Abstract
Introduction
Chronic hepatitis B (CHB) affects 257 million individuals worldwide with an annual estimated mortality rate of 880,000 individuals. Accurate diagnosis of the stage of disease is difficult, and there is considerable uncertainty concerning the optimal point in time, when treatment should be started.
Objectives
By analyzing and comparing the metabolomes of patients at different stages of CHB and comparing them to healthy individuals, we want to determine the metabolic signature of disease progression and develop a more accurate metabolome-based method for diagnosis of disease progression ultimately giving a better basis for treatment decisions.
Methods
In this study, we used the combination of transient elastography and serum metabolomics of 307 serum samples from a group of 90 patients with CHB before and under treatment (with a follow-up time up to 10 years) at different progression stages over the clinical phases and 43 healthy controls..
Results
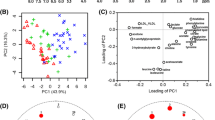
Our data show that the metabolomics approach can successfully discover CHB changing from the immune tolerance to the immune clearance phase and show distinctive metabolomes from different medical treatment stages. Perturbations in ammonia detoxification, glutamine and glutamate metabolism, methionine metabolism, dysregulation of branched-chain amino acids, and the tricarboxylic acid (TCA) cycle are the main factors involved in the progression of the disease. Fluctuations increasing in aspartate, glutamate, glutamine, methionine and 13 other metabolites are fingerprints of progression.
Conclusions
The metabolomics approach may expand the diagnostic armamentarium for patients with CHB. This method can provide a more detailed decision basis for starting medical treatment.







Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Ahn, H., Weaver, M., Lyon, D., Choi, E., & Fillingim, R. B. (2017). Inhibiting pyrimidine biosynthesis impairs Ebola virus replication through depletion of nucleoside pools and activation of innate immune responses. Physiology & Behavior, 176(10), 139–148.
Ardawi, M. S. M., & Newsholme, E. A. (1982). Maximum activities of some enzymes of glycolysis, the tricarboxylic acid cycle and ketone-body and glutamine utilization pathways in lymphocytes of the rat. Biochemical Journal, 208(3), 743–748.
Cano, A., Mariño, Z., Millet, O., Martínez-Arranz, I., Navasa, M., Falcón-Pérez, J. M., et al. (2017). A metabolomics signature linked to liver fibrosis in the serum of transplanted hepatitis C patients. Scientific Reports, 7(1), 1–10.
Clemmesen, J. O., Kondrup, J., & Ott, P. (2000). Splanchnic and leg exchange of amino acids and ammonia in acute liver failure. Gastroenterology, 118(6), 1131–1139.
Cruzat, V., Rogero, M. M., Keane, K. N., Curi, R., & Newsholme, P. (2018). Glutamine: Metabolism and immune function, supplementation and clinical translation. Nutrients, 10(11), 1–31.
Curi, R., Newsholme, P., & Newsholme, E. A. (1986). Intracellular distribution of some enzymes of the glutamine utilisation pathway in rat lymphocytes. Biochemical and Biophysical Research Communications, 138(1), 318–322.
Curi, R., Newsholme, P., Marzuca-Nassr, G. N., Takahashi, H. K., Hirabara, S. M., Cruzat, V., et al. (2016). Regulatory principles in metabolism—Then and now. Biochemical Journal, 473(13), 1845–1857.
Dumas, M. E., & Davidovic, L. (2013). Metabolic phenotyping and systems biology approaches to understanding neurological disorders. F1000Prime Reports, 5, 1–7.
EbbelsJ, T. M. D., Lindon, J. C., & Coen, M. (2010). Processing and modeling of nuclear magnetic Resonance (NMR) metabolic profiles. In Metabolic profilling (pp. 365–388). Springer Nature.
El-Serag, H., Lok, A. S. F., & Thomas, D. L. (2010). The dawn of a new era: Transforming our domestic response to hepatitis B & C. Gastroenterology, 138(4), 17–22.
Gleeson, M., & Bishop, N. C. (2000). Modification of immune responses to exercise by carbohydrate, glutamine and anti-oxidant supplements. Immunology and Cell Biology, 78(5), 554–561.
Gowda, G. A. N., Zhang, S., Gu, H., Asiago, V., Shanaiah, N., & Raftery, D. (2008). Metabolomics-based methods for early disease diagnostics. Expert Review of Molecular Diagnostics, 8(5), 617–633. https://doi.org/10.1586/14737159.8.5.617.
Helling, G., Wahlin, S., Smedberg, M., Pettersson, L., Tjäder, I., Norberg, Å., et al. (2016). Plasma glutamine concentrations in liver failure. PLoS ONE, 11(3), 1–10.
Horowitz, J. H., Rypins, E. B., Henderson, J. M., Heymsfield, S. B., Moffitt, S. D., Bain, R. P., et al. (1981). Evidence for impairment of transsulfuration pathway in cirrhosis. Gastroenterology, 81(4), 668–675. https://doi.org/10.1016/0016-5085(81)90489-3.
Lampertico, L. P., Agarwal, K., Berg, T., Buti, M., Janssen, H. L. A., Papatheodoridis, G., & Zoulim, F. T. F. (2017). EASL 2017 Clinical Practice Guidelines on the management of hepatitis B virus infection. Journal of Hepatology, 67, 370–398.
Lê Cao, K. A., Boitard, S., & Besse, P. (2011). Sparse PLS discriminant analysis: Biologically relevant feature selection and graphical displays for multiclass problems. BMC Bioinformatics, 12(1), 253. https://doi.org/10.1186/1471-2105-12-253.
Lê Cao, K. A., Rossouw, D., Robert-Granié, C., & Besse, P. (2008). A sparse PLS for variable selection when integrating omics data. Statistical Applications in Genetics and Molecular Biology. https://doi.org/10.2202/1544-6115.1390.
Leweling, H., Breitkreutz, R., Behne, F., Staedt, U., Striebel, J. P., & Holm, E. (1996). Hyperammonemia-induced depletion of glutamate and branched-chain amino acids in muscle and plasma. Journal of Hepatology, 25(5), 756–762.
Lu, S. C., Alvarez, L., Huang, Z., Chen, L., An, W., Corrales, F. J., & Kanel, G. (2011). Methionine adenosyltransferase 1A knockout mice are predisposed to liver injury and exhibit increased expression of genes involved. PNAS, 98(10), 5560–5565.
Maestri, N. E., McGowan, K. D., & Brusilow, S. W. (1992). Plasma glutamine concentration: A guide in the management of urea cycle disorders. The Journal of Pediatrics, 121(2), 259–261.
Masson, J. J. R., Billings, H. W. W., & Palmer, C. S. (2017). Metabolic reprogramming during hepatitis B disease progression offers novel diagnostic and therapeutic opportunities. Antiviral Chemistry and Chemotherapy, 25(2), 53–57.
Murphy, C., & Newsholme, P. (1998). Importance of glutamine macrophages and human monocytes to L-arginine biosynthesis and rates of nitrite or urea production. Clinical Science, 95(4), 397–407.
Nagamani, S. C. S., & Erez, A. (2016). A metabolic link between the urea cycle and cancer cell proliferation. Molecular and Cellular Oncology, 3(2), 1–3. https://doi.org/10.1080/23723556.2015.1127314.
Newsholme, P., Curi, R., Gordon, S., & Newsholme, E. A. (1986). Metabolism of glucose, glutamine, long-chain fatty acids and ketone bodies by murine macrophages. Biochemical Journal, 239(1), 121–125.
Newsholme, P. (2001). Why is L-glutamine metabolism important to cells of the immune system in health, postinjury, surgery or infection? The Journal of Nutrition, 131(9), 2515S-2522S.
Nicholson, J. K., Holmes, E., Kinross, J. M., Darzi, A. W., Takats, Z., & Lindon, J. C. (2012). Metabolic phenotyping in clinical and surgical environments. Nature, 491(7424), 384–392. https://doi.org/10.1038/nature11708.
Park, H. S., Choe, W. H., Han, H. S., Yu, M. H., Kim, Y. J., Jung, S. I., et al. (2019). Assessing significant fibrosis using imaging-based elastography in chronic hepatitis B patients: Pilot study. World Journal of Gastroenterology, 25(25), 3256–3267.
Perazzo, H., Veloso, V. G., Grinsztejn, B., Hyde, C., & Castro, R. (2015). Factors that could impact on liver fibrosis staging by transient elastography. International Journal of Hepatology. https://doi.org/10.1155/2015/624596.
Puchades-Carrasco, L., Palomino-Schätzlein, M., Pérez-Rambla, C., & Pineda-Lucena, A. (2016). Bioinformatics tools for the analysis of NMR metabolomics studies focused on the identification of clinically relevant biomarkers. Briefings in Bioinformatics, 17(3), 541–552. https://doi.org/10.1093/bib/bbv077.
Regev, A., Berho, M., Jeffers, L. J., Milikowski, C., Molina, E. G., Pyrsopoulos, N. T., et al. (2002). Sampling error and intraobserver variation in liver biopsy in patients with chronic HCV infection. American Journal of Gastroenterology, 97(10), 2614–2618.
Sands, C. J., Guha, I. N., Kyriakides, M., Wright, M., Beckonert, O., Holmes, E., et al. (2015). Metabolic phenotyping for enhanced mechanistic stratification of chronic hepatitis C-induced liver fibrosis. The American Journal of Gastroenterology, 110(1), 159–169. https://doi.org/10.1038/ajg.2014.370%5Cn.
Schoeman, J. C., Hou, J., Harms, A. C., Vreeken, R. J., Berger, R., Hankemeier, T., & Boonstra, A. (2016). Metabolic characterization of the natural progression of chronic hepatitis B. Genome Medicine, 8(1), 1–13. https://doi.org/10.1186/s13073-016-0318-8.
Sitryawan, A., Hawes, J. W., Harris, R. A., Shimomura, Y., Jenkins, A. E., & Hutson, S. M. (1998). A molecular model of human branched-chain amino acid metabolism. American Journal of Clinical Nutrition, 68(1), 72–81.
Tajiri, K., & Shimizu, Y. (2013). Branched-chain amino acids in liver diseases. World Journal of Gastroenterology, 19(43), 7620–7629.
Wishart, D. S., Tzur, D., Knox, C., Eisner, R., Guo, A. C., Young, N., et al. (2007). HMDB: The human metabolome database. Nucleic Acids Research, 35(Suppl. 1), 521–526.
Word Health Organization. (2017). Global hepatitis report 2017. http://www.who.int/hepatitis/publications/global-hepatitis-report2017/en
Worley, B., & Powers, R. (2013). Multivariate analysis in metabolomics. Current Metabolomics, 1(1), 92–107.
Xia, J., Broadhurst, D. I., Wilson, M., & Wishart, D. S. (2013). Translational biomarker discovery in clinical metabolomics: An introductory tutorial. Metabolomics, 9(2), 280–299. https://doi.org/10.1007/s11306-012-0482-9.
Yue, D., Zhang, Y., Cheng, L., Ma, J., Xi, Y., Yang, L., et al. (2016). Hepatitis B virus X protein (HBx)-induced abnormalities of nucleic acid metabolism revealed by 1 H-NMR-based metabonomics. Scientific Reports, 6, 1–13. https://doi.org/10.1038/srep24430.
Acknowledgements
We wish to thank the laboratory technicians Anne Bentzen-Petersen and Camilla Rosenkilde Larsen for their help.
Funding
This work was funded by the Innovation Fund Denmark. The NMR laboratory at Aalborg University is supported by the Obel Family, SparNord and Carlsberg Foundations.
Author information
Authors and Affiliations
Contributions
HTTN wrote the manuscript, contributed to study design, performed and analyzed samples with NMR, NMR data processing, statistical analysis and data interpretation. RW contributed to study design, NMR data analysis, data interpretation, and critical revision of the manuscript. VQL contributed to study design, statistical analysis, data analysis feature enhancement, debugging and code revising. HBK designed the study, clinical data advising, data interpretation, and critical revision of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
None
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. The study was approved by The North Denmark Region Committee on Health Research Ethics (Protocol N-20120028: Liver fibrosis – A translational study on the mechanisms of hepatic fibrogenesis).
Informed consent
Informed consent was obtained from all individual participants included in the study. All of the authors have read and approved the paper and it has not been published previously nor is it being considered by any other peer-reviewed journal.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nguyen, H.T.T., Wimmer, R., Le, V.Q. et al. Metabolic fingerprint of progression of chronic hepatitis B: changes in the metabolome and novel diagnostic possibilities. Metabolomics 17, 16 (2021). https://doi.org/10.1007/s11306-020-01767-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-020-01767-y