Abstract
Key message
This study presents the results of a systematic genetic analysis between wild and cultivated chestnuts in an orchard in southern Spain, highlighting a complex structure and considerable genetic diversity and opening the possibility to generalize this approach to other Mediterranean orchards.
Context
Tree genetic monitoring offers a good opportunity to evaluate populations and preserve their long-term adaptive evolutionary potential. Chestnut is a multipurpose species of high economic importance in the Mediterranean basin and considered an example of integration between natural and man-driven distribution of diversity under changing environmental and historical conditions. Due to its multipurpose characteristics, man influenced its populations (grafting/sexual propagation) and a complex genetic structure is expected.
Aims
We monitored the trees of a chestnut orchard for studying the genetic diversity and relationship in grafts and rootstocks and detecting possible response in its adaptive potential.
Methods
For this, morphological traits and genomic and genic microsatellite markers were used.
Results
Chestnut trees showed considerable genetic structure, with high level of clonality in the varieties and genetic diversity in rootstocks. The similarity analysis revealed a different clustering pattern for varieties, detecting higher variability for genomic microsatellite markers. Rootstocks harboured a high level of diversity, not previously described, and not contained in the genetic information from populations and varieties from the same region.
Conclusion
Results contribute to understanding the human role in the management of chestnut and demonstrate that rootstocks constitute an unexploited reservoir of variation valuable for conservation strategies against stress factors and future and unpredictable environmental changes.
Similar content being viewed by others
1 Introduction
Genetic monitoring offers a good opportunity for evaluating the status of tree population genetic resources over time, preserving their long-term adaptive evolutionary potential and detecting possible critical signals that demand management action (Schwatz et al. 2006; Graudal et al. 2014). This implies a systematic survey of the amount of genetic variation, genetic composition and spatial genetic structure to detect potential changes in these parameters that may result in loss of gene-level variability (Laikre et al. 2008).
Tree genetic resources exist at different levels of domestication, and the landscapes within which they are located are themselves domesticated to a greater or lesser extent (Michon 2005). In this respect, traditional agroecosystems involve the integration of trees with crops and livestock production and exhibit common features such as a high diversity of species, the use of diversified traditional varieties and low inputs associated with traditional farming practises (Garrity 2004). In these systems, on farm conservation is practised to preserve landraces in areas in which they were originally cultivated and with traditional technologies that have been practised by farmers for millennia (Maxted et al. 1997). Nevertheless, this conservation will only be effective if it is possible to assess the genetic diversity conserved in these agroecosystems and how they can evolve (Graudal et al. 2014).
The Mediterranean basin still harbours traditional agroecosystems of particular importance for preserving biodiversity, and sweet chestnut (Castanea sativa Miller) is a good example. This multipurpose species of high economic importance is valued not only for fruit and timber production but also for its contribution to the landscape and environment. It is accepted that domestication events in chestnut started several millennia ago and were characterized by clonal propagation and selection of the best genotypes that resulted in a population structure far from what would be expected in a purely natural situation. Thus, chestnut genetic structure is complex and depends on the type of management: orchards (dedicated to fruit production), coppices (dedicated to timber production) and naturalized populations (Grossmann and Romane 2004). In particular, orchards are constituted by trees of advanced and heterogeneous age, and chestnut cultivation involves grafting of traditional cultivars onto rootstocks. Thus, the trees from a given variety are clones, whereas rootstocks are the result of the germination of nuts selected by growers for its superior traits from different chestnut varieties or from wild populations (Martín et al. 2009; Marinoni et al. 2013). Furthermore, trees can show one or more varieties in the aerial part and branches without grafts (Martín et al. 2007).
These traditional farming systems are common throughout the Mediterranean region and constitute important elements of production systems and the farmers’ livelihood strategies. The majority of this germplasm is rich in excellent cultivars resulting from the selection for specific nut traits by farmers. Thus, the landraces are highly adapted to the local environment and are likely to contain locally adapted alleles of gene complexes (Martín et al. 2016). In this respect, molecular studies indicate the existence of several local domestication events in the species supported by the association between chestnut cultivars and their origin areas and also by the high diversity of different clonally propagated cultivars found in all countries where chestnut is traditionally grown (Gobbin et al. 2007; Martín et al. 2009, 2010; Pereira-Lorenzo et al. 2010).
Nevertheless, in the last decades, there has been a strong decline in chestnut cultivation closely associated with ink disease caused by Phytophthora cinnamomi Rands. This oomycete infects the root system, causing the wilting and death of chestnut trees (Crandall et al. 1945). Its impact on the production has dramatic consequences in local economies that discourage farmers to continue growing chestnut. In some parts in Europe, the impact of P. cinnamomi has been mitigated by the use of rootstocks that come from the hybridization between C. sativa and two Asian tolerant species (Castanea crenata and Castanea mollissima) (Fernández-López et al. 2001). However, these hybrids displayed some problems as graft incompatibility reactions between rootstock and cultivar, different agronomic traits not accepted by growers and/or consumers and poor adaptation to climatic conditions. In particular, in southern Europe, the use of these hybrids has proved to be unsuitable due to difficulties of adaptation of this resistant material to southern latitudes. For this reason, local rootstocks from C. sativa, developed by growers in different edaphoclimatic regions from central and southern Spain, are usually the ones which are best adapted to the warmer and drier local conditions and to the traditional varieties (Pereira-Lorenzo et al. 2010; Dinis et al. 2011).
Genetic variation in orchards and naturalized populations has been evaluated separately (Martín et al. 2012; Mattioni et al. 2013); thus, there are no genetic data on their possible relationship. Furthermore, the role of rootstocks in this genetic diversity has not been addressed in any study. It has been speculated that the genetic structure contained in rootstocks could be similar to that contained in the varieties or in the populations although there are no empirical data that support this hypothesis.
Molecular markers provide valuable information about the genetic status and biological processes of populations, making genetic monitoring increasingly feasible and cost-effective (Allendorf et al. 2010). Microsatellite markers (simple sequence repeats (SSRs)) have become the most used markers for studying forest genetics, because they are highly polymorphic, codominant and widespread across the genome (Glaubitz and Moran 2000). However, there has been renewed interest in complementing the analysis of neutral markers with assessment of loci that may be directly involved in responses to processes such as environmental changes (Hoffman and Willi 2008). The increased availability of DNA sequences has permitted the development of expressed sequence tag (EST)-based SSR markers from EST sequences expressed in different physiologic conditions of plants. Their main advantage compared with genomic SSRs is that they are present in expressed regions of the genome, thus potentially having known functions (Varshney et al. 2005).
The aim of this study was the genetic monitoring of the productive system of traditional chestnut orchards. For this purpose, an orchard was selected in which previous observations suggested the existence of genetic diversity and the use of grafting. It is important to note that the situation of the orchard described in the study is an archetype of those chestnuts dedicated to fruit production in the Mediterranean basin. The specific objectives were to (1) study the current status of the genetic diversity harboured in the orchard, in the aerial part and rootstocks; (2) confirm the existence of clonal varieties; (3) test the potential of EST-SSR markers to detect possible response in the adaptive potential of chestnut varieties and conduct genetic monitoring; (4) study the genetic relationship between clonal varieties vs. rootstocks; and (5) compare the genetic diversity obtained with that for the rest of Andalusia.
2 Material and methods
2.1 Plant material
The study was conducted in the farm field ‘La Chaparra’, located in the Natural Park Sierra Norte de Sevilla (Andalusia, southern Spain). This orchard covers an area of 1.19 ha and has a density of 35 trees/ha (Fig. 1). For the analysis, 25 large and vigorous trees were catalogued and data of girth of rootstocks and the type of male catkins were recorded (Table 1).
Chestnut cultivation for fruit production is based on a set of traditional varieties grafted onto seedling rootstocks. Thus, trees can present two or more genotypes: (i) the rootstock part that comes from sexual reproduction and (ii) the productive part that comes from vegetative multiplication. Thus, the set of rootstocks may be comparable with a natural population (considering that this traditional orchard do not use hybrids as rootstocks) and the varieties with clones of identical genotypes.
To determine whether trees were grafted, and considering the information above, different samples of leaves were collected for DNA extraction and posterior molecular analysis. Thus, for each tree, one sample was taken from the aerial part and another from the rootstock. Furthermore, in those cases in which trees were suspected to be grafted with more than one variety, two different leaves were collected from the aerial part (named with A and B, Table 1).
2.2 Morphological and molecular characterization
According to flowering, four different types of male catkins are described by UPOV (1989): longistaminate, mesostaminate, braquistaminate and astaminate. These inflorescences produce different amount of pollen, from longistaminate type that generate large amounts of pollen to astaminate type where pollen is absent.
The type of male catkin served as first indicator to detect the possible existence of more than one variety in the same tree. In the study, two different situations were found: (i) trees that showed only one type of male catkin and (ii) trees with two types of male catkins. In this case, two different samples of leaves were taken that were distinguished with A/B (Table 1).
For molecular analysis, DNA was extracted from 20 mg of lyophilized leaves according to DNeasy Plant Mini Kit protocol (Qiagen, CA). A set of eight neutral microsatellites (CsCAT1, CsCAT2, CsCAT3, CsCAT6, CsCAT14, CsCAT16, EMCs25 and EMCs38) developed in C. sativa (Marinoni et al. 2003; Buck et al. 2003) was tested. Furthermore, nine functional microsatellite primer pairs (FIR030, GOT014, PIE227, PIE228, PIE233, PIE260, POR009, POR026 and WAG004) were used. These primers were selected from EST expressed in bud tissue of Quercus robur and Quercus petraea (EVOLTREE database http://www.evoltree.soton.ac.uk/portal). The EST-SSRs were previously mapped on a F1 intraspecific cross (C. sativa × C. sativa) and interspecific cross (Q. robur × Q. petraea), and each locus belongs to a different linkage group (Durand et al. 2010).
PCR reaction mixture consisted of 12.5 μL total volume containing 20 ng of genomic DNA following the Qiagen Type-it protocol. Cycling parameters were 15 min at 95 °C; 30 cycles of 30 s at 94 °C, 90 s at 60 °C and 1 min at 72 °C; and a final step of 30 min at 72 °C. Amplification products (0.1–1 μL) were added to 20 μL formamide and 0.3 μL Genescan-500 ROX and denatured at 95 °C for 5 min and run on an ABI PRISM 3100 DNA sequencer. Allele scoring was performed using the GeneScan 3.5 and Genotyper 3.7 softwares (Applied Biosystems).
2.3 Statistical analysis
To address genetic monitoring, we selected the operational indicator ‘trends in population condition’ proposed by Graudal et al. (2014) and the verifiable indicators related to the diversity in adaptive traits and population genetic structure. Nevertheless, it must be noted that we conducted an approximation to genetic monitoring considering that chestnut orchards are not natural populations. In this respect, genetic diversity parameters such as the number of alleles per locus (Na), the observed heterozygosity (Ho), the expected heterozygosity (He) and the number of private alleles (specific alleles detected in a concrete variety) were estimated using GenAlEx 6 (Peakall and Smouse 2005). The inbreeding coefficient F IS (Weir and Cockerham 1984) was calculated using Arlequin 3.1 software (Schneider et al. 2000) and its deviation from zero tested by 10,000 allele permutations. Differentiation between varieties and rootstocks was calculated by F ST according to Weir and Cockerham (1984) and its analogue, R ST, according to Slatkin (1995). Furthermore, analysis of molecular variance (amova) was conducted to quantify the proportion of genetic variation due to differences among rootstocks vs. varieties using Arlequin 3.1 software (Schneider et al. 2000). Significance was tested using the nonparametric approach described in Excoffier et al. (1992) with 1000 permutations.
ntsys 2.1 software (Exeter Software, Setauker, NY, USA) was used to (i) identify different varietal genotypes for the whole set of SSRs and EST-SSRs and to (ii) detect possible relationships between varieties vs. rootstocks. The band similarity coefficient of Lynch (1990) was calculated for paired comparison of all samples. This was tested by cophenetic matrix correlation during the reconstruction of a cophenetic matrix based on tree matrix (Rohlf and Fisher 1986).
The possible introgression between rootstocks and varieties was assessed using a Bayesian approach with structure v.2.3.4 software (Pritchard et al. 2000). This attempts to reveal the genetic structure by placing individuals in K number of clusters. Structure was run using the admixture model on the whole dataset and the correlated allele frequencies (Falush et al. 2007; Hubisz et al. 2009). Based on the initial results, six independent runs (from 1 to 6) were performed for each K value, with a burn-in period of 10,000 steps followed by 105 MCMC replicates. To identify the number of clusters (K) that best explained the data, the rate of change on L(K) (ΔK) between successive K values was calculated according to Evanno et al. (2005) using structure harvester (Earl and vonHoldt 2012). The six runs for each simulation were averaged using clumpp software (Jakobsson and Rosenberg 2007) and represented graphically with distruct (Rosenberg 2004).
Finally, varietal information obtained in this study was compared with the Andalusian catalogue of traditional chestnut varieties. In this region, the main orchards dedicated to nut production appear in Huelva and Malaga provinces and represent approximately the 70% of the species surface in the region. For the analysis, 12 genotypes previously identified in Huelva and 19 in Malaga (Martín et al. 2009) were selected and structure v.2.3.4 software (Pritchard et al. 2000) was used. In this case, the software was run with the option of including prior information on the spatial location of varieties and using the admixture model on the whole dataset and the correlated allele frequencies (Falush et al. 2007; Hubisz et al. 2009).
3 Results
3.1 Graft detection
A total of 22 individuals displayed only one type of male catkin in the aerial part, and within them, molecular markers indicated that 20 were grafted (Table 1). There was a strong relationship between grafted chestnut and the type of male catkin, with more than 80% of them showing astaminate catkins (i.e. those that do not produce pollen). Furthermore, the two individuals that results showed were not grafted (SE-01-1.16var and SE-01-1.17var) displayed mesostaminate and longistaminate catkins (i.e. those that produce great amounts of pollen).
Three individuals with two different types of male catkin were detected (SE-01-1.3var, SE-01-1.4var and SE-01-1.5var). Results showed that all three trees were grafted, and samples A and B were different, and one of them had the same microsatellite profile as the basal part. This result can be explained by the fact that one aerial sample corresponded to the graft, while the other sample was from a branch of the rootstock.
3.2 SSR and EST-SSR-based varietal identification
After cataloguing grafted trees, we evaluated the varieties. The identification and genetic relationships among the varieties were depicted by two neighbour-joining dendrograms according to the two types of markers (Fig. 2).
The similarity analysis conducted using genomic SSRs detected nine genotypes among the 23 individuals analysed (Fig. 2a). Moreover, the level of similarity between groups of varieties was 0.18, and the cophenetic value was high and significant (r = 0.953; P < 0.001), indicating a good fit for the results obtained (Fig. 2a). The dendrogram showed two dominant genotypes shared by nine and seven individuals, respectively, whereas the remaining seven trees showed their own different genotype (Fig. 2a). Furthermore, there were two individuals (SE-01-1.23var and SE-01-1.29var) with a high degree of genetic similarity to the genotype shared by seven other trees.
A different grouping pattern was found with functional markers (Fig. 2b). In this case, EST-SSR loci displayed a lower level of discrimination, detecting only four genotypes in contrast to the nine found with SSRs, at a similarity level of 0.48 (r = 0.969; P < 0.001). In this case, the dendrogram also showed a clear separation between two principal genotypes with a high level of similarity (0.75) and with two individuals (SE-01-1.1var and SE-01-1.22var) of different genotypes, clustered further from the rest. It should be noted that the dendrogram discriminated individuals according to the type of male catkin, showing two similar groups characterized by the presence of astaminate catkins and two differentiated genotypes corresponding to mesostaminate and longistaminate catkins (Table 1; Fig. 2b).
The presence of exclusive alleles was analysed considering the four genotypes found using EST-SSRs. Thus, one private allele in locus FIR030 was detected for the accession SE-01-1.1var corresponding to longistaminate and four private alleles (PIE227, PIE228, POR09 and WAG004) for the accession SE-01-1.22var corresponding to mesostaminate catkins. Moreover, for the two dominant genotypes with astaminate catkins, four private alleles (PIE227, PIE228, POR09 and WAG004) were found in the first group (represented by nine trees) and three alleles (PIE233, GOT014 and WAG004) in the second group.
3.3 Genetic diversity in SSRs and EST-SSRs
Both genomic SSR and EST-SSR markers were polymorphic in the material evaluated with 53 and 36 alleles detected, respectively (Table 2). Allelic variation was in the range of 4–11 alleles in the genomic SSRs and 3–5 in EST-SSRs, with means of 6.6 and 4.0 alleles, respectively (Table 2). The level of gene diversity was higher for genomic SSRs than EST-SSRs (0.61 vs. 0.52), although this difference was not significant (Table 2). The inbreeding coefficient (F IS), which is the measure of heterozygote deficit, showed significant deviation from zero in five loci, two in genomic SSR (CsCAT2 and EMCs25) and three in EST-SSR markers (FIR030, GOT014 and PIE260) (Table 2).
The polymorphism obtained in genomic SSRs within the rootstock population was higher than that obtained in varieties, with significant differences in the number of alleles and expected diversity (49 and 0.72 vs. 39 and 0.50, respectively). The same results were obtained for EST-SSRs, where rootstocks displayed higher levels of genetic diversity, although these differences were less pronounced (33 and 0.59 vs. 27 and 0.45). The values of F IS were negative in rootstocks both for SSRs and EST-SSRs, and there were no cases in which this value deviated significantly from zero.
The coefficient of differentiation R ST showed higher values than F ST coefficient for both types of markers, although this difference was higher in the case of SSRs (0.25 vs. 0.14 for SSRs and 0.17 vs. 0.11 for EST-SSRs) (Table 2).
3.4 Genetic structure between rootstocks and varieties
To study the possible genetic relationship and structure between rootstocks and varieties, a similarity analysis was performed with a new cluster analysis using SSR markers (Fig. 3). The dendrogram distinguished two different groups at a similarity level of 0.30 (r = 0.953; P < 0.001), generally corresponding to rootstocks and varieties. Thus, one group included all rootstocks except accessions SE-01-1.8 and SE-01-1.16 and the other varieties except SE-01-1.1var, SE-01-1.22var and SE-01-1.27var (Fig. 3). Likewise, amova analysis detected significant genetic differentiation between rootstocks vs. varieties in both types of markers, being this difference more pronounced in the case of neutral markers (22.01 vs. 17.98%, respectively) (Table 3). These results were corroborated by analysis of differences in the presence of private alleles: Rootstocks showed 23 exclusive alleles not found in varieties (14 for SSRs and 9 for EST-SSRs), and varieties displayed 7 alleles not found in rootstocks (4 for SSRs and 3 for EST-SSRs) (Table 2). The results obtained with structure were congruent with the clustering pattern obtained with the dendrogram, showing the most probable division at K = 2 that corresponded to rootstock and variety groups (Online Resource 1). Furthermore, possible introgression between rootstocks and varieties was detected in only five individuals (four rootstocks and one variety). In the case of rootstocks, the admixture value was between 0.4232 ≤ q ≤ 0.6202, and for the variety, this value was q = 0.3743.
The genetic information found in the evaluated material, both in rootstocks and varieties, was compared with that reported in other studies in chestnut natural populations and traditional varieties from the same region (Andalusia), using Bayesian analysis with structure. In the case of varieties, we assumed from previous results that varieties from Malaga and Huelva provinces have their own genetic integrity and were adapted to local conditions of the areas in which they were grown (Martín et al. 2007, 2009). In this respect, each province was considered a population containing the different genotypes identified therein. Thus, the Huelva population was consisted of 12 genotypes corresponding to 12 clonal varieties, Malaga of 19 and Sevilla of 9 (Online Resource 2). The results indicated that the most probable division with the strongest support in terms of log-likelihood values was for K = 2. This level of structure separated the samples into two groups: The first group (green cluster) comprised mainly varieties from Sevilla and Malaga and the second group (red cluster) varieties from Huelva (Fig. 4a).
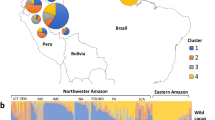
Bayesian clustering of the chestnut accessions obtained with structure software (Pritchard et al. 2000). a Genetic relationship among traditional varieties for K = 2. b Genetic relationship among rootstocks and chestnut populations for K = 3
In the comparison of populations and rootstocks, the most probable division was at K = 3, whereas for K > 3, the results were not consistent and membership analyses were unstable among runs. For K = 3, three clusters were identified with limited admixture among clusters (Fig. 4b). The first group comprised populations from southeastern Spain (SP01, SP05 and SP09, blue cluster); the second was made by populations from southwestern Spain (SP04, SP11 and SP14, yellow cluster), and the third group was only formed by samples from rootstocks (cluster red). Furthermore, only population SP12 displayed a high degree of admixture of clusters blue and yellow (Fig. 4b).
4 Discussion
This study presents the results of a systematic genetic structure analysis of chestnut traditional varieties, rootstocks and populations in an orchard in southern Spain. This material was surveyed using neutral and functional microsatellite markers with the main goal of enhancing our knowledge of cultivar identity, genetic variability and phylogenetic relationships.
The results indicate that despite being a region where chestnut is mostly used for timber production, the traditional fruit production system is based in clonal varieties, considering that only 2 of 25 individuals were not grafted. In this respect, nine varieties of clonal nature were identified using neutral microsatellite markers, with two of them considered dominant. It is possible that these results can be extrapolated to information provided by farmers in the area, concerning the existence of two varietal types, called ‘Fina’ and ‘Boronda’ (Gallardo 2002), but confirming this hypothesis will require additional studies. Furthermore, there were two individuals with a high degree of genetic similarity to these dominant genotypes. This similarity may be the result of the germination of a nut from an original variety that, due to good fruit traits, was selected as a new variety. Such an observation was previously described in chestnut from southern Spain, a region that favours short juvenile period of the species and with high ability of farmers to practice grafting (Martín et al. 2009). The resemblance between these groups of varieties could indicate that they were brought from other areas, possibly as cutting material, and sexual reproduction processes have led to the current situation. Results obtained with structure could support this hypothesis, since varieties from Sevilla and Malaga were grouped together. Processes of this nature were also proposed for the evolution of wild olive (Muñoz et al. 2015). In addition, the fact that one of the rootstocks (SE-01-1.08) was included in the group of varieties suggests that this process could operate in both directions. Conversely, those varieties clearly different from the dominant groups could have emerged from rootstock population, and the best genotypes were used for grafting.
The EST-SSR loci results showed considerably lower polymorphism and diversity than that detected using genomic SSRs in chestnut, in agreement with reports for other plant species (Scott et al. 2000; Woodhead et al. 2005). Nevertheless, this low variation in EST-SSRs contrasts with their efficient discrimination capability for male catkin identification. Thus, the EST-SSR dendrogram separated the individuals according to the presence of astaminate, mesostaminate and longistaminate catkins. Such a result was unexpected because it was speculated that strong local selection due to human management would mask the possible signal of functional markers (Martín et al. 2016). Nevertheless, some of these markers demonstrated significant differences in the level of differentiation among chestnut populations from contrasting climatic environments in relation to bud burst, with northern populations flushing and forming winter buds later, and growing more than Mediterranean populations (Martín et al. 2010).
Our results highlighted that the chestnut production system is characterized by a complex structure and considerable genetic diversity, confirming results of previous studies on chestnut populations (Martín et al. 2012; Mattioni et al. 2013) and traditional varieties (Gobbin et al. 2007; Martín et al. 2009; Pereira-Lorenzo et al. 2010). Likewise, results of the present study showed that rootstocks harboured a high level of diversity, not previously described, and not contained in the genetic information from populations and varieties. In this respect, the most remarkable finding was the genetic integrity found between rootstocks and traditional varieties. The results indicate that rootstocks come from sexual reproduction and constitute a genetic population that harbour a high genetic diversity. Furthermore, only three varieties (SE-01-1.1var, SE-01-1.22var and SE-01-1.27var) could have emerged from this population. Considering the advanced age of the trees, it could be assumed that the rootstock population is native, or at least adapted to the area.
Furthermore, rootstocks and populations displayed different genetic composition. Thus, structure analysis detected a clear structure among rootstocks and populations, showing absence of admixture between them (see Q admixture value in Online Resource 1). Likewise, different alleles were detected in these materials, including 23 private alleles in the rootstocks and 16 in the populations (data not shown). Similarly, a clear structure was also detected in the varietal germplasm, where Bayesian analysis evidenced that traditional chestnut varieties from Sevilla were clustered with those from Malaga, whereas varieties from Huelva had a definite identity. Nevertheless, despite being grouped with varieties from Malaga, seven different alleles not catalogued previously in southern material were found in varieties from Sevilla (Martín et al. 2009).
It is assumed that the wise use of genetic resources in trees maintained on farm is one of the real options available to support sustainable production systems (Graudal et al. 2014; Alfaro et al. 2014). Nevertheless, there is a substantial lack of field studies on the extent of conservation activities and on the reasons for maintaining landraces. It is widely accepted that without monitoring, it is not possible to verify effectiveness of such conservation (Schwatz et al. 2006). In this respect, this study provides valuable baseline data concerning the complex genetic relationships between wild and cultivated chestnuts that should contribute to understanding the human role in the management of the species, expand capacities to manage its genetic resources and open the possibility to generalize this approach to the rest of orchards in the Mediterranean basin. Furthermore, temporal changes in the reported results on the amount of diversity and its distribution in space could be assessed as relevant state indicator in these chestnut production systems. According to Aravanopoulos (2011), a frequency of one evaluation per decade should be adequate, given the current levels of anthropogenic exploitation and environmental changes. Nevertheless, taking into account that chestnut orchards are highly human-managed, we consider that these measurements should be conducted in a shortened period of time. Furthermore, the advance age of the farmers that are the real maintainers of these traditional systems supports this fact (Martín et al. 2007).
On farm conservation of traditional chestnut varieties is dynamic and sustains the ongoing evolution of the crop, simulated by human management of varietal mixtures, graft exchange and mutations. Nevertheless, the implication of the public administration to encourage the producer associations and small farmers to maintain this system is necessary, complementing it with strategies of ex situ maintenance of this germplasm. At regional scale, the Natural Park of Sierra de Aracena y Picos de Aroche (Huelva province) is conducting an initiative to establish an arboretum with the most important genotypes from this region. The traditional varieties identified in this study will be included in it. Furthermore, these genotypes will be incorporated in a core collection at national level within the research project AGL2013-48017-C2-1-R from the Spanish Ministry of Economy and Competitiveness. Finally, our results demonstrate that rootstocks could constitute an unexploited reservoir of variation that would be valuable to counteract stress factors and future and unpredictable environmental changes in the species.
References
Alfaro R, Fady B, Vendramin GG, Dawson I, Fleming RA, Sáenz-Romero C, Linding-Cisneros RA, Murdok T, Vicenti B, Navarro CM, Skroppa T, Baldenelli G, El-Kassaby Y, Loo J (2014) The role of forest genetic resources in responding to biotic and abiotic factors in the context of anthropogenic climate change. Forest Ecol Manag 333:76–87
Allendorf FW, Hohenlohe PA, Luikart G (2010) Genomics and the future of conservation genetics. Nat Rev Genet 11:697–709
Aravanopoulos FA (2011) Genetic monitoring in natural perennial plant populations. Botany 89:75–81
Buck EJ, Russell K, Hadonou M, James CJ, Blakesley D (2003) Isolation and characterization of polymorphic microsatellites in European chestnut (Castanea sativa Mill.). Mol Ecol Notes 3:239–241
Crandall BS, Gravatt GF, Ryan MM (1945) Root disease of Catanea species and some coniferous and broadleaf nursery stocks, caused by Phytophthora cinnamomi. Phytopathology 35:162e80
Dinis LT, Peixoto F, Pinto T, Costa R, Bennett RN, Gomes-Laranjo J (2011) Study of morphological and phenological diversity in chestnut trees (Judia’ variety) as a function of temperature sum. Environ Exp Bot 70:110–120
Durand J, Bodénès C, Chancerel E, Frigerio JM, Vendramin G, Sebastiani F et al (2010) A fast and cost-effective approach to develop and map EST-SSR markers: oak as a case study. BMC Genom 11:570
Earl DA, vonHoldt BM (2012) Structure Harvester: a website and program for visualizing structure output and implementing the Evanno method. Conserv Genet Res 4:359–361
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of cluster of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Falush D, Stephens M, Pritchard JK (2007) Inference of population structure using multilocus genotype data: dominant markers and null alleles. Mol Ecol Notes 7:574–578
Fernandez-Lopez J, Vasquez-Ruiz-de-Occenda R, Diaz-Vasquez R, Pereira-Lorenzo S (2001) Evaluation of resistance of Castanea sp. clones to Phytophthora sp. using excised chestnut shoots. For Snow Landscape Res 76:451–454
Gallardo F (2002) Aprovechamiento tradicional del Castaño (Castanea sativa Mill.) en el Parque Natural Sierra Norte de Sevilla. Su Problemática. Constantina. III Congreso Forestal Español
Garrity DP (2004) Agroforestry and the achievement of the millennium development goals. Agroforest Syst 61:5–17
Glaubitz JC, Moran GF (2000) Genetic tools: the use of biochemical and molecular markers. In: Young A, Boshier D, Boyle T (eds) Forest conservation genetics. CSIRO Publishing, Australia, pp 39–52
Gobbin D, Hohl L, Conza L, Jermini M, Gessler C, Conedera M (2007) Microsatellite-based characterization of the Castanea sativa cultivar heritage of southern Switzerland. Genome 50:1089–1103
Graudal L, Aravanopoulos F, Bennadji Z, Changtragoon S, Fady B, Kjær ED, Loo J, Ramamonjisoa L, Vendramin GG (2014) Global to local genetic diversity indicators of evolutionary potential in tree species within and outside forests. Forest Ecol Manag 333:35–51
Grossmann A, Romane F (2004) Final report EU project CASCADE EVK-2-CT-1999–00000. http://soi.cnr.it/chestnut/. Accessed 4 April 2016
Hoffman AA, Willi Y (2008) Detecting genetic responses to environmental change. Nature Rev Genet 9:421–432
Hubisz MJ, Falush D, Stephen M, Pritchard JK (2009) Inferring weak population structure with the assistance of sample group information. Mol Ecol Res 9:1322–1332
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodal in analysis of population structure. Bioinformatics 14:1801–1806
Laikre L, Larsson LC, Palmé A, Charlier J, Josefsson M, Ryman N (2008) Potentials for monitoring gene level biodiversity: using Sweden as an example. Biodivers Conserv 17:893–910
Lynch M (1990) The similarity index and DNA fingerprint. Mol Biol Evol 7:478–484
Marinoni D, Akkak A, Bounous G, Edwards KJ, Botta R (2003) Development and characterization of microsatellite markers in Castanea sativa (Mill.). Mol Breeding 11:127–136
Marinoni D, Akkak A, Beltramo C, Guaraldo P, Boccacci P, Bounous G, Ferrara AM, Ebone A, Viotto E, Botta R (2013) Genetic and morphological characterization of chestnut (Castanea sativa Mill.) germplasm in Piedmont (north-western Italy). Tree Genet Genomes 9:1017–1030
Martín MA, Moral A, Martín LM, Alvarez JB (2007) The genetic resources of European sweet chestnut (Castanea sativa Miller) in Andalusia, Spain. Genet Resou Crop Evol 54:379–387
Martín MA, Alvarez JB, Mattioni C, Cherubini M, Villani F, Martín LM (2009) Identification and characterisation of traditional chestnut varieties of southern Spain using morphological and simple sequence repeats SSR markers. Ann Appl Biol 154:389–398
Martín MA, Mattioni C, Cherubini M, Taurchini D, Villani F (2010) Genetic characterisation of traditional chestnut varieties in Italy using microsatellites (SSRs) markers. Ann Appl Biol 157:37–44
Martín MA, Mattioni C, Molina JR, Alvarez JB, Cherubini M, Herrera MA, Villani F, Martín LM (2012) Landscape genetic structure of chestnut (Castanea sativa Mill.) in Spain. Tree Genet Genomes 8:127–136
Martín MA, Mattioni C, Cherubini M, Villani F, Martín LM (2016) A comparative study of European chestnut varieties in relation to adaptive markers. Agroforest Syst. doi:10.1007/s10457-016-9911-5
Mattioni C, Martín MA, Pollegioni P, Cherubini M, Villani F (2013) Microsatellite markers reveal a strong geographical structure in European populations of Castanea sativa (Fagaceae): evidence for multiple glacial refugia. Am J Bot 100:1–11
Maxted N, Ford-Lloyd BV, Hawkes JG (1997) Complementary conservation strategies. In: Maxted N, Ford-Lloyd BV, Hawkes JG (eds) Plant genetic conservation. Chapman and Hall, London, pp 15–39
Michon G (2005) Domesticating forests: how farmers manage forest resources. Institut de Richerche pour le Dèveloppement, Paris, France, the Centre for International Forestry Research, Borgor, Indonesia, and the World Agroforestry Centre, Nairobi, Kenya
Muñoz C, Trujillo I, Martínez-Urdioz Barranco D, Rallo L, Marfil P, Gaut BS (2015) Olive domestication and diversification in the Mediterranean Basin. New Phytol 1:436–447
Peakall R, Smouse PE (2005) GenAlEx 6: genetic analysis in excell. Population genetic software for teaching and research. Australian National University, Canberra http://www.anu.edu.au/BoZo/GenAlEx
Pereira-Lorenzo S, Costa R, Ramos-Cabrer A, Ribeiro C, da Silva M, Manzano G, Barreneche T (2010) Variation in grafted european chestnut and hybrids microsatellite reveals two main origins in the Iberian Peninsula. Tree Genet Genomes 5:701–715
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rohlf FJ, Fisher DL (1986) Test for hierarchical structure in random data sets. Syst Zool 17:407–412
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Schneider S, Roessli D, Excoffier L (2000) Arlequin: a software for population genetics data analysis, version 3.1. genetics and Biometry Laboratory, Department of Anthropology, University of Geneva, Switzerland
Schwatz MK, Luikart G, Waples R (2006) Genetic monitoring as a promising tool for conservation and management. Trends Ecol Evol 22:25–33
Scott KD, Eggler P, Seaton G, Rossetto M, Ablett EM, Lee LS, Henry RJ (2000) Analysis of SSRs derived from grape ESTs. Theor Appl Genet 100:723–726
Slatkin M (1995) A measure of population subdivision based on microsatellite allele frequencies. Genetics 139:457–462
UPOV (1989) Guidelines for the conduct of test for distinctness, homogeneity and stability (Chestnut, Castanea sativa Mill.). TG/124/3, 23 pp
Varshney RK, Graner A, Sorrells ME (2005) Genic microsatellite markers in plants: features and applications. Trends Biotechnol 23:48–55
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of populations structure. Evolution 38:1358–1370
Woodhead M, Russell J, Squirrell J, Hollingsworth PM, Mackenzie K, Gibby M, Powell W (2005) Comparative analysis of population genetic structure in Athyrium distentifolium (Pteridophyta) using AFLPs and SSRs from anonymous and transcribed gene regions. Mol Ecol 14:1681–1695
Acknowledgements
The first author is grateful to the Secretaría General de Ciencia, Tecnología e Innovación de la Consejería de Economía e Infraestructuras from the Regional Government of Extremadura (Spain) for the financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This research was supported by grants AGL2013-48017-C2-1-R and AGL-2014-53822-C2-1-R from the Spanish Ministry of Economy and Competitiveness and the European Regional Development Fund (FEDER) from the European Union.
Additional information
Handling Editor: Bruno Fady
Contribution of the co-authors M. Angela Martín participated in the design of the experiment, ran the data analysis, led the discussion and wrote the paper. Elvira Monedero participated in the collection of data in the field and conducted the laboratory analyses. Luis Miguel Martín designed the experiment, conducted the collection of data in the field, supervised the work and coordinated the research project.
Rights and permissions
About this article
Cite this article
Martín, M., Monedero, E. & Martín, L. Genetic monitoring of traditional chestnut orchards reveals a complex genetic structure. Annals of Forest Science 74, 15 (2017). https://doi.org/10.1007/s13595-016-0610-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13595-016-0610-1