Abstract
Background
DNA copy number variation (CNV) has been recognized as an important source of genetic variation. Array comparative genomic hybridization (aCGH) is commonly used for CNV detection, but the microarray platform has a number of inherent limitations.
Results
Here, we describe a method to detect copy number variation using shotgun sequencing, CNV-seq. The method is based on a robust statistical model that describes the complete analysis procedure and allows the computation of essential confidence values for detection of CNV. Our results show that the number of reads, not the length of the reads is the key factor determining the resolution of detection. This favors the next-generation sequencing methods that rapidly produce large amount of short reads.
Conclusion
Simulation of various sequencing methods with coverage between 0.1× to 8× show overall specificity between 91.7 – 99.9%, and sensitivity between 72.2 – 96.5%. We also show the results for assessment of CNV between two individual human genomes.
Similar content being viewed by others
Background
DNA copy number variation (CNV) has long been known as a source of genetic variation, but its importance has only been recognized recently [1, 2]. In a landmark study in 2006, Redon and colleagues found that 1,447 CNV regions cover at least 12% of the human genome, with no large stretches exempt from CNV [3]. The CNV regions cover more nucleotide content per genome than single nucleotide polymorphisms (SNPs), suggesting the importance of CNV in genetic diversity [3]. A common way to detect CNV is to utilize microarray-based methods [4]. The most commonly used method, array comparative genomic hybridization (aCGH) was first used to detect CNV a decade ago [5, 6].
Microarray-based methods have revolutionized the way of how large-scale genome studies are carried out. Today, the next-generation sequencing technologies are transforming biology research [7]. The rapid development of new sequencing technologies is continuously increasing the speed of sequencing and decreasing the cost. The next-generation sequencing, such as 454 [8], Solexa [9] and SOLiD [10] have already showed advantages over microarrays in several aspects. Apart from being rapid and cheap, data produced by sequencing can be re-used for varied purposes as opposed to data from microarray-based methods that can usually solely be used by one specific study. In addition, reproducibility has been one of the major challenges for microarray technology [11]. The once revolutionizing microarray-based ChIP-Chip technology is being replaced by ChIP-Seq, in which the DNA fragments are sequenced instead of being hybridized to an array [12]. Sequencing-based methods are also used to produce genome-wide DNA methylation profiles, detect SNP, study chromosome translocations and RNA transcriptome profiling [13–20].
Variation in sequencing coverage in genome assemblies has been used as an indicator for potential CNV between an assembled genome and shotgun data from another genome [21, 22]. This is analogous to a comparison of copy number between microarray probes and a single set of DNA fragments. There are two major problems with this kind of approach. Given a certain hybridization condition, hybridization efficiency varies among microarray probes. Likewise, given a certain alignment threshold, sequencing errors in combination with differences between genomes may result in erroneous distribution of the reads.
Secondly, the number of probes on a microarray does not represent the real copy number of probe sequences in a genome. Likewise, the copy number of DNA segments in an assembled genome may not represent the true one. Notably, the regions containing multiple copies are the most difficult to assemble correctly and is still the key unsolved problem in shotgun assembly [23]. Assembly errors like these cause false variation in the sequencing coverage and thus yield erroneous indication of CNV.
In this paper we describe an efficient solution based on a robust model that combines the advantages of aCGH and high-throughput sequencing. We also assessed CNV between two individuals (Dr. J. Craig Venter [24], Dr. James Watson [21]). An implementation of our method is freely available at http://tiger.dbs.nus.edu.sg/CNV-seq.
Results and discussion
The Model
We have developed a method to detect CNV by shotgun sequencing, CNV-seq. The method is based on a robust statistical model that allows confidence assessment of observed copy number ratios and is conceptually derived from aCGH (Figure 1). The microarray-based procedure, aCGH involves a whole genome microarray where two sets of labeled genomic fragments are hybridized. Instead of a microarray, CNV-seq uses a sequence as a template and two sets of shotgun reads, one set from each target individual, X and Y (Figure 1). The two sets of shotgun reads are mapped by sequence alignment on a template genome. We use a sliding window approach to analyze the mapped regions and CNVs are detected by computing the number of reads for each individual in each of the windows, yielding ratios. These observed ratios are assessed by the computation of a probability of a random occurrence, given no copy number variation.
A comparison of the conceptual steps in aCGH and CNV-seq methods. 1. Starting material in both cases is genomic fragments from two genomes. 2. In CNV-seq the fragments are samples and sequenced. 3. Genomic fragments are directly hybridized on to an array. In CNV-seq the mapping is performed by sequence alignment. 4. In microarray the light intensities reflect the number of hybridized fragments. In CNV-seq the number of mapped reads are counted directly. 5. Data analysis, including estimation of copy number ratios, confidence values, etc. 6. Output of the results.
The random sampling in shotgun sequencing results in uneven coverage that may lead to observed coverage ratios that falsely imply CNV. Therefore, a statistical model is essential for the assessment of the probability of false positive ratios. The average number of reads in a region of a genome is dependent on the total number of reads sampled, the length of the genome and the length of the region. We use this relationship to compute a minimum sliding window size to achieve a desired minimum confidence level of the observations.
The mean number of reads for X and Y genomes in a sliding window determines the distribution of the ratios. The number of reads in a window is approximately distributed according to Poisson, Po(λ), where the mean number of reads per window is λ, given by
where N is the total number of sequenced reads, G is the size of the genome and W is the size of the sliding window, and W < <G. We use the Gaussian distribution to approximate the Poisson distribution with mean and variance λ = μ = σ2. This approximation is good when the mean number of reads per window is greater than 10 with continuity correction.
The predicted copy number ratio, r in each sliding window can be computed by
where z is the ratio of read counts in the window and N X and N Y are the total number of reads in the genomes X and Y respectively. Assuming an independent distribution of the read counts, we can obtain a probability, p of a copy number ratio being r or divergent from 1:1 ratio by a random chance. For this purpose, we need the distribution of the read count ratio z. This distribution is given by Gaussian ratio distribution [25]. The computation of this distribution is cumbersome, but it can be transformed to another variable, t, by Geary-Hinkley transformation [26]:
where μ X , , μ Y and are the means and the variances for X and Y respectively. The new variable t approximately have a standard Gaussian distribution when the mean number of reads per window is greater than 6 in Y and less than 40,000 in X. The p-value can be computed by
where Φ (t) is the cumulative standard Gaussian distribution function. The probability p decreases with increasing sliding window size (Figure 2) and we would like p to be as low as possible. Conversely, increased sliding window size leads to a decreased resolution of CNV regions. Therefore it is advantageous to compute a window size, which yields the best possible resolution according to a preset threshold of p for r. Based on the above equations, We can calculate the best possible resolution, or the theoretical minimum window size, W by
Dependencies of p in CNV-seq. The relation of p and sliding window size is shown on 0.1× to 8× sequence coverage for log2(r') = 0.6 and average read length 250 bases. The values are computed using equation (5). Increased window length results in decreased probability, p of observing ratio r' or higher by cheer chance. It is possible to compensate lack of coverage by increasing the window size, but this results in lowering the resolution.
and
where p' is the desired significance level, and r' is the CNV detection threshold ratio. Φ-1 is the inverse function of Φ. The number of reads sampled will affect the minimum window size. For example, if one wants to detect CNV with ratio ≥ 3 : 2 at significance level 0.002, a genome size of 3 G bases and 10 M reads in both genomes will yield the minimum window size of 37,243 bases, while 1 M reads will yield the window size of 372,431 bases. The use larger number of reads allows detection of ten times shorter CNV.
An alternative approach is to calculate the range of copy number ratios that can be detected at a certain significance level p', with a predefined window size W':
where
and
Validation
In order to assess the performance of CNV-seq, we used simulated and real human data. For the simulation of shotgun data, in total of 101 genomes were constructed, containing varied number, sizes and locations of CNV regions, SNP and short insertions/deletions (indels). We simulated three sequencing methods, Solexa, 454 and Sanger [27] for each constructed genome on 0.1× to 8× coverage. This resulted in the total of 8,400 simulations.
The Figure 3 shows the results of the simulations on varied coverage and varied p' for constant log2(r') = 0.6. Each dot represents an average of 100 simulations and the sizes of the dots reflect the sizes of the lengths of the sliding windows that are the theoretical minimum lengths, given by equation (5). The overall specificity for our method is between 91.7 – 99.9%, the sensitivity between 72.2 – 96.5% with the median of 99.4% and 89.9% respectively. The mean sequence length is dependent on the technology simulated. Thus, in order to reach the same coverage, a larger number of fragments need to be sequenced when sequencing is performed with Solexa, which produces short reads compared to the Sanger and 454 methods. According to our model, the largest number of sequenced reads yields the shortest length of the sliding window and thus the best resolution. The range of window sizes in our simulations varies from 1,103 bases to 2,951,792 bases, decreasing with increasing average sequencing coverage. The results show that our model performs well in the presence of errors. Despite of increased resolution due to shortening of the sliding window size, the sensitivity is increased together with increased sequencing coverage. Slight drop in specificity with increasing sequencing coverage can be observed (Figure 3). This is likely to be due to SNPs, short indels, and read mapping errors, that are not considered in our statistical model and have a more profound effect on small windows. The specificity does not drop in error free data. The effect of errors may be reduced by using a window size that is larger than the theoretical minimum. For example, the theoretical minimum window for 8× Solexa sequencing at p = 0.001 is 1947 bases. This window size gives a specificity of 95.4%, while a 2 times larger window yields specificity of 97.8% (Figure 4).
Performance of CNV-seq. The performance of CNV-seq on data simulating 454, Sanger and Solexa methods. Results are shown for 0.1×–8× coverages (right) and p-value range of 10-5-10-2 (top). Each dot represents an average of 100 simulations and the size of the dots represents the window (log10) size, i.e. resolution used. The window sizes are calculated using equation (5).
Specificity vs window size. In order to increase specificity, a larger than the theoretical minimum window size can be used by sacrificing resolution. The specificities using 1×, 1.5×, 2×, 3×, 4×, and 5× of the theoretical minimum window size are shown, for simulated Solexa sequencing data at 8× coverage.
Analysis of human data
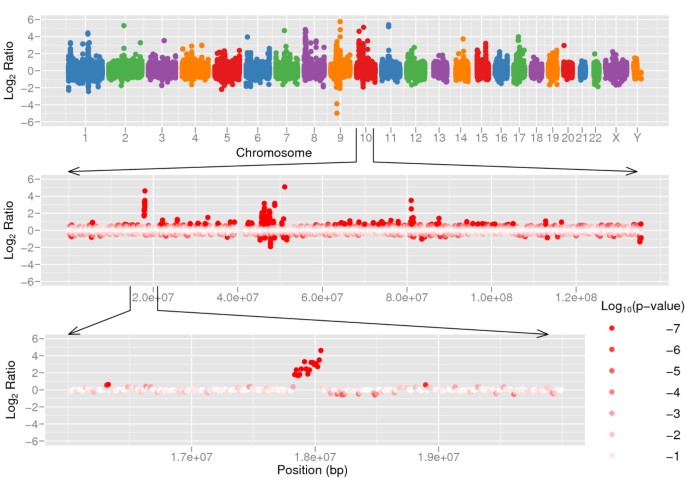
The genomes of two individuals, Dr. Craig J. Venter and Dr. J. Watson were recently sequenced on 7.5× and 7.4× coverage respectively [21, 24]. The genome of Dr. Craig J. Venter is sequenced using Sanger method and Dr. J. Watson's genome using 454 technology. We compared the two genomes using CNV-seq (Figure 5 and Additional File 1). The thresholds p' = 10-5 and log2(r') = 0.6 yield sliding window size of 26,481 bases for autosomal chromosomes. The sex chromosomes have a lower sequencing coverage than autosomal chromosomes, therefore larger window sizes are used: 72,044 bases for chromosome X and 269,032 bases for chromosome Y. We identified 174 contiguous regions covered by four or more consecutive windows. The sizes of these regions range from 66,202 bases to 941,612 bases.
Copy number variation between two human individuals. Copy number variation detected by CNV-seq using shotgun sequence data from two individuals, Venter and Watson. The top panel shows a genome level log 2 ratio plot. The middle panel shows the plot for chromosome 10. The bottom panel shows detailed view of a CNV region in chromosome 10. The red color gradient in the middle and bottom sections represents log10 p calculated on each of ratios.
The comparison of the 174 CNV calls with those in the Database of Genomic Variants (DGV) [2] revealed 142 of the CNV calls to overlap more than 50% with previously reported CNV regions. In order to asses the significance of CNV calls, we performed 5,000 permutation tests, using 174 randomly distributed CNV regions of the same sizes as in the original experiment. In average, only 56 and maximum 78 of 174 regions overlap more than 50% with CNV in DGV (Figure 6) 5,000 random sets. The real CNV calls have significantly larger overlap with DGV (p = 0).
Permutation test of CNV calls. A permutation test was performed in order to test the significance of our CNV calls. The histogram shows the number of CNV calls overlapping with CNV in DGV. The X-axis shows the number of calls overlapping with DGV. The Y-axis shows the frequency of the overlapping number in 5,000 sets of permuted CNV calls.
We also intersected the CNV calls with the CNVs identified by aCGH in the two genomes. There are 23 and 45 CNV regions reported in Watson's and Venter's genome respectively [21, 24]. We found 15 of our CNV calls overlap with 10 of previously reported Watson's CNV regions, and only 11 of our CNV calls overlap with 5 of Venter's. The low overlap with Venter's CNV calls made by aCGH is not surprising, for the reason that the majority of the CNV regions were detected by only one of three microarray platforms [24]. There are 121 CNV calls that made by CNV-seq but not aCGH and overlap with DGV data, suggesting that CNV-seq can detect CNV regions that were missed by aCGH. One of these regions is shown in Figure 5 (bottom panel), a 238 kb region (copy number ratio 6:1, p = 0) containing two genes (FAM23B, MRC1L1) and one miRNA (hsa-mir-511-2). We have used stringent thresholds in our analysis, thus by lowering thresholds, such as p-value and the number of consecutive windows, will increase the number of reported CNV calls.
A major assumption in CNV-seq is that shotgun sampling of DNA fragments is random, and therefore the CNV calls made by CNV-seq are not due to different sequencing bias between the two sets of data compared. When the two sets of data are prepared in the same way, this assumption is valid. However, when the shotgun sequences are generated using two different sequencing methods, the assumption may not hold any more. Solexa sequencing reads are recently reported to be GC-biased dependent on a library preparation procedure [28]. Venter's genome was sequenced using 454 and Watson's genome was sequenced using the Sanger method. We compared the distribution of GC frequencies in the shotgun reads in both genomes. There are no significant differences between the two distributions (p = 0.2106, Kolmogorov-Smimrov test).
Conclusion
We have developed a method to detect CNV using shotgun data. Our approach not only combines the advantages of microarray methods and high-throughput sequencing, but is also based on a robust statistical model allowing confidence assessment. We tested the approach on both simulated and real data and the results show that the method can be applied to relatively low sequencing coverage with good specificity and sensitivity. We have also developed a model to compute the theoretical limit of resolution for given data at a desired confidence level.
We expect the continued rapid development of sequencing technologies to further lower the cost and increase the speed of sequencing. Thus, sequencing-based approaches are likely to gain increased advantage over microarrays. Next-generation sequencing methods mostly produce a large number of short reads and our results show that the number of reads sequenced – not the length of the reads, is the most important factor that determines the resolution, i.e. larger number of sequenced fragments results in increased resolution. Alternatively, given a constant resolution an increase in the number of sequenced reads will result in increased sensitivity and specificity. Therefore, a large number of short reads is an advantage as opposed to a small number of long reads.
Methods
Simulations
The human chromosome 1 (NCBI build 36) was used to construct one diploid reference genome and 100 diploid test genomes. The unmodified chromosome 1 sequence was used as the template genome. The test individual genomes are constructed by the introducing CNV, SNPs and short indels. The CNV is introduced into each of the test genomes by concatenating the two chromosomes and by selecting nine source sequences at random positions to replace 26 target sequences at random positions. Four of the nine source sequences are used four times each to replace four random target sequences and the remaining five of the nine sequences are used to replace two random target sequences each. The procedure results in the total of 35 segments in each of the 100 simulated test genomes with the following copy number ratios: 26 with ratio 1:2, five with ratio 4:2 and four with ratio 6:2. The length of the source sequences is 10k, where k is a random number between log10 500 and log10 2 M, yielding the median length of 26,464 bases and the mean 234,065.7 bases. In addition, each test genome is modified by randomly introducing 5 SNPs/kb and short, 1–3 bp insertions/deletions with the frequency of 0.5 indels/kb.
The reference genome is constructed the same way as the individual test genomes, except no CNV was introduced.
We simulated the shotgun sequencing process for test and reference genomes by using real sequence quality files, specific for each sequencing method. The quality files used for Sanger and 454 sequencing were downloaded from the personal genome projects of Venter [24] and Watson [21] in Trace Archive [29], respectively. For the simulation of Solexa method we used quality files from the project SRA000261 in Trace Archive. The lengths of the quality files define the read lengths at a random starting position. The errors were introduced according to quality values given in the quality files. Both Sanger and 454 methods use Phred quality values [30], q and the error probabilities, e are given by e = 10q/-10. The errors are introduced by generating a random number R between 0 and 1. If R <e, then one of the following errors will be introduced: Substitution to one of the three remaining bases, an insertion or a deletion. The probability of an indel is 10% of all introduced errors with the equal ratio of indels. The base frequency in the source genome is used to calculate the frequency of each base, which is in turn used to give the insertion and substitution probability. The Solexa quality values, q s can be converted to Phred quality scores as follows
We simulated the shotgun process for 0.1×, 0.2×, 0.5×, 1×, 2×, 5× and 8× coverages.
The performance is measured by counting the number of sliding windows giving a correct alternatively an incorrect prediction. Our model describes the theoretical limit of detection for given data with given r' and p'. The true copy number ratio of each window is known in the simulated data, i.e. the true r. All windows where true r ≥ r' or r ≤ 1/r' should be classified as CNV in order to achieve 100% sensitivity. Similarly, all windows where true r ≤ r' or r ≥ 1/r' should not be classified as CNV in order to achieve 100% specificity.
CNV detection in human data
The shotgun sequencing data were downloaded from the personal genome projects of Venter and Watson in Trace Archive. The template genome was downloaded from Ensembl [31], human genome assembly, NCBI Build 36. The thresholds p' = 10-5 and log2(r') = 0.6 are used. Given the data these thresholds yield the window size, W = 26, 481 bases for autosomal chromosomes, 72,044 bases for chromosome X and 269,032 bases for chromosome Y.
CNV-seq
All calculations are performed using R [32] and sequences aligned by BLAT [33]. The whole procedure is automated by Perl http://www.perl.org scripts.
References
Sebat J, Lakshmi B, Troge J, Alexander J, Young J, Lundin P, Månér S, Massa H, Walker M, Chi M, Navin N, Lucito R, Healy J, Hicks J, Ye K, Reiner A, Gilliam TC, Trask B, Patterson N, Zetterberg A, Wigler M: Large-scale copy number polymorphism in the human genome. Science 2004, 305(5683):525–528. 10.1126/science.1098918
Iafrate AJ, Feuk L, Rivera MN, Listewnik ML, Donahoe PK, Qi Y, Scherer SW, Lee C: Detection of large-scale variation in the human genome. Nat Genet 2004, 36(9):949–951. 10.1038/ng1416
Redon R, Ishikawa S, Fitch KR, Feuk L, Perry GH, Andrews TD, Fiegler H, Shapero MH, Carson AR, Chen W, Cho EK, Dallaire S, Freeman JL, González JR, Gratacòs M, Huang J, Kalaitzopoulos D, Komura D, MacDonald JR, Marshall CR, Mei R, Montgomery L, Nishimura K, Okamura K, Shen F, Somerville MJ, Tchinda J, Valsesia A, Woodwark C, Yang F, Zhang J, Zerjal T, Zhang J, Armengol L, Conrad DF, Estivill X, Tyler-Smith C, Carter NP, Aburatani H, Lee C, Jones KW, Scherer SW, Hurles ME: Global variation in copy number in the human genome. Nature 2006, 444(7118):444–454. 10.1038/nature05329
Carter NP: Methods and strategies for analyzing copy number variation using DNA microarrays. Nat Genet 2007, 39(7 Suppl):S16-S21. 10.1038/ng2028
Solinas-Toldo S, Lampel S, Stilgenbauer S, Nickolenko J, Benner A, Döhner H, Cremer T, Lichter P: Matrix-based comparative genomic hybridization: biochips to screen for genomic imbalances. Genes Chromosomes Cancer 1997, 20(4):399–407. 10.1002/(SICI)1098-2264(199712)20:4<399::AID-GCC12>3.0.CO;2-I
Pinkel D, Segraves R, Sudar D, Clark S, Poole I, Kowbel D, Collins C, Kuo WL, Chen C, Zhai Y, Dairkee SH, Ljung BM, Gray JW, Albertson DG: High resolution analysis of DNA copy number variation using comparative genomic hybridization to microarrays. Nat Genet 1998, 20(2):207–211. 10.1038/2524
Schuster SC: Next-generation sequencing transforms today's biology. Nat Methods 2008, 5: 16–18. 10.1038/nmeth1156
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA, Berka J, Braverman MS, Chen YJ, Chen Z, Dewell SB, Du L, Fierro JM, Gomes XV, Godwin BC, He W, Helgesen S, Ho CH, Ho CH, Irzyk GP, Jando SC, Alenquer MLI, Jarvie TP, Jirage KB, Kim JB, Knight JR, Lanza JR, Leamon JH, Lefkowitz SM, Lei M, Li J, Lohman KL, Lu H, Makhijani VB, McDade KE, McKenna MP, Myers EW, Nickerson E, Nobile JR, Plant R, Puc BP, Ronan MT, Roth GT, Sarkis GJ, Simons JF, Simpson JW, Srinivasan M, Tartaro KR, Tomasz A, Vogt KA, Volkmer GA, Wang SH, Wang Y, Weiner MP, Yu P, Begley RF, Rothberg JM: Genome sequencing in microfabricated high-density picolitre reactors. Nature 2005, 437(7057):376–380.
Bentley DR: Whole-genome re-sequencing. Curr Opin Genet Dev 2006, 16(6):545–552. 10.1016/j.gde.2006.10.009
Valouev A, Ichikawa J, Tonthat T, Stuart J, Ranade S, Peckham H, Zeng K, Malek JA, Costa G, McKernan K, Sidow A, Fire A, Johnson SM: A high-resolution, nucleosome position map of C. elegans reveals a lack of universal sequence-dictated positioning. Genome Res 2008, 18(7):1051–1063. 10.1101/gr.076463.108
Shendure J: The beginning of the end for microarrays? Nat Methods 2008, 5(7):585–587. 10.1038/nmeth0708-585
Johnson DS, Mortazavi A, Myers RM, Wold B: Genome-wide mapping of in vivo protein-DNA interactions. Science 2007, 316(5830):1497–1502. 10.1126/science.1141319
Cokus SJ, Feng S, Zhang X, Chen Z, Merriman B, Haudenschild CD, Pradhan S, Nelson SF, Pellegrini M, Jacobsen SE: Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature 2008, 452(7184):215–219. 10.1038/nature06745
Hillier LW, Marth GT, Quinlan AR, Dooling D, Fewell G, Barnett D, Fox P, Glasscock JI, Hickenbotham M, Huang W, Magrini VJ, Richt RJ, Sander SN, Stewart DA, Stromberg M, Tsung EF, Wylie T, Schedl T, Wilson RK, Mardis ER: Whole-genome sequencing and variant discovery in C. elegans. Nat Methods 2008, 5(2):183–188. 10.1038/nmeth.1179
Van Tassell CP, Smith TPL, Matukumalli LK, Taylor JF, Schnabel RD, Lawley CT, Haudenschild CD, Moore SS, Warren WC, Sonstegard TS: SNP discovery and allele frequency estimation by deep sequencing of reduced representation libraries. Nat Methods 2008, 5(3):247–252. 10.1038/nmeth.1185
Chen W, Kalscheuer V, Tzschach A, Menzel C, Ullmann R, Schulz MH, Erdogan F, Li N, Kijas Z, Arkesteijn G, Pajares IL, Goetz-Sothmann M, Heinrich U, Rost I, Dufke A, Grasshoff U, Glaeser B, Vingron M, Ropers HH: Mapping translocation breakpoints by next-generation sequencing. Genome Res 2008, 18(7):1143–1149. 10.1101/gr.076166.108
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B: Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 2008, 5(7):621–628. 10.1038/nmeth.1226
Marioni J, Mason C, Mane S, Stephens M, Gilad Y: RNA-seq: An assessment of technical reproducibility and comparison with gene expression arrays. Genome Res 2008.
Nagalakshmi U, Wang Z, Waern K, Shou C, Raha D, Gerstein M, Snyder M: The transcriptional landscape of the yeast genome defined by RNA sequencing. Science 2008, 320(5881):1344–1349. 10.1126/science.1158441
Wilhelm BT, Marguerat S, Watt S, Schubert F, Wood V, Goodhead I, Penkett CJ, Rogers J, Bähler J: Dynamic repertoire of a eukaryotic transcriptome surveyed at single-nucleotide resolution. Nature 2008, 453(7199):1239–1243. 10.1038/nature07002
Wheeler DA, Srinivasan M, Egholm M, Shen Y, Chen L, McGuire A, He W, Chen YJ, Makhijani V, Roth GT, Gomes X, Tartaro K, Niazi F, Turcotte CL, Irzyk GP, Lupski JR, Chinault C, Song Xz, Liu Y, Yuan Y, Nazareth L, Qin X, Muzny DM, Margulies M, Weinstock GM, Gibbs RA, Rothberg JM: The complete genome of an individual by massively parallel DNA sequencing. Nature 2008, 452(7189):872–876. 10.1038/nature06884
Sherwood E: Methods and applications in DNA sequence alignments. PhD thesis. Karolinska Institutet; 2007.
Tammi MT, Arner E, Kindlund E, Andersson B: Correcting errors in shotgun sequences. Nucleic Acids Res 2003, 31(15):4663–4672. 10.1093/nar/gkg653;
Levy S, Sutton G, Ng PC, Feuk L, Halpern AL, Walenz BP, Axelrod N, Huang J, Kirkness EF, Denisov G, Lin Y, MacDonald JR, Pang AWC, Shago M, Stockwell TB, Tsiamouri A, Bafna V, Bansal V, Kravitz SA, Busam DA, Beeson KY, McIntosh TC, Remington KA, Abril JF, Gill J, Borman J, Rogers YH, Frazier ME, Scherer SW, Strausberg RL, Venter JC: The diploid genome sequence of an individual human. PLoS Biol 2007, 5(10):e254. 10.1371/journal.pbio.0050254
Hinkley DV: On the ratio of two correlated normal random variables. Biometrika 1969, 56(3):635–639. 10.1093/biomet/56.3.635
Hayya J, Armstrong D, Gressis N: A note on the ratio of two normally distributed variables. Manage Sci 1975, 21(11):1338–1341. 10.1287/mnsc.21.11.1338
Sanger F, Coulson AR: A rapid method for determining sequences in DNA by primed synthesis with DNA polymerase. J Mol Biol 1975, 94(3):441–448. 10.1016/0022-2836(75)90213-2
Quail MA, Kozarewa I, Smith F, Scally A, Stephens PJ, Durbin R, Swerdlow H, Turner DJ: A large genome center's improvements to the Illumina sequencing system. Nat Methods 2008, 5(12):1005–1010. 10.1038/nmeth.1270
Wheeler DL, Barrett T, Benson DA, Bryant SH, Canese K, Chetvernin V, Church DM, Dicuccio M, Edgar R, Federhen S, Feolo M, Geer LY, Helmberg W, Kapustin Y, Khovayko O, Landsman D, Lipman DJ, Madden TL, Maglott DR, Miller V, Ostell J, Pruitt KD, Schuler GD, Shumway M, Sequeira E, Sherry ST, Sirotkin K, Souvorov A, Starchenko G, Tatusov RL, Tatusova TA, Wagner L, Yaschenko E: Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 2008, (36 Database):D13-D21.
Ewing B, Green P: Base-calling of automated sequencer traces using phred. II. Error probabilities. Genome Res 1998, 8(3):186–194.
Birney E, Andrews TD, Bevan P, Caccamo M, Chen Y, Clarke L, Coates G, Cuff J, Curwen V, Cutts T, Down T, Eyras E, Fernandez-Suarez XM, Gane P, Gibbins B, Gilbert J, Hammond M, Hotz HR, Iyer V, Jekosch K, Kahari A, Kasprzyk A, Keefe D, Keenan S, Lehvaslaiho H, McVicker G, Melsopp C, Meidl P, Mongin E, Pettett R, Potter S, Proctor G, Rae M, Searle S, Slater G, Smedley D, Smith J, Spooner W, Stabenau A, Stalker J, Storey R, Ureta-Vidal A, Woodwark KC, Cameron G, Durbin R, Cox A, Hubbard T, Clamp M: An overview of Ensembl. Genome Res 2004, 14(5):925–928. 10.1101/gr.1860604
R Development Core Team:R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria; 2008. [http://www.R-project.org]
Kent WJ: BLAT-the BLAST-like alignment tool. Genome Res 2002, 12(4):656–664.
Acknowledgements
This work was supported by National University of Singapore FRC grant number R154000265112. CX acknowledges support from the National University of Singapore Research Scholarship. The authors thank Dr. Yap Von Bing for providing valuable advice on statistics.
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors' contributions
CX and MT contributed to all aspects of this research. Both authors read and approved the final manuscript.
Electronic supplementary material
12859_2008_2810_MOESM1_ESM.txt
Additional File 1: CNV regions identified between Venter's and Watson's genomes. The 174 identified CNV regions, including the length, location, log 2 ratio, and p-value for each of the regions. (TXT 10 KB)
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under license to BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Xie, C., Tammi, M.T. CNV-seq, a new method to detect copy number variation using high-throughput sequencing. BMC Bioinformatics 10, 80 (2009). https://doi.org/10.1186/1471-2105-10-80
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2105-10-80