telencephlaic (t) and epiphyseal (e) neurons. (JPEG 1 MB)
12861_2010_631_MOESM5_ESM.JPEG
Additional file 5: Figure S3. Neural plate patterning is unaffected upon blocking Her8a function. Whole-mount in situ hybridization for barhl2 (A,B), her5 (C,D) and her9 (E,F) in embryos injected with her8a capped mRNA (right column) compared to control embryos (left column). Dorsal views of whole-mount embryos are shown, anterior to the top. All three markers highlight defined neural plate territories (barhl2: transverse diencephalic domain; her5: prospective midbrain-hindbrain boundary; her9: prospective eye field, midbrain-hindbrain boundary and lateral rhombencephalic stripes, see Figure 5) and appear identically expressed in the wild-type and morphant neural plate. Abbreviations: d: diencephalon; e: eye field; mhb: midbrain-hindbrain boundary; sc: presumptive spinal cord. (JPEG 1 MB)
12861_2010_631_MOESM6_ESM.JPEG
Additional file 6: Figure S4. her8a MO1 and her8a MO2 have identical effects on neurog1 expression. Whole-mount in situ hybridization for neurog1 expression in embryos injected with her8aMO1 (B) or her8aMO2 (C) compared to control embryos (whole-mount views of 3 somite-embryos, anterior to the top). neurog1 expression is ectopically induced between the clusters of motoneurons and lateral neurons in rhombomeres 2 and 4 (blue arrowheads in B,C), a location normally devoid of neurog1 transcripts (white arrowheads in A). The phenotype is highly reproducible and identical in both morphant groups. (JPEG 2 MB)
12861_2010_631_MOESM7_ESM.JPEG
Additional file 7: Table S3. List of the sequences used for the molecular phylogeny (Figure 1) and their genomic locations. (JPEG 995 KB)
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under license to BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Webb, K.J., Coolen, M., Gloeckner, C.J. et al. The Enhancer of split transcription factor Her8a is a novel dimerisation partner for Her3 that controls anterior hindbrain neurogenesis in zebrafish. BMC Dev Biol 11, 27 (2011). https://doi.org/10.1186/1471-213X-11-27
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-213X-11-27