Abstract
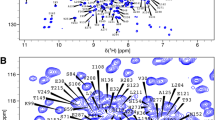
The widespread use of β-lactam antibiotics has given rise to a dramatic increase in clinically-relevant β-lactamases. Understanding the structure/function relation in these variants is essential to better address the ever-growing incidence of antibiotic resistance. We previously reported the backbone resonance assignments of a chimeric protein constituted of segments of the class A β-lactamases TEM-1 and PSE-4 (Morin et al. in Biomol NMR Assign 4:127–130, 2010. doi:10.1007/s12104-010-9227-8). That chimera, cTEM17m, held 17 amino acid substitutions relative to TEM-1 β-lactamase, resulting in a well-folded and fully functional protein with increased dynamics. Here we report the 1H, 13C and 15N backbone resonance assignments of chimera cTEM-19m, which includes 19 substitutions and exhibits increased active-site perturbation, as well as one of its deconvoluted variants, as the first step in the analysis of their dynamic behaviours.




Similar content being viewed by others
References
Ambler RP et al (1991) A standard numbering scheme for the class A beta-lactamases. Biochem J 276(Pt 1):269–270
Cavanagh J, Fairbrother WJ, Palmer III AG, Skelton NJ, Rance M (2006) Protein NMR spectroscopy: principles and practice, 2nd edn. Academic Press Inc., San Diego
Clouthier CM et al (2012) Chimeric beta-lactamases: global conservation of parental function and fast time-scale dynamics with increased slow motions. PloS One 7:e52283. doi:10.1371/journal.pone.0052283
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Gobeil SM, Clouthier CM, Park J, Gagné D, Berghuis AM, Doucet N, Pelletier JN (2014) Maintenance of native-like protein dynamics may not be required for engineering functional proteins. Chem Biol. doi:10.1016/j.chembiol.2014.07.016
Goddard TD, Kneller DG (2006) SPARKY 3, University of California, San Francisco
Johnson BA, Blevins RA (1994) NMR view: a computer program for the visualization and analysis of NMR data. J Biomol NMR 4:603–614. doi:10.1007/BF00404272
Lim D, Sanschagrin F, Passmore L, De Castro L, Levesque RC, Strynadka NC (2001) Insights into the molecular basis for the carbenicillinase activity of PSE-4 beta-lactamase from crystallographic and kinetic studies. Biochemistry 40:395–402
Meyer MM, Silberg JJ, Voigt CA, Endelman JB, Mayo SL, Wang ZG, Arnold FH (2003) Library analysis of SCHEMA-guided protein recombination. Protein Sci 12:1686–1693. doi:10.1110/ps.0306603
Meyer MM, Hochrein L, Arnold FH (2006) Structure-guided SCHEMA recombination of distantly related beta-lactamases protein engineering. Des Sel PEDS 19:563–570. doi:10.1093/protein/gzl045
Minasov G, Wang X, Shoichet BK (2002) An ultrahigh resolution structure of TEM-1 beta-lactamase suggests a role for Glu166 as the general base in acylation. J Am Chem Soc 124:5333–5340
Morin S, Gagné SM (2009) NMR dynamics of PSE-4 beta-lactamase: an interplay of ps-ns order and mus-ms motions in the active site. Biophys J 96:4681–4691. doi:10.1016/j.bpj.2009.02.068
Morin S, Levesque RC, Gagne SM (2006) 1H, 13C, and 15N backbone resonance assignments for PSE-4, a 29.5 kDa class A beta-lactamase from Pseudomonas aeruginosa. J Biomol NMR 36(Suppl 1):11. doi:10.1007/s10858-005-5343-7
Morin S, Clouthier CM, Gobeil S, Pelletier JN, Gagné SM (2010) Backbone resonance assignments of an artificially engineered TEM-1/PSE-4 Class A beta-lactamase chimera. Biomol NMR Assign 4:127–130. doi:10.1007/s12104-010-9227-8
Pimenta AC, Fernandes R, Moreira IS (2014) Evolution of drug resistance: insight on TEM beta-lactamases structure and activity and beta-lactam antibiotics. Mini Rev Med Chem 14:111–122
Savard PY, Gagné SM (2006) Backbone dynamics of TEM-1 determined by NMR: evidence for a highly ordered protein. Biochemistry 45:11414–11424. doi:10.1021/bi060414q
Savard PY, Sosa-Peinado A, Levesque RC, Makinen MW, Gagné SM (2004) 1H, 13C and 15N backbone resonance assignments for TEM-1, a 28.9 kDa beta-lactamase from E. coli. J Biomol NMR 29:433–434. doi:10.1023/B:JNMR.0000032503.96942.68
Studier FW (2005) Protein production by auto-induction in high density shaking cultures. Protein Expr Purif 41:207–234
Voigt CA, Martinez C, Wang ZG, Mayo SL, Arnold FH (2002) Protein building blocks preserved by recombination. Nat Struct Biol 9:553–558. doi:10.1038/nsb805nsb805
Acknowledgments
The authors thank Michelle M. Meyer and Frances H. Arnold for providing the original cTEM-19m construct; Tara Sprules and Sameer Al-Abdul-Wahid from the Québec/Eastern Canada High Field NMR Facility for NMR technical assistance; and the Concordia University BIOFINS platform for access to the circular dichroism apparatus. This work was supported by Natural Sciences and Engineering Research Council of Canada (NSERC) Discovery Grants RGPIN 227853 and 402623 (to J.N.P and N.D, respectively), in addition to a grant from the National Institute of General Medical Sciences (NIGMS) of the National Institutes of Health (NIH) under Award No. R01GM105978 (to N.D.). S.G. is the recipient of a FRQNT Graduate Scholarship and D.G. is the recipient of an NSERC Alexander Graham Bell Canada Graduate Scholarship. N.D. holds a Fonds de Recherche Québec – Santé (FRQS) Research Scholar Junior 1 Career Award.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gobeil, S.M.C., Gagné, D., Doucet, N. et al. 15N, 13C and 1H backbone resonance assignments of an artificially engineered TEM-1/PSE-4 class A β-lactamase chimera and its deconvoluted mutant. Biomol NMR Assign 10, 93–99 (2016). https://doi.org/10.1007/s12104-015-9645-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-015-9645-8