Abstract
Cluster analysis aims at classifying objects into categories on the basis of their similarity and has been widely used in many areas such as pattern recognition and image processing. In this paper, we propose a novel clustering algorithm called QCC mainly based on the following ideas: the density of a cluster center is the highest in its K nearest neighborhood or reverse K nearest neighborhood, and clusters are divided by sparse regions. Besides, we define a novel concept of similarity between clusters to solve the complex-manifold problem. In experiments, we compare the proposed algorithm QCC with DBSCAN, DP and DAAP algorithms on synthetic and real-world datasets. Results show that QCC performs the best, and its superiority on clustering non-spherical data and complex-manifold data is especially large.
Similar content being viewed by others
1 Introduction
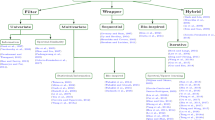
Clustering is one of primary methods in data mining and data analysis. It aims at classifying objects into categories or clusters, on the basis of their similarity. The clusters are collections of objects whose intra-class similarity is high and inter-class similarity is low. Up to present the study on clustering algorithms has been very active. Several different clustering methods have been proposed (Xu and Wunsch 2005), and they can be roughly divided into partitioning methods (Han and Kamber 2001; Kaufman and Rousseeuw 2009; Ng and Han 2002; Ordonez and Omiecinski 2002), hierarchical clustering (Zhang et al. 1996; Guha et al. 1998, 1999; Karypis et al. 1999), density-based clustering (Ester et al. 1996; Hinneburg and Keim 1998; Ankerst et al. 1999), grid-based clustering (Wang et al. 1997, 1999; Agrawal et al. 1998), model-based clustering (Moore 1999; Smith et al. 2013; Rhouma and Frigui 2001), and so on (Ling et al. 2007).
Given a dataset \(X=\{x_1,x_2,\ldots ,x_n\}\), the basic idea of a partitioning method is to partition the dataset into k clusters \((k<n)\). This kind of clustering algorithm generally starts with an initial partition of X and then uses an iterative control strategy to optimize an objective function. Among the proposed partitioning methods, k-means (Han and Kamber 2001) and k-medoids (Kaufman and Rousseeuw 2009) are the primary representatives. However, these methods are not applicable to non-spherical clusters. The model-based clustering method (a.k.a., distribution-based method) assumes that the objects in a specified cluster are most likely to be derived from a unique model. Expectation Maximization (EM)-based methods (Mclanchan and Krishan 1997) are the well-known branch of such methods, which adopt a fixed number of models, such as GMM, to approximate the object distribution. However, it is usually difficult to know the model or describe the distribution of real datasets before clustering. Moreover, this kind of clustering algorithm is still not applicable to non-spherical datasets (Jain 2010).
The hierarchical clustering organizes the objects as a hierarchical structure and assumes that the objects close to each other are more likely to be in the same cluster than the objects far away from each other. Single-Link (Sneath and Sokal 1962), Complete-Link (King 1967) and Ward method (Ward 1963) are the representative algorithms of this kind. The key idea of density-based clustering is that the clusters are defined as areas with higher density. This kind of method can correctly cluster the non-spherical datasets. The most popular example of density-based clustering is DBSCAN (Ester et al. 1996). The spectral clustering algorithm does not make assumptions on the forms of the clusters; it utilizes the spectrum of the similarity matrix to map the data into a lower-dimensional space in which the objects can be easily clustered by traditional clustering techniques (Von Luxburg 2007; Donath and Hoffman 1973; Hagen and Kahng 1992). This kind of clustering algorithm performs well on non-convex datasets. However, the density-based and spectral clustering algorithms cannot successfully cluster datasets containing structures with different densities or complex manifold.
In order to solve the problems mentioned above, we propose a new clustering method, called QCC. The proposed algorithm is based on the following ideas: the density of a cluster center is the maximum among its neighbors or reverse neighbors, and clusters are divided by sparse areas. At first, we introduce a new concept of local density of each object. Then QCC finds the quasi-cluster centers which correspond to initial clusters. Afterwards, we define a new metric to evaluate the similarity between initial clusters. Finally, we obtain the final clusters by merging the initial clusters between which the similarity is greater than \(\alpha \). The experimental results on the synthetic and real-world datasets show that the proposed algorithm is more effective than DP, DAAP and DBSCAN.
The rest of this paper is organized as follows. Section 2 is the related work of clustering. Section 3 introduces some related concepts and definitions of clustering algorithms. Section 4 presents the details of the proposed clustering algorithm QCC. Section 5 gives the experimental results of comparing QCC with DP, DAAP and DBSCAN. Section 6 concludes our work.
2 Related work
As one of primary methods in data mining and data analysis, clustering has got more and more attention from the industry and academia. A great number of clustering algorithms have been proposed. For many of traditional clustering algorithms, a key step is to find cluster centers. For example, k-means (Han and Kamber 2001) and k-medoids (Kaufman and Rousseeuw 2009) methods classify the objects into a cluster based on the distances to the cluster center. An objective function, typically the sum of the distance to a set of putative cluster centers, is optimized until the best cluster centers candidates are found. In 2007, Brendan and Delbert proposed a new clustering algorithm by passing messages between data points in Science, called affinity propagation (AP) (Frey and Dueck 2007). The main purpose of AP clustering is to find the optimal representative points. Different from k-means, the AP algorithm does not need to specify the initial cluster centers in advance. In contrast, it regards all data points as potential cluster centers, and therefore avoids the arbitrariness in selecting the initial cluster centers. However, AP cannot directly specify the final number of clusters. In order to generate K clusters, Zhang et al. (2010) propose K-AP clustering algorithm.
The above mentioned center-based methods are not able to detect non-spherical clusters (Jain 2010), since data points are always assigned to the nearest center. In 2014, Rodriguez and Laio proposed a new clustering algorithm in Science, called DP. The DP algorithm is based on the following idea: cluster centers are surrounded by neighbors with lower local density, and they are at a relatively large distance from any points with a higher local density. Non-spherical clusters can be easily detected by DP clustering algorithm. However, DP suffers from identifying the ideal number of clusters. In 2016, Wang and Song proposed an improved clustering algorithm called STClu that is insensitive to parameters. However the result of DP and STClu is undesirable when clustering complex-manifold datasets. In 2014, Hong et al. proposed a clustering algorithm called DAAP, and it can solve complex-manifold problems by computing a similarity which is defined as the sum of the Edge-Weight of shortest path (Jia et al. 2014). However, the time complexity of DAAP is much higher than AP, K-AP and DP, for computing the specifically-defined similarity. Moreover, DAAP needs too many parameters to set such as damping coefficient, the maximal iteration, the number of neighbors and clusters, and generally, the clustering effect of DAAP is undesirable on datasets that contain structures with different densities (Jia et al. 2014).
3 Preliminaries
Most of the existing density-based clustering algorithms, such as DBSCAN and DP, define the density of each point \(p_i\) as the number of neighbors with distance to \(p_i\) less than the cutoff distance \(d_c\), as shown in Eq. 1
In Eq. 1 if \(d_{ij}-d_c<0\), \(\varphi (d_{ij}-d_c )=1\) and otherwise \(\varphi (d_{ij}-d_c )=0\). \(\rho _i\) is equal to the number of points that are closer to point i than \(d_c\). However, once the inter-class density variations is great, the value of \(d_c\) is hard to set. For example, as shown in Fig. 1, if the value of \(d_c\) is set inappropriately, then there are no neighbors in the neighborhood of points a and b within \(d_c\), but the neighbors of \(C_1\)’s center include all points in \(C_1\). Moreover, although points a and b are normal points, they will be regarded as noise by DBSCAN. However, points a and b will be regarded as the cluster center by DP algorithm, since the local density of points a and b is the largest in their neighborhood.
Since the proposed method is center-based, like DP and DAAP clustering algorithms, it also needs to compute the density of every point. In order to avoid the above problem, we introduce the following definitions. Let D be a dataset, p and q be two objects in D, and k be a positive integer. We use d(p, q) to denote the Euclidean distance between objects p and q.
Definition 1
(K-distance) The K-distance of p, denoted as \( Dist _K(p)\), is the distance d(p, o) between p and o in D, such that:
-
(1)
At least K objects \(o' \in D{/}\{p\}\) satisfy \(d(p,o') \le d(p,o)\), and
-
(2)
At most \((K-1)\) objects \(o' \in D{/}\{p\}\) satisfy \(d(p,o')<d(p,o)\)
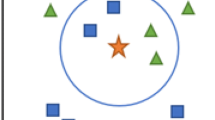
\( Dist _K(p)\) is the distance between p and the Kth nearest neighbor of p. For example, as shown in Fig. 2, when \(K=3\), the \( Dist _3(p)=d(p,q)\). The \( Dist _K(p)\) can represent the density of the object p. The smaller \( Dist _K(p)\) is, the much denser the area around p is. Therefore, like paper Jin et al. (2006), we define the density of p, denoted as \( Den (p)\), as the following equation:
Definition 2
(K Nearest Neighbor and Reverse K Nearest Neighbor) If \(d(p,q) \le Dist _K(p)\), then the object q is called the K Nearest Neighbor of p. All the K Nearest Neighbors compose the K Nearest Neighborhood, denoted as \( KNN (p)\). If \(d(q,p) \le Dist _K(q)\), then the object q is called the Reverse K Nearest Neighbor of p, and all the Reverse K Nearest Neighbors compose the Reverse K Nearest Neighborhood, denoted as \( RKNN (p)\). The formulation of \( KNN (p)\) and \( RKNN (p)\) is given as follows.
For example, as shown in Fig. 2, when \(K=3\), \(3 NN (p)=\{a,b,q\}\), \(3 NN (a)=\{b,p,q\}\), \(3 NN (b)=\{a,p,q\}\), \(3 NN (q)=\{d,p,b\}\), \(3 NN (d)=\{q,p,b\}\) and we can obtain that \( R 3 NN (p)=\{a,b,q,d\}\).
4 The QCC algorithm
In this paper we divide the neighbors of every point into Dense Neighbors and Sparse Neighbors, which are defined in Definition 3.
Definition 3
(Dense and Sparse Neighbor) If the density of q is greater than the density of p and \(q\in KNN (p)\), then the object q is called the Dense Neighbor of p, denoted as \( DN (p)\). On the contrary, if the density of q is smaller than or equal to the density of p and \(q\in KNN (p)\), then q is called the Sparse Neighbor of p, denoted as SN(p).
Definition 4
(Exemplar) We define point q as the Exemplar of p where \(q \in Q\) and \(d(p,q)= min _{i=1}^n\{d(p,q_i)\}\). Q is a collection defined as follows:
In Definition 4, n is the number of points in Q. From the definition of Exemplar, we can know that each point of the dataset possesses at most one Exemplar. If the density of p is greater than the density of all k nearest neighbors or reverse k nearest neighbors of p, p is the Exemplar of itself. Then we call p a Quasi-Cluster Center.
Definition 5
(Quasi-Cluster Center) If object p satisfies one of the following two conditions, then we call p a Quasi-Cluster Center.
-
1.
\(\forall q \in KNN (p), Den (p)\ge Den (q)\) or
-
2.
\(\forall q \in RKNN (p), Den (p)\ge Den (q)\)
Figure 3 is the Exemplar Graph (EG) which can be comprised by connecting each point p to its Exemplar. As shown in Fig. 3, the parameter K is set to 30, and the red points \(c_1,c_2,\ldots ,c_7\) are Quasi-Cluster Centers. Other red points will be treated as outliers that will be explained in Algorithm 1.

As shown in Fig. 3, the top-right class is divided into 6 small clusters. However, from the result of Fig. 4, we can see that the Quasi-Cluster Centers are the real cluster centers, when we set an appropriate value of k. We also find that the number of Quasi-Cluster Centers appears to decrease as the value of parameter k increases. For example, QCC finds 6 cluster centers in the top-right cluster in Fig. 3, when \(K=30\). However, as shown in the left figure of Fig. 4, QCC only finds one cluster center that is the real cluster center of this cluster, when \(K=50\). Hence, QCC can cluster these datasets by constantly spreading out from the cluster centers, and obtain the accurate clustering result when the value of K is appropriate. However, in this way, it is hard to correctly cluster the complex-manifold datasets. In order to solve the complex-manifold problem, we define the similarity between clusters as follows.
Definition 6
(Similarity between clusters) Similarity between clusters \(C_i\) and \(C_j\), denoted as \( Sim (C_i,C_j)\), is defined as the ratio of the number of points in \(C_i \cap C_j\) and K. The formulation of similarity between clusters is shown as follows:
As shown in Fig. 5, the set of red points is the intersection of C1 and C2. We consider the ratio of the number of these red points and K as the similarity between two adjacent initial clusters. The value of \( Sim (C_{i},C_{j} )\) is no less than 0. If these two adjacent initial clusters are divided by a sparse area, the similarity between these two clusters will be small. Then these two clusters are two individual clusters. On the contrary, if these two adjacent initial clusters are connected by a density area, the similarity between these two adjacent clusters will be large. Then these two clusters will be merged into one cluster. In this way, even if one large cluster is divided into many small clusters because the value of k is small, as shown in Fig. 3, these small clusters \(C1, C2,\ldots , C7\) will be finally merged into one cluster.
Based on the above definitions, we propose a novel clustering algorithm, called QCC. The procedure of QCC algorithm is minutely described in Algorithm 1.
Firstly, the proposed clustering algorithm QCC uses the KNN searching algorithm to obtain KNN and RKNN of each point in D, and computes the density of each point. Secondly, in step 5 of Algorithm 1, QCC finds the Exemplar of each point using Definition 4, and obtains all Quasi-Cluster Centers using Definition 5. After that, QCC obtains the initial clusters via the following steps.
-
1.
QCC arbitrarily finds a Quasi-Cluster Center, and classifies it and its Sparse Neighbors to the same cluster \(C_i\).
-
2.
Then QCC arbitrarily finds a point p in this cluster and classifies the Sparse Neighbors of p to cluster \(C_i\), until all points of this cluster have been visited.
-
3.
Afterwards, QCC finds an unvisited Quasi-Cluster Center and repeats the above steps, until all Quasi-Cluster Centers have been visited.
By doing so, the clusters spread from dense areas to sparse areas. As shown in Fig. 5, the red points are classified to C1 and C2 at the same time. Then QCC merges all the clusters between which the similarity is greater than \(\alpha \) into one cluster. If the similarity between clusters C1 and C2 is smaller than \(\alpha \), then the red points will be classified into the cluster that its Exemplar belongs to. \(\alpha \) is a user-defined parameter. A larger value of \(\alpha \) leads to more clusters. Actually, QCC algorithm is robust with respect to the choice of parameter K and \(\alpha \), and this will be analysed in the next section.
After the above steps, QCC regards the clusters with \(|c_i|<K\) (i.e., the number of points in \(C_i\) smaller than K) as outlier clusters. In other words, the point in these clusters is marked as outlier. So the red points in Fig. 3 will be regarded as outliers except \(C1, C2,\ldots , C7\). Finally, QCC outputs the final clusters. Note that the value of K is preferably smaller than the number of points in the smallest normal cluster of the dataset, otherwise the smallest normal cluster may be merged to an adjacent big cluster. QCC can correctly cluster the dataset, as long as the above conditions are met. The complexity of the QCC is \(O(n^2)\). If we use the K-D tree to search the neighbors of each point, the complexity of QCC would be decreased to \(O(n*\log n)\).
5 Experimental analysis
5.1 Clustering on synthetic datasets
In order to demonstrate the effectiveness of QCC, we compare the proposed clustering algorithm QCC with DBSCAN, DAAP and DP algorithms. DBSCAN is a famous density-based clustering algorithm devoted to solve the manifold problem. We conduct experiments on four synthetic datasets and Olivetti Face Database. Four synthetic datasets are illustrated in Fig. 6. Data1, taken from Ester et al. (1996), consists of two spherical classes, two manifold classes and a few outliers, a total of 582 points. Data2, taken from Ha et al. (2014), consists of three spherical classes, one complex-manifold class and some noise points, a total of 1400 points. Data3, taken from Cassisi et al. (2013), is composed of six high density manifold classes and some noise points, a total of 8000 points. Data4, taken from Zhu et al. (2014), consists of 159 points and has one dense spherical class and one sparse manifold class.
For DP, we decide on the right number of clusters to Data1, Data3 and Data4, and show the best clustering result in repeated tests on Data2. Hence, we don’t show the decision graph, deciding the number of the clusters, of DP in all results. For DAAP, we set the density factor \(\rho \) to 2, the maximal iteration maxits to 1000, convergence of iteration coefficient convits to 100.
Figure 7 shows the clustering results of each approach on Data1. For DAAP, the number (k) of neighbors used for constructing the adjacency matrix is set to 6, the value of damping coefficient (lam) is set to 0.9. From this figure, we can see that DP algorithm can correctly cluster the spherical class and simple manifolds class, but can’t correctly cluster the complex-manifold class. Data1 is correctly clustered by DAAP, DBSCAN (\( eps =15\), \( minpoints =5\)) and QCC (\(K=20\), \(\alpha =0.3\)). DBSCAN and QCC algorithms detect out the noise points in Data1, but DAAP can’t.
Figure 8 shows the clustering results of each algorithm on Data2. For DAAP, the number (k) of neighbors used for constructing the adjacency matrix is set to 6, the value damping coefficient (lam) is 0.9. DP fails to cluster the complex-manifold data that is grouped into 6 clusters. Owing to specifically-defined similarity, DAAP has certain capacity to cluster complex-manifold dataset. However, DAAP fails to cluster the manifold class in Data2. Since the shortest path is too long, the end region (marked by red square) of the manifold class was wrongly clustered. Data2 is correctly clustered by DBSCAN (\( eps =0.2\), \( minpoints =40\)) and QCC (\(K=20\), \(\alpha =0.3\)) algorithms, and most of the noise points in Data2 are detected out by DBSCAN and QCC.
Figure 9 shows the clustering results of each algorithm on Data3. Although DP obtains the right number of clusters in Data3 by manually selecting the cluster centers in the decision graph, three clusters are incorrectly clustered. DAAP obtains the right number of clusters, but some clusters are incorrectly clustered, too. Moreover, DAAP mistakenly regards a part of noise as a small normal cluster. The clustering result of DBSCAN is obviously superior to DP and DAAP, and DBSCAN detects out the noise points in Data3. However, some points in normal clusters are treated as noise points by DBSCAN. Although QCC (\(K=80\), \(\alpha =6\)) fails to detect out the noise in Data3, QCC obtains the right number of clusters and correctly clusters all normal points.
Figure 10 shows the clustering results of each algorithm on Data4. Same as for the results on Data3, although DP and DAAP obtain the right number of clusters by manually select or set, the results of DP and DAAP are undesirable. Since the density variations of the two clusters of Data4 is great, DBSCAN (\( eps =2\), \( minpoints =5\)) fails to correctly cluster Data4. DBSCAN does not obtain the right number of clusters in Data4, and mistakenly treats some normal points as noise. The performance of QCC (\(K=5\), \(\alpha =0.2\)) is obviously superior to that of DP, DAAP and DBSCAN on Data4.
From the above results and analysis, we can see that the DP algorithm has a certain capacity to cluster non-spherical data. However the DP algorithm can hardly correctly cluster the complex-manifold datasets. DAAP has a certain capacity to cluster the datasets with complex manifold. However, DAAP fails to cluster those datasets containing long manifold class (Data2), lots of noise (Data3) or great density variations between clusters (Data4). Although the performance of DBSCAN is superior to that of DP and DAAP, DBSCAN fails to correctly cluster on Data3 and Data4. Therefore, from the results with the synthetic datasets, we can see that the proposed clustering algorithm QCC can get the right number of final clusters without human intervention, and the scope of QCC’s application is wider than other clustering algorithms. Whether the datasets contain complex-manifold data or the density variations of inter-class are great, QCC can get satisfactory clustering results. In order to demonstrate the effectiveness of QCC, we also conduct experiments on real datasets in the following section.
5.2 Clustering on Olivetti Face Database
Like paper Rodriguez and Laio (2014), we also apply the QCC algorithm to the Olivetti Face Database (Samaria and Harter 1994), a widespread benchmark for machine learning algorithms, with the aim of identifying, without any previous training, the number of subjects in the database. Similar to the experiment on synthetic datasets, we compare QCC with DP, DAAP and DBSCAN algorithms. In this experiment, we use 10 clusters of Olivetti Face Database. Each cluster is composed of 10 face pictures. The size of each picture is \(M*N\) with \(M=112\), \(N=92\). We regard the correlation of picture A and B as the similarity between two images, denoted as S(A, B), and it is computed by the following equation.
Here A and B are the subjects of the Olivetti Face Database. \(A_{mn}\) and \(B_{mn} (m=\{1,2,\ldots ,M\},n=\{1,2,\ldots ,N\})\) represent the pixels of the two subject pictures. The value of S is scaled between 0 and 1. The larger the value of S is, the more similar the two pictures are. We define the distance between two pictures, denoted as d(A, B), as follows.
The density is estimated via Eq. 2. In order to intuitively describe the efficiency of QCC, we use the criteria of Purity and Recall to evaluate the clustering performance, which are defined as follows:
Here, let D be a dataset containing TMc classes \(TC=\{tc_1,tc_2,\ldots ,tc_{ TMc }\}\). The result of clustering algorithm contains Mc clusters \(C=\{c_1,c_2,\ldots ,c_{ Mc }\}\). \(|c_i|\) is the number of points in \(c_i\). The range of Purity and Recall is [0,1]. The larger the value of Purity or Recall is, the better the clustering performance is.
The results of DP, DAAP, DBSCAN and QCC are shown in Figs. 11, 12, 13 and 14, respectively. In all results, faces with the same color belong to the same cluster.
Figure 11 shows the clustering result of DP on the Olivetti Face Database. We choose 12 points as cluster centers through decision graph (Rodriguez and Laio 2014), so DP detects 12 clusters. From Table 1, we can see that the Recall of DP is 0.82 and the Purity of DP is 0.88, which indicates that some recalled faces are not correctly clustered.
Figure 12 shows the clustering result of DAAP on the Olivetti Face Database. Although DAAP correctly detects 10 clusters by manually setting parameter k (i.e., the final number of clusters), the Recall and Purity of DAAP are undesirable (only 0.67 and 0.61). Specially, the Purity of DAAP is the lowest among the four algorithms.
Figure 13 shows the clustering result of DBSCAN on the Olivetti Face Database. DBSCAN only detects out 8 clusters. Two clusters, marked with red spots, are regarded as noise points. Moreover, even in these 8 clusters, some faces marked with red spots are not recalled. Hence the Recall of DBSCAN is only 0.63, and it is the lowest among the four algorithms, although the Purity of DBSCAN is 0.98 (one recalled face marked with green spot is incorrectly clustered).
The clustering results of QCC on the Olivetti Face Database is shown in Fig. 14. The results show that the Olivetti Face Database is grouped into 11 clusters by QCC, because one of the real 10 clusters that within the red border is divided into two clusters. Six images marked with a red spot are considered as outliers by QCC. However, among the 11 clusters, 6 clusters are really correct, 2 clusters identifies 9 face images, and one cluster identifies 6 face images in all 10 face images. Moreover, all the 11 clusters remain pure, namely including only images of the same cluster. Hence, the Purity of QCC is 1 (the best one of all), and the Recall of QCC is 0.9.
Through above experiments and analysis, we can get to the conclusion that the proposed algorithm QCC outperforms DP, DAAP and DBSCAN algorithms. QCC algorithm has a broader application than DAAP, DP and DBSCAN algorithms, as QCC is applicable to datasets containing different density or complex-manifold clusters.
5.3 Robustness analysis
In order to demonstrate that QCC is robust with respect to the choice of parameter K and \(\alpha \), we do the following experiments on two synthetic datasets (Data1 and Data5). As shown in Figs. 17 and 18, Data5 consists of one spherical class, two complex-manifold classes, a total of 2374 points.
First, for Data1, we set \(\alpha \) to 0.2 with K changing from 10 to 30, and the experimental results are shown in Fig. 15. For Data5, we set \(\alpha \) to 0.3 with K changing from 15 to 35, and the experimental results are shown in Fig. 17. We can learn from Figs. 15, 17 and Table 2 that the number of Quasi-Cluster Center gets smaller as the parameter K becomes larger. When processing the cluster with complex manifold, QCC will obtain many Quasi-Cluster Centers. Each Quasi-Cluster Center represents an initial cluster. Then QCC computes the similarity between these initial clusters. The values of similarity of these initial clusters that belong to the same class must be great. Finally, QCC merges these initial clusters, and obtains the final cluster with complex manifold. Hence, QCC can correctly cluster the datasets with different values of K. Moreover, theoretically QCC is applicable to datasets with any complex manifold.
Then, for Data1, we set K to 25 with \(\alpha \) changing from 0.1 to 0.8, and the experimental results are shown in Fig. 16. Figure 16a shows the result of Quasi-Cluster Centers of Data1. Figure 16b–i shows the clustering results of Data1. For Data5, we set K to 40 with \(\alpha \) changing from 0.1 to 0.8, and the experimental results are shown in Fig. 18. Figure 18a shows the result of Quasi-Cluster Centers of Data5. Figure 18b–i shows the clustering results of Data5 with different value of \(\alpha \). Through lots of experiments, the good value range of \(\alpha \) is [0.2, 0.5].
From these above results, we can see that QCC can obtain the correct clustering results with different values of \(\alpha \). Hence, we can conclude that QCC is robust to parameters K and \(\alpha \).
6 Conclusion
In this paper, we propose a new clustering algorithm called QCC. The core idea of QCC is that clusters are divided by sparse regions. Based on this idea, firstly, we introduce a new metric to measure the local density of each object. Then we define the Quasi-Cluster Centers. Remarkably, the real cluster centers must be included in the set of Quasi-Cluster Centers. After that, QCC obtains the initial clusters through spreading from dense areas to sparse areas. Afterwards, we define the similarity between initial clusters, and obtain the final clusters by merging the initial clusters for which the similarity is greater than \(\alpha \). Therefore, QCC applies to complex-manifold dataset. Through the experiments on the four synthetic datasets and the Olivetti Face Database, we confirm that the proposed clustering algorithm QCC can correctly cluster complex-manifold datasets and datasets with large density variation, and QCC is more effective than DP, DAAP and DBSCAN.
References
Agrawal, R., et al. (1998). Automatic subspace clustering of high dimensional data for data mining applications. In ACM (Vol. 27).
Ankerst, M., et al. (1999). OPTICS: Ordering points to identify the clustering structure. In ACM Sigmod record. ACM.
Cassisi, C., et al. (2013). Enhancing density-based clustering: Parameter reduction and outlier detection. Information Systems, 38(3), 317–330.
Donath, W. E., & Hoffman, A. J. (1973). Lower bounds for the partitioning of graphs. IBM Journal of Research and Development, 17(5), 420–425.
Ester, M., et al. (1996). A density-based algorithm for discovering clusters in large spatial databases with noise. In Kdd.
Frey, B. J., & Dueck, D. (2007). Clustering by passing messages between data points. Science, 315(5814), 972–976.
Guha, S., Rastogi, R., & Shim, K. (1998). CURE: An efficient clustering algorithm for large databases. In ACM SIGMOD record. ACM.
Guha, S., Rastogi, R., & Shim, K. (1999). ROCK: A robust clustering algorithm for categorical attributes. In Data engineering, 1999. Proceedings of the 15th international conference on. IEEE.
Han, J., & Kamber, M. (2001). Data mining: Concepts and techniques. San Francisco, CA: Morgan Kaufmann.
Hagen, L., & Kahng, A. B. (1992). New spectral methods for ratio cut partitioning and clustering. IEEE Transactions on Computer-Aided Design of Integrated Circuits and Systems, 11(9), 1074–1085.
Ha, J., Seok, S., & Lee, J.-S. (2014). Robust outlier detection using the instability factor. Knowledge-Based Systems, 63, 15–23.
Hinneburg, A. & Keim, D.A. (1998). An efficient approach to clustering in large multimedia databases with noise. In KDD.
Jain, A. K. (2010). Data clustering: 50 years beyond K-means. Pattern Recognition Letters, 31(8), 651–666.
Jia, H., et al. (2014). A density-adaptive affinity propagation clustering algorithm based on spectral dimension reduction. Neural Computing and Applications, 25(7–8), 1557–1567.
Jin, W., et al. (2006). Ranking outliers using symmetric neighborhood relationship. In Advances in knowledge discovery and data mining (pp. 577–593). Berlin: Springer.
Kaufman, L., & Rousseeuw, P. J. (2009). Finding groups in data: An introduction to cluster analysis. New York: Wiley.
Karypis, G., Han, E.-H., & Kumar, V. (1999). Chameleon: Hierarchical clustering using dynamic modeling. Computer, 32(8), 68–75.
King, B. (1967). Step-wise clustering procedures. Journal of the American Statistical Association, 62, 86–101.
Li, T., Ma, S., & Ogihara, M. (2004). Document clustering via adaptive subspace iteration. In Proceedings of the 27th annual international ACM SIGIR conference on research and development in information retrieval (pp. 218–225).
Ling, H., Lingda, W., & Yi-chao, C. (2007). Survey of clustering algorithms in data mining. Application Research of Computers, 1, 10–13.
Moore, A. W. (1999). Very fast EM-based mixture model clustering using multiresolution kd-trees. Advances in Neural Information Processing Systems, 543–549.
Mclanchan, G., & Krishan, T. (1997). The em algorithm and extensions. Series in Probability and Statistics, 15(1), 154–156.
Ng, R. T., & Han, J. (2002). Clarans: A method for clustering objects for spatial data mining. IEEE Transactions on Knowledge and Data Engineering, 14(5), 1003–1016.
Ordonez, C. & Omiecinski, E. (2002). FREM: Fast and robust EM clustering for large data sets. In Proceedings of the eleventh international conference on Information and knowledge management. ACM.
Rodriguez, A., & Laio, A. (2014). Clustering by fast search and find of density peaks. Science, 344(6191), 1492–1496.
Rhouma, M. B. H., & Frigui, H. (2001). Self-organization of pulse-coupled oscillators with application to clustering. IEEE Transactions on Pattern Analysis and Machine Intelligence, 23(2), 180–195.
Samaria, F. S., & Harter, A. C. (1994). Parameterisation of a stochastic model for human face identification. In Applications of computer vision. Proceedings of the second IEEE workshop on. IEEE.
Smith, A., et al. (2013). Sequential Monte Carlo methods in practice. Berlin: Springer.
Sneath, P. H. A., & Sokal, R. R. (1962). Numerical taxonomy. Nature, 193, 855–860.
Von Luxburg, U. (2007). A tutorial on spectral clustering. Statistics and Computing, 17(4), 395–416.
Wang, G. & Song, Q. (2016). Automatic clustering via outward statistical testing on density metrics.
Wang, W., Yang, J., & Muntz, R. (1997). STING: A statistical information grid approach to spatial data mining. In VLDB.
Wang, W., Yang, J., & Muntz, R. (1999). STING+: An approach to active spatial data mining. In Data engineering, 1999. Proceedings of the 15th international conference on. IEEE.
Ward, J. H, Jr. (1963). Hierarchical grouping to optimize an objective function. Journal of the American statistical Association, 58(301), 236–244.
Xu, R., & Wunsch, D. (2005). Survey of clustering algorithms. IEEE Transactions on Neural Networks, 16(3), 645–678.
Zhang, X., et al. (2010). K-AP: Generating specified K clusters by efficient affinity propagation. In Data mining (ICDM), 2010 IEEE 10th international conference on. IEEE.
Zhang, T., Ramakrishnan, R., & Livny, M. (1996). BIRCH: An efficient data clustering method for very large databases. In ACM SIGMOD record. ACM.
Zhu, Q., et al. (2014). A clustering algorithm based on natural nearest neighbor. Journal of Computational Information Systems, 10(13), 5473–5480.
Acknowledgements
The authors would like to greatly thank the editors and the anonymous reviewers for their insightful and helpful comments, which resulted in substantial improvements to this work. This research was supported by the National Nature Science Foundation of China (No. 61272194).
Author information
Authors and Affiliations
Corresponding author
Additional information
Editor: João Gama.
Rights and permissions
About this article
Cite this article
Huang, J., Zhu, Q., Yang, L. et al. QCC: a novel clustering algorithm based on Quasi-Cluster Centers. Mach Learn 106, 337–357 (2017). https://doi.org/10.1007/s10994-016-5608-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10994-016-5608-2