Abstract
Acute myeloid leukemia (AML), the most common acute leukemia in adults, increases exponentially with age. While a number of recent advances have improved treatment, high cure rates have not yet been achieved. Nucleophosmin (NPM1) is found mutated in nearly one-third of newly diagnosed cases and leads to NPM1 protein that is mislocalized to the cytoplasm instead of the nucleolus. If the mechanistic basis through which this mislocalization leads to malignancy could be revealed, this AML subtype may be targetable with new drugs. Here, we review the structure and functions of the normal and mutant forms of nucleophosmin. We discuss several recent studies that have shed light on the pathophysiology of NPM1 mutations. We discuss the importance of HOX gene misregulation in NPM1-mutated leukemias, as well as evidence for the reliance of mutated NPM1 on its continued nuclear export. Together, these aspects, as well as new tools to manipulate and study NPM1, open the door to new therapeutic strategies that may ultimately improve treatment of this common subtype of AML.
Similar content being viewed by others
Introduction
Acute myeloid leukemia (AML) is the most common acute leukemia in adults. Currently, about 40–50% of AML patients (age 18–60 years) can be cured using conventional chemotherapy [1]. However, the incidence of AML increases exponentially with age (median age at diagnosis about 70 years) [2] and the cure rate is significantly lower (5–10%) in older adults (> 60 years) [1]. Although high levels of supportive care combined with an accurate relapse risk prediction have improved patient outcomes [1], the goal of a high cure rate in AML has not yet been achieved. Thus, increasing our understanding of the cellular and molecular biology of AML subtypes is necessary for the development of new therapeutic approaches to meet this urgent medical need.
Recent large-scale genomic studies have outlined the genetic landscape of adult AML, paving the way for the development of new therapeutic strategies [3, 4]. Nucleophosmin (NPM1) is among the most frequently mutated genes in AML, found in nearly one-third of newly diagnosed cases both in younger and older adults. AML with mutated NPM1 was described for the first time more than 10 years ago [5]. Since then, individual and cooperative studies have answered many of the relevant clinical and biological questions [6], contributing to the classification of this AML subtype as a distinct independent clinical entity in the WHO classification of hematopoietic tumors [7]. However, many critical questions remain open, including the molecular mechanisms by which mutant NPM1 promotes leukemogenesis and whether mutant NPM1 is necessary for the maintenance of leukemia.
Here, we review the structure and functions of the normal and mutant forms of nucleophosmin, with a particular emphasis on recent studies that have shed light on the pathophysiology of NPM1 mutations and opened the door to the development of new therapies for one of the most common subtypes of AML.
Structure of NPM1 gene and protein
NPM1 is a nucleolar multifunctional phosphoprotein (hence the name nucleophosmin) that shuttles between the nucleolus and the cytoplasm [8]. The human NPM1 gene spans about 23 kb at chromosome 5q35 and contains 12 exons and can be transcribed into three major transcript variants (Fig. 1a). Transcript variant 1 is the longest and the most represented among the variants and encodes for isoform 1, with 11 exons (1–9 and 11–12) and 294 amino acids (Fig. 1a).
NPM1 is a member of the nucleophosmin/nucleoplasmin (NPM) family of nuclear chaperones, which also includes NPM2 and NPM3 [9] (Fig. 1b). The three NPM family members share significant sequence and structural homology. All NPM proteins contain an N-terminal “core” region, which is the most conserved among the NPM family (Fig. 1b). This region is responsible for protein oligomerization [10, 11] and is crucial for its chaperone activity by preventing protein misfolding in the nucleoli [10, 12]. Additionally, the N-terminal domain contains two nuclear export signal (NES) motifs, both of which are critical for the nuclear-cytoplasmic shuttling of NPM1 [13, 14]. One of the two NES motifs has been demonstrated to enable the association of NPM1 with the centrosome, preventing its duplication. Removing or mutating this NES motif leads to nuclear retention of NPM1 and causes genome instability [14].
The central region of NPM proteins is highly disordered and contains two ~ 10–20 amino acid stretches of acidic aspartic and glutamic acid residues as well as a nuclear localization signal (NLS) (Fig. 1b). The negative charge associated with the aspartic and glutamic acids is involved in the binding of NPM proteins to histones, possibly by mimicking the negative charge of RNA/DNA [15, 16]. This intrinsically disordered region also has been demonstrated to drive homo- and heterotypic liquid–liquid phase separation of NPM1 [17, 18], which may be a key biophysical characteristic driving the localization and function of the protein in the nucleolus.
Between the acidic region and the C terminus, NPM1 contains a 55 amino acid sequence enriched in basic residues. This basic domain is suggested to be important for the interactions between NPM1 and nucleic acids [19] and mediates NPM1 binding to p53, enhancing its stability and tumor suppressor functions [20]. This domain also contains the second NLS of NPM1 (Fig. 1b).
The C-terminal region of NPM1 is unique to the isoform 1 of NPM1 [21] (Fig. 1b). This region contains several aromatic residues, including W288 and W290. These two amino acids are critical for proper three-helix folding of the C-terminal domain, and are necessary for nucleolar localization of NPM1 (nucleolar localization signal or NoLS) (Fig. 1b) [22, 23].
NPM1 is a multifunctional protein
The discovery of NPM1 mutations in cancer has significantly increased research in NPM1, resulting in a plethora of data supporting numerous putative functions for WT NPM1 (Table 1). Here, we summarize in brief the role of NPM1 in the following processes: histone chaperoning, ribosome biogenesis, centrosome duplication and the DNA damage response.
Similar to other members of the NPM family [24], NPM1 has been shown to act as a histone chaperone and participate in nucleosome formation (Fig. 2) through interactions with both core histones H2A/H2B/H3 [25] and the linker histone H1 [16]. These interactions appear to be important for maintaining both active and repressed genomic regions [26,27,28,29].
The structure and functional domains of NPM1 strongly support its role in ribosome processing and assembly (Fig. 2). Indeed, NPM1 is abundant in the nucleolus, where ribosomal biogenesis takes place, and its nucleocytoplasmic shuttling activity has been shown to facilitate transport of ribosomal components from the nucleolus to the cytoplasm [13, 30, 31] and vice versa [32]. Additionally, NPM1 has intrinsic ribonuclease activity and is capable of cleaving the pre-ribosome into small and large subunits [33,34,35]. Curiously, despite its localization in the granular compartment of the nucleolus, the role that NPM1 plays in regulating ribosomal biogenesis is unclear. Loss of NPM1 triggers a number of cellular responses, including activation of the DNA damage response and increased genome instability, but no significant changes in ribosomal biogenesis have been observed [36]. Therefore, further studies are needed to establish the exact role of NPM1 in ribosome biogenesis.
NPM1 is a key player in the control of genome stability. Specifically, NPM1 directly binds to centrosomes during interphase and its presence is required to prevent spurious centrosome duplication (Fig. 2). NPM1 knock-out mice die between embryonic day E9 and E16 due to severe anemia resulting from aberrant centrosome duplication and subsequent genomic instability [36, 37]. Upon cell cycle progression, the CDK2-cyclin E complex phosphorylates NPM1 on Thr-199, causing its dissociation from centrosomes and thus allowing for proper duplication and mitotic spindle formation [38].
The most well-characterized function of NPM1 involves the regulation of tumor suppressor proteins and the DNA damage response. NPM1 is known to regulate the stability of several tumor suppressors such as ARF and p53 (Fig. 2). Specifically, NPM1 binds to ARF, protecting it from degradation [39, 40], which in turn represses cell proliferation by stabilizing p53 [41]. Consistently, reducing the level of NPM1 accelerates tumorigenesis [36], probably as a result of the reduced stability of ARF. Additionally, NPM1 is known to increase p53 stability by directly binding and inhibiting the p53 E3-ligase MDM2 (Fig. 2). The NPM1-MDM2 interaction is particularly enhanced upon NPM1 nuclear relocalization following nucleolar stress [42] and this pathway is currently being investigated for the development of novel therapies in NPM1-mutated AML through nucleolar stress induction [43].
In summary, NPM1 appears to be involved in several critical biological processes. However, most of its putative functions have been inferred based on either indirect observations or in vitro experiments. As of today, the only well-characterized function of NPM1 is the regulation of centrosome duplication. More in vivo studies are needed to validate the major roles of this multifunctional protein.
NPM1 mutations in AML
The WHO classification of hematopoietic tumors recognizes AML with mutated NPM1 as a distinct entity [7]. NPM1-mutated AML typically occur as de novo myelomonocytic or monocytic acute leukemia (M4 or M5 of the French–American–British classification) [7]. Strikingly, about half of NPM1-mutated AML cases have mutations in DNMT3A and about 40% have co-occurring FLT3 mutations [44]. This association strongly indicates cooperation between these mutations, and double and triple-mutant mouse models have confirmed this hypothesis [45]. The prognosis of NPM1/FLT3 double-mutant and NPM1/DNMT3A/FLT3 triple-mutant AML cases is still very poor and new therapeutic strategies for these AML genotypes are urgently needed.
The discovery of NPM1 mutations in AML was unexpected. In the early 2000s, translocations involving NPM1 in lymphoid and myeloid malignancies had been reported [46]. At that time, detection of aberrant cytoplasmic localization of NPM1 through immunohistochemistry was widely used to predict these genetic lesions in patient samples. In fact, in lymphomas bearing NPM1-ALK translocation, NPM1 is delocalized from the nucleoli to the cytoplasm [47]. Using the same staining on bone marrow biopsies, researchers were looking for novel NPM1 rearrangements in other hematological malignancies. Surprisingly, NPM1 was localized to the cytoplasm (NPM1 cytoplasmic positive or NPM1c+) in about 30% of AML cases. However, the vast majority of NPM1c+ AML cases had a normal karyotype and none had NPM1 rearrangements. Subsequently, the entire NPM1 gene was sequenced and NPM1 mutations were identified for the first time [5].
NPM1 mutations in AML are for the most part restricted to exon 12 [5]. More than 50 mutations in exon 12 in NPM1 have been described so far [6] and they usually consist of 4 bp insertions, occurring between nucleotides 960 and 961 [5]. Of these, the most common is mutation A, found in about 80% of cases, which consists of a duplication of TCTG at position 960 (Table 2). Mutations B and D, though less common, also produce a 4 bp insertion at position 960 (Table 2). Other mutations, including a few outside of exon 12, are rare, occurring in less than 1% of cases (Table 2) [6].
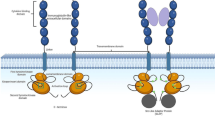
All exon 12 variants cause a frameshift in the last few C-terminal amino acids of NPM1. The frameshift eliminates one or both tryptophan residues responsible for the nucleolar localization of NPM1 and generates a new C-terminal NES motif (Fig. 3) [5, 48, 49]. From a structural standpoint, all exon 12 variants lead to C-terminal misfolding and loss of the NoLS. Additionally, the frameshift results in the generation of a novel NES that drastically increases the ability of NPM1 to bind the nuclear exporter CRM1/XPO1 [5, 48, 49]. Therefore, mutations in NPM1 result in a stronger overall nuclear export signal, which re-localizes NPM1 mutant protein from the nucleolus to the cytoplasm (NPM1 cytoplasmic or NPM1c) [5, 48, 49] (Fig. 3).
Top: representation of the wild type and mutant NPM1 alleles. Changes in the C-terminal aminoacid sequence are indicated. While the wild-type protein (red) is localized to the nucleus, mutant NPM1 (green) is in the cytoplasm of leukemic cells. Bottom: schematic representation of the interaction between mutant NPM1 and XPO1, that results in cytoplasmic localization of the mutant protein
The role of mutant NPM1 in leukemogenesis
The role of NPM1 in leukemogenesis and leukemia maintenance is unclear. Considering the myriad roles of the protein, NPM1 mutations likely have a complex pathophysiology in malignant hematopoietic cells.
Knock-out mice have demonstrated that the loss of one or both copies of NPM1 leads to uncontrolled centrosome duplication and aneuploidy [36]. While the complete absence of NPM1 (NPM1−/−) leads to embryonic lethality, heterozygous NPM1 (NPM1+/−) mice are viable. However, NPM1+/− mice show features of genomic instability and suffer from abnormal hematopoiesis with anemia, which resembles human myelodysplastic syndromes (MDS) [36]. Additionally, NPM+/− mice develop myeloid and lymphoid malignancies, albeit with a long latency [36, 50]. Altogether these studies demonstrate that the functions of NPM1 in centrosome duplication and maintenance of genomic integrity are dose dependent and that NPM1 can act as a haploinsufficient tumor suppressor in malignant hematopoiesis. Further, these findings suggest that loss of one copy of NPM1 in MDS with 5q35 deletions likely contributes to disease pathogenesis. However, given that 80% of AML with mutated NPM1 patients display normal karyotype and do not acquire 5q35 deletions [5, 7], whether haploinsufficiency in NPM1 contributes to disease in these patients remains unclear. Considering its putative role in ribosome biogenesis, we can hypothesize that the reduction of NPM1 dosage in the nucleoli in NPM1c+ AML could itself alter ribosome synthesis and promote leukemogenesis, by interfering with protein synthesis in leukemic cells. This remains to be shown.
NPM1 is responsible for the stabilization and the nucleolar localization of the tumor suppressor ARF. While the interaction of NPM1 and ARF is not disrupted by NPM1 mutations, Colombo et al. demonstrated that ARF is relocated to the cytoplasm where its half-life is significantly reduced compared to in the nucleoli [51]. This report indicated reduced ARF levels may promote cell cycle progression and leukemogenesis. However, as these studies were performed in mouse embryonic fibroblasts and NIH3T3 cells, the consequences of increased shuttling of ARF to the cytoplasm need to be verified using hematopoietic progenitor cells or primary leukemias.
The F-box protein Fbw7 is the substrate recognition component of an E3 ubiquitin-protein ligase complex that is dependent on NPM1 for its nucleolar localization. In AML with mutated NPM1, Fbw7 is delocalized to the cytoplasm along with NPM1c [52], resulting in increased turnover of the protein. Since Fbw7 is responsible for c-Myc degradation, reduced levels of this protein result in higher levels of the oncogene c-Myc [52]. The combination of higher c-Myc levels and lower ARF may synergistically promote leukemogenesis in NPM1c+ cells. However, this hypothesis requires further in vitro and in vivo experimental validation.
Transgenic mouse models have demonstrated that NPM1c+ mice develop clear signs of myeloproliferation in bone marrow and spleen [53]. Additionally, along with an increase in mature myeloid cells, transgenic mice show a striking block of megakaryocyte differentiation that closely resembles the megakaryocyte abnormalities seen in patients with AML with mutated NPM1 [54]. Similar results have been reported in knock-in models, where mutant NPM1 induced myeloproliferation as well as other features characteristic of human NPM1-mutated AML, such as increased expression of HOX genes in hematopoietic progenitors [55]. However, none of these models developed short-latency AML, suggesting that co-operating mutations are necessary for AML development and progression in mice [56]. Indeed, NPM1/FLT3 and NPM1/NRAS double-mutant mouse models both develop AML [56, 57]. Despite the fact that the molecular mechanisms behind these phenotypes are still obscure, both the knock-in and transgenic mouse models suggest that NPM1 mutations may act in a gain-of-function manner. This hypothesis is further supported by recent studies, as detailed below.
NPM1c facilitates HOX expression in a gain-of-function manner
AML with mutated NPM1 has a unique gene expression [58, 59] profile that is independent from other co-occurring mutations. Particularly, homeobox (HOX) genes HOXA and HOXB genes are consistently overexpressed compared to AML with wild type NPM1 [58, 59]. As these genes represent the major stem cell-related genes expressed in this AML, they are thought they play an important role in maintaining the self-renewal capacity of leukemic cells. Below, we specifically discuss the relevance of HOX genes in AML with mutated NPM1.
HOX genes are highly expressed during embryogenesis and encode for transcription factors involved in the regulation of anatomical development [60, 61]. Multiple HOX genes are found grouped in clusters (named A to D) within the same genomic locus. Interestingly, the HOXA and HOXB gene clusters are highly expressed in normal adult hematopoietic stem cells (HSCs), but they are silenced in mature blood cells [62], suggesting a role in HSC self-renewal. Additionally, ectopic expression of HOX genes transforms hematopoietic progenitor cells into leukemia [62]. As described above, one of the characterizing features of NPM1-mutated AML is the consistent high expression of HOXA and HOXB [63]. While this feature was first reported in 2005 [58, 59], whether high HOX expression depends directly on NPM1c or just reflects a specific differentiation state of AML cells remains unclear.
Our group recently reported that acute loss of NPM1c results in dramatic drop (within 6 h) in the expression of multiple HOX genes and that AML cells differentiate only few days after this loss [64]. In the same work, we demonstrated that ectopic expression of HOXA9 and the HOX co-factor MEIS1 largely rescues differentiation upon loss of NPM1c [64]. These observations demonstrate that high levels of HOX is directly dependent on NPM1c and that NPM1c acts upstream of HOX to maintain the undifferentiated state of AML cells.
While our results establish that NPM1c has a critical role in shaping the transcriptome of AML cells and acts in a gain-of-function manner, the mechanisms through which mutant NPM1 promotes HOX expression remain elusive. HOX expression is known to be strongly regulated by selected activating (e.g., H3K4me3, H3K79me2, H3K27Ac) and repressive (e.g., H3K27me3) histone marks [65,66,67,68]. Interestingly, targeting H3K4 and H3K79 methylation levels through inhibition of the MLL–Menin interaction and DOT1L, respectively, have been proven to be effective therapeutic strategies in models of AML with mutated NPM1 [69]. Kühn et al. showed that pharmacologic reduction of H3K4me3 and H3K79me2 levels resulted in lower HOXA, HOXB and MEIS1 expression levels, followed by differentiation [69]. Our group investigated the impact of nuclear relocalization of NPM1c on H3K27Ac, demonstrating that loss of NPM1 from the cytoplasm results in a significant decrease of acetylated H3K27, particularly at HOX loci [64]. Loss of H3K27Ac was coupled with lower HOX expression and was followed by differentiation [64]. While our data emphasize the role of active histone marks in supporting high HOX expression and maintaining the undifferentiated state of AML cells, more studies are necessary to shed light on putative direct connections between NPM1c and HOX genes.
One recent study suggested that NPM1c may bind to the myeloid transcription factor PU.1, relocating it to the cytoplasm [70]. This study proposed that reduced nuclear levels of PU.1 may result in HOX overexpression, as ectopic PU.1 expression led to HOX downregulation [70]. An alternative hypothesis involves the architectural protein CTCF. NPM1c has been recently reported to bind and displace CTCF to the cytoplasm, altering the landscape of CTCF binding sites and promoting HOX [71]. Additionally, Luo et al. reported that the CTCF binding site between HOXA7 and HOXA9 is essential for maintaining high expression of multiple HOXA genes in leukemic cells [72], further tightening the association between CTCF and HOX genes in AML. Both these hypotheses are intriguing; however, further experiments in appropriate models are needed to definitively clarify how HOX genes are regulated by PU.1 and CTCF in AML cells with mutated NPM1.
Recent studies have also investigated downstream targets of HOX transcription factors in leukemia. ChIP-sequencing studies performed in HOXA9/MEIS1-transformed murine bone marrow cells have shown that HOXA9 mostly binds to active distal enhancers [73, 74]. HOXA9 binding sites were enriched for consensus sequences of several myeloid transcription factors, including GATA, CEBP and PU.1, but also HOXA9, likely resulting in an autoregulatory circuit [73, 74]. Importantly, distal enhancer-associated genes were upregulated upon binding of HOXA9 and downregulated upon HOXA9 inactivation, suggesting a direct role in regulating transcription at target gene [74]. We are just beginning to understand the functions of HOX transcription factors in leukemia, and more work is required to understand which HOX downstream targets are crucial for leukemic transformation and whether HOX transcription factors, such as HOXA9, could be directly targeted for therapeutic purposes. Understanding the exact mechanisms through which mutant NPM1 promotes HOX expression and how HOX genes support leukemic transformation and maintenance are both critical steps that will open new therapeutic avenues for a large fraction of AML patients.
Nuclear export of NPM1c is a therapeutic vulnerability in AML with mutated NPM1
Evidence suggests that NPM1 mutations are founding genetic lesions in AML. NPM1 mutations, which are frequent events in AML, are never found in clonal hematopoiesis or pre-leukemic clones but are present in all leukemic cells at AML diagnosis [75, 76]. In addition, while NPM1 mutants are undetectable in patients in complete remission, they are present at relapse in about 90% of patients previously diagnosed with AML with mutated NPM1 [77], suggesting that these mutations occur in AML founding clones. Therefore, in contrast with subclonal late-occurring mutations (i.e., mutations present only in a fraction of AML cells at diagnosis, e.g., FLT3-ITD), NPM1 mutations could be an ideal candidate for targeted therapy as they are present in all leukemic cells.
All NPM1 mutations in AML described so far result in the generation of a novel C-terminal NES. Exportin-1 (XPO1) is a nuclear exporter mainly localized at the nuclear pore, responsible for cytoplasmic shuttling of proteins that contain a NES. As consequence, the interaction between XPO1 and mutant NPM1 results in the cytoplasmic localization of NPM1c. Two recent studies have demonstrated that the interaction between XPO1 and NPM1c is essential for AML maintenance [64, 70]. In both studies, pharmacological inhibition of nuclear export by Selinexor, a selective XPO1 inhibitor, resulted in nuclear relocalization of NPM1c, differentiation of AML cells and antileukemic effect in vivo.
As XPO1 inhibition is predicted to result in nuclear accumulation of multiple tumor suppressors, including TP53 and pRB [78], the antileukemic effects exerted by Selinexor in the above studies could not be attributed solely to the relocalization of NPM1c. However, using specific gene editing techniques, our group demonstrated that restoring the nuclear localization of NPM1c by itself is sufficient to induce differentiation and inhibit growth of NPM1-mutated AML cells [64]. Therefore, we believe that the cytoplasmic localization of NPM1c creates a therapeutic vulnerability that can be exploited through the inhibition of nuclear export. We also speculate that loss of the XPO1-NPM1c interaction coupled with increased nuclear concentrations of key tumor suppressors such as TP53 produces synergistic antileukemic effects in NPM1-mutated AML cells.
Selinexor has a defined toxicity profile, characterized by nausea and anorexia as major side effects, and published clinical trials in both solid [79, 80] and hematological cancers [81, 82] administered it one to three times per week. Selinexor has been also tested as single agent in a phase I trial in AML [83]. Surprisingly, NPM1-mutated AML patients treated with the drug did not show better outcomes compared to those with WT NPM1 [83]. Unpublished in vitro data from our group suggest that constant XPO1 inhibition is needed to obtain significant differentiation and growth arrest of AML cells. Current administration schedules are therefore unlikely to result in constant relocalization of NPM1c to the nucleus in leukemic cells. Thus, infrequent administration of Selinexor in patients may not reproduce what we observed upon genetic disruption of nuclear export or in vitro pharmacologic XPO1 inhibition. An alternative strategy could be to administer lower doses more frequently. In this regard, Gu et al. successfully treated one single NPM1-mutant patient-derived xenograft mouse model using low-dose Selinexor, administered four times per week [70]. While this strategy seems promising, further validation is needed.
Newer nuclear export inhibitors are currently being tested pre-clinically in leukemia, with promising results [84, 85]. While maintaining the cytotoxic effects of current XPO1 inhibitors, these newer compounds have demonstrated more favorable toxicity profiles in mouse models [84, 85]. Before next-generation nuclear export inhibitor is tested in clinical trials specifically designed for AML patients with mutated NPM1, further pre-clinical and clinical data are needed.
Conclusions
AML has one of the least complex genomic landscapes of any cancer, with an average of fewer than 15 somatic coding mutations per AML genome [86] compared to the hundreds of coding mutations observed in cancers of the breast, lung, and colon [87]. This simplicity enables the prioritization of targets for intervention and supports rational drug design. As nearly a third of patients with AML have a mutation in NPM1 and these mutations are gatekeepers for the maintenance of the malignancy, NPM1 could be a promising therapeutic target.
Recent work has demonstrated that AML with mutated NPM1 is addicted to nuclear export of the mutant protein, and that the leukemic activity of the mutant protein is linked to overactivity of the HOX program [64]. The finding that AML with mutated NPM1 is dependent on NPM1c should lead to further investment into the discovery of drugs and small molecules that interfere with the function of NPM1c or downstream effectors, such as HOX genes. The identification of such compounds would lead to a significant development in the AML field. Further, due to the fact that NPM1 mutations represent founder events, rather than subclonal mutations such as FLT3, the risk of relapse with these agents would potentially be much lower. While we are likely decades away from meaningful intervention with these types of compounds, a number of candidates for intervention have already been found.
Menin–MLL inhibitors and DOT1L inhibitors both display significant antileukemic activity in in vitro and in vivo models of NPM1-mutant leukemia [69]. Furthermore, these inhibitors synergistically act to suppress HOX gene expression and reduce leukemia burden [69]. As we continue to uncover the mechanisms that link mutant NPM1 to HOX activation, more targets for therapeutic intervention may be uncovered leading to targeted therapies for AML with mutated NPM1.
Small molecule selective inhibitors of nuclear export, which specifically inhibit XPO1 and thereby disrupt the interaction of XPO1 with mutant NPM1, effectively relocalize mutant NPM1 to the nucleus, reduce the activity of the HOX program, and hinder the self-renewal ability of leukemic cells. These features provide significant rationale for the clinical use of XPO1 inhibitors, both as single agents and in combination with other therapies, with the caveat that optimized administration scheduled may be necessary to achieve the highest antileukemic effects.
The emergence of CRISPR technologies allowing for endogenous tagging of both WT and mutant forms of NPM1 ([64] and unpublished data) will enable a more complete understanding of the functions of WT NPM1 in the nucleolus and mutant NPM1 in the cytoplasm and during nuclear export. For example, we now have the tools to screen for selective degraders of mutant NPM1 through the use of endogenous fluorescent tagging approaches. CRISPR dropout screens and small molecule screens may identify proteins that regulate the stability of mutant NPM1 in the cytoplasm. These studies will be crucial for identifying new therapeutic approaches to target mutant NPM1.
In summary, mutations in NPM1 have now been shown unequivocally to maintain leukemia and therefore represent one of the most important targets for therapeutic intervention in AML. Further research into this disease will undoubtedly lead to new targeted therapies for AML and a better understanding of the mechanisms through which the HOX program drives leukemogenesis.
References
Döhner H, Weisdorf DJ, Bloomfield CD. Acute Myeloid Leukemia. N Engl J Med. 2015;373:1136–52. https://doi.org/10.1056/NEJMra1406184.
Deschler B, Lubbert M. Acute myeloid leukemia: epidemiology and etiology. Cancer. 2006;107:2099–107. https://doi.org/10.1002/cncr.22233.
Network C. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med. 2013;368:2059–74. https://doi.org/10.1056/NEJMoa1301689.
Papaemmanuil E, et al. Genomic classification and prognosis in acute myeloid leukemia. N Engl J Med. 2016;374:2209–21. https://doi.org/10.1056/NEJMoa1516192.
Falini B, et al. Cytoplasmic nucleophosmin in acute myelogenous leukemia with a normal karyotype. N Engl J Med. 2005;352:254–66. https://doi.org/10.1056/NEJMoa041974.
Heath EM, et al. Biological and clinical consequences of NPM1 mutations in AML. Leukemia. 2017;31:798–807. https://doi.org/10.1038/leu.2017.30.
Swerdlow SH. International Agency for Research on Cancer & World Health Organization. WHO classification of tumours of haematopoietic and lymphoid tissues, 4th edn. International Agency for Research on Cancer; 2008. ISBN-10: 9283224310
Borer RA, Lehner CF, Eppenberger HM, Nigg EA. Major nucleolar proteins shuttle between nucleus and cytoplasm. Cell. 1989;56:379–90.
Olson MOJ. The nucleolus. New York: Springer; 2011.
Namboodiri VM, Akey IV, Schmidt-Zachmann MS, Head JF, Akey CW. The structure and function of Xenopus NO38-core, a histone chaperone in the nucleolus. Structure. 2004;12:2149–60. https://doi.org/10.1016/j.str.2004.09.017.
Lee HH, et al. Crystal structure of human nucleophosmin-core reveals plasticity of the pentamer–pentamer interface. Proteins. 2007;69:672–8. https://doi.org/10.1002/prot.21504.
Szebeni A, Olson MO. Nucleolar protein B23 has molecular chaperone activities. Protein Sci. 1999;8:905–12. https://doi.org/10.1110/ps.8.4.905.
Maggi LB Jr, et al. Nucleophosmin serves as a rate-limiting nuclear export chaperone for the Mammalian ribosome. Mol Cell Biol. 2008;28:7050–65. https://doi.org/10.1128/MCB.01548-07.
Wang W, Budhu A, Forgues M, Wang XW. Temporal and spatial control of nucleophosmin by the Ran-Crm1 complex in centrosome duplication. Nat Cell Biol. 2005;7:823–30. https://doi.org/10.1038/ncb1282.
Swaminathan V, Kishore AH, Febitha KK, Kundu TK. Human histone chaperone nucleophosmin enhances acetylation-dependent chromatin transcription. Mol Cell Biol. 2005;25:7534–45. https://doi.org/10.1128/MCB.25.17.7534-7545.2005.
Gadad SS, et al. The multifunctional protein nucleophosmin (NPM1) is a human linker histone H1 chaperone. Biochemistry. 2011;50:2780–9. https://doi.org/10.1021/bi101835j.
Mitrea DM, et al. Nucleophosmin integrates within the nucleolus via multi-modal interactions with proteins displaying R-rich linear motifs and rRNA. Elife; 2016. https://doi.org/10.7554/eLife.13571 (2016).
Mitrea DM, et al. Self-interaction of NPM1 modulates multiple mechanisms of liquid–liquid phase separation. Nat Commun. 2018;9:842. https://doi.org/10.1038/s41467-018-03255-3.
Hingorani K, Szebeni A, Olson MO. Mapping the functional domains of nucleolar protein B23. J Biol Chem. 2000;275:24451–7. https://doi.org/10.1074/jbc.M003278200.
Colombo E, Marine J-C, Danovi D, Falini B, Pelicci P. Nucleophosmin regulates the stability and transcriptional activity of p53. Nat Cell Biol. 2002;4:529–33. https://doi.org/10.1038/ncb814.
Chang JH, Olson MO. A single gene codes for two forms of rat nucleolar protein B23 mRNA. J Biol Chem. 1989;264:11732–7.
Nishimura Y, Ohkubo T, Furuichi Y, Umekawa H. Tryptophans 286 and 288 in the C-terminal region of protein B23.1 are important for its nucleolar localization. Biosci Biotechnol Biochem. 2002;66:2239–42. https://doi.org/10.1271/bbb.66.2239.
Grummitt CG, Townsley FM, Johnson CM, Warren AJ, Bycroft M. Structural consequences of nucleophosmin mutations in acute myeloid leukemia. J Biol Chem. 2008;283:23326–32. https://doi.org/10.1074/jbc.M801706200.
Dingwall C, Laskey RA. Nucleoplasmin: the archetypal molecular chaperone. Semin Cell Biol. 1990;1:11–7.
Okuwaki M, Matsumoto K, Tsujimoto M, Nagata K. Function of nucleophosmin/B23, a nucleolar acidic protein, as a histone chaperone. FEBS Lett. 2001;506:272–6.
Shandilya J, et al. Acetylated NPM1 localizes in the nucleoplasm and regulates transcriptional activation of genes implicated in oral cancer manifestation. Mol Cell Biol. 2009;29:5115–27. https://doi.org/10.1128/MCB.01969-08.
Malik-Soni N, Frappier L. Nucleophosmin contributes to the transcriptional activation function of the Epstein-Barr virus EBNA1 protein. J Virol. 2014;88:2323–6. https://doi.org/10.1128/JVI.02521-13.
Holmberg Olausson K, Nister M, Lindstrom MS. Loss of nucleolar histone chaperone NPM1 triggers rearrangement of heterochromatin and synergizes with a deficiency in DNA methyltransferase DNMT3A to drive ribosomal DNA transcription. J Biol Chem. 2014;289:34601–19. https://doi.org/10.1074/jbc.M114.569244.
Liu H, et al. Nucleophosmin acts as a novel AP2alpha-binding transcriptional corepressor during cell differentiation. EMBO Rep. 2007;8:394–400. https://doi.org/10.1038/sj.embor.7400909.
Yu Y, et al. Nucleophosmin is essential for ribosomal protein L5 nuclear export. Mol Cell Biol. 2006;26:3798–809. https://doi.org/10.1128/MCB.26.10.3798-3809.2006.
Pelletier CL, et al. TSC1 sets the rate of ribosome export and protein synthesis through nucleophosmin translation. Cancer Res. 2007;67:1609–17. https://doi.org/10.1158/0008-5472.CAN-06-2875.
Lindström MS, Zhang Y, Ribosomal Protein S. 9 Is a novel B23/NPM-binding protein required for normal cell proliferation. J Biol Chem. 2008;283:15568–76. https://doi.org/10.1074/jbc.M801151200.
Savkur RS, Olson MO. Preferential cleavage in pre-ribosomal RNA by protein B23 endoribonuclease. Nucleic Acids Res. 1998;26:4508–15.
Pan WA, et al. The RNA recognition motif of NIFK is required for rRNA maturation during cell cycle progression. RNA Biol. 2015;12:255–67. https://doi.org/10.1080/15476286.2015.1017221.
Itahana K, et al. Tumor suppressor ARF degrades B23, a nucleolar protein involved in ribosome biogenesis and cell proliferation. Mol Cell. 2003;12:1151–64.
Grisendi S, et al. Role of nucleophosmin in embryonic development and tumorigenesis. Nature. 2005;437:147–53. https://doi.org/10.1038/nature03915.
Colombo E, et al. Nucleophosmin is required for DNA integrity and p19Arf protein stability. Mol Cell Biol. 2005;25:8874–86. https://doi.org/10.1128/MCB.25.20.8874-8886.2005.
Mukhopadhyay A, et al. 14-3-3gamma prevents centrosome amplification and neoplastic progression. Sci Rep. 2016;6:26580. https://doi.org/10.1038/srep26580.
Kuo ML, den Besten W, Thomas MC, Sherr CJ. Arf-induced turnover of the nucleolar nucleophosmin-associated SUMO-2/3 protease Senp3. Cell Cycle. 2008;7:3378–87. https://doi.org/10.4161/cc.7.21.6930.
Kuo ML, den Besten W, Bertwistle D, Roussel MF, Sherr CJ. N-terminal polyubiquitination and degradation of the Arf tumor suppressor. Genes Dev. 2004;18:1862–74. https://doi.org/10.1101/gad.1213904.
Sherr CJ. The INK4a/ARF network in tumour suppression. Nat Rev Mol Cell Biol. 2001;2:731–7. https://doi.org/10.1038/35096061.
Rubbi CP, Milner J. Disruption of the nucleolus mediates stabilization of p53 in response to DNA damage and other stresses. EMBO J. 2003. https://doi.org/10.1093/emboj/cdg579.
Falini B, Brunetti L, Martelli MP. Dactinomycin in NPM1-mutated acute myeloid leukemia. N Engl J Med. 2015;373:1180–2. https://doi.org/10.1056/NEJMc1509584.
Brunetti L, Gundry MC, Goodell MA. DNMT3A in leukemia. Cold Spring Harb Perspect Med; 2017. https://doi.org/10.1101/cshperspect.a030320.
Guryanova OA, et al. DNMT3A mutations promote anthracycline resistance in acute myeloid leukemia via impaired nucleosome remodeling. Nat Med. 2016;22:1488–95. https://doi.org/10.1038/nm.4210.
Falini B, Mason DY. Proteins encoded by genes involved in chromosomal alterations in lymphoma and leukemia: clinical value of their detection by immunocytochemistry. Blood. 2002;99:409–26.
Falini B, et al. Translocations and mutations involving the nucleophosmin (NPM1) gene in lymphomas and leukemias. Haematologica. 2007;92:519–32.
Falini B, et al. Altered nucleophosmin transport in acute myeloid leukaemia with mutated NPM1: molecular basis and clinical implications. Leukemia. 2009;23:1731–43. https://doi.org/10.1038/leu.2009.124.
Falini B, et al. Both carboxy-terminus NES motif and mutated tryptophan(s) are crucial for aberrant nuclear export of nucleophosmin leukemic mutants in NPMc + AML. Blood. 2006;107:4514–23. https://doi.org/10.1182/blood-2005-11-4745.
Sportoletti P, et al. Npm1 is a haploinsufficient suppressor of myeloid and lymphoid malignancies in the mouse. Blood. 2008;111:3859–62. https://doi.org/10.1182/blood-2007-06-098251.
Colombo E, et al. Delocalization and destabilization of the Arf tumor suppressor by the leukemia-associated NPM mutant. Cancer Res. 2006;66:3044–50. https://doi.org/10.1158/0008-5472.CAN-05-2378.
Bonetti P, et al. Nucleophosmin and its AML-associated mutant regulate c-Myc turnover through Fbw7 gamma. J Cell Biol. 2008;182:19–26. https://doi.org/10.1083/jcb.200711040.
Cheng K, et al. The cytoplasmic NPM mutant induces myeloproliferation in a transgenic mouse model. Blood. 2010;115:3341–5. https://doi.org/10.1182/blood-2009-03-208587.
Sportoletti P, et al. The human NPM1 mutation A perturbs megakaryopoiesis in a conditional mouse model. Blood. 2013;121:3447–58. https://doi.org/10.1182/blood-2012-08-449553.
Vassiliou GS, et al. Mutant nucleophosmin and cooperating pathways drive leukemia initiation and progression in mice. Nat Genet. 2011;43:470–5. https://doi.org/10.1038/ng.796.
Sportoletti P, et al. Mouse models of NPM1-mutated acute myeloid leukemia: biological and clinical implications. Leukemia. 2015;29:269–78. https://doi.org/10.1038/leu.2014.257.
Dovey OM, et al. Molecular synergy underlies the co-occurrence patterns and phenotype of NPM1-mutant acute myeloid leukemia. Blood. 2017. https://doi.org/10.1182/blood-2017-01-760595.
Alcalay M, et al. Acute myeloid leukemia bearing cytoplasmic nucleophosmin (NPMc + AML) shows a distinct gene expression profile characterized by up-regulation of genes involved in stem-cell maintenance. Blood. 2005;106:899–902. https://doi.org/10.1182/blood-2005-02-0560.
Verhaak RG, et al. Mutations in nucleophosmin (NPM1) in acute myeloid leukemia (AML): association with other gene abnormalities and previously established gene expression signatures and their favorable prognostic significance. Blood. 2005;106:3747–54. https://doi.org/10.1182/blood-2005-05-2168.
Mallo M, Alonso CR. The regulation of Hox gene expression during animal development. Development. 2013;140:3951–63. https://doi.org/10.1242/dev.068346.
Pearson JC, Lemons D, McGinnis W. Modulating Hox gene functions during animal body patterning. Nat Rev Genet. 2005;6:893–904. https://doi.org/10.1038/nrg1726.
Argiropoulos B, Humphries RK. Hox genes in hematopoiesis and leukemogenesis. Oncogene. 2007;26:6766–76. https://doi.org/10.1038/sj.onc.1210760.
Spencer DH, et al. Epigenomic analysis of the HOX gene loci reveals mechanisms that may control canonical expression patterns in AML and normal hematopoietic cells. Leukemia. 2015;29:1279–89. https://doi.org/10.1038/leu.2015.6.
Brunetti L, et al. Mutant NPM1 maintains the leukemic state through HOX expression. Cancer Cell. 2018;34(499):499–512. https://doi.org/10.1016/j.ccell.2018.08.005. e.
Cao K, et al. SET1A/COMPASS and shadow enhancers in the regulation of homeotic gene expression. Genes Dev. 2017;31:787–801. https://doi.org/10.1101/gad.294744.116.
Deshpande AJ, et al. AF10 regulates progressive H3K79 methylation and HOX gene expression in diverse AML subtypes. Cancer Cell. 2014;26:896–908. https://doi.org/10.1016/j.ccell.2014.10.009.
Yokoyama A, et al. Leukemia proto-oncoprotein MLL forms a SET1-like histone methyltransferase complex with menin to regulate Hox gene expression. Mol Cell Biol. 2004;24:5639–49. https://doi.org/10.1128/MCB.24.13.5639-5649.2004.
Schuettengruber B, Chourrout D, Vervoort M, Leblanc B, Cavalli G. Genome regulation by polycomb and trithorax proteins. Cell. 2007;128:735–45. https://doi.org/10.1016/j.cell.2007.02.009.
Kuhn MW, et al. Targeting chromatin regulators inhibits leukemogenic gene expression in NPM1 mutant leukemia. Cancer Discov. 2016;6:1166–81. https://doi.org/10.1158/2159-8290.CD-16-0237.
Gu X, et al. Leukemogenic nucleophosmin mutation disrupts the transcription factor hub that regulates granulomonocytic fates. J Clin Invest. 2018;128:4260–79. https://doi.org/10.1172/JCI97117.
Wang A, Han Y, Chen P, Jia N, Minden MD. AACR annual meeting 2018 abstract #2991. Chicago, IL: American Association for Cancer Research; 2018.
Luo H, et al. CTCF boundary remodels chromatin domain and drives aberrant HOX gene transcription in acute myeloid leukemia. Blood. 2018;132:837–48. https://doi.org/10.1182/blood-2017-11-814319.
Huang Y, et al. Identification and characterization of Hoxa9 binding sites in hematopoietic cells. Blood. 2012;119:388–98. https://doi.org/10.1182/blood-2011-03-341081.
Sun Y, et al. HOXA9 reprograms the enhancer landscape to promote leukemogenesis. Cancer Cell. 2018;34:643–58. https://doi.org/10.1016/j.ccell.2018.08.018 (e645).
Ivey A, et al. Assessment of minimal residual disease in standard-risk AML. N Engl J Med. 2016. https://doi.org/10.1056/NEJMoa1507471.
Shlush LI, et al. Identification of pre-leukaemic haematopoietic stem cells in acute leukaemia. Nature. 2014;506:328–33. https://doi.org/10.1038/nature13038.
Krönke J, et al. Clonal evolution in relapsed NPM1-mutated acute myeloid leukemia. Blood. 2013;122:100–8. https://doi.org/10.1182/blood-2013-01-479188.
Gravina GL, et al. Nucleo-cytoplasmic transport as a therapeutic target of cancer. J Hematol Oncol. 2014;7:85. https://doi.org/10.1186/s13045-014-0085-1.
Gounder MM, et al. Phase IB study of selinexor, a first-in-class inhibitor of nuclear export, in patients with advanced refractory bone or soft tissue sarcoma. J Clin Oncol. 2016;34:3166–74. https://doi.org/10.1200/JCO.2016.67.6346.
Abdul Razak AR, et al. First-in-class, first-in-human phase I study of selinexor, a selective inhibitor of nuclear export, in patients with advanced solid tumors. J Clin Oncol. 2016;34:4142–50. https://doi.org/10.1200/JCO.2015.65.3949.
Chen C, et al. Safety and efficacy of selinexor in relapsed or refractory multiple myeloma and Waldenstrom macroglobulinemia. Blood. 2018;131:855–63. https://doi.org/10.1182/blood-2017-08-797886.
Alexander TB, et al. Phase I study of selinexor, a selective inhibitor of nuclear export, in combination with fludarabine and cytarabine, in pediatric relapsed or refractory acute leukemia. J Clin Oncol. 2016;34:4094–101. https://doi.org/10.1200/JCO.2016.67.5066.
Garzon R, et al. A phase 1 clinical trial of single-agent selinexor in acute myeloid leukemia. Blood. 2017;129:3165–74. https://doi.org/10.1182/blood-2016-11-750158.
Etchin J, et al. KPT-8602, a second-generation inhibitor of XPO1-mediated nuclear export, is well tolerated and highly active against AML blasts and leukemia-initiating cells. Leukemia. 2017;31:143–50. https://doi.org/10.1038/leu.2016.145.
Hing ZA, et al. Next-generation XPO1 inhibitor shows improved efficacy and in vivo tolerability in hematological malignancies. Leukemia. 2016;30:2364–72. https://doi.org/10.1038/leu.2016.136.
Cancer Genome Atlas Research, N. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med. 2013;368:2059–74. https://doi.org/10.1056/NEJMoa1301689.
Alexandrov LB, et al. Signatures of mutational processes in human cancer. Nature. 2013;500:415–21. https://doi.org/10.1038/nature12477.
Li J, Zhang X, Sejas DP, Bagby GC, Pang Q. Hypoxia-induced nucleophosmin protects cell death through inhibition of p53. J Biol Chem. 2004;279:41275–9. https://doi.org/10.1074/jbc.C400297200.
Liu WH, Yung BY. Mortalization of human promyelocytic leukemia HL-60 cells to be more susceptible to sodium butyrate-induced apoptosis and inhibition of telomerase activity by down-regulation of nucleophosmin/B23. Oncogene. 1998;17:3055–64. https://doi.org/10.1038/sj.onc.1202234.
Gao H, et al. B23 regulates GADD45a nuclear translocation and contributes to GADD45a-induced cell cycle G2-M arrest. J Biol Chem. 2005;280:10988–96. https://doi.org/10.1074/jbc.M412720200.
Okuda M, et al. Nucleophosmin/B23 is a target of CDK2/cyclin E in centrosome duplication. Cell. 2000;103:127–40.
Federici L, et al. Nucleophosmin C-terminal leukemia-associated domain interacts with G-rich quadruplex forming DNA. J Biol Chem. 2010;285:37138–49. https://doi.org/10.1074/jbc.M110.166736.
Arcovito A, et al. Synergic role of nucleophosmin three-helix bundle and a flanking unstructured tail in the interaction with G-quadruplex DNA. J Biol Chem. 2014;289:21230–41. https://doi.org/10.1074/jbc.M114.565010.
Scognamiglio PL, et al. G-quadruplex DNA recognition by nucleophosmin: new insights from protein dissection. Biochim Biophys Acta. 2014;1840:2050–9. https://doi.org/10.1016/j.bbagen.2014.02.017.
Scott DD, Oeffinger M. Nucleolin and nucleophosmin: nucleolar proteins with multiple functions in DNA repair. Biochem Cell Biol. 2016;94:419–32. https://doi.org/10.1139/bcb-2016-0068.
Poletto M, Lirussi L, Wilson DM, 3rd & Tell G. Nucleophosmin modulates stability, activity, and nucleolar accumulation of base excision repair proteins. Mol Biol Cell 25, 1641–52, https://doi.org/10.1091/mbc.E13-12-0717 (2014).
Ziv O, et al. Identification of novel DNA-damage tolerance genes reveals regulation of translesion DNA synthesis by nucleophosmin. Nat Commun. 2014;5:5437. https://doi.org/10.1038/ncomms6437.
Okuwaki M. The structure and functions of NPM1/Nucleophosmin/B23, a multifunctional nucleolar acidic protein. J Biochem. 2008;143:441–8. https://doi.org/10.1093/jb/mvm222.
Ahn JY, et al. Nucleophosmin/B23, a nuclear PI(3,4,5)P(3) receptor, mediates the antiapoptotic actions of NGF by inhibiting CAD. Mol Cell. 2005;18:435–45. https://doi.org/10.1016/j.molcel.2005.04.010.
Bolli N, et al. A dose-dependent tug of war involving the NPM1 leukaemic mutant, nucleophosmin, and ARF. Leukemia. 2009;23:501–9. https://doi.org/10.1038/leu.2008.326.
Dumbar TS, Gentry GA, Olson MO. Interaction of nucleolar phosphoprotein B23 with nucleic acids. Biochemistry. 1989;28:9495–501.
Sipos K, Olson MO. Nucleolin promotes secondary structure in ribosomal RNA. Biochem Biophys Res Commun. 1991;177:673–8.
Amin MA, Matsunaga S, Uchiyama S, Fukui K. Depletion of nucleophosmin leads to distortion of nucleolar and nuclear structures in HeLa cells. Biochem J. 2008;415:345–51. https://doi.org/10.1042/BJ20081411.
Holmberg Olausson K, Elsir T, Moazemi Goudarzi K, Nister M, Lindstrom MS. NPM1 histone chaperone is upregulated in glioblastoma to promote cell survival and maintain nucleolar shape. Sci Rep. 2015;5:16495. https://doi.org/10.1038/srep16495.
Lin J, Kato M, Nagata K, Okuwaki M. Efficient DNA binding of NF-kappaB requires the chaperone-like function of NPM1. Nucleic Acids Res. 2017;45:3707–23. https://doi.org/10.1093/nar/gkw1285.
Liu CD, et al. The nuclear chaperone nucleophosmin escorts an Epstein-Barr Virus nuclear antigen to establish transcriptional cascades for latent infection in human B cells. PLoS Pathog. 2012;8:e1003084. https://doi.org/10.1371/journal.ppat.1003084.
Destouches D, et al. Implication of NPM1 phosphorylation and preclinical evaluation of the nucleoprotein antagonist N6L in prostate cancer. Oncotarget. 2016;7:69397–411. https://doi.org/10.18632/oncotarget.8043.
Acknowledgements
This work has been supported by the NIH (DK092883, CA183252) and the Samuel Waxman Cancer Research Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
About this article
Cite this article
Brunetti, L., Gundry, M.C. & Goodell, M.A. New insights into the biology of acute myeloid leukemia with mutated NPM1. Int J Hematol 110, 150–160 (2019). https://doi.org/10.1007/s12185-018-02578-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12185-018-02578-7