Abstract
Multiple myeloma cells acquire the resistance to anti-cancer drugs through physical and functional interactions with the bone marrow microenvironment via two overlapping mechanisms. First, bone marrow stromal cells (BMSCs) produce soluble factors, such as interleukin-6 and insulin-like growth factor-1, to activate signal transduction pathways leading to drug resistance (soluble factor-mediated drug resistance). Second, BMSCs up-regulate the expression of cell cycle inhibitors, anti-apoptotic members of the Bcl-2 family and ABC drug transporters in myeloma cells upon direct adhesion [cell adhesion-mediated drug resistance (CAM-DR)]. Elucidation of the mechanisms underlying drug resistance may greatly contribute to the advancement of cancer therapies. Recent investigations, including ours, have revealed the involvement of epigenetic alterations in drug resistance especially CAM-DR. For example, we found that class I histone deacetylases (HDACs) determine the sensitivity of proteasome inhibitors and the histone methyltransferase EZH2 regulates the transcription of anti-apoptotic genes during the acquisition of CAM-DR by myeloma cells. In addition, another histone methyltransferase MMSET was shown to confer drug resistance to myeloma cells by facilitating DNA repair. These findings provide a rationale for the inclusion of epigenetic drugs, such as HDAC inhibitors and histone methylation modifiers, in combination chemotherapy for MM patients to increase the therapeutic index.
Similar content being viewed by others
Introduction
Multiple myeloma (MM) is the second most common hematologic malignancy and one of the leading causes of death in the field of hematology. Its average incidence is two in 100,000 people in Japan, but is constantly increasing with age; the incidence rises to 20 in 100,000 people in their seventies. Unrelated to a striking increase of the elderly population, the age-adjusted incidence of MM increased from 0.92 to 5.2 and from 0.81 to 4.8 per 100,000 men and women, respectively, during 1975–2010 in Japan [1]. The introduction of high-dose chemotherapy with stem cell support and novel therapeutic agents, including proteasome inhibitors and immunomodulatory drugs, has significantly improved the treatment outcome of MM patients [2–4]. According to the recent data published by the Japanese Society of Myeloma, the median survival and 5-year overall survival were 38.9 months and 31.2 %, respectively, in 1208 patients mainly treated with a combination of alkylating agents and corticosteroids during 1990–2000, whereas these values improved to 60.0 months and 50.3 %, respectively, in 2234 patients who received novel treatments during 2001–2012 [5]. However, the prognosis of MM is still worse than that of other hematologic malignancies, such as acute myeloid leukemia and diffuse large B cell lymphoma. Therefore, novel strategies for prevention, early detection and better treatment are in high demand, which should be based on the molecular pathophysiology of the disease.
Recent extensive investigations with advanced technologies, such as high-density array-based gene expression profiling and next-generation sequencing (NGS), have a great contribution to the identification of genetic abnormalities in MM [6–11]. In brief, normal plasma cells are transformed to myeloma cells by the deregulation of Cyclin D family proteins due to recurrent chromosomal translocations involving the immunoglobulin heavy chain (IgH) locus at 14q32, such as t(11;14)(q13;q32) and t(6;14)(p21;q32), and hyperdiploidy including trisomy 11, which increases the dosage of the cyclin D1 gene at 11q13, at the initial step of myelomagenesis [12]. Further progression of the disease is mediated via point mutations of Ras-family genes and overexpression of the c-Myc oncogene. The terminal stage of myeloma progression is characterized by stroma-independent growth, which results in extramedullary diseases and plasma cell leukemia. Genetic abnormalities associated with this stage include mutational inactivation of TP53 [13], the protocadherin family [14], and inhibitory components of the NF-κB pathway [15]. In addition, NGS analyses uncovered non-synonymous point mutations of epigenetic regulators such as KDM6A/UTX (10 %), KDM6B/JMJD3, MMSET (8 %), MLL (1 %) and HOXA9 [6, 10]. These achievements have been gradually translated to the clinic, which is best exemplified by the development of molecular targeting agents such as histone deacetylase (HDAC) inhibitors and BRAF inhibitors [16, 17]. In this review, we summarize the recent advances in basic and translational research on the mechanisms underlying drug resistance of MM cells with special attention to epigenetic aspects.
Drug resistance of multiple myeloma cells via interaction with the bone marrow microenvironment
Multiple myeloma is caused by neoplastic transformation of terminally differentiated B lymphocytes, called plasma cells, with capability of producing immunoglobulins [18, 19]. Like their normal counterpart, MM cells mainly proliferate and survive within a bone marrow milieu via physical and functional interactions with bone marrow stromal cells (BMSCs) and surrounding microenvironment [20, 21]. The interaction with BMSCs also confers anti-cancer drug resistance to myeloma cells via two overlapping mechanisms. First, BMSCs produce soluble factors, such as interleukin-6 (IL-6), to activate signal transduction pathways leading to drug resistance in myeloma cells [22]. Second, BMSCs up-regulate the expression of cell cycle inhibitors (CDK inhibitors p21Cip1/Waf1 and p27Kip1), anti-apoptotic members of the Bcl-2 family (Bcl-2 and Mcl-1) and ABC-family drug transporters (ABCB1/P-glycoprotein, ABCC3 and NEK2) in myeloma cells upon direct contact through adhesion molecules and/or extracellular matrix (ECM) proteins that transduce signals to modify the gene expression program favoring cell survival [23, 24]. These forms of drug resistance are termed soluble factor-mediated drug resistance (SFM-DR) and cell adhesion-mediated drug resistance (CAM-DR), respectively. In addition to the innate or de novo resistance conferred by the bone marrow microenvironment, long-term exposure to anti-cancer drugs results in acquired resistance due to the enhanced ability of DNA damage repair and drug metabolism in MM cells (Fig. 1). The innate/de novo and acquired resistance are causally related each other to make MM one of the most reluctant malignancies [25].
Mechanisms of drug resistance in multiple myeloma. In general, the resistance to anti-cancer drugs is classified into two forms: de novo resistance and acquired resistance. In multiple myeloma, the former is further divided into soluble factor-mediated drug resistance (SFM-DR), which is caused by cytokines such as interleukin-6 (IL-6) and insulin-like growth factor-1 (IGF-1), and cell adhesion-mediated drug resistance (CAM-DR), which is mediated via direct contact with bone marrow stromal cells and ECM proteins. Effectors of SFM- and CAM-DR include CDK inhibitors (p21Cip1/Waf1, p27Kip1, p57Kip2) and anti-apoptotic members of the Bcl-2 family (Bcl-x, Bcl-2). Long-term exposure to anti-cancer drugs results in acquired resistance due to the enhanced ability of DNA damage repair and drug metabolism
Following the adhesion of MM cells to BMSCs or fibronectin, IL-6 is secreted by BMSCs and/or by MM cells in an autocrine manner [26]. IL-6 not only contributes to the growth of MM cells, but also implicated in the resistance to various apoptotic insults including Fas ligand and chemotherapeutic agents [27]. These responses are mainly controlled through the JAK/STAT signaling pathway [28]. For example, it has been reported that primary MM cells with the capability of autocrine IL-6 production are more resistant to dexamethasone than those without autocrine IL-6 signal [29]. A blocking antibody against IL-6 receptors increases the sensitivity of MM cells to bortezomib, which highlights the particular importance of IL-6 in SFM-DR [30]. Obviously, SFM-DR is caused not only by IL-6 but also by insulin-like growth factor-1 (IGF-1) [31, 32], stroma-derived factor-1 (SDF-1), hepatocyte growth factor (HGF) [33], vascular endothelial growth factor (VEGF) and epithelial growth factor (EGF) [34]. The IGF-1/IGF-1 receptor signaling pathway is constitutively activated in bortezomib-resistant MM cell lines and serum IGF-1 concentration is significantly higher in MM patients refractory to bortezomib than in those who responded to the drug [32]. Blocking of IGF-1/IGF-1 receptor signaling could induce apoptotic cell death in MM, which is enhanced by the combination with bortezomib [31]. BMSCs were also shown to induce bortezomib-resistant NF-κB activity in MM cells by producing soluble factors including IL-6, IL-8, VEGF and tumor necrosis factor-β (TNF-β) from BMSCs [35].
It is widely accepted that CAM-DR to conventional anti-MM agents, such as melphalan, doxorubicin and vincristine, is mainly mediated through VLA-4, a heterodimer of CD49d/α4 and CD29/β1 integrins, on MM cells [36, 37]. Other integrin molecules, including β7 integrin and VLA-5 (α5β1), may also play a role in CAM-DR in MM cells [38, 39]. The ITGB7 gene encoding β7-integrin is a transcriptional target of MAF family proteins, whose expression is detected in up to 50 % of MM patients and is particularly high in those carrying t(14;16), t(14;20) and t(8;14) due to IgH enhancer-driven transactivation [40]. Primary MM cells with a higher expression of these adhesion molecules are generally more resistant to therapeutics and tend to be selected during treatment, tipping the contribution of CAM-DR to acquired drug resistance [41]. A comparison of the gene expression profiles of isogenic cell lines that acquired melphalan resistance via adhesion to fibronectin and via long-term exposure to melphalan revealed that CAM-DR is mostly achieved through post-transcriptional mechanisms and the acquired drug resistance is associated with a specific transcriptome change [42]. VLA-4-mediated CAM-DR is characterized by G1 cell cycle arrest accompanied by an increase in the expression levels of CDK inhibitors p21Cip1/Waf1 and p27Kip1, a decrease in Cyclin A and Cyclin E activity, and down-regulation of Bim, a proapoptotic member of the Bcl-2 family, all of which is accomplished through the modulation of ubiquitin-dependent degradation. Furthermore, VLA-4-mediated adhesion also induces resistance to apoptotic signals such as Fas ligand in various hematologic malignancies including MM via post-transcriptional mechanisms. For example, an increase in solubility and redistribution of c-FLIP inhibit DISC (death-inducing signaling complex) to block Fas-induced apoptosis [43].
It is obvious that SFM-DR and CAM-DR are not completely separate pathways but rather are closely related to each other. In line with this notion, the synergistic anti-apoptotic effects of VLA-4 and IL-6 signaling was shown to converge on the STAT3 activation pathway in MM cells [28]. Similarly, VLA-4 activation synergistically promotes drug resistance with HGF through the PI3K/Akt and NF-κB signaling pathways in MM cells [33].
Innate drug resistance of cancer stem cell in multiple myeloma
An emerging concept of cancer stem cell (CSC) proposes that a small population of tumor cells are particularly resistant to chemotherapy due to their ability to be quiescent and efflux anti-cancer agents via drug transporters, which hamper the effects of cytotoxic drugs typically targeting rapidly growing cells, and ultimately cause relapse via continuous self-renewal, asymmetric division, and differentiation into mature and growing cancer cells [44]. Cancer stem cells also possess epithelial–mesenchymal transition (EMT)-like phenotypes, which underlie local invasion, extramedullary involvement, distant metastasis, and drug resistance. Several lines of evidence suggest that this is also the case with MM [45, 46].
The CD138 antigen is a transmembrane (type I) heparan sulfate proteoglycan, called syndecan-1, and is expressed almost exclusively on mature plasma cells and their malignant counterpart but not on mature B lymphocytes including post-germinal center and memory B cells [47]. Surface CD138 expression is detected in nearly 100 % of MM cells and is used as a marker for diagnosis and isolation for research purposes. Cancer stem cells of MM are believed to reside in a CD138-negative population, which expresses phenotypic markers of memory B cells (CD138−/CD19+/CD20+/CD27+) and possess stemness features including drug resistance [48]. A novel 3D-model also reveals the expression of CD20 on drug-resistant MM cells [49]. CD19-positive clonotypic B cells are detectable in the peripheral blood and bone marrow of MM patients [50, 51]. On the contrary, Hosen et al. [52] argued that MM-initiating cells are enriched in the CD138−/CD19−/CD38++ component, whereas CD19-positive fractions fail to engraft in SCID mice. CD138-negative myeloma stem cells are enriched in a side population (SP) and resistant to dexamethasone, lanalidomide, bortezomib and carfilzomib due to the enhanced activity of aldehyde dehydrogenase 1 and drug efflux pumps (ABCB1/P-glycoprotein and ABCC3) [53–55]. A seminal report by Leung-Hagesteijn et al. [56] identified that CD138−/CD38+ subsets show innate resistance to bortezomib because they lack XBP-1, a transcription factor essential for plasma cell development as well as an unfolded protein response and subsequent endoplasmic reticulum (ER) stress-induced apoptosis.
Brief overview of epigenetics
Chromatin is a highly ordered structure composed of DNA, histones and other chromosomal proteins. Histones are proteins found in all eukaryotic cells and form a unit known as the nucleosome, which consists of a histone octamer (H2A, H2B, H3 and H4, two pairs of each), around which 147 bp of DNA are wrapped, resulting in a tight DNA packaging. The nucleosome complex forms the chromatin, which is further condensed to chromosomes during mitosis. Changes in the chromatin structure enable the rapid and reversible regulation of gene expression and subsequent biological events, including drug responses, without a heritable genetic alteration. This process, called epigenetic regulation, is accomplished through post-translational modifications of histone tails and DNA methylation [57]. It is now known that the positioning of nucleosomes offers another layer of epigenetic regulation of gene expression and genomic stability [58]. Due to space limitation, we do not refer to the role of DNA methylation for epigenetic regulation in MM, which is a subject of other excellent review articles [59, 60].
N-terminal histone tails are rich in lysine, arginine, serine and threonine, which serve as substrates for post-translational modifications such as methylation, acetylation, phosphorylation, ubiquitination, sumoylation and deamination. These modifications affect the interaction of DNA with histones and other DNA-binding protein complexes, thus playing a pivotal role in the epigenetic regulation of several biological events [61]. Among the four core histones, histone H3 is most important for chromatin organization; the methylation of lysine on position 4 of histone H3 (H3K4) and H3K36 is generally associated with open chromatin structures, whereas the methylation of H3K27 and H3K9 represents a hallmark of condensed chromatin at silent loci (see below). Alterations in these modifications entail various pathological conditions; however, the epigenetic mechanisms underlying drug resistance have not been fully elucidated. Yet, some studies have suggested the involvement of an altered chromatin state by H3K4 demethylation in anti-cancer drug tolerance [62, 63].
Roles of histone deacetylases in drug resistance of multiple myeloma
Chromatin organization is spatiotemporally regulated by the acetylation status of histone tails, which is determined by the dynamic balance between two counteracting enzymes: histone acetyltransferases (HATs) and histone deacetylases (HDACs). HATs are composed of more than 10 enzymes that belong to at least four structurally distinct families (p300/CGP, GNAT, MYST and others) and transfer an acetyl group from acetyl-CoA to the lysine residues of histones, resulting in neutralization of their positive charge and dissociation from the negatively charged phosphate groups of DNA [64]. As a result, the chromatin structure becomes less condensed to allow the binding of transcription factors and auxiliary activators such as BET (bromodomain and extra-terminal) family proteins BRD2, BRD3, BRD4 and BRDT. BRD4 and BRD2 directly bind to ε-N-acetylated lysine residues of core histones at promoter regions and facilitate transcription and elongation of downstream genes by recruiting Mediator and P-TEFb complexes [65]. Overall, histone acetylation is a histone mark associated with active gene expression, which is best exemplified by p300-mediated acetylation of H3K27 at enhancer regions [66]. In general, HATs tend to be inactivated in cancer cells. In MM, overexpression of the miR-106b-25 cluster down-regulates the expression of PCAF (p300/CBP-associated factor), a histone acetyltransferase involved in transcriptional control of TP53 [67]. HDACs mediate the reverse process by removing acetyl groups from histone lysine residues back to acetyl-CoA, resulting in a condensed chromatin structure and transcriptional inactivation [68]. Histone acetylation-mediated chromatin dynamics not only affect transcription but also modify the sensitivity to anti-cancer drugs, because DNA damaging agents like alkylators and topoisomerase inhibitors act less efficiently on DNA within condensed chromatin.
Histone deacetylases (HDACs) regulate a plethora of biological phenomenon via deacetylation of histone and non-histone proteins. HDACs, 11 HDACs and 7 SIRT, are divided into four classes (I, IIa, IIb, III and IV) based on structural and functional differences (Fig. 2). Class I HDACs (HDAC1, 2, 3 and 8) serve as universal regulators of cell growth and death via transcriptional repression of negative cell cycle regulators including p21Cip1/Waf1 [69] and deacetylation of the pro-apoptotic protein p53 [70]. Class IIa HDACs (HDAC4, 5, 7 and 9) directly act on tissue-specific transcription factors, such as MEF2, Runx2 and FoxP3, to govern the development and functions of the musculoskeletal and nervous systems [71] as well as specific cell types such as regulatory T-lymphocytes [72]. HDAC6, composes class IIb HDACs with HDAC10, is involved in the clearance of misfolded/unfolded proteins by facilitating aggresome formation, transport of aggregated proteins to microtubule organizing centers, and autophagolysosome formation (autophagy) via deacetylation of tubulin and chaperone proteins [73]. Recent investigations revealed that HDAC1, 2, 3 and 6 are highly expressed in myeloma cells and contribute to their malignant behavior [74], suggesting that pan-HDAC inhibitors, such as panobinostat and vorinostat, are effective for MM compared with class I-specific HDAC inhibitors such as romidepsin and valproic acid. It is anticipated that the perturbation of HDAC1 and HDAC3 activity causes growth arrest and cell death in myeloma cells. In addition, HDAC6 inhibition may enhance the cytotoxic effect of proteasome inhibitors by increasing protein overload in myeloma cells, leading to ER stress-mediated apoptosis [75]. These preclinical assumptions were substantiated by recent randomized clinical trials called PANORAMA studies, in which the combination of panobinostat, bortezomib and dexamethasone was more effective for relapsed and/or refractory myeloma patients than the combination of placebo, bortezomib and dexamethasone in terms of overall response and progression-free survival [76, 77]. Because class I HDACs are essential for the maintenance of normal proliferating cells, hematological and gastrointestinal toxicities are inevitably associated with HDAC inhibitor-containing regimens. Therefore, toxicity management holds a key to the success of MM treatment with HDAC inhibitors. Nonetheless, HDAC inhibitors are promising options for improving the treatment outcome in not only relapsed and/or refractory cases but also in newly diagnosed patients in the near future.
The structure and function of histone deacetylases. Class I HDACs (HDAC1, 2, 3 and 8) are usually localized in the nucleus and deacetylate histones to negatively regulate transcription. Class IIa HDACs (HDAC4, 5, 7 and 9) shuttle between the cytoplasm and nucleus, and bind to tissue-specific transcription factors, such as MEF2, Runx2 and FoxP3, via long N-termini to modify their function and localization. HDAC6, composes class IIb HDACs with HDAC10, essentially resides in the cytoplasm and is involved in autophagy via deacetylation of tubulin and chaperone proteins. The catalytic domains of each HDAC are indicated in red. The sizes (amino acids) and chromosomal locations are shown on the right in each column
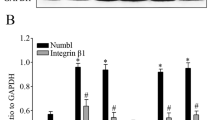
Given the feasibility of therapeutic application, we have focused on the epigenetic aspect of CAM-DR in MM. We have found that proteasome inhibitors are able to overcome CAM-DR by disrupting VLA-4-mediated interaction of MM cells with BMSCs [36]. Proteasome inhibitor-induced down-regulation of VLA-4 is not NF-κB-dependent but rather occurs via epigenetic mechanisms because genetic and pharmacologic inhibition of HDACs reproduced this process [78]. Indeed, proteasome inhibitors repress the transcription of class I HDAC genes through caspase-8-mediated cleavage of Sp1 transcriptional activator [79, 80]. Furthermore, class I HDACs (HDAC1, 2, 3) determine the sensitivity of MM cells to proteasome inhibitors and other anti-MM agents. HDAC1 is strongly expressed in CD138-negative fractions of MM (Fig. 3) and represses the transcription of Xbp1 and CHOP, which may underlie the innate resistance of myeloma stem cells to ER stress-induced apoptosis elicited by proteasome inhibitors [56, 81]. Therefore, HDAC inhibitors could sensitize myeloma stem cells to proteasome inhibitors by restoring the expression of Xbp1 and CHOP [82]. This theory at least partly explains the synergistic effects of bortezomib with panobinostat in the PANORAMA trials [76, 77] as well as the effectiveness of retreatment of MM patients who achieved remission with bortezomib and relapsed more than 6 months after the initial treatment [83].
Roles of histone methyltransferases in drug resistance of multiple myeloma
Histone methylation provides more complex epigenetic codes than acetylation-mediated on–off type determination. Histone methylation exists as the mono-, di- or tri-methylation of lysine residues and as the mono- and di-methylation of arginine residues on each core histone. The position and depth of methylation (mono-, di- or tri-) individually define the biological output of each mark (Table 1). In sum, transcriptional activation is associated with H3K4me1 at enhancer regions, H3K4me2/me3 at promoter regions as well as H3K36me3, H3K79me1/me2/me3, H4R3me1 and H4K20me1 at gene bodies, whereas gene silencing is marked with H3K9me2/me3 at enhancer regions and H3K27me3 at promoter regions [84]. In addition, H3K9me2/me3 and H4K20me3 are responsible for the repression of transcription from repetitive and transposable elements, thereby contributing to genomic stability. The position- and methylation depth-dependent difference in functional outcome is at least partly explained by reader proteins bind to each modification. For example, ATP-dependent DNA helicase CHD1 binds to di- and tri-methylated H3K4 via its chromodomain and remodels the chromatin to an active configuration. For transcriptional repression, heterochromatin protein 1 (HP1) binds to H3K9me3 via its chromodomain to constitute deeply silenced heterochromatic regions. Similarly, the reader protein CBX binds to H3K27me3 and recruits other components of Polycomb repressive complex 1 (PRC1) and subsequently PRC2 for gene silencing [59, 85].
Mammalian cells possess more than 30 histone methyltransferases also called as lysine methyltransferases (KMTs) that belong to 8 families composed of the Su(var)3-9, Enhance of zest and Trithorax (SET) family and KMT4/DOT1L, which lacks the SET domain and exclusively methylates H3K79 [86]. Each KMT catalyzes the methylation of different positions at three depths, namely 1, 2 or 3 methyl groups at each residue (Table 1). Recent data support the theory that individual histone changes are not sufficient to influence gene expression, but their combinatorial patterns are more important. The initial investigation revealed the presence of 51 distinct chromatin states to govern the epigenetic modulation of gene expression across the human genome [87]. Another study has defined over 800 different combinatorial patterns of histone changes, confirming the joint effect of histone modifications [88]. Moreover, the landscape of histone modifications is dynamic and constantly changes with the functional interactions of specific histone modifications, such as the requirement of H2BK123 ubiquitination for subsequent H3K4me3 and H3K79me2 [89].
Abnormal patterns of histone modifications are commonly observed in hematological malignancies. Mutations of histone-modifying enzymes and other epigenetic regulators have been reported in numerous cancers including MM [90]. An initial study using NGS has discovered recurrent non-synonymous mutations in histone-modifying enzymes KDM6A/UTX, KMT2A/MLL, KMT2B/MLL2, KMT2C/MLL3 and KMT3G/MMSET, which commonly cause an increased expression of HoxA9, proposing a potential therapeutic target in MM [6]. In addition to aberrant expression of HoxA9, deregulated MMSET, which stands for Multiple Myeloma SET domain, drastically alters the gene expression program, which may be responsible for malignant behavior of MM harboring t(4;14). MMSET is overexpressed in an aggressive form of MM carrying t(4;14), which is deteced in 10–15 % of all MM patients [12]. The MMSET/WHSC1/NSD2 gene is ubiquitously expressed in all tissues and generates MMSET-1, MMSET-2 and other minor transcripts [91]. The longest transcript MMSET-2 is composed of four domains: PWWP (proline-tryptophan-tryptophan-proline) domain, a HMG (high mobility group) box, a PHD (plant-homeodomain) zinc finger and a SET domain [92]. MMSET mainly catalyzes H3K36 dimethylation and H4K20 trimethylation, which mediate transcriptional activation of oncogenic loci and repair of DNA double-strand breaks, respectively [93, 94]. Drug resistance of MM cases carrying t(4;14) is at least partly explained by H4K20me3-mediated recruitment of 53BP1 and enhanced repair of DNA damage induced by anti-cancer agents [94]. In addition, the accumulation of H3K36me2 marks causes global reduction and focal accumulation of H3K27me3 via redistribution of KMT6A/EZH2, resulting in altered gene expression program favoring cell proliferation and survival [95, 96]. Increased expression of MMSET underlies the constitutive activation of NF-κB, a hallmark of MM in advanced stages [97]. MMSET knockdown leads to a decrease in cell proliferation, activation of caspase-3 and caspase-9, and reduced expression of adhesion molecules (DSG2 and ADAM9) in MM cells harboring t(4;14) [98]. These findings make MMSET amenable to therapeutic interventions.
A reduction in H3K27me3 is also caused by misregulation and/or mutations in the H3K27 methyltransferase KMT6A/EZH2 and H3K27 demethylases KDM6A/UTX and KDM6B/JMJD3 in numerous cancers, which results in altered chromatin states linked to malignant transformation [99]. Furthermore, somatic mutations in H3K27 have been observed in aggressive gliomas [100, 101]. These findings indicate that H3K27 methylation is profoundly implicated in oncogenesis; however, relatively little is known about its specific role in MM biology including CAM-DR. Recently, we identified H3K27me3 as a critical histone modification for CAM-DR in MM [102]. Cell adhesion counteracts anti-cancer drug-induced hypermethylation of H3K27 via Akt-mediated inactivating phosphorylation of EZH2, which sustains the expression of anti-apoptotic genes, such as IGF1, BCL2 and HIF1A, to promote drug resistance in MM cells. Inhibition of the IGF-1R/PI3 K/Akt pathway was able to reverse CAM-DR by promoting EZH2 dephosphorylation and H3K27 hypermethylation both in vitro and in refractory murine myeloma models. This is the first report of the epigenetic mechanism for CAM-DR involving a regulatory circuit from the membrane to the nucleus (Fig. 4). In accordance with this model, small compounds counteracting EZH2 phosphorylation, such as PI3K/Akt, CDK and IGF-1R inhibitors, were found to be effective for the reversal of CAM-DR in vitro and in vivo. Of these, the IGF-1R inhibitor OSI-906 appears to be particularly effective in overcoming CAM-DR, which may be translated to the clinic to improve the treatment outcome of myeloma patients in combination with conventional anti-MM agents. In support of this view, the potent and selective cytotoxicity of IGR-1R inhibitors against MM cells was clearly demonstrated by a recent systematic pharmacological screen using 119 cell lines from >10 different types of hematologic malignancies [103]. As anticipated, PI3K and Akt inhibitors were also effective for MM cells albeit with lower selectivity than IGF-1R inhibitors [102, 103].
EZH2 is frequently overexpressed and dysregulated in multiple types of cancer. The expression level of EZH2 is correlated with the higher proliferation rates and aggressive behavior of cancer cells as well as the poor prognosis of patients with breast, prostate and bladder cancers [104]. It has been reported that EZH2 up-regulation is mediated by IL-6, through c-Myc activation, or is a consequence of EZH2-targeting miR-26a down-regulation in MM [105]. Together with the aforementioned inactivating mutations of H3K27 demethylases (UTX and JMJD3), all of these mechanisms can explain the increased levels of the H3K27me3 silencing signature at PRC2-targeted genes in MM cells [106, 107]. On the contrary, EZH2 functions as a tumor suppressor in some types of cancer. Inactivating mutations of EZH2 were found in patients with myeloid malignancies including myelodysplastic syndrome and myeloproliferative neoplasms, and have been associated with poor patient survival [108, 109]. In line with this observation, mice with conditional deletions of EZH2 and TET2 in hematopoietic stem cells develop myelodysplastic syndrome and myeloproliferative neoplasms [110, 111]. Loss-of-function mutations and deletions of the EZH2 and SUZ12 genes have been reported in 25 % of T-cell acute lymphoblastic leukemia [112]. Consistent with this finding, the conditional deletion of EZH2 in bone marrow cells causes T-cell leukemia [113]. These results suggest that the oncogenic role of EZH2 is cell context-dependent and differs between solid tumors and hematological malignancies. EZH2 is only rarely mutated or deleted in MM [6]; therefore, it can integrate environmental cues to biological output enhancing malignant phenotypes in this disease. From a mechanistic standpoint, it is interesting that EZH2 methylates cellular proteins other than histone H3 and modulates their functions. For example, Kim et al. [114] showed that EZH2 methylated STAT3 and up-regulated its transcriptional activity, contributing to the maintenance of stem cell phenotypes and tumor progression of glioblastoma and possibly in MM [28, 34]. Taken together, EZH2 is central to tumor biology, from the initiation of cancer to drug resistance, and thus is a major therapeutic target in various cancers. Overall, these findings provide a rationale for the inclusion of epigenetic drugs, such as HDAC inhibitors and KMT/KDM inhibitors, in combination chemotherapy to increase the therapeutic index in MM (see below).
Therapeutic Implications
As mentioned above, panobinostat, a pan-HDAC inhibitor, has already been approved for the treatment of relapsed and refractory MM in combination with bortezomib and dexamethasone [76, 77]. It is noteworthy that the combination of panobinostat and bortezomib is effective for bortezomib-resistant cases, as anticipated from the results of in vitro experiments. This “one-drug addition” effect is well explained by preclinical evidence that HDAC6 inhibition enhances bortezomib-induced ER stress leading to proteotoxic cell death [81] and HDAC1 inhibition could overcome the innate resistance of myeloma stem cells to bortezomib by restoring the expression of Xbp1 and CHOP, both of which are indispensable for ER stress-induced apoptosis [82]. In this context, another interesting target is HDAC6, a class IIb cytoplasmic HDAC that plays a central role in the degradation of misfolded proteins through autophagy [73]. A selective HDAC6 inhibitor, ACY-1215, is expected to be more effective for MM in combination with proteasome inhibitors and less toxic than pan-HDAC inhibitors [115]. The efficacy of ACY-1215 has been demonstrated in in vitro and in vivo experiments [116] and is now tested in ongoing phase I/II clinical trials.
Histone methylation-modifying drugs (KMT inhibitors and/or KDM inhibitors) are under development, but their efficacy has not been fully studied in vivo yet [117]. One of the most important targets in MM is KMT3G/MMSET because of its universal expression in MM and several other cancers as well as its contribution to drug resistance and the poor prognosis of patients who overexpress this enzyme due to t(4;14). MMSET is believed to exhibit its activity through the PHD domain, as mutagenesis in this domain abrogates the methyltransferase activity of MMSET [118]. Thus, targeting the PHD domain of MMSET is an attractive strategy for the development of new epigenetic drugs, which could improve the prognosis of MM patients harboring t(4;14). EZH2 is another good candidate for anti-MM drug development. Given the role of EZH2 inactivation for CAM-DR, small molecular inhibitors of KDM6A/UTX or KDM6B/JMJD3, which counteracts EZH2 to increase H3K27 trimethylation, may be useful for repressing anti-apoptotic genes that operate for the acquisition of drug resistance in MM.
Finally, novel drugs targeting reader proteins of epigenetic marks are also under development. A promising example is BET inhibitors, which disrupt the binding of the bromodomain protein BRD4 to oncogenic super-enhancers, resulting in down-regulation of c-Myc in MM cells [119, 120]. Epigenetic drugs would pave a new avenue to more effective treatments because of their unique mode of action.
References
Cancer Information Service, National Cancer Center, Japan 2015.
Gay F, Larocca A, Wijermans P, et al. Complete response correlates with long-term progression-free and overall survival in elderly myeloma treated with novel agents: analysis of 1175 patients. Blood. 2011;117:3025–31.
Kumar SK, Rajkumar SV, Dispenzieri A, et al. Improved survival in multiple myeloma and the impact of novel therapies. Blood. 2008;111:2516–20.
Kumar SK, Dispenzieri A, Lacy MQ, et al. Continued improvement in survival in multiple myeloma: changes in early mortality and outcomes in older patients. Leukemia. 2014;28:1122–8.
Ozaki S, Handa H, Saitoh T, et al. Trends of survival in patients with multiple myeloma in Japan: a multicenter retrospective collaborative study of the Japanese Society of Myeloma. Blood Cancer J. 2015;5:e349.
Chapman MA, Lawrence MS, Keats JJ, et al. Initial genome sequencing and analysis of multiple myeloma. Nature. 2011;471:467–72.
Egan JB, Shi C-X, Tembe W, et al. Whole-genome sequencing of multiple myeloma from diagnosis to plasma cell leukemia reveals genomic initiating events, evolution, and clonal tides. Blood. 2012;120:1060–6.
Keats JJ, Chesi M, Egan JB, et al. Clonal competition with alternating dominance in multiple myeloma. Blood. 2012;120:1067–76.
Walker BA, Wardell CP, Melchor L, et al. Intraclonal heterogeneity and distinct molecular mechanisms characterize the development of t(4;14) and t(11;14) myeloma. Blood. 2012;120:1077–86.
Lohr JG, Stojanov P, Carter LS, et al. Widespread genetic heterogeneity in multiple myeloma: implications for targeted therapy. Cancer Cell. 2014;25:91–101.
Bolli N, Avet-Loiseau H, Wedge DC, et al. Heterogeneity of genomic evolution and mutational profiles in multiple myeloma. Nat Commun. 2014;5:2997.
Furukawa Y, Kikuchi J. Molecular pathogenesis of multiple myeloma. Int J Clin Oncol. 2015;20:413–22.
de Haart SJ, Willems SM, Mutis T, et al. Comparison of intramedullary myeloma and corresponding extramedullary soft tissue plasmacytomas using genetic mutational panel analyses. Blood Cancer J. 2016;6:e426.
Cifola I, Lionetti M, Pinatel E, et al. Whole-exome sequencing of primary plasma cell leukemia discloses heterogeneous mutational patterns. Oncotarget. 2015;6:17543–58.
Demchenko YN, Glebov OK, Zingone A, et al. Classical and/or alternative NF-κB pathway activation in multiple myeloma. Blood. 2010;115:3541–52.
West AC, Johnstone RW. New and emerging HDAC inhibitors for cancer treatment. J Clin Invest. 2014;124(1):30–9.
Andrulis M, Lehners N, Capper D, et al. Targeting the BRAF V600E mutation in multiple myeloma. Cancer Discov. 2013;3:862–9.
Braggio E, Kortüm KM, Stewart AK. SnapShot: multiple myelomas. Cancer Cell. 2015;28(678):e1.
Chesi P, Bergsagel L. Molecular pathogenesis of multiple myeloma: basic and clinical updates. Int J Hematol. 2013;97:313.
Meads MB, Hazlehurst LA, Dalton WS. The bone marrow microenvironment as a tumor sanctuary and contributor to drug resistance. Clin Cancer Res. 2008;14:2519–26.
Abe M. Targeting the interplay between myeloma cells and the bone marrow microenvironment in myeloma. Int J Hematol. 2011;94:334–43.
Cheung W-C, Van Ness B. Distinct IL-6 signal transduction leads to growth arrest and death in B cells or growth promotion and cell survival in myeloma cells. Leukemia. 2002;16:1182–8.
Abdi J, Chen G, Chang H. Drug resistance in multiple myeloma: latest findings and new concepts on molecular mechanisms. Oncotarget. 2013;4:2186–207.
Zhou W, Yang Y, Xia J, et al. NEK2 induces drug resistance mainly through activation of efflux drug pumps and is associated with poor prognosis in myeloma and other cancers. Cancer Cell. 2013;23:48–62.
Hazlehurst LA, Argilagos RF, Emmons M, et al. Cell adhesion to fibronectin (CAM-DR) influences acquired mitoxantrone resistance in U937 cells. Cancer Res. 2006;66:2338–45.
Matthes T, Manfroi B, Zeller A, Dunand-Sauthier I, Bogen B, Huard B. Autocrine amplification of immature myeloid cells by IL-6 in multiple myeloma-infiltrated bone marrow. Leukemia. 2015;29:1882–1890.
Chauhan D, Kharbanda S, Ogata A, et al. Interleukin-6 inhibits Fas-induced apoptosis and stress-activated protein kinase activation in multiple myeloma cells. Blood. 1997;89:227–34.
Shain KH, Yarde DN, Meads MB, et al. β1 integrin adhesion enhances IL-6-mediated STAT3 signaling in myeloma cells: implications for microenvironment influence on tumor survival and proliferation. Cancer Res. 2009;69:1009–15.
Frassanito MA, Cusmai A, Iodice G, Dammacco F. Autocrine interleukin-6 production and highly malignant multiple myeloma: relation with resistance to drug-induced apoptosis. Blood. 2001;97:483–9.
Voorhees PM, Chen Q, Kuhn DJ, et al. Inhibition of interleukin-6 signaling with CNTO 328 enhances the activity of bortezomib in preclinical models of multiple myeloma. Clin Cancer Res. 2007;13:6469–78.
Kuhn DJ, Berkova Z, Jones RJ, et al. Targeting the insulin-like growth factor-1 receptor to overcome bortezomib resistance in preclinical models of multiple myeloma. Blood. 2012;120:3260–70.
Chiron D, Maiga S, Surget S, et al. Autocrine insulin-like growth factor 1 and stem cell factor but not interleukin 6 support self-renewal of human myeloma cells. Blood Cancer J. 2013;3:e120.
Holt RU, Baykov V, Ro TB, et al. Human myeloma cells adhere to fibronectin in response to hepatocyte growth factor. Haematologica. 2005;90:479–88.
Zhang X-D, Baladandayuthapani V, Lin H, et al. Tight junction protein 1 modulates proteasome capacity and proteasome inhibitor sensitivity in multiple myeloma via EGFR/JAK1/STAT3 signaling. Cancer Cell. 2016;29:639–52.
Markovina S, Callander NS, O’Connor SL, et al. Bone marrow stromal cells from multiple myeloma patients uniquely induce bortezomib resistant NF-kB activity in myeloma cells. Mol Cancer. 2010;9:176.
Noborio-Hatano K, Kikuchi J, Takatoku M, et al. Bortezomib overcomes cell adhesion-mediated drug resistance through downregulation of VLA-4 expression in multiple myeloma. Oncogene. 2009;28:231–42.
Kikuchi J, Koyama D, Mukai H, Furukawa Y. Suitable drug combination with bortezomib for multiple myeloma under stroma-free conditions and in contact with fibronectin or bone marrow stromal cells. Int J Hematol. 2014;99:726–36.
Damiano JS, Cress AE, Hazlehurst LA, Shtil AA, Dalton WS. Cell adhesion mediated drug resistance (CAM-DR): role of integrins and resistance to apoptosis in human myeloma cell lines. Blood. 1999;93:1658–67.
Neri P, Ren L, Azab AK, et al. Integrin β7-mediated regulation of multiple myeloma cell adhesion, migration, and invasion. Blood. 2011;117:6202–13.
Hurt EM, Wiestner A, Rosenwald A, et al. Overexpression of c-maf is a frequent oncogenic event in multiple myeloma that promotes proliferation and pathological interactions with bone marrow stroma. Cancer Cell. 2004;5:191–9.
Schmidmaier R, Morsdorf K, Baumann P, Emmerich B, Meinhardt G. Evidence for cell adhesion-mediated drug resistance of multiple myeloma cells in vivo. Int J Biol Markers. 2006;21:218–22.
Hazlehurst LA, Enkemann SA, Beam CA, et al. Genotypic and phenotypic comparisons of de novo and acquired melphalan resistance in an isogenic multiple myeloma cell line model. Cancer Res. 2003;63:7900–6.
Shain KH, Landowski TH, Dalton WS. Adhesion-mediated intracellular redistribution of c-Fas-associated death domain-like IL-1-converting enzyme-like inhibitory protein-long confers resistance to CD95-induced apoptosis in hematopoietic cancer cell lines. J Immunol. 2002;168:2544–53.
Hosen N. Multiple myeloma-initiating cells. Int J Hematol. 2013;97:306–12.
Paiva B, Paino T, Sayagues J-M, et al. Detailed characterization of multiple myeloma circulating tumor cells shows unique phenotypic, cytogenetic, functional, and circadian distribution profile. Blood. 2013;122:3591–8.
Roccaro AM, Mishima Y, Sacco A, et al. CXCR4 regulates extra-medullary myeloma through epithelial-mesenchymal-transition-like transcriptional activation. Cell Rep. 2015;12:622–35.
Reijmers RM, Spaargaren M, Pals ST. Heparan sulfate proteoglycans in the control of B cell development and the pathogenesis of multiple myeloma. FEBS J. 2013;280:2180–93.
Matsui W, Huff CA, Wang Q. Characterization of clonogenic multiple myeloma cells. Blood. 2004;103:2332–6.
Kirshner J, Thulien KJ, Martin LD, et al. A unique three dimensional model for evaluating the impact of therapy on multiple myeloma. Blood. 2008;112:2935–45.
Pilarski LM, Seeberger K, Coupland RW, et al. Leukemic B cells clonally identical to myeloma plasma cells are myelomagenic in NOD/SCID mice. Exp Hematol. 2002;30:221–8.
Pilarski LM, Belch AR. Clonotypic myeloma cells able to xenograft myeloma to nonobese diabetic severe combined immunodeficient mice copurify with CD34+ hematopoietic progenitors. Clin Cancer Res. 2002;8(10):3198–204.
Hosen N, Matsuoka Y, Kishida S, et al. CD138-negative clonogenic cells are plasma cells but not B cells in some multiple myeloma patients. Leukemia. 2012;26(9):2135–41.
Matsui W, Wang Q, Barber JP, et al. Clonogenic multiple myeloma progenitors, stem cell properties, and drug resistance. Cancer Res. 2008;68:190–7.
Hawley TS, Riz I, Yang W, et al. Identification of an ABCB1 (P-glycoprotein)-positive carfilzomib resistant myeloma subpopulation by the pluripotent stem cell fluorescent dye CDy1. Am J Hematol. 2013;88:265–72.
Nara M, Teshima K, Watanabe A, et al. Bortezomib reduces the tumorigenicity of multiple myeloma via downregulation of upregulated targets in clonogenic side population cells. PLoS One. 2013;8:e56954.
Leung-Hagesteijn C, Erdmann N, Cheung G, et al. Xbp1s-negative tumor B cells and pre-plasmablasts mediate therapeutic proteasome inhibitor resistance in multiple myeloma. Cancer Cell. 2013;24:289–304.
Lalonde M-E, Cheng X, Cote J. Histone target selection within chromatin: an exemplary case of teamwork. Genes Dev. 2014;28:1029–41.
Struhl K, Segal E. Determinants of nucleosome positioning. Nat Struct Mol Biol. 2013;20:267–73.
Oshima M, Iwama A. Epigenetics of hematopoietic stem cell aging and disease. Int J Hematol. 2014;100:326–34.
Celik H, Kramer A, Challen GA. DNA methylation in normal and malignant hematopoiesis. Int J Hematol. 2016;103:617–26.
Black JC, Van Rechem C, Whetstine JR. Histone lysine methylation dynamics: establishment, regulation, and biological impact. Mol Cell. 2012;48:491–507.
Sharma SV, et al. A chromatin-mediated reversible drug-tolerant state in cancer cell subpopulations. Cell. 2010;141:69–80.
Roesch A, et al. Overcoming intrinsic multidrug resistance in melanoma by blocking the mitochondrial respiration chain of slow-cycling JARID1Bhigh cells. Cancer Cell. 2013;23:811–25.
Farria A, Li W, Dent SYR. KATs in cancer: functions and theraoies. Oncogene. 2015;34:4901–13.
Settleman J. Bet on drug resistance. Nature. 2016;529:289–90.
Ong C-T, Corces VG. Enhancers: emerging roles in cell fate specification. EMBO Rep. 2012;13:423–30.
Pichiorri F, Suh S-S, Ladetto M, et al. MicroRNAs regulate critical genes associated with multiple myeloma pathogenesis. Proc Natl Acad Sci USA. 2008;105:12885–90.
Barneda-Zahonero B, Parra M. Histone deacetylases and cancer. Mol. Oncol. 2012;6:579–89.
Lagger G, O’Carroll D, Rembold M, et al. Essential function of histone deacetylase 1 in proliferation control and CDK inhibitor repression. EMBO J. 2002;21:2672–81.
Murphy M, Ahn J, Walker KK, et al. Transcriptional repression by wild-type p53 utilizes histone deacetylases, mediated by interaction with mSin3a. Genes Dev. 1999;13:2490–501.
Moresi V, Williams AH, Meadows E, et al. Myogenin and class II HDACs control neurogenic muscle atrophy by inducing E3 ubiquitin ligases. Cell. 2010;143:35–45.
Tao R, de Zoeten EF, Ozkaynak E, et al. Deacetylase inhibition promotes the generation and function of regulatory T cells. Nat Med. 2007;13:1299–307.
Lam HC, Cloonan SM, Bhashyam AR, et al. Histone deacetylase 6–mediated selective autophagy regulates COPD-associated cilia dysfunction. J Clin Invest. 2013;123:5212–30.
Mithraprabhu S, Kalff A, Chow A, et al. Dysregulated class I histone deacetylases are indicators of poor prognosis in multiple myeloma. Epigenetics. 2014;9:1511–20.
White E. The role for autophagy in cancer. J Clin Invest. 2015;125:42–6.
Richardson PG, Schlossman RL, Alsina M, et al. PANORAMA2: panobinostat in combination with bortezomib and dexamethasone in patients with relapsed and bortezomib-refractory myeloma. Blood. 2013;122:2331–7.
San-Miguel JF, Hungria VTM, Yoon S-S, et al. Panobinostat plus bortezomib and dexamethasone versus placebo plus bortezomib and dexamethasone in patients with relapsed or relapsed and refractory multiple myeloma: a multicentre, randomised, double-blind phase 3 trial. Lancet Oncol. 2014;15:1195–206.
Sripayap P, Nagai T, Hatano K, et al. Romidepsin overcomes cell adhesion-mediated drug resistance in multiple myeloma cells. Acta Haematol. 2014;132:1–4.
Kikuchi J, Wada T, Shimizu R, et al. Histone deacetylases are critical targets of bortezomib-induced cytotoxicity in multiple myeloma. Blood. 2010;116:406–17.
Kikuchi J, Yamada S, Koyama D, et al. The novel orally active proteasome inhibitor K-7174 exerts anti-myeloma activity in vitro and in vivo by down-regulating the expression of class I histone deacetylases. J Biol Chem. 2013;288:25593–602.
Chaidos A, Barnes CP, Cowan G, et al. Clinical drug resistance linked to interconvertible phenotypic and functional states of tumor-propagating cells in multiple myeloma. Blood. 2013;121:318–28.
Stessman HAF, Baughn LB, Sarver A, et al. Profiling bortezomib resistance identifies secondary therapies in a mouse myeloma model. Mol Cancer Ther. 2013;12:1140–50.
Petrucci MT, Giraldo P, Corradini P, et al. A prospective, international phase 2 study of bortezomib retreatment in patients with relapsed multiple myeloma. Br J Haematol. 2013;160:649–59.
Greer EL, Shi Y. Histone methylation: a dynamic mark in health, disease and inheritance. Nat Rev Genet. 2012;13:343–57.
Di Croce L, Helin K. Transcriptional regulation by Polycomb group proteins. Nat Struct Mol Biol. 2013;20:1147–55.
Tram Nguyen A, Zhang Y. The diverse functions of Dot1 and H3K79 methylation. Genes Dev. 2011;25:1345–58.
Ernst J, Kellis M. Discovery and characterization of chromatin states for systematic annotation of the human genome. Nat Biotechnol. 2010;28:817–25.
Linghu C, Zheng H, Zhang L, Zhang J. Discovering common combinatorial histone modification patterns in the human genome. Gene. 2013;518:171–8.
Kouzarides T. Chromatin modifications and their function. Cell. 2007;128:693–705.
Brien GL, Valerio DG, Armstrong SA. Exploiting the epigenome to control cancer-promoting gene-expression programs. Cancer Cell. 2016;29:464–76.
Stec I, Wright TJ, van Ommen GJ, et al. WHSC1, a 90 kb SET domain-containing gene, expressed in early development and homologous to a Drosophila dysmorphy gene maps in the Wolf-Hirschhorn syndrome critical region and is fused to IgH in t(4;14) multiple myeloma. Hum Mol Genet. 1998;7:1071–82.
Marango J, Shimoyama M, Nishio H, et al. The MMSET protein is a histone methyltransferase with characteristics of a transcriptional corepressor. Blood. 2008;111:3145–54.
Martinez-Garcia E, Popovic R, Min D-J, et al. The MMSET histone methyl transferase switches global histone methylation and alters gene expression in t(4;14) multiple myeloma cells. Blood. 2011;117:211–20.
Pei H, Zhang L, Luo K, et al. MMSET regulates histone H4K20 methylation and 53BP1 accumulation at DNA damage sites. Nature. 2011;470:124–8.
Kuo AJ, Cheung P, Chen K, Zee BM, Kioi M, Lauring J, et al. NSD2 links dimethylation of histone H3 at lysine 36 to oncogenic programming. Mol Cell. 2011;44:609–20.
Popovic R, Martinez-Garcia E, Giannopoulou EG, et al. Histone methyltransferase MMSET/NSD2 alters EZH2 binding and reprograms the myeloma epigenome through global and focal changes in H3K36 and H3K27 methylation. PLoS Genet. 2014;10:e1004566.
Yang P, Guo L, Duan ZJ, et al. Histone methyltransferase NSD2/MMSET mediates constitutive NF-kB signaling for cancer cell proliferation, survival, and tumor growth via a feed-forward loop. Mol Cell Biol. 2012;32:3121–31.
Brito JL, Walker B, Jenner M, et al. MMSET deregulation affects cell cycle progression and adhesion regulons in t(4;14) myeloma plasma cells. Haematologica. 2009;94:78–86.
Plass C, Pfister SM, Lindroth AM, Bogatyrova O, Claus R, Lichter P. Mutations in regulators of the epigenome and their connections to global chromatin patterns in cancer. Nat Rev Genet. 2013;14:765–80.
Schwartzentruber J, et al. Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature. 2012;482:226–31.
Bender S, et al. Reduced H3K27me3 and DNA hypomethylation are major drivers of gene expression in K27 M mutant pediatric high-grade gliomas. Cancer Cell. 2013;24:660–72.
Kikuchi J, Koyama D, Wada T, et al. Phosphorylation-mediated EZH2 inactivation promotes drug resistance in multiple myeloma. J Clin Invest. 2015;125:4375–90.
Rahal R, et al. Pharmacological and genomic profiling identifies NF-κB-targeted treatment strategies for mantle cell lymphoma. Nat Med. 2014;20:87–92.
Yamaguchi H, Hung M-C. Regulation and role of EZH2 in cancer. Cancer Res Treat. 2014;46:209–22.
Kim E, Ilagan JO, Liang Y, et al. SRSF2 mutations contribute to myelodysplasia by mutant-specific effects on exon recognition. Cancer Cell. 2015;27:617–30.
Croonquist PA, Van Ness B. The polycomb group protein enhancer of zeste homolog 2 (EZH 2) is an oncogene that influences myeloma cell growth and the mutant ras phenotype. Oncogene. 2005;24:6269–80.
Sander S, Bullinger L, Klapproth K, et al. MYC stimulates EZH2 expression by repression of its negative regulator miR-26a. Blood. 2008;112:4202–12.
Nikoloski G, Langemeijer SM, Kuiper RP, et al. Somatic mutations of the histone methyltransferase gene EZH2 in myelodysplastic syndromes. Nat Genet. 2010;42:665–7.
Ernst T, Chase AJ, Score J, et al. Inactivating mutations of the histone methyltransferase gene EZH2 in myeloid disorders. Nat Genet. 2010;42:722–6.
Muto T, Sashida G, Oshima M, et al. Concurrent loss of Ezh2 and Tet2 cooperates in the pathogenesis of myelodysplastic disorders. J Exp Med. 2013;210:2627–39.
Sashida G, Harada H, Matsui H, et al. Ezh2 loss promotes development of myelodysplastic syndrome but attenuates its predisposition to leukaemic transformation. Nat Commun. 2014;5:4177.
Ntziachristos P, Tsirigos A, Van Vlierberghe P, et al. Genetic inactivation of the polycomb repressive complex 2 in T cell acute lymphoblastic leukemia. Nat Med. 2012;18:298–301.
Simon C, Chagraoui J, Krosl J, et al. A key role for EZH2 and associated genes in mouse and human adult T-cell acute leukemia. Genes Dev. 2012;26:651–6.
Kim E, et al. Phosphorylation of EZH2 activates STAT3 signaling via STAT3 methylation and promotes tumorigenicity of glioblastoma stem-like cells. Cancer Cell. 2013;23:839–52.
Santo L, Hideshima T, Kung AL, et al. Preclinical activity, pharmacodynamic, and pharmacokinetic properties of a selective HDAC6 inhibitor, ACY-1215, in combination with bortezomib in multiple myeloma. Blood. 2012;119:2579–89.
Mishima Y, Santo L, Eda H, et al. Ricolinostat (ACY-1215) induced inhibition of aggresome formation accelerates carfilzomib-induced multiple myeloma cell death. Br J Haematol. 2015;169:423–34.
Kim KH, Roberts CWM. Targeting EZH2 in cancer. Nat Med. 2016;22:128–34.
Huang Z, Wu H, Chuai S, Xu F, Yan F, Englund N, et al. NSD2 is recruited through its PHD domain to oncogenic gene loci to drive multiple myeloma. Cancer Res. 2013;73:6277–88.
Delmore JE, Issa GC, Lemieux ME, et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell. 2011;146:904–17.
Loven J, Hoke HA, Lin CY, et al. Selective inhibition of tumor oncogenesis by disruption of super-enhances. Cell. 2013;153:320–34.
Acknowledgments
Y.F. and J.K. received the Award in Aki’s Memory and the Kano Foundation Research Grant, respectively, from the International Myeloma Foundation Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
YF received research funding and honoraria from Janssen Pharmaceutical K.K., Takeda Pharmaceutical Co., Novartis Pharmaceutical Co., Fujimoto Pharmaceutical Co., Eisai Pharmaceutical Co., Celgene Co. and Bristol-Myers Squibb Co. JK declares no conflict of interest.
About this article
Cite this article
Furukawa, Y., Kikuchi, J. Epigenetic mechanisms of cell adhesion-mediated drug resistance in multiple myeloma. Int J Hematol 104, 281–292 (2016). https://doi.org/10.1007/s12185-016-2048-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12185-016-2048-5