Abstract
Background
Antimicrobial resistance among Pseudomonas aeruginosa (P. aeruginosa), a leading cause of nosocomial infections worldwide, is escalating. This study investigated the prevalence of extended-spectrum β-lactamases (ESBLs) and metallo-β-lactamases (MBLs) among 104 P. aeruginosa clinical isolates from Alexandria Main University Hospital, Alexandria, Egypt.
Methods
Antimicrobial susceptibility testing was performed using agar dilution technique, or broth microdilution method in case of colistin. ESBL and MBL prevalence was assessed phenotypically and genotypically using polymerase chain reaction (PCR). The role of plasmids in mediating resistance to extended-spectrum β-lactams was studied via transformation technique using plasmids isolated from ceftazidime-resistant isolates.
Results
Antimicrobial susceptibility testing revealed alarming resistance rates to carbapenems, cephalosporins, and fluoroquinolones. Using PCR as the gold standard, phenotypic methods underestimated ESBL production while overestimating MBL production. Eighty-five isolates (81.7%) possessed only ESBL encoding genes, among which 69 isolates harbored a single ESBL gene [blaOXA-10 (n = 67) and blaPER (n = 2)]. Four ESBL-genotype combinations were detected: blaPER + blaOXA-10 (n = 8), blaVEB-1 + blaOXA-10 (n = 6), blaPSE + blaOXA-10 (n = 1), and blaPER + blaVEB-1 + blaOXA-10 (n = 1). Three isolates (2.9%) possessed only the MBL encoding gene blaVIM. Three ESBL + MBL- genotype combinations: blaOXA-10 + blaAIM, blaOXA-10 + blaVIM, and blaPER + blaOXA-10 + blaAIM were detected in 2, 1 and 1 isolate(s), respectively. Five plasmid preparations harboring blaVEB-1 and blaOXA-10 were successfully transformed into chemically competent Escherichia coli DH5α with transformation efficiencies ranging between 6.8 × 10 3 and 3.7 × 10 4 CFU/μg DNA plasmid. Selected tested transformants were ceftazidime-resistant and harbored plasmids carrying blaOXA-10.
Conclusions
The study highlights the importance of the expeditious characterization of ESBLs and MBLs using genotypic methods among P. aeruginosa clinical isolates to hinder the development and dissemination of multidrug resistant strains.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Pseudomonas aeruginosa (P. aeruginosa), a clinically significant Gram-negative bacterium belonging to the ESKAPE organisms, has been recognized as a leading cause of serious nosocomial infections worldwide [1]. P. aeruginosa can cause both acute and chronic infections such as ventilator-associated pneumonia, abscesses, skin and soft tissue infections, septic arthritis, bacteremia, meningitis, ulcerative keratitis, and conjunctival erythema. P. aeruginosa infections are more serious in immunocompromised individuals, such as acquired immunodeficiency syndrome patients, cystic fibrosis patients, and patients receiving chemotherapies [2,3,4].
Carbapenems, aminoglycosides, fluoroquinolones, and cephalosporins have been used for treatment of P. aeruginosa infections. However, resistance of P. aeruginosa to various antimicrobial agents is increasing in many countries. Ceftazidime is the most routinely prescribed cephalosporin for combating P. aeruginosa infections owing to its distinctive anti-pseudomonal activity, yet, ceftazidime resistance is escalating [5].
Antimicrobial resistance has rendered P. aeruginosa infections life-threatening and hard to treat [6], because of the limited therapeutic options remaining. This results in worse outcomes and higher morbidity and mortality, particularly in those with severe P. aeruginosa infections [7]. Previous treatment with antibiotics showing high antipseudomonal activity as well as prolonged duration of antibiotic treatment are among the major risk factors contributing to the emergence of resistant P. aeruginosa strains [8].
P. aeruginosa can develop resistance to different antibiotics though intrinsic or acquired resistance mechanisms [9]. Intrinsic resistance occurs due to expression of efflux pump systems, decreased outer membrane permeability, and production of antibiotic-inactivating enzymes [10]. In addition, P. aeruginosa acquires resistance to antimicrobials through mutational changes and acquisition of resistance genes via horizontal gene transfer. Transfer of genes encoding extended-spectrum β-lactamases (ESBLs) and metallo-β-lactamases (MBLs), through conjugation, transduction, and transformation, is the main contributor to the dissemination of antimicrobial resistance among P. aeruginosa strains [9].
Various ESBLs, belonging to molecular classes A and D, have been identified in P. aeruginosa [11]. Class A ESBLs, e.g. TEM, VEB, PER, SHV, GES, and BEL [12], mediate resistance to penicillins as well as narrow spectrum and third generation cephalosporins, but not the carbapenems [11]. The activity of Class A ESBLs may be inhibited by clavulanic acid as well as tazobactam [13]. Class D ESBLs, known as OXA-type ESBLs, resist inactivation by β-lactamase inhibitors except for OXA-18 and OXA-45 [13]. P. aeruginosa also produces different MBLs, including AIM, FIM, GIM, IMP, NDM, SPM, and VIM. They belong to class B β-lactamases, and can break down most β-lactam antibiotics, including carbapenems [5, 14, 15].
An alarming widespread detection of ESBL genes among Gram-negative pathogens in Egypt and surrounding countries; such as Saudi Arabia and Sudan; has been recorded [16]. Also, a recent meta-analysis in Egypt, published in 2022, has revealed an elevated prevalence of MBL-producing P. aeruginosa reaching about 33.7% [17]. Furthermore, the co-expression of ESBLs and MBLs has been reported among clinical isolates, emphasizing the necessity of their rapid detection to establish a suitable policy focusing on restricting empirical antibiotics’ prescription [5, 6].
This study aimed to investigate the prevalence of ESBLs and MBLs, both phenotypically and genotypically, among P. aeruginosa clinical isolates collected from Alexandria Main University Hospital (AMUH), the largest tertiary hospital in Alexandria, Egypt. In addition, the role of plasmids in mediating resistance to extended-spectrum β-lactams was studied via curing and transformation experiments using plasmids isolated from selected ceftazidime-resistant P. aeruginosa clinical isolates, a nationally and globally reported challenging and disseminating pathogen.
Methods
Bacterial isolates
In the present study, 104 P. aeruginosa clinical isolates previously collected from different clinical specimens (pus n = 56, bronchial lavage n = 25, urine n = 13, sputum n = 8, and blood cultures n = 2) from the medical microbiology lab at AMUH, Alexandria, Egypt, between September 2017 and November 2017 and from June 2018 to October 2018 [18] were included. The collected isolates were previously identified [18] through Gram staining, growth on cetrimide agar plates (Himedia, India), triple sugar iron (TSI) test, growth at 42 °C, and the oxidase test (Himedia, India) [19].
Antimicrobial susceptibility testing
The minimum inhibitory concentrations (MICs) of piperacillin-tazobactam, ceftazidime, cefepime, imipenem, meropenem, gentamicin, ciprofloxacin, levofloxacin, and moxifloxacin against the tested isolates were determined using the agar dilution technique, whereas MIC of colistin was determined using the broth microdilution technique [20] and the results were interpreted according to CLSI, 2018 [21]. The antibiotic powders/solutions of pharmaceutical grade were purchased from the Egyptian market as follows: piperacillin-tazobactam (Tazocin® 4.5 g, Pfizer pharmaceutical Co.); ceftazidime (Inzad®1g, Rameda & Marcyrl Co.); cefepime (Cefepime® 1g, Pharco B International); imipenem (Tienam®500 mg, Merck Sharp & Dohme B.V.); meropenem (Meronem® 500 mg, Astrazeneca, UK); ciprofloxacin (Ciprofloxacin® 200 mg/100 ml infusion, AMRYIA. IND); levofloxacin (Tavanic® 500 mg/100 ml infusion, Sanofi Aventis); and moxifloxacin (Vigamox® 5 mg/ml eye drops, Alcon, Geneva). Colistin was obtained as colistin sulfate powder (Amoun Pharmaceutical Co., Egypt) and gentamicin was obtained as gentamicin sulfate powder (Sigma -Alderich, Germany).
The resistance score (R score), designated as the number of antibiotics to which the isolate showed resistance, was calculated for each tested strain. Each resistant call was given a score of 1, while an intermediate resistance towards the tested antibiotic was given a score of 0.5 [22].
Phenotypic detection of β-lactamases
Detection of ESBLs
Double disc synergy test (DDST)
A disc of amoxicillin-clavulanate (20/10 µg) was placed 30 mm apart (center to center) from a disc containing ceftazidime (30 µg) or cefepime (30 µg) [Himedia, India] on a Müller-Hinton agar (LAB M, UK) plate swabbed with the overnight culture of each tested isolate and the plate was incubated at 37 °C for 24 h. ESBL production was considered positive if the inhibition zone around cephalosporin discs (ceftazidime or cefepime) was extended on the side towards the amoxicillin-clavulanate disc [23].
Phenotypic confirmatory disc diffusion test (PCDDT)
Discs containing ceftazidime (30 µg) alone and in combination with clavulanic acid (10 µg) [Himedia, India] were placed 30 mm apart (center to center) on Müller-Hinton agar plate swabbed with the overnight culture of the tested isolate. The plates were incubated at 37 °C for 24 h. An increase of 5 mm in the inhibition zone diameter around the ceftazidime-clavulanate disc relative to the ceftazidime disc was indicative of ESBL production [21, 24].
Detection of MBLs
Imipenem-EDTA combined disc test (IPM-EDTA CDT)
Overnight cultures of the tested isolates were swabbed on Müller-Hinton agar plates. Discs of imipenem (10 μg) and imipenem-EDTA (10/750 μg) [Himedia, India] were placed on the plates 20 mm apart (center to center). The inhibition zone diameters of the imipenem and imipenem-EDTA discs were compared after 24 h of incubation at 37 °C. An increased inhibition zone of ≥ 7 mm with the imipenem-EDTA disc compared to the imipenem disc alone was considered a positive indication for MBL production [25].
Molecular detection of β-lactamases
Detection of genes encoding selected β-lactamases
For the preparation of DNA templates from the tested clinical isolates, four colonies of each isolate were suspended in 200 μl of sterile water for injection in an Eppendorf tube. The suspension was heated at 95 °C for 30 min and then frozen at – 20 °C for 30 min. After thawing, the tube was centrifuged at 14,000 rpm for 10 min, then preserved as aliquots at -20 °C for future use [26, 27]. To isolate DNA from transformed cells, a single colony was suspended in 5 μl sterile water for injection and heated at 95 °C for 10 min to lyse the cells [28]. The presence of selected β-lactamase genes: class A ESBL genes (blaPER, blaPSE and blaVEB-1), class B MBL genes (blaIMP, blaVIM, blaAIM and blaNDM), and class D ESBL genes (blaOXA-10) was tested in the extracted DNA by the polymerase chain reaction (PCR) using previously published primers [29,30,31,32] purchased from Thermo-Scientific (USA). The used primers’ sequences and the applied thermocycling conditions are listed in Additional file 1: Tables S1 and S2, respectively. The amplified PCR products were resolved using 1% agarose gel electrophoresis in Tris Acetate EDTA (TAE) buffer (40 mM Tris, 1 mM EDTA, and 20 mM acetic acid, pH 8.5), and the bands were visualized at 254 nm, using a UV trans-illuminator (Entela, USA). A 100 bps DNA ladder was run alongside the PCR products to detect band size (GeneDirex, Taiwan).
Plasmid isolation and characterization
Plasmid isolation from the ceftazidime-resistant P. aeruginosa clinical isolates harboring both blaOXA-10 and blaVEB-1 (P23, P78, P100, P101, P108, P121, and P123) was performed using the plasmid isolation kit "QIAprep® Plasmid Miniprep Kit" (Qiagen, Germany) according to the manufacturer’s instructions. The concentrations of the obtained plasmids were measured using Genova nano micro-volume spectrophotometer (Jeneway, UK). The plasmid extract was used as a DNA template for PCR detection of the plasmid mediated genes, blaVEB-1 and blaOXA-10, as previously described. The extracted plasmids were resolved on 0.8% agarose gel electrophoresis in the presence of 1 Kbp DNA ladder to describe the plasmid profile (GeneDirex, Taiwan).
Transformation of chemically competent Escherichia coli cells with plasmids harboring bla VEB-1 and bla OXA-10
The extracted plasmid preparations, carrying both blaVEB-1 and blaOXA-10, were transformed into chemically competent E. coli cells using heat shock technique [33]. E. coli DH5α chemically competent cells were prepared as previously described [33]. Briefly, frozen Eppendorf tubes containing 50 μl of chemically competent E. coli DH5α cells were thawed on ice for 15 min. Five microliters of each plasmid preparation were added to a tube containing the thawed competent cells, mixed thoroughly, and incubated on ice for 15 min. The cells were incubated for 90 s in a pre-adjusted heat block at 42 °C, and then transferred back to ice for 5 min. Nine hundred microliters of Luria–Bertani (LB) broth were added to each Eppendorf tube and the cells were incubated at 37 °C with shaking for 1 h at 160 rpm to recover the bacterial cells. From each culture, 50 μl were aseptically plated onto sterile LB agar plate containing 0.5 × MIC of ceftazidime for each selected isolate. After incubation at 37 °C for 24 h, the plates were checked for any transformants. For further confirmation, the susceptibility of selected resultant transformants as well as E. coli DH5α to ceftazidime was detected by the disc diffusion method [21], using ceftazidime (CAZ, 30) disc (Himedia, India), as well as MIC determination by agar dilution method [20]. Selected transformants were subjected to plasmid isolation, then the plasmid extracts were used as templates for PCR amplification of blaVEB-1 and blaOXA-10 genes as previously described.
Transformation efficiency (no. of transformants per μg of plasmid DNA) was calculated as follows [34]:
Curing of resistance plasmids from selected P. aeruginosa clinical isolates
Plasmid curing experiment was performed for the seven selected P. aeruginosa isolates using ethidium bromide (EtBr) [100 μg/ml and 300 μg/ml] and sodium dodecyl sulfate (SDS) [5% and 10%] as curing agents. These agents were sterilized by filtration using cellulose acetate syringe filters (Pore size: 0.45 µm, Diameter: 13 mm) [Filter-bio, China]. The selected isolates were grown in LB broth at 37 ºC for 24 h. One hundred microliters of each culture were added to 2 ml LB broth containing the different concentrations of the curing agents and incubated at 37 ºC for 24 h with constant shaking at 180 rpm. The cultures were ten-fold serially diluted with 0.9% sterile saline and then an aliquot of 100 μl of each dilution was plated onto a sterile nutrient agar plate to obtain separate colonies. The plates were incubated at 37 ºC for 24 h. About 150–200 representative colonies were picked up onto control LB agar plate as well as LB agar plates containing sub-inhibitory concentrations (0.5 × MIC) of the marker antibiotics (ceftazidime, imipenem, gentamicin, and levofloxacin) for each selected isolate. The plates were incubated at 37 ºC for 24 h, and then examined for the growth of the cured colonies [35, 36]. The cells that failed to grow on antibiotic-containing plates while still being able to grow on the antibiotic-free plate were considered cured.
The curing rate was calculated as follows [37]:
The cured colonies were picked from the control plates and checked for their susceptibility towards the tested antimicrobial agents by the disc diffusion method [21] using the following antibiotic discs (Himedia, India): ceftazidime (CAZ, 30), imipenem (IPM, 10), gentamicin (GEN, 10), and levofloxacin (LE, 5).
Sequencing of the bla VEB-1 gene
blaVEB-1 was amplified from the seven selected P. aeruginosa clinical isolates using PCR. The PCR fragments were purified, and then sequenced using the same forward and reverse primers used for the PCR protocol (Additional file 1: Table S1). PCR fragment purification and nucleotide sequencing was conducted at GATC Biotech DNA sequence company (Germany, https://www.genomeweb.com/companies/gatc-biotech) using Sanger sequencing technology. The sequences were aligned against the sequence of blaVEB-1a (GenBank accession no. HM370393.1) using the Basic Local Alignment Search Tool (BLAST) to confirm the sequence identity.
Statistical analysis
Data were analyzed using the Chi-square test analysis using SPSS version 25 for windows (SPSS Inc., Chicago, IL, USA). To study the association between susceptibility to different pairs of antimicrobial agents, data were analyzed using IBM SPSS software package version 20.0. (IBM Corp., Armonk, NY, USA). Spearman’s correlation was applied to construct the correlation matrix. After calculating the p-values, the significance of the obtained results was judged at the 5% level.
Results
Antimicrobial susceptibility testing
A total of 104 P. aeruginosa clinical isolates, collected from AMUH, were included in this study. The antimicrobial susceptibility testing revealed that the collected isolates displayed elevated resistance to imipenem (86.5%), the fluoroquinolones: ciprofloxacin, levofloxacin, and moxifloxacin (85.6, 84.6, and 83.7%, respectively), cefepime and gentamicin (82.7% each), followed by meropenem (76%) and ceftazidime (67.3%). On the other hand, colistin resistance was minimal where 94.2% of the tested isolates were colistin-susceptible. For piperacillin-tazobactam, 27.9% of the isolates were susceptible and the remaining isolates showed intermediate (33.7%) to complete resistance (38.5%) (Table 1). Among the tested isolates, 3.8% (n = 4) recorded the highest R score of 10 while 28.8% of the isolates (n = 30) exhibited R score of 9. On the other hand, 4 isolates (3.8%) were susceptible to all of the tested antibiotics recording R score of 0. The antibiotic resistance profile and the R score of each of the tested isolates are illustrated in Additional file 1: Table S3.
Among β-lactam antibiotics, piperacillin- tazobactam showed the highest MIC range (4 to >1024 µg/ml). The three tested fluoroquinolones: ciprofloxacin, levofloxacin, and moxifloxacin had the same MIC range (< 0.5 to >128 µg/ml). The MIC values of gentamicin and colistin ranged from 1 to >1024 µg/ml, and from <0.5 to 256 µg/ml, respectively (Table 1).
Gentamicin showed the highest MIC50 and MIC90 (1024 and > 1024 µg/ml, respectively). On the contrary, colistin exhibited the lowest MIC50 and MIC90 (< 0.5 and 2 µg/ml, respectively). Regarding β-lactam antibiotics, piperacillin-tazobactam, ceftazidime, cefepime, and meropenem possessed equal MIC50 values (64 µg/ml). The highest MIC90 was recorded for imipenem (> 512 µg/ml), while the lowest was observed in case of piperacillin-tazobactam (256 µg/ml). Amongst the tested fluoroquinolones, ciprofloxacin possessed the lowest MIC50 and MIC90 (16 and 64 µg/ml, respectively). Both levofloxacin and moxifloxacin had the same MIC50 (64 µg/ml) (Table 1).
Based on the susceptibility data of the collected P. aeruginosa clinical isolates, illustrated in Additional file 1: Table S3, a correlation matrix relating the ten tested antimicrobial agents included in this study was established (Fig. 1). The matrix indicated that the highest correlations were observed in the resistance pattern of moxifloxacin and each of ciprofloxacin, meropenem, and levofloxacin (Spearman’s correlation coefficient rs = 0.929, 0.84 and 0.82, respectively) as well as in case of the resistance pattern of gentamicin and cefepime (rs = 0.802). All these relations were highly significant (p-value < 0.001). Six pairs of antibiotics showed highly significant correlations (p-value < 0.001) with rs ranging between 0.78 and 0.725. These correlations were between the resistance patterns of ciprofloxacin and meropenem; imipenem and levofloxacin; imipenem and meropenem; imipenem and moxifloxacin; ciprofloxacin and levofloxacin; and cefepime and levofloxacin. All remaining Spearman’s correlation coefficients were also greater than 0.5 (rs ranging between 0.695 and 0.501) showing significance at 0.001 level except for the correlation between the resistance pattern of ciprofloxacin and piperacillin-tazobactam (rs = 0.484; p-value < 0.001) as well as the correlations between the resistance pattern of colistin and all other tested antibiotics (non-significant correlations with rs ≤ 0.124).
Correlation matrix showing Spearman’s correlation coefficients (rs) for each pair of antibiotics calculated according to the resistance patterns of 104 tested P. aeruginosa clinical isolates. The boldness of the blue color refers to the strength of the relationship between antibiotics, with stronger correlations having bolder colors. Numbers within boxes indicate correlation coefficient (rs) values. Spearman’s correlation coefficients (rs) written in bold indicate statistically significant levels of correlation at p-value ≤ 0.001. PIT piperacillin-tazobactam, CAZ ceftazidime, CPM cefepime, IPM imipenem, MRP meropenem, GEN gentamicin, CIP ciprofloxacin, LE levofloxacin, MO moxifloxacin, COL colistin
The highest percentage of antimicrobial resistance among pus isolates was towards gentamicin, ciprofloxacin, and levofloxacin (85.7% each). The least effective antibiotics against bronchial lavage and urine isolates were imipenem (96 and 92.3% resistance, respectively), followed by the tested fluoroquinolones (88 and 84.6% resistance for each, respectively). Cefepime showed a similar activity (88% resistance) as the fluoroquinolones against bronchial lavage isolates. Both fluoroquinolones, ciprofloxacin, and moxifloxacin, exhibited the highest percentage of antimicrobial resistance (75% for each) among sputum isolates. Both blood isolates were resistant to cefepime, gentamicin as well as the tested carbapenems and fluoroquinolones. On the contrary, colistin was the most effective antibiotic against isolates from all clinical sources (Additional file 1: Table S3).
Phenotypic detection of β-lactamases
None of the tested isolates showed positive results when phenotypically tested for ESBL production using DDST or PCDDT. The production of MBL was phenotypically detected among 87.5% of the isolates using the IPM-EDTA CDT. All imipenem-resistant isolates were found to be MBL producers.
Molecular detection of ESBL- and MBL-encoding genes
The three tested genes belonging to class A ESBLs; blaPER, blaVEB-1 and blaPSE were detected among 11.5, 6.7 and 1% of the tested isolates, respectively. Among class B, only blaVIM and blaAIM MBLs were detected with prevalence rates of 3.8 and 2.9%, respectively. The tested member of class D β-lactamases, blaOXA-10, showed the highest occurrence rate (83.7%) among the tested isolates. Only two isolates: P23 and P93 harbored three genes each: P23 harbored blaPER, blaVEB, and blaOXA-10; and P93 harbored blaPER, blaAIM, and blaOXA-10. On the other hand, 12 isolates possessed none of the tested genes (Additional file 1: Table S3).
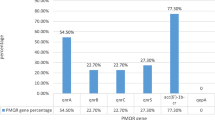
Table 2 illustrates the frequency of ESBL and MBL genes among P. aeruginosa clinical isolates. Eighty-five isolates (81.7%) possessed only ESBL encoding genes, among which 69 isolates harbored a single ESBL gene while 16 isolates harbored a combination of two or more ESBL genes. Regarding the single ESBL genotype, blaOXA-10 and blaPER were detected among 64.4 and 1.9% of the isolates, respectively. The most common ESBL-genotype combinations were blaPER + blaOXA-10 followed by blaVEB-1 + blaOXA-10 (recorded among 7.7 and 5.8% of the isolates, respectively). Each of blaPSE + blaOXA-10 and blaPER + blaVEB-1 + blaOXA-10 ESBL-genotype combinations was observed in one isolate (1%). Three isolates (2.9%) possessed only an MBL encoding gene ( blaVIM). Three ESBL + MBL- genotype combinations: blaOXA-10 + blaAIM, blaOXA-10 + blaVIM, and blaPER + blaOXA-10 + blaAIM were detected in 2, 1 and 1 isolate(s), respectively. blaVEB-1, blaPSE and blaAIM were always detected in the presence of other beta-lactamases.
More than half of the ESBL producers were isolated from pus (52.9%) while the rest were obtained from various sources: bronchial lavage (22.4%), urine (14.1%), sputum (9.4%), and blood (1.2%). Two of the three isolates possessing only MBL encoding genes were pus isolates and the third was obtained from bronchial lavage. All isolates harboring ESBL + MBL genotype combinations were pus isolates. Non-ESBL/non-MBL producers were mainly isolated from pus and bronchial lavage (41.7% each), followed by urine and blood culture (8.3% each) (Additional file 1: Table S3).
All non-ESBL/non-MBL producing isolates were resistant to the tested carbapenems and quinolones and susceptible to colistin. Piperacillin-tazobactam, cefepime, and gentamicin showed similar activity against non-ESBL/non-MBL producers (8.3% susceptibility for each). The least active antibiotic against ESBL producers was imipenem followed by ciprofloxacin (11.8 and 16.5% susceptibility, respectively) while the most active agent was colistin followed by piperacillin-tazobactam (92.9 and 31.8% susceptibility, respectively). MBL producers exhibited 100% resistance rate to all of the tested antibiotics, except for colistin. Also, all the ESBL + MBL producers were susceptible to colistin, however, they were resistant to cefepime and gentamicin. Among the tested isolates, a non-significant association (p-value > 0.05) has been detected between the antimicrobial susceptibility and the ESBL/MBL profile (Table 3 & Additional file 1: Table S3). MBL producers displayed the highest R score mean value followed by the non-ESBL/non-MBL producers (R score mean values = 9 and 8.3, respectively). The same R score mean value of 7 was recorded for both ESBL producers and ESBL + MBL producers (Fig. 2).
Plasmid isolation
Plasmids were isolated from ceftazidime-resistant P. aeruginosa isolates (P23, P78, P100, P101, P108, P121, and P123) that co-harbored both blaVEB-1 and blaOXA-10. After gel electrophoresis, two plasmid bands were detected among each of five (out of seven tested) plasmid preparations belonging to isolates P23, P100, P101, P108, and P121 (Fig. 3). However, plasmid preparations of P78 and P123 could not be visualized by gel electrophoresis. Positive PCR amplification of blaVEB-1 and blaOXA-10 from the seven plasmid preparations confirmed that both genes were plasmid-borne (Fig. 4a, b). The amplified PCR products of blaVEB-1 were sequenced and showed 100% similarity to the publically available sequence of VEB-1a (GenBank accession no. HM370393.1).
PCR products of a blaVEB-1 gene and b blaOXA-10 gene using plasmid preparations as templates. a blaVEB-1 gene (643 bps); M: a 100 bps DNA ladder, lane 1: P23, lane 2: P78, lane 3: P100, lane 4: P101, lane 5: P108, lane 6: P121, lane 7: P123, b blaOXA-10 gene (276 bps); M: a 100 bps DNA ladder, lane 1: P23, lane 2: P78, lane 3: P100, lane 4: P101, lane 5: P108, lane 6: P121, and lane 7: P123
Curing of the plasmids from the isolates carrying bla VEB-1 and bla OXA-10
Curing of the plasmids from the seven selected P. aeruginosa isolates was attempted using SDS and EtBr as curing agents. After selection on LB agar plates containing 0.5 × MIC of marker antimicrobial agents (ceftazidime, imipenem, gentamicin, and levofloxacin), all the colonies selected after treatment showed resistance to the tested antibiotics.
Transformation of bla VEB-1 and bla OXA-10—carrying plasmids into chemically competent E. coli
The transformation of 5 plasmid preparations from P23, P100, P101, P108, and P121, visualized by gel electrophoresis, into chemically competent E. coli DH5α was successful. Plasmid preparation of isolate P101 showed the highest transformation efficiency (3.7 × 104 CFU/μg plasmid DNA), whilst the lowest transformation efficiency was observed in case of plasmid preparation of isolate P108 (6.8 × 103 CFU/μg plasmid DNA). All the selected tested transformants were ceftazidime-resistant showing more than 512-fold increase in MIC values except for transformants obtained from plasmid preparation of isolate P108 that showed more than 64-fold increase compared to the wild E. coli DH5α (Table 4). Furthermore, plasmids were isolated from selected transformants and were used as templates for PCR detection of both blaVEB-1 and blaOXA-10. Only blaOXA-10 was detected among the five plasmid preparations that were successfully transformed into E. coli DH5α (Additional file 2: Fig. S1).
Discussion
Owing to the extensive use of antimicrobial agents and the continuous rise in the number of immunosuppressed patients, P. aeruginosa has become a leading cause of life-threatening Gram-negative hospital-acquired infections [38]. Among the most worrisome features of this pathogen is its remarkable ability to inherently resist various classes of antibiotics and acquire resistance to nearly all effective antimicrobial agents [39]. Consequently, this hampers the available treatment options, increases the health care costs, and elevates the burden of morbidity and mortality [40].
Resistance to each of cefepime, gentamicin, moxifloxacin, levofloxacin, ciprofloxacin, and imipenem was greater than 80% among the tested isolates. Higher resistance rates to cefepime (98.2%) and ceftazidime (91.2%) among P. aeruginosa isolates were formerly reported in Egypt [41]. In contrast, Farhan et al. [42] and Diab et al. [43] reported lower rates of resistance in Egypt to cefepime and ceftazidime, respectively. MIC values comparable to those obtained in the current study for both cephalosporins have been previously recorded in Egypt and Saudi Arabia [44, 45]. The 38.5% resistance rate to piperacillin-tazobactam reported here is lower than that documented by Diab et al. [43]. Yet, the MIC values of piperacillin-tazobactam among the tested isolates are close to those described by Basha et al. [44] and Ochoa et al. [46].
Despite the fact that carbapenems are among the most effective treatment choices against P. aeruginosa [5], imipenem was the least effective antibiotic in this study. This result is in line with that reported by Khorvash et al. [47], in Iran, who found imipenem resistance of 97.9% among the tested P. aeruginosa isolates. In Egypt, 78.3 and 72% resistance to imipenem were previously reported by Abaza et al. [40] and Diab et al. [43], respectively. On the contrary, our finding disagrees with Farhan et al. [42] and Elmaraghy et al. [48], from Egypt, who found that imipenem showed resistance rates of only 8 and 14.9%, respectively. In our study, meropenem also exhibited an elevated resistance rate reaching 76%. Similar to imipenem, high prevalence of meropenem resistance was formerly observed in Egypt [40, 43]. The obtained ranges of MIC values of imipenem and meropenem are consistent, to some extent, with those reported by Basha et al. [44].
Although aminoglycosides have been recognized as a significant treatment option for P. aeruginosa infections, the development of aminoglycoside resistance has been widely reported [49]. The current study showed that the resistance rate to gentamicin was 82.7%. This result is higher than the previously reported rates globally [50,51,52] (ranging between 16.8 and 28.5%) and in Egypt [42, 48, 53] (ranging from 6 to 44.8%). In the current study, gentamicin possessed the highest MIC50 and MIC90 values among all tested antibiotics. A comparable finding was reported by Basha et al. [44].
In this study, elevated resistance rates to the three tested fluoroquinolones: ciprofloxacin, levofloxacin, and moxifloxacin (85.6, 84.6 and 83.7%, respectively) were found. Similarly, in Egypt, Abaza et al. [40] reported 76.6% resistance rate to ciprofloxacin and Basha et al. [44] recorded 67% resistance level to levofloxacin. On the contrary, Abbas et al. [53] reported lower resistance rates to ciprofloxacin and levofloxacin (8 and 6%, respectively). Also, in contrast with our findings, a Chinese study recorded that more than 60% of the P. aeruginosa isolates were susceptible to the same three fluoroquinolones tested here [54]. Such discrepancies in resistance levels might be due to the difference in antibiotic consumption rates and selective pressure in distinct geographical regions.
Colistin was the most effective antimicrobial agent against the present collection of isolates recording a susceptibility rate of 94.2%, and the lowest MIC50 and MIC90 values (< 0.5 and 2 µg/ml, respectively). Similarly, in an Iranian study conducted by Malekzadegan et al. [55], the tested P. aeruginosa isolates showed susceptibility to colistin with MIC50 and MIC90 of 0.5 and 1 µg/mL, respectively.
The pairwise correlation between the susceptibility patterns of the tested clinical isolates to the antimicrobial agents under investigation was studied. Highly significant correlations have been noticed between all of the tested pairs of β-lactam antibiotics (rs ≥ 0.523; p-value < 0.001) as well as the studied pairs of fluoroquinolones (rs ≥ 0.735; p-value < 0.001). Such highly significant, but partial, correlations could be attributed to the potential common resistance mechanisms to these antimicrobial agents belonging to the same class, while highlighting strain variation, differences in antibiotic chemical structures and enzyme specificities [22]. In addition, highly significant correlations (p-value < 0.001), with rs ranging between 0.84 and 0.64, have been detected between the three tested members of fluoroquinolones and both carbapenems. Such fluoroquinolone-carbapenem significant correlations could be explained according to the fact that, in P. aeruginosa, exposure to a fluoroquinolone stimulates carbapenem resistance owing to transcriptional downregulation of the porin OprD as well as efflux pumps upregulation [56]. On the contrary, the correlations between colistin and all the other tested antibiotics were non-significant with rs ≤ 0.124. This could be due to the different underlying mechanisms by which bacteria acquire resistance to colistin relative to other classes of antibiotics.
Despite the high level of resistance to ceftazidime and cefepime, none of the tested isolates showed positive results for ESBL production when phenotypically screened using conventional DDST and PCDDT. Similarly, Mansour et al. [57] demonstrated that none of the tested P. aeruginosa isolates were ESBL producers by the conventional DDST test. On the contrary, a noticeable level of ESBL detection (36%), by phenotypic DDST, was reported by Farhan et al. [42]. Generally, the available methods for the phenotypic detection of ESBLs are unreliable. This issue has been previously discussed by Zafer et al. [58], Jiang et al. [59], and Poulou et al. [60] who documented the uncertainty and unreliability of the current phenotypic methods for detection of ESBLs in P. aeruginosa and that these methods may give false-negative results. The presence of a combination of ESBL and ampC genes may lead to failure in the phenotypic detection of ESBLs by conventional DDST. Also, the false-negative results obtained from DDST despite the phenotypic resistance observed could be explained by the presence of other resistance mechanism, such as efflux and impermeability [24].
The phenotypic detection of class B MBLs (carbapenemases), using the combined imipenem-EDTA disc test, revealed that all the imipenem-resistant isolates were MBL producers. In Egypt, a similar finding was reported by Basha et al. [44] who found that all of the tested carbapenem resistant P. aeruginosa isolates, displaying the highest MIC values to imipenem and meropenem, showed positive results using this phenotypic test.
Molecular detection of selected β-lactamase encoding genes of different Ambler classes was performed using the PCR technique. The isolates were screened for class A β-lactamases; blaPER, blaPSE, and blaVEB-1. blaPER, an ESBL first reported in Turkey in 1991, showed a prevalence of 11.5% among the tested isolates; a finding comparable to that previously recorded by Gaballah et al. [61]. In contrast, Strateva et al. [32] noted the absence of blaPER among Bulgarian P. aeruginosa isolates. blaPSE, was detected only in one isolate. Also, Gaballah et al. [61] found that only 2 out of 30 P. aeruginosa isolates harbored blaPSE. On the other hand, high occurrence rate of blaPSE was previously reported in Egypt [53] and Taiwan [62]. Among our isolates, 6.7% harbored blaVEB-1. Such low prevalence of blaVEB-1 is comparable to that reported in an Egyptian study conducted by Zafer et al. [58]. blaVEB was also detected in Egypt by Gaballah et al. [61], yet at a prevalence rate of 33% among the tested P. aeruginosa isolates. On the other hand, in another Egyptian study, Abbas et al. [53] noticed the absence of blaVEB among the tested P. aeruginosa isolates. Among numerous acquired β-lactamase enzymes, the blaVEB-1 possesses the most considerable clinical significance because it mediates resistance to oxyimino β -lactams [63]. Furthermore, blaVEB-1 is one of the most commonly detected ESBLs among P. aeruginosa from the Middle East including Saudi Arabia, Kuwait, and Iran [64].
Regarding class B MBLs, four genes were investigated among the P. aeruginosa isolates: blaIMP, blaVIM, blaNDM and blaAIM. VIM β-lactamases are the most common type of MBLs in P. aeruginosa [65]. However, in the current study, blaVIM was detected only among 3.8% of the tested isolates. In Egypt, a higher prevalence rate of blaVIM was recorded by Farhan et al. [42]. In contrast, other Egyptian studies reported the absence of blaVIM among P. aeruginosa isolates [40, 53]. blaIMP was not detected at all. A similar finding has been reported in several studies globally [32, 45, 62, 66, 67] and in Egypt [43, 44, 61]. On the contrary, Abaza et al. [40] and Farhan et al. [42] reported 36.7 and 42.8% prevalence of blaIMP among their tested P. aeruginosa isolates, respectively. None of the tested isolates in this study possessed blaNDM. The absence of that gene among P. aeruginosa strains has been previously recorded in Egypt [61] and Saudi Arabia [45]. Yet, Basha et al. [44] and Mukaya et al. [68] reported that blaNDM was the most prevalent MBL (90.9 and 51.9%, respectively) among the tested P. aeruginosa isolates in their studies. To the best of the authors’ knowledge, this is the first study of the prevalence of blaAIM in Egypt. Few reports are available worldwide about the occurrence of this newly identified MBL in P. aeruginosa. In the current investigation, only 3 isolates (2.9%) harbored this gene. Adelaide imipenemase 1 (AIM-1) was first reported, in 2008, in Adelaide, Australia. It has been suggested that blaAIM-1 gene, in P. aeruginosa, originated from Pseudoxanthomonas mexicana, a non- pathogenic environmental bacterium [69]. Although this gene has been associated with a high-level resistance phenotype as well as a wide prevalence among environmental, non-pathogenic bacteria, it is scarcely detected among pathogenic bacterial communities. P. aeruginosa harboring blaAIM-1 gene has been infrequently, though repeatedly, detected in certain region; Adelaide [69]. Nevertheless, in Iran, Neyestanaki et al. [31] reported 1% prevalence of blaAIM among the tested P. aeruginosa isolates. In Iraq, a 37.5% prevalence rate of blaAIM has been recorded among the tested isolates [70]. However, in another Iranian study, the complete absence of blaAIM was recorded [47].
The results of the phenotypic detection of MBLs revealed that 87.5% of the tested isolates were MBL producers. Based on the PCR results, only 7 isolates carried MBL genes. This might suggest that the MBL enzymatic activity observed by phenotypic IMP-EDTA CDT might be due to other MBL genes not investigated in this study. Similarly, Abbas et al. [53] and Gaballah et al. [61] found that some IMP-EDTA CDT-positive isolates did not harbor the tested MBL gene(s). However, it is worthy to note that EDTA has been recognized as an effective permeabilizer against P. aeruginosa [71] and, owing to its metal chelating properties, EDTA can increase the susceptibility of Acinetobacter spp. to various antibiotics including imipenem [72]. This might result in false-positive results of IMP-EDTA CDT. Marra et al. [73] documented that EDTA may cause false-positive detection rate of MBLs reaching 69.6%. According to Chu et al. [74], the IMP-EDTA disc method should be applied with caution for the detection of MBLs where this test is suitable for preliminary screening. However, it should not be utilized as the only indicator for detecting MBLs.
blaOXA-10, a class D ESBL, displayed the highest occurrence rate (83.7%) among the tested isolates. blaOXA-10 is one of the most prevalent ESBL genes in P. aeruginosa in several countries, including Iran, Bulgaria, Korea, and Palestine [31, 32, 75, 76]. In Egypt, Zafer et al. [58] revealed that blaOXA-10 -like genes were the most common ESBLs among the tested P. aeruginosa isolates with a prevalence rate exceeding 40%.
It was noted that the molecular detection of ESBL genes was not consistent with the phenotypic screening where 85.6% of the isolates were shown molecularly to carry ESBL genes, yet none of the isolates were found to be ESBL producers phenotypically. In accordance, Aktaş et al. [77] mentioned that DDST might fail in the detection of isolates producing PER-1- and OXA-derived enzymes. Also, in their study, only 37% of the PER-1-positive strains were DDST-positive. Moreover, Zafer et al. [58] found that, upon phenotypic screening for ESBL production, only 9 isolates showed positive results while 20 and 5 isolates, respectively, were OXA-10-positive and VEB-positive by PCR. Similar differences between the phenotypic and genotypic prevalence of β-lactamases among Acinetobacter baumannii isolates was noted by our group [78]. Such findings confirm the unreliability of the phenotypic methods singly and/or alone for the detection of β-lactamases as they may lead to false-negative or false-positive results, thus affecting the choice of the proper antibiotic therapy and in turn resulting in treatment failure.
In this study, four ESBL-genotype combinations were detected among 16 (15.4%) of the tested isolates. The co-existence of more than one ESBL encoding gene among P. aeruginosa isolates has been previously reported [12, 79]. In addition, in the current investigation, ESBL + MBL- genotype combinations were detected among 4 isolates (3.8%). Similarly, the co-harboring of ESBL and MBL encoding genes among P. aeruginosa isolates have been recorded in Egypt [61], India [5], and Saudi Arabia [80]. The co-production of various classes of β-lactamases by a single clinical isolate may give rise to serious diagnostic challenges and result in therapeutic failure. Accordingly, expeditious recognition of β-lactamase production might help in the implementation of suitable antibiotic therapy as well as infection control strategies [81, 82].
In an Indian study conducted by Chaudhary and Payasi [5], the susceptibility to different antibiotics among P. aeruginosa isolates with various ESBL/MBL profiles was studied. In comparison to their study, lower susceptibility rates to imipenem and meropenem were detected among our ESBL producers while lower percentages of resistance to both carbapenems were observed among our ESBL + MBL producers. Also, our ESBL producers showed lower percentages of susceptibility to piperacillin-tazobactam. Compared to the findings of Chaudhary and Payasi [5], our ESBL producers and ESBL + MBL producers showed a higher resistance rate to cefepime. For ceftazidime, a higher resistance rate was detected among our ESBL producers while a lower resistance rate was noticed among the ESBL + MBL producers. Also, a lower percentage of resistance against piperacillin-tazobactam was detected among our ESBL + MBL producers. In line with our findings, Chaudhary and Payasi [5] reported that all MBL producers were resistant to piperacillin-tazobactam, ceftazidime, cefepime, imipenem, and meropenem. Against these five antibiotics, our non-ESBL/non-MBL producers showed resistance rates ranging between 58.3 and 100%. On the contrary, in their study, the non-ESBL producers showed 100% susceptibility to these antibiotics [5].
Upon comparing the resistance profiles of MBL producers and ESBL + MBL producers in our study, MBL producers showed higher resistance rates against piperacillin-tazobactam, ceftazidime, imipenem, meropenem, and the three tested fluoroquinolones. Similarly, Chaudhary and Payasi [5] reported higher percentages of resistance against piperacillin-tazobactam, ceftazidime, imipenem and meropenem among MBL producers compared to ESBL + MBL producers. Although a higher resistance rate against cefepime among MBL producers was recorded in their study [5], in ours, equal percentages of susceptibility towards cefepime, gentamicin and colistin were detected among both groups. In our study, higher resistance levels against piperacillin-tazobactam, cefepime, imipenem, meropenem, gentamicin, and the three tested fluoroquinolones were detected in case of non-ESBL/non MBL producers when compared to ESBL producers. On the contrary, Chaudhary and Payasi [5] recorded higher susceptibility rates (100%) to piperacillin-tazobactam, cefepime, imipenem and meropenem among non-ESBL producers compared to ESBL producers. In our study, lower percentages of resistance against both ceftazidime and colistin were noticed in case of non-ESBL/non MBL producers compared to ESBL producers. In comparison with ESBL producers, a higher susceptibility rate (100%) to ceftazidime among non-ESBL producers was reported by Chaudhary and Payasi [5].
Ceftazidime, a third-generation cephalosporin, has been commonly used for the treatment of P. aeruginosa infections. However, the rising rate of ceftazidime resistance has led to poor therapeutic outcomes [83]. The widespread of ceftazidime-resistant P. aeruginosa clinical isolates has been reported globally [45, 46, 83, 84] and in Egypt [41, 42, 44]. Mainly, ceftazidime resistance can be mediated by the production of various β-lactamases including ESBL, MBL and infrequently AmpC-β-lactamases [83]. High prevalence rate of blaOXA-10 and blaVEB-1 -like genes among ceftazidime-resistant P. aeruginosa clinical isolates has been previously reported [84, 85], in accordance with the present findings.
In this study, plasmid isolation from the seven ceftazidime-resistant isolates harboring blaVEB-1 and blaOXA-10 was done. After gel electrophoresis, two plasmid bands were detected in 5 out of 7 isolates. The PCR results revealed that all the plasmid preparations harbored both blaVEB-1 and blaOXA-10. This agrees with Maurya et al. [86] and Behbahani et al. [87] who documented that both blaVEB and blaOXA-10 genes, respectively, are plasmid mediated. Furthermore, to gain more insight about the role of P. aeruginosa plasmids in mediating antibiotic resistance, plasmid curing and transformation experiments were carried out to confirm that the antibiotic resistance genes were plasmid mediated. In the current work, EtBr and SDS, at different tested concentrations, were used as curing agents. All seven isolates remained resistant to the tested antimicrobial agents. This could be attributed to the high copy number of these plasmids [36], or the contribution of chromosomal mediated resistance mechanisms among the tested isolates. In contrast, Thomas et al. [36] reported successful attempts of curing with 10% SDS and loss of resistance to ceftazidime, imipenem, and gentamicin among all the tested strains. However, using EtBr as curing agent, ceftazidime resistance was only lost at a concentration of 200 µg/ml among 68.7% of the tested strains.
The plasmid preparations from the seven isolates harboring blaVEB-1 and blaOXA-10 were transformed into chemically competent E. coli DH5α. The transformation of only five plasmid preparations was successful with high transformation efficiencies. The selected tested transformants showed high resistance to ceftazidime and harbored plasmids carrying blaOXA-10. In a previous investigation, Maurya et al. [88] reported the successful transformation of E. coli JM107 cells with a plasmid harboring blaOXA-10. The high level of acquired resistance to ceftazidime in the tested transformants, in this investigation, may be due to the uptake of plasmid encoding blaOXA-10 and/or the co-presence of other ESBLs genes not explored in this study. The failure of the detection of blaVEB-1 in the obtained transformants of E. coli may be due to the inability of the plasmids carrying the gene to replicate into the recipient cells, or due to the large size of the plasmid carrying the gene leading to unsuccessful transfer of that plasmid. The impact of increasing plasmid size on the decline of the transformation efficiency has been previously reported [89, 90].
Conclusions
The current study shows that resistance levels to carbapenems, cephalosporins, and fluoroquinolones among P. aeruginosa clinical isolates are alarming, thus rendering these infections hard to treat. This necessitates the implementation of proper antibiotic usage policies to prevent the injudicious use of antimicrobial agents in hospitals. Moreover, ESBL-genotype combinations as well as ESBL + MBL- genotype combinations have been detected among the tested isolates, thus leading to poor therapeutic outcomes. The expeditious characterization of ESBLs and MBLs is mandatory to hinder their dissemination, impede the spread of multidrug resistant pathogens, and reduce the associated mortality rates among affected patients.
Availability of data and materials
Data generated or analysed during this study are included in this published article and its supplementary information files.
Abbreviations
- ESBLs:
-
Extended-spectrum β-lactamases
- MBLs:
-
Metallo-β-lactamases
- PCR:
-
Polymerase chain reaction
- AMUH:
-
Alexandria Main University Hospital
- TSI:
-
Triple sugar iron
- MIC:
-
Minimum inhibitory concentration
- CLSI:
-
Clinical and Laboratory Standards Institute
- R score:
-
Resistance score
- DDST:
-
Double disc synergy test
- PCDDT:
-
Phenotypic confirmatory disc diffusion test
- IPM-EDTA CDT:
-
Combined imipenem-EDTA disc test
- DNA:
-
Deoxyribonucleic acid
- TAE:
-
Tris Acetate EDTA
- EDTA:
-
Ethylenediamine tetraacetic acid
- UV:
-
Ultraviolet
- LB:
-
Luria–Bertani
- EtBr:
-
Ethidium bromide
- SDS:
-
Sodium dodecyl sulfate
- BLAST:
-
Basic Local Alignment Search Tool
- CFU:
-
Colony forming unit
References
Peterson LR. Bad bugs, no drugs: no ESCAPE revisited. Clin Infect Dis. 2009;49:992–3.
Driscoll JA, Brody SL, Kollef MH. The epidemiology, pathogenesis and treatment of Pseudomonas aeruginosa infections. Drugs. 2007;67:351–68.
Gellatly SL, Hancock RE. Pseudomonas aeruginosa: new insights into pathogenesis and host defenses. Pathog Dis. 2013;67:159–73.
Chatterjee M, Anju C, Biswas L, Kumar VA, Mohan CG, Biswas R. Antibiotic resistance in Pseudomonas aeruginosa and alternative therapeutic options. Int J Med Microbiol. 2016;306:48–58.
Chaudhary M, Payasi A. Rising antimicrobial resistance of Pseudomonas aeruginosa isolated from clinical specimens in India. J Proteomics Bioinform. 2013;6:005–9.
Kotwal A, Biswas D, Kakati B, Singh M. ESBL and MBL in cefepime resistant Pseudomonas aeruginosa: an update from a rural area in Northern India. J Clin Diagn Res. 2016;10:109–11.
Lister PD, Wolter DJ, Hanson ND. Antibacterial-resistant Pseudomonas aeruginosa: clinical impact and complex regulation of chromosomally encoded resistance mechanisms. Clin Microbiol Rev. 2009;22(4):582–610.
Paramythiotou E, Lucet JC, Timsit JF, Vanjak D, Paugam-Burtz C, Trouillet JL, et al. Acquisition of multidrug-resistant Pseudomonas aeruginosa in patients in intensive care units: role of antibiotics with antipseudomonal activity. Clin Infect Dis. 2004;38:670–7.
Breidenstein EB, de la Fuente-Núñez C, Hancock RE. Pseudomonas aeruginosa: all roads lead to resistance. Trends Microbiol. 2011;19:419–26.
Hancock RE, Speert DP. Antibiotic resistance in Pseudomonas aeruginosa: mechanisms and impact on treatment. Drug Resist Updat. 2000;3:247–55.
Paterson DL, Bonomo RA. Extended-spectrum β-lactamases: a clinical update. Clin Microbiol Rev. 2005;18:657–86.
Laudy AE, Rog P, Smolinska-Krol K, Cmiel M, Sloczynska A, Patzer J, et al. Prevalence of ESBL-producing Pseudomonas aeruginosa isolates in Warsaw, Poland, detected by various phenotypic and genotypic methods. PLoS ONE. 2017;12: e0180121.
Bush K, Jacoby GA, Medeiros AA. A functional classification scheme for beta-lactamases and its correlation with molecular structure. Antimicrob Agents Chemother. 1995;39:1211–33.
Hong DJ, Bae IK, Jang IH, Jeong SH, Kang HK, Lee K. Epidemiology and characteristics of metallo-β-lactamase-producing Pseudomonas aeruginosa. Infect Chemother. 2015;47:81–97.
Jabalameli F, Taki E, Emaneini M, Beigverdi R. Prevalence of metallo-β-lactamase-encoding genes among carbapenem-resistant Pseudomonas aeruginosa strains isolated from burn patients in Iran. Rev Soc Bras Med Trop. 2018;51:270–6.
Azab KSM, Abdel-Rahman MA, El-Sheikh HH, Azab E, Gobouri AA, Farag MMS. Distribution of extended-spectrum β-lactamase (ESBL)-encoding genes among multidrug-resistant Gram-negative pathogens collected from three different countries. Antibiotics (Basel). 2021;10:247. https://doi.org/10.3390/antibiotics10030247.
Azzam A, Khaled H, Hesham M. The prevalence of metallo-β-lactamase-producing Pseudomonas aeruginosa in Egypt: a systematic review and meta-analysis. J Adv Pharm Res. 2022;6:38–248.
Edward EA, El Shehawy MR, Abouelfetouh A, Aboulmagd E. Prevalence of different virulence factors and their association with antimicrobial resistance among Pseudomonas aeruginosa clinical isolates from Egypt. BMC Microbiol. 2023;23:161.
De Vos P, Garrity GM, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer K-H, Whitman B, Eds. Bergey’s Manual of Systematic Bacteriology. In: Volume three: the Firmicutes. 2nd ed. Springer-Verlag, Dordrecht Heidelberg London New York; 2012.
Andrews JM. Determination of minimum inhibitory concentrations. J Antimicrob Chemother. 2001;48:5–16.
Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing. Wayne: Clinical and Laboratory Standards Institute; 2018.
Ramadan AA, Abdelaziz NA, Amin MA, Aziz RK. Novel blaCTX-M variants and genotype-phenotype correlations among clinical isolates of extended spectrum beta lactamase-producing Escherichia coli. Sci Rep. 2019;9(1):4224.
Khalil MA, Sonbol FI, Badr A, Ali SS. Comparative study of virulence factors among ESβL-producing and nonproducing Pseudomonas aeruginosa clinical isolates. Turk J Med Sci. 2015;45(1):60–9.
Lin S, Liu M-F, Lin C-F, Shi Z-Y. Phenotypic detection and polymerase chain reaction screening of extended-spectrum β-lactamases produced by Pseudomonas aeruginosa isolates. J Microbiol Immunol Infect. 2012;45(3):200–7.
Kazeminezhad B, Rad AB, Gharib A, Zahedifard S. blaVIM and blaIMP genes detection in isolates of carbapenem resistant P. aeruginosa of hospitalized patients in two hospitals in Iran. Iran J Pathol. 2017;12:392–6.
Gupta R, Malik A, Rizvi M, Ahmed M. Presence of metallo-beta-lactamases (MBL), extended-spectrum beta-lactamase (ESBL) & AmpC positive non-fermenting Gram-negative bacilli among Intensive Care Unit patients with special reference to molecular detection of blaCTX-M & blaAmpC genes. Indian J Med Res. 2016;144:271–5.
Sepehriseresht S, Boroumand MA, Pourgholi L, Sotoudeh AM, Habibi E, Sattarzadeh TM. Detection of vim-and ipm-type metallo-beta-lactamases in Pseudomonas aeruginosa clinical isolates. Arch Iran Med. 2012;15:670–3.
Škulj M, Okršlar V, Jalen Š, Jevševar S, Slanc P, Štrukelj B, et al. Improved determination of plasmid copy number using quantitative real-time PCR for monitoring fermentation processes. Microb Cell Fact. 2008;7:6.
Yakupogullari Y, Poirel L, Bernabeu S, Kizirgil A, Nordmann P. Multidrug-resistant Pseudomonas aeruginosa isolate co-expressing extended-spectrum β-lactamase PER-1 and metallo-β-lactamase VIM-2 from Turkey. J Antimicrob Chemother. 2008;61:221–2.
Fiett J, Baraniak A, Mrówka A, Fleischer M, Drulis-Kawa Z, Naumiuk Ł, et al. Molecular epidemiology of acquired-metallo-β-lactamase-producing bacteria in Poland. Antimicrob Agents Chemother. 2006;50:880–6.
Neyestanaki DK, Mirsalehian A, Rezagholizadeh F, Jabalameli F, Taherikalani M, Emaneini M. Determination of extended spectrum beta-lactamases, metallo-beta-lactamases and AmpC-beta-lactamases among carbapenem resistant Pseudomonas aeruginosa isolated from burn patients. Burns. 2014;40(8):1556–61.
Strateva T, Ouzounova-Raykova V, Markova B, Todorova A, Marteva-Proevska Y, Mitov I. Problematic clinical isolates of Pseudomonas aeruginosa from the university hospitals in Sofia, Bulgaria: current status of antimicrobial resistance and prevailing resistance mechanisms. J Med Microbiol. 2007;56:956–63.
Russell DW, Sambrook J. Molecular cloning: a laboratory manual. 3rd ed. Cold Spring Harbor Laboratory Cold Spring Harbor; 2001.
Rahimzadeh M, Sadeghizadeh M, Najafi F, Arab S, Mobasheri H. Impact of heat shock step on bacterial transformation efficiency. Mol Biol Res Commun. 2016;5:257–61.
Albaayit SFA, Al-Khafaji ASK, Radif HM. Investigation of plasmid-associated fluoroquinolone resistance in nosocomial Pseudomonas aeruginosa isolated from infected burn wounds. J Biol Sci. 2018;18:514–9.
Thomas B, Agu G, Makanjuola S, Davies A. Plasmid profiling and antibiotic resistance of extended spectrum beta lactamases producing Pseudomonas aeruginosa expressing Ampc beta-lactamase enzyme. Am-Eurasian J Sci Res. 2015;10:109–17.
Fanaki NH, Omar HM, Khalil AM, Edward EA. Prevalence of carbapenem resistant Klebsiella and Proteus clinical isolates: a real threat to the Egyptian health care system. Int J Curr Microbiol Appl Sci. 2018;7:2041–57.
Pachori P, Gothalwal R, Gandhi P. Emergence of antibiotic resistance Pseudomonas aeruginosa in intensive care unit; a critical review. Genes Dis. 2019;6(2):109–19.
Sonbol FI, Khalil MA, Mohamed AB, Ali SS. Correlation between antibiotic resistance and virulence of Pseudomonas aeruginosa clinical isolates. Turk J Med Sci. 2015;45:568–77.
Abaza AF, El Shazly SA, Selim HS, Aly GS. Metallo-beta-lactamase producing Pseudomonas aeruginosa in a healthcare setting in Alexandria. Egypt Pol J Microbiol. 2017;66:297–308.
Mahmoud AB, Zahran WA, Hindawi GR, Labib AZ, Galal R. Prevalence of multidrug-resistant Pseudomonas aeruginosa in patients with nosocomial infections at a university hospital in Egypt, with special reference to typing methods. J Virol Microbiol. 2013;13:165–259.
Farhan SM, Ibrahim RA, Mahran KM, Hetta HF, Abd El-Baky RM. Antimicrobial resistance pattern and molecular genetic distribution of metallo-β-lactamases producing Pseudomonas aeruginosa isolated from hospitals in Minia. Egypt Infect Drug Resist. 2019;12:2125–33.
Diab M, Fam N, El-Said M, El-Defrawy EE-DI, Saber M. Occurrence of VIM-2 Metallo- ß-Lactamases in imipenem resistant and susceptible Pseudomonas aeruginosa clinical isolates from Egypt. Afr J Microbiol Res. 2013;7:4465–72.
Basha AM, El-Sherbiny GM, Mabrouk MI. Phenotypic characterization of the Egyptian isolates “extensively drug-resistant Pseudomonas aeruginosa” and detection of their metallo-β-lactamases encoding genes. Bull Natl Res Cent. 2020;44:117–27.
Al-Agamy MH, Shibl AM, Zaki SA, Tawfik AF. Antimicrobial resistance pattern and prevalence of metallo-β-lactamases in Pseudomonas aeruginosa from Saudi Arabia. Afr J Microbiol Res. 2011;5(30):5528–33.
Ochoa SA, Cruz-Córdova A, Rodea GE, Cázares-Domínguez V, Escalona G, Arellano-Galindo J, et al. Phenotypic characterization of multidrug-resistant Pseudomonas aeruginosa strains isolated from pediatric patients associated to biofilm formation. Microbiol Res. 2015;172:68–78.
Khorvash F, Yazdani M, Shabani S, Soudi A. Pseudomonas aeruginosa-producing metallo-β-lactamases (VIM, IMP, SME, and AIM) in the clinical isolates of intensive care units, a university hospital in Isfahan, Iran. Adv Biomed Res. 2017;6:147–52.
Elmaraghy N, Abbadi S, Elhadidi G, Hashem A, Yousef A. Virulence genes in Pseudomonas aeruginosa strains isolated at Suez Canal University Hospitals with respect to the site of infection and antimicrobial resistance. Int J Clin Microbiol. 2019;2:8–19.
Atassi G, Scheetz MH, Nozick S, Rhodes NJ, Murphy-Belcaster M, Murphy KR, et al. Genomics of aminoglycoside resistance in Pseudomonas aeruginosa bloodstream infections at a United States Academic Hospital. medRxiv. 2021:1–19.
Teixeira B, Rodulfo H, Carreno N, Guzman M, Salazar E, Donato MD. Aminoglycoside resistance genes in Pseudomonas aeruginosa isolates from Cumana, Venezuela. Rev Inst Med Trop Sao Paulo. 2016;58:13.
Asghar AH, Ahmed OB. Prevalence of aminoglycoside resistance genes in Pseudomonas aeruginosa isolated from a tertiary care hospital in Makkah, KSA. Clin Pract. 2018;15:541–7.
Georgescu M, Gheorghe I, Curutiu C, Lazar V, Bleotu C, Chifiriuc M-C. Virulence and resistance features of Pseudomonas aeruginosa strains isolated from chronic leg ulcers. BMC Infect Dis. 2016;16(Suppl 1):92.
Abbas HA, El-Ganiny AM, Kamel HA. Phenotypic and genotypic detection of antibiotic resistance of Pseudomonas aeruginosa isolated from urinary tract infections. Afr Health Sci. 2018;18(1):11–21.
Yang X, Xing B, Liang C, Ye Z, Zhang Y. Prevalence and fluoroquinolone resistance of Pseudomonas aeruginosa in a hospital of South China. Int J Clin Exp Med. 2015;8:1386–90.
Malekzadegan Y, Abdi A, Heidari H, Moradi M, Rastegar E, Sedigh E-S. In vitro activities of colistin, imipenem and ceftazidime against drug-resistant Pseudomonas aeruginosa and Acinetobacter baumannii isolates in the south of Iran. BMC Res Notes. 2019;12:301.
Hakki M, Humphries RM, Hemarajata P, Tallman GB, Shields RK, Mettus RT, et al. Fluoroquinolone prophylaxis selects for meropenem-nonsusceptible Pseudomonas aeruginosa in patients with hematologic malignancies and hematopoietic cell transplant recipients. Clin Infect Dis. 2019;68:2045–52.
Mansour W, Dahmen S, Poirel L, Charfi K, Bettaieb D, Boujaafar N, et al. Emergence of SHV-2a extended-spectrum β-lactamases in clinical isolates of Pseudomonas aeruginosa in a university hospital in Tunisia. Microb Drug Resist. 2009;15:295–301.
Zafer MM, Al-Agamy MH, El-Mahallawy HA, Amin MA, Ashour MSE-D. Antimicrobial resistance pattern and their beta-lactamase encoding genes among Pseudomonas aeruginosa strains isolated from cancer patients. Biomed Res Int. 2014;2014:101635–42.
Jiang X, Zhang Z, Li M, Zhou D, Ruan F, Lu Y. Detection of extended-spectrum β-lactamases in clinical isolates of Pseudomonas aeruginosa. Antimicrob Agents Chemother. 2006;50:2990–5.
Poulou A, Grivakou E, Politi L, Dimitroulia E, Tsakris A. Performance of the modified CLSI extended-spectrum β-lactamase (ESBL) confirmatory test for detecting ESBLs in Pseudomonas aeruginosa. Diagn Microbiol Infect Dis. 2017;90:70–1.
Gaballah A, Elbaradei A, Elsheredy A, Kader O. Emergence of blaVEB and blaGES among VIM-producing Pseudomonas aeruginosa clinical isolates in Alexandria, Egypt. Acta Microbiol Immunol Hung. 2019;66:131–42.
Du S, Kuo H, Cheng C, Fei A, Wei H, Chang S. Molecular mechanisms of ceftazidime resistance in Pseudomonas aeruginosa isolates from canine and human infections. Vet Med. 2010;55:172–82.
Davodian E, Sadeghifard N, Ghasemian A, Noorbakhsh S. Molecular detection of blaVEB-1 beta-lactamase encoding gene among extended spectrum B-lactamase-positive wound isolates of Pseudomonas aeruginosa. Arch Pediatr Infect Dis. 2015;3: e26362.
Sid Ahmed MA, Khan FA, Sultan AA, Soderquist B, Ibrahim EB, Jass J, et al. β-lactamase-mediated resistance in MDR-Pseudomonas aeruginosa from Qatar. Antimicrob Resist Infect Control. 2020;9:170.
Zhao WH, Hu ZQ. Acquired metallo-β-lactamases and their genetic association with class 1 integrons and IS CR elements in Gram-negative bacteria. Future Microbiol. 2015;10:873–87.
Jácome PRLdA, Alves LR, Cabral AB, Lopes ACS, Maciel MAV. Phenotypic and molecular characterization of antimicrobial resistance and virulence factors in Pseudomonas aeruginosa clinical isolates from Recife, State of Pernambuco, Brazil. Rev Soc Bras Med Trop. 2012;45:707–12.
Manoharan A, Chatterjee S, Mathai D, Group SS. Detection and characterization of metallo beta lactamases producing Pseudomonas aeruginosa. Indian J Med Microbiol. 2010;28:241–4.
Mukaya KJ, Maina J, Museve B, Nyerere A, Kiiru J. Antimicrobial resistance profile and genetic profiling of Pseudomonas aeruginosa strains obtained from different inpatient wards at Kenyatta National Hospital. IOSR J Pharm Biol Sci. 2018;13:1–9.
Amsalu A, Sapula SA, Whittall JJ, Hart BJ, Bell JM, Turnidge J, et al. Worldwide distribution and environmental origin of the Adelaide imipenemase (AIM-1), a potent carbapenemase in Pseudomonas aeruginosa. Microb Genom. 2021;7:000715. https://doi.org/10.1099/mgen.0.000715.
Hussein Al-abedi KJ, Abd A-M. Molecular detection of metallo-β-lactamase genes in carbapenem-resistant isolates of Pseudomonas aeruginosa recovered from patients in Al-Diwaniyah province. Iraq QJPS. 2019;24:6–11.
Ayres HM, Furr JR, Russell AD. Effect of permeabilizers on antibiotic sensitivity of Pseudomonas aeruginosa. Lett Appl Microbiol. 1999;28:13–6.
Denny B, West P, Panigrahi D. Effects of permeabilizers on antimicrobial susceptibility of Stenotrophomonas maltophilia and Acinetobacter spp. J Microbiol Immunol Infect. 2003;36:72–6.
Marra AR, Pereira CAP, Gales AC, Menezes LC, Cal RGR, de Souza JMA, et al. Bloodstream infections with metallo-β-lactamase-producing Pseudomonas aeruginosa: epidemiology, microbiology, and clinical outcomes. Antimicrob Agents Chemother. 2006;50:388–90.
Chu YW, Cheung T, Ngan J, Kam KM. EDTA susceptibility leading to false detection of metallo-beta-lactamase in Pseudomonas aeruginosa by Etest and an imipenem-EDTA disk method. Int J Antimicrob Agents. 2005;26:340–1.
Lee S, Park Y, Kim M, Lee HK, Han K, Kang CS, et al. Prevalence of Ambler class A and D β-lactamases among clinical isolates of Pseudomonas aeruginosa in Korea. J Antimicrob Chemother. 2005;56:122–7.
Essawi T, Farraj MA, Sabri I. Extended Spectrum B-lactamases and antimicrobial susceptibility among clinical isolates of Pseudomonas aeruginosa in the West Bank, Palestine. J Mar Island Cult. 2013;3:56–60.
Aktaş Z, Poirel L, Midilli K, Nordmann P. PER-1-and OXA-10-like β-lactamases in ceftazidime-resistant Pseudomonas aeruginosa isolates from intensive care unit patients in Istanbul, Turkey. Clin Microbiol Infect. 2005;11:193–8.
Abouelfetouh A, Torky AS, Aboulmagd E. Phenotypic and genotypic characterization of carbapenem-resistant Acinetobacter baumannii isolates from Egypt. Antimicrob Resist Infect Control. 2019;8:185.
Hosu MC, Vasaikar SD, Okuthe GE, Apalata T. Detection of extended spectrum beta-lactamase genes in Pseudomonas aeruginosa isolated from patients in rural Eastern Cape Province, South Africa. Sci Rep. 2021;11:7110.
Ahmed OB, Asghar AH. The coexistence of extended-spectrum β-lactamase and metallo-β-lactamase genes in Gram-negative bacteria. Arch Pharm Pract. 2021;12:22–8.
Oberoi L, Singh N, Sharma P, Aggarwal A. ESBL, MBL and Ampc beta lactamases producing superbugs - havoc in the intensive care units of Punjab India. J Clin Diagn Res. 2013;7:70–3.
Salimi F, Eftekhar F. Coexistence of AmpC and extended-spectrum β-lactamases in metallo-β-lactamase producing Pseudomonas aeruginosa burn isolates in Tehran, Jundishapur. J Microbiol. 2013;6: e7178.
Umadevi S, Joseph NM, Kumari K, Easow JM, Kumar S, Stephen S, et al. Detection of extended spectrum beta lactamases, ampc beta lactamases and metallobetalactamases in clinical isolates of ceftazidime resistant Pseudomonas aeruginosa. Braz J Microbiol. 2011;42:1284–8.
Sultana S, Paul S, Naher A, Akter R, Khan T. Phenotypic and molecular detection of extended-spectrum beta-lactamase among ceftazidime resistant Pseudomonas aeruginosa isolated from wound swab. Bangladesh J Med Microbiol. 2018;12:15–9.
Girlich D, Naas T, Leelaporn A, Poirel L, Fennewald M, Nordmann P. Nosocomial spread of the integron-located veb-1-like cassette encoding an extended-pectrum beta-lactamase in Pseudomonas aeruginosa in Thailand. Clin Infect Dis. 2002;34:603–11.
Maurya AP, Talukdar AD, Chanda DD, Chakravarty A, Bhattacharjee A. Integron-borne transmission of VEB-1 extended-spectrum β-lactamase in Pseudomonas aeruginosa in a tertiary care hospital in India. Antimicrob Agents Chemother. 2014;58:6966–9.
Behbahani MR, Keshavarzi A, Pirbonyeh N, Javanmardi F, Khoob F, Emami A. Plasmid-related β-lactamase genes in Pseudomonas aeruginosa isolates: a molecular study in burn patients. J Med Microbiol. 2019;68:1740–6.
Maurya AP, Dhar D, Basumatary MK, Paul D, Ingti B, Choudhury D, et al. Expansion of highly stable bla OXA-10 β-lactamase family within diverse host range among nosocomial isolates of Gram-negative bacilli within a tertiary referral hospital of Northeast India. BMC Res Notes. 2017;10:145–50.
Chan V, Dreolini LF, Flintoff KA, Lloyd SJ, Mattenley AA. The effect of increasing plasmid size on transformation efficiency in Escherichia coli. J Exp Microbiol Immunol. 2002;2:207–23.
Hanahan D. Studies on transformation of Escherichia coli with plasmids. J Mol Biol. 1983;166:557–80.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Contributions
The basic idea and the study design was from EA. MRE carried out the practical work. EAE, MRE and AA analyzed and interpreted the results and wrote the original draft of the manuscript. AA and EA reviewed and edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethics approval and consent to participate
The study was carried out in accordance with the guidelines and regulations of the Institutional Review Board (IRB) Committee, Faculty of Medicine, Alexandria University (IRB number: 00012098-FWA number: 00018699, serial number: 0306097). Consent to participate is not applicable for this study because this study does not contain any experiments with human participants and animals and the clinical isolates included in this study were obtained from existing clinical collections routinely assembled as part of laboratory practices of the medical microbiology lab at Alexandria Main University Hospital, Alexandria, Egypt. The need of consents was waived as neither the diagnosis nor the treatment was altered. Moreover, the data of the patients were not exposed.
Consent for publication
Not applicable.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Edward, E.A., El Shehawy, M.R., Abouelfetouh, A. et al. Phenotypic and molecular characterization of extended spectrum- and metallo- beta lactamase producing Pseudomonas aeruginosa clinical isolates from Egypt. Infection (2024). https://doi.org/10.1007/s15010-024-02297-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s15010-024-02297-8