Abstract
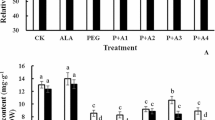
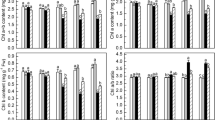
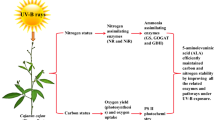
5-aminolevulinic acid (ALA) is an essential precursor for the biosynthesis of tetrapyrrols such as heme and chlorophyll (Chl). Previous studies have focused mainly on promotive effects of exogenous ALA on plant growth, while regulatory mechanisms affecting Chl biosynthesis have been only partially discussed. In the present study, the ameliorative role of exogenous ALA was investigated on Chl and endogenous ALA biosynthesis in six-day-old etiolated oilseed rape (Brassica napus L.) cotyledons during the de-etiolation stage. We showed that exogenously applied ALA of a low dosage enhanced Chl and ALA accumulation in cotyledons, while 600 µM ALA treatment inhibited the accumulation of Chl and ALA severely. However, the gene expression levels of glutamyl-tRNA reductase (HEMA) and glutamate-1-semialdehyde aminotransferase (GSA) were not affected under either low or high ALA concentrations. Furthermore, water deficit induced by polyethylene glycol 6000 (PEG) suppressed the Chl and ALA accumulation in cotyledons, while the inhibition was partially alleviated in the cotyledons pretreated with ALA. The decrease in Chl biosynthesis induced by PEG stress was assumed to be related to downregulation of HEMA and Mg-chelatase ChlH (ChlH), and upregulation of ferrochelatase (FC) genes. Moreover, exogenously applied ALA did not show any effect on the expression of Chl synthesis-related genes under the PEG treatment. These results showed a difference in suppressing ALA synthesis due to the high concentration of ALA and PEG. Exogenously applied ALA did not affect the expression of HEMA and GSA, thus exogenous ALA regulated Chl synthesis not via the regulation of transcriptional level in ALA biosynthesis. However, the inhibition in Chl and endogenous ALA accumulation by the PEG treatment may be attributed to downregulation of HEMA and ChlH, and upregulation of FC.
Similar content being viewed by others
Abbreviations
- ALA:
-
5-aminolevulinic acid
- Chl:
-
chlorophyll
- ChlH :
-
gene encoding Mg-chelatase ChlH subunit
- FC :
-
gene encoding ferrochelatase
- FLU:
-
fluorescent
- FM:
-
fresh mass
- GluTR:
-
glutamyl-tRNA reductase
- GSA :
-
gene encoding glutamate-1-semialdehyde aminotransferase
- HEMA :
-
gene encoding glutamyl-tRNA reductase
- Lhcb:
-
LHC protein of PSII type III chlorophyll a/b-binding protein
- MgCh:
-
Mg-chelatase
- Pchlide:
-
protochlorophyllide
- PEG:
-
polyethylene glycol
- POR:
-
protochlorophyllide oxidoreductase
- qRT-PCR:
-
quantitative real-time PCR
- URO :
-
gene encoding uroporphyrinogen decarboxylase
References
Aarti D., Tanaka R., Ito H. et al.: High light inhibits chlorophyll biosynthesis at the level of 5-aminolevulinate synthesis during de-etiolation in cucumber (Cucumis sativus) cotyledons.–Photochem. Photobiol. 83: 171–176, 2007.
Aarti P.D., Tanaka R., Tanaka A.: Effects of oxidative stress on chlorophyll biosynthesis in cucumber (Cucumis sativus) cotyledons.–Physiol. Plantarum 128: 186–197, 2006.
Ali B., Wang B., Ali S. et al.: 5-Aminolevulinic acid ameliorates the growth, photosynthetic gas exchange capacity, and ultrastructural changes under cadmium stress in Brassica napus L.–J. Plant Growth Regul. 32: 604–614, 2013.
Arnon D.I., Hoagland D.R.: Crop production in artificial solution with special reference to factors affecting yield and absorption of inorganic nutrients.–Soil Sci. 50: 463–485, 1940.
Beale S.I.: Enzymes of chlorophyll biosynthesis.–Photosynth. Res. 60: 43–73, 1999.
Beale S.I., Castelfranco P. A.: The biosynthesis of d-aminolevulinic acid in higher plants. I. Accumulation of d-aminolevulinic acid in greening plant tissues.–Plant Physiol. 53: 291–296, 1974.
Castelfranco P.A., Jones O.T.G.: Protoheme turnover and chlorophyll synthesis in greening barley tissue.–Plant Physiol. 55: 485–490, 1975.
Goslings D., Meskauskiene R., Kim C.H. et al.: Concurrent interactions of heme and FLU with Glu tRNA reductase (HEMA1), the target of metabolic feedback inhibition of tetrapyrrole biosynthesis, in dark-and light-grown Arabidopsis plants.–Plant J. 40: 957–967, 2004.
Granick S.: Magnesium porphyrins formed by barley seedlings treated with 5-aminolevulinic acid.–Plant Physiol. 34: S XVIII, 1959.
Hotta Y., Tanaka T., Takaoka H. et al.: New physiological effects of 5-aminolevulinic acid in plants: The increase of photosynthesis, chlorophyll content, and plant growth.–Biosci. Biotech. Bioch. 61: 2025–2028, 1997.
Ilag L.L., Kumar A.M., Söll D.: Light regulation of chlorophyll biosynthesis at the level of 5-aminolevulinic formation in Arabidopsis.–Plant Cell 6: 265–275, 1994.
Kittsteiner U., Mostowska A., Rüdiger W.: The greening process in cress seedlings. I. Pigment accumulation and ultrastructure after application of 5-aminolevulinic and complexing agents.–Physiol. Plantarum 81: 139–147, 1991.
Kim J.G., Back K., Lee H.Y. et al.: Increased expression of Fe-chelatase leads to increased metabolic flux into heme and confers protection against photodynamically induced oxidative stress.–Plant Mol. Biol. 86: 271–287, 2014.
Liu D., Pei Z.F., Naeem M.S. et al.: 5-Aminolevulinic acid activates antioxidative defence system and seedling growth in Brassica napus L. under water-deficit stress.–J. Agron. Crop Sci. 197: 284–295, 2011.
Livak K.J., Schmittgen T.D.: Analysis of relative gene expression data using real-time quantitative PCR and the 2-CT method.–Methods 25: 402–408, 2001.
Masuda T.: Recent overview of the Mg branch of the tetrapyrrole biosynthesis leading to chlorophylls.–Photosynth. Res. 96: 121–143, 2008.
Matsumoto F., Obayashi T., Sasaki-Sekimoto Y. et al.: Gene expression profiling of the tetrapyrrole metabolic pathway in Arabidopsis with a mini-array system.–Plant Physiol. 135: 2379–2391, 2004.
Maximová N., Slováková L.: Accumulation of photosynthetic pigments in Larix decidua Mill. and Picea abies (L.) Karst. cotyledons treated with 5-aminolevulinic acid under different irradiation.–Photosynthetica 52: 203–210, 2014.
McCormac A.C., Terry M.J.: Light-signalling pathways leading to the co-ordinated expression of HEMA1 and Lhcb during chloroplast development in Arabidopsis thaliana.–Plant J. 32: 549–559, 2002.
Memon S.A., Hou X., Wang L. et al.: Promotive effect of 5-aminolevulinic acid on chlorophyll, antioxidative enzymes and photosynthesis of Pakchoi (Brassica campestris ssp. chinensis var. communis Tsen et Lee).–Acta Physiol. Plant. 31: 51–57, 2009.
Meskauskiene R., Nater M., Goslings D. et al.: FLU: A negative regulator of chlorophyll biosynthesis in Arabidopsis thaliana.–P. Natl. Acad. Sci. USA 98: 12826–12831, 2001.
Mohanty S., Grimm B., Tripathy B.C.: Light and dark modulation of chlorophyll biosynthetic genes in response to temperature.–Planta 224: 692–699, 2006.
Naeem M.S., Warusawitharana H., Liu H. et al.: 5-aminolevulinic acid alleviates the salinity-induced changes in Brassica napus as revealed by the ultrastructural study of chloroplast.–Plant Physiol. Bioch. 57: 84–92, 2012.
Papenbrock J., Grimm B.: Regulatory network of tetrapyrrole biosynthesis–studies of intercellular signaling involved in metabolic and developmental control of plastids.–Planta 213: 667–681, 2001.
Pavlovic A., Demko V., Durchan M., Hudák J.: Feeding with aminolevulinic acid increased chlorophyll content in Norway spruce (Picea abies) in the dark.–Photosynthetica 47: 631–634, 2009.
Pontoppidan B., Kannangara C.G.: Purification and partial characterization of barley glutamate-tRNAGlu reductase, the enzyme that directs glutamate to chlorophyll biosynthesis.–Eur. J. Biochem. 225: 529–537, 1994.
Porra R.J., Thompson W.A., Kriedemann P.E.: Determination of accurate extinction coefficients and simultaneous-equations for assaying chlorophyll a and chlorophyll b extracted with 4 different solvents verification of the concentration of chlorophyll standards by atomic absorption spectroscopy.–BBABioenergetics 975: 384–394, 1989.
Schoefs B., Franck F.: Protochlorophyllide reduction: Mechanisms and evolution.–Photochem. Photobiol. 78: 543–557, 2003.
Sisler E.C., Klein W.H.: The effect of age and various chemicals on the lag phase of chlorophyll synthesis in dark grown bean seedlings.–Physiol. Plantarum 16: 315–322, 1963.
Sundqvist C.: Transformation of protochlorophyllide, formed from exogenous d-aminolevulinic acid, in continuous light and in flashlight.–Physiol. Plantarum 22: 147–156, 1969.
Tanaka R., Tanaka A.: Tetrapyrrole biosynthesis in higher plants.–Annu. Rev. Plant Biol. 58: 321–346, 2007.
Tanaka R., Yoshida K., Nakayashiki T. et al.: Differential expression of two hemA mRNAs encoding glutamyl-tRNA reductase proteins in greening cucumber seedlings.–Plant Physiol. 110: 1223–1230, 1996.
Tanaka R., Yoshida K., Nakayashiki T. et al.: The third member of the hemA gene family encoding glutamyl-tRNA reductase is primarily expressed in roots in Hordeum vulgare.–Photosynth. Res. 53: 161–171, 1997.
Tewari A.K., Tripathy B.C.: Temperature-stress-induced impairment of chlorophyll biosynthetic reactions in cucumber and wheat.–Plant Physiol. 117: 851–858, 1998.
Tewari A.K., Tripathy B.C.: Acclimation of chlorophyll biosynthetic reactions to temperature stress in cucumber (Cucumis sativus L.).–Planta 208: 431–437, 1999.
Author information
Authors and Affiliations
Corresponding author
Additional information
Acknowledgments: This work was supported by the Special Fund for Agro-scientific Research in the Public Interest (201303022), Jiangsu Collaborative Innovation Center for Modern Crop Production, the National Natural Science Foundation of China (31170405, 31301678, 31570434), and the Science and Technology Department of Zhejiang Province (2012C12902-1, 2011R50026-5).
Rights and permissions
About this article
Cite this article
Liu, D., Kong, D.D., Fu, X.K. et al. Influence of exogenous 5-aminolevulinic acid on chlorophyll synthesis and related gene expression in oilseed rape de-etiolated cotyledons under water-deficit stress. Photosynthetica 54, 468–474 (2016). https://doi.org/10.1007/s11099-016-0197-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11099-016-0197-7