Abstract
Winter oilseed rape (Brassica napus) is an important crop in the Czech Republic and Poland. Clubroot disease caused by the pathogen Plasmodiophora brassicae is a serious and still-growing problem for oilseed rape growers in both countries. The aim of this study was to evaluate the pathotype composition of P. brassicae populations from the Czech Republic and Poland, according to the three evaluation systems, and to determine soil inoculum loads for representative fields via traditional end-point PCR as well as quantitative PCR analysis. There were considerable differences between the populations of P. brassicae from both countries, and the number of pathotypes varied depending on the evaluation system and the threshold used to distinguish susceptible vs. resistant plant reactions. This is the first study comparing the effect of different thresholds. Using an index of disease (ID) of 25 % to distinguish susceptible vs. resistant reactions, there was a total of seven pathotypes identified based on the differentials of Williams, five with the system of Somé et al., and 18 with the European Clubroot Differential (ECD) set. However, based on a threshold of 50 %, there were nine pathotypes according to the evaluation system by Williams, four based on the differentials of Somé et al., and 15 with the ECD set. Changing of the thresholds led to the reclassification of some pathotypes. Several pathotypes were common in both countries. High amounts of pathogen DNA were found in many of the field soils analysed by quantitative PCR. There was a weak correlation between soil pH and infestation of P. brassicae for the Polish soils.
Similar content being viewed by others
Introduction
Clubroot, caused by Plasmodiophora brassicae Woronin, is a serious soilborne disease of the Brassicaceae family. For many years the disease occurred worldwide on vegetable brassicas (Dixon 2009). In recent years clubroot has become a serious problem on farms growing spring and winter oilseed rape (Brassica napus L.) (Dixon 2009). Clubroot occurs throughout the Czech Republic, mainly on cruciferous vegetables (Rod 1994; Kopecky et al. 2012; Chytilova and Dusek 2007), as well as in Poland, where it was found on vegetables and oilseed rape (Korbas et al. 2014).
Winter oilseed rape is the second most important crop in the Czech Republic and is grown on about 400 000 ha (Czech Statistical Office 2014). The first occurrence of clubroot on winter oilseed rape was recorded in 2011, with a serious infestation found on 44 farms, mainly in the north and northeast of the country. Since then, the disease has spread in the oilseed rape growing areas (Kazda et al. 2013). In Poland, the acreage of oilseed rape has averaged about 800 000 ha in recent years (Dmochowska 2014). For years, clubroot was observed in vegetable crops, but was noticed only incidentally in oilseed rape. However, clubroot has been found with increased frequency on oilseed rape, causing concern among growers (Korbas et al. 2009). Currently, the disease occurs in large areas cultivated to oilseed rape, primarily in the northern and south-western regions of the country; however, clubroot infestations extend to the borders with Belarus and the Ukraine (Jedryczka et al. 2014; Korbas et al. 2014). The main agricultural plain (Mazovia, Greater Poland) is relatively free of the disease, likely because of lower rainfall, better regulation of soil pH, and good agricultural practices including appropriate crop rotation (Jedryczka et al. 2013).
The symptoms of clubroot include gall formation on the roots, leaf discoloration in the autumn, and stunting and wilting of the shoots in spring. These symptoms can result in lower yields and crop quality (Karling 1968; Wallenhammar 1999). Many management methods are used to curtail pathogen spread and reduce disease severity, but none of these is sufficiently effective to eliminate the pathogen. An integrated approach is therefore needed. Breeding of cultivars for resistance to clubroot is one of the most desirable disease management strategies. These activities should be based on screening of pathotypes and knowledge of their distribution, to obtain varieties resistant to the prevalent pathotypes (Diederichsen et al. 2009).
Several systems have been developed to classify pathotypes of P. brassicae. Among the most commonly used are the classification system of Williams (1966), which consists of four differential hosts that can distinguish a maximum of 16 pathotypes, the European Clubroot Differential (ECD) set (Buczacki et al. 1975) consisting of 15 differentials in three subsets of hosts (B. rapa, B. napus, and B. oleracea), and the differential system of Somé et al. (1996), with three hosts.
Pathotype screening in the Czech Republic was initiated by Rod (1994) with inoculum originating from vegetable production. The study revealed seven pathotypes as classified by Williams (1966), with pathotype 7 as most common. Use of the ECD set (Buczacki et al. 1975) allowed identification of 35 pathotypes in the Czech Republic, with 16/15/31 as the most frequent pathotype (Rod 1994). Similarly, a study on B. oleracea performed in Poland by Robak (1991) showed pathotype 7 as most common, according to the classification of Williams (1966), followed by race 2 and race 4 in minor quantities.
The Czech Republic and Poland share a common border, oilseed rape is intensively cultivated in both countries, and the problem of clubroot is shared as well. However, no detailed information is available on the pathotype composition of the P. brassicae populations from Czech and Polish oilseed rape fields. The objective of this study was to characterize populations of the pathogen from various soil samples collected from oilseed rape fields across the Czech Republic and Poland. It was hypothesized that the pathotype structure of P. brassicae might differ between the countries. The types of soil, the climate and the species and cultivars of different brassicas differed greatly between the two countries over many years, and might have influenced the distribution of the pathogen and the development of different pathotypes. Furthermore, the pH and P. brassicae inoculum loads of the sampled soils were measured to study the correlation between these two parameters.
Materials and methods
Terminology
The terminology used is as defined by Strelkov et al. (2006). A population of P. brassicae refers to a collection of pathogen resting spores obtained from an infested field and used to inoculate a set of differential hosts.
Pathogen collection
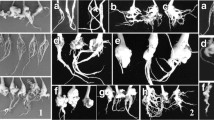
Soil samples were collected from 14 clubroot-infested fields of oilseed rape in the Czech Republic (CZ), selected based on damage and yield losses in 2012 (Table 1, Fig. 1a). Samples consisted of five random subsamples of ca. 5 kg each, collected from within infested patches. Subsamples were collected to a depth of 20 cm. The soil samples were air dried for two weeks, thoroughly homogenized, sieved (2 mm mesh) and used for DNA extraction. Polish soil samples (ca. 5 kg each) were collected in 2010–2012 from 16 fields (Table 1), 14 of which represented the areas where clubroot symptoms were observed most frequently on oilseed rape (Fig. 1b). The remaining two samples originated from the central-west region of the agricultural plain, where clubroot symptoms on oilseed rape are much less common. Prior to testing, the soil pH was measured with soil pH-meters (Hanna instruments, HI 98128 for the Czech soil samples; Draminski Ltd. for the Polish soil samples). Briefly, 100 g of soil was diluted in 200 ml of distilled water and the pH was measured. Twenty seeds of the Chinese cabbage cv. ‘Granaat’ (Brassica rapa subsp. pekinensis), considered as universally susceptible host to clubroot, were seeded in 5 kg of each soil sample to trap P. brassicae populations. Plants were grown in four 6 × 6 cm plastic pots per each sample, with five seeds per pot (Poland) in two series or in 25 4 × 4 cm plastic pots with 1 seed per pot in two series (Czech Republic). They were kept for 6 weeks under greenhouse conditions at 21 °C, with frequent watering and fertilization (‘Florovit’ applied once per 2 weeks at 0.2 % concentration). The galled roots were harvested, slowly dried at room temperature and then used for inoculation of the differential hosts according to Strelkov et al. 2006.
Differential hosts
Pathogen populations from the Czech Republic and Poland were evaluated for their pathotype designation on the differentials of the ECD set (Buczacki et al. 1975), Somé et al. (1996), and Williams (1966) (Tables 1 and 2). Some of the host genotypes, including ECD 10, 11, and 13, are common to both the Williams (1966) and ECD differential sets. Similarly, two of the differentials of Somé et al. (1996) are part of the ECD set (ECD 06 and ECD 10). The rutabaga (Brassica napus var. napobrassica) ‘Laurentian’ and spring oilseed rape cultivar ‘Brutor’ are unique to the differential sets of Williams (1966) and Somé et al. (1996), respectively. Seeds of all differentials, except ‘Laurentian’ and ‘Brutor’, were obtained from Horticulture Research International, Genetic Resources Unit (Wellesbourne, Warwick, UK). Seeds of ‘Laurentian’ and ‘Brutor’ were acquired from the Crucifer Genetics Cooperative (Madison, WI, USA) and the Leibniz Institute of Plant Genetics and Crop Plant Research (IPK) Genebank (Gatersleben, Germany), respectively.
Host inoculation
Resting spores were extracted from galled root tissue by the method of Tewari et al. (2005) with some modifications, as described by Strelkov et al. (2006). Briefly, 4 g of clubbed roots were ground in 50 ml of distilled water with a blender. The suspension was then filtered through six layers of cheese-cloth. Resting spore extractions were conducted at room temperature (20 °C). The spore suspension was examined by haemocytometer (VWR, Mississauga, Ontario) and the resting spore concentration adjusted to a final concentration of 1 × 107 spores/ml, which was used for inoculation of the differential hosts. Seeds of the differentials were germinated in Petri dishes for five days. Seedlings were inoculated by dipping the rootlets for 30 s in spore suspension and planted in 4 × 4 cm plastic pots, which were filled with either Sunshine Professional Growing Mix (Sun Gro Horticulture Canada Ltd) or soil mixed with peat with pH 5.5 (Biovita Ltd., Poland) in a 2:1 ratio. Pots were kept in a greenhouse for 5 weeks; the soil was saturated with water for the first week after inoculation and then fertilized and watered as needed, with the temperature maintained at 21 °C. Twelve seedlings of each host line were inoculated with each population of P. brassicae, with treatments replicated three times for a total of 36 plants of each differential per treatment.
Disease assessment
Five weeks after inoculation, plants were removed from the pots and washed with water. The roots were evaluated for clubroot severity on a 0–3 scale (Kuginuki et al. 1999), where: 0 - no galling, 1 - a few small galls, 2 - moderate galling on the main and lateral roots, and 3 - severe galling, the root is totally deformed. An index of disease (ID) was calculated for each differential host using the formula of Horiuchi and Hori (1980) as modified by Strelkov et al. (2006):
Where: ∑ is the total sum, n is the number of plants in class, N is the total number of plants, 0, 1, 2, 3 and are the symptom severity classes.
The results shown represent the mean ID of each host over the three replicates for each P. brassicae population. Two thresholds were compared; ID 25 % as described by Somé et al. (1996) and ID < 50 % with the 95 % confidence interval (CI) not exceeding 50 %, as proposed by LeBoldus et al. (2012). Statistical analysis was carried out using Statistica v. 7.0 (StatSoft Inc., USA) and Microsoft Excel 2010 (Microsoft, Inc., USA).
PCR and quantitative PCR analysis of infested soil samples
Total genomic DNA was extracted from all soil samples collected from infested fields with a FastDNA Spin kit for soil (MP Biomedicals) as per the manufacturer’s instructions. An aliquot of 250 mg of soil was used per sample, with three replicates.
Infestation by P. brassicae was detected by conventional PCR analysis as described by Cao et al. (2007). The amplification products were run on 1 % agarose gels stained with SYBR Safe (Invitrogen) and visualized under UV light in a Syngene BioImaging System (Synoptics Inc.).
Quantitative PCR analysis was performed on diluted DNA (1/10 v/v) with sd-H2O to avoid inhibition of the reaction. The quantitative PCR amplification was conducted according Rennie et al. (2011). Briefly, qPCR amplifications were carried out in a StepOnePlus Real Time PCR System (Applied Biosystems) in a 10 μl volume reaction (2.5 μl template DNA, 2.5 μl of 3.2 mM DR1F and DR1R primer mix, and 5 μl Dynamite qPCR Mastermix (Molecular Biology Service Unit, University of Alberta, Edmonton, CA), which contained SYBR Green (Molecular Probes) as the detection dye. The reaction conditions consisted of 95 °C for 2 min (initial denaturation step), followed by 35 cycles of 95 °C for 15 s (denaturation step) and 60 °C for 60 s (annealing, extension and fluorescence measurement step). A standard curve for the quantification of P. brassicae spores was generated with DNA isolated from known quantities of spores, which had been purified as per Castlebury et al. (1994) with modifications made by Cao et al. (2007). Briefly, 100 g of galled tissue and 500 ml of water were blended, and the homogenate passed through a 450 μm metallic sieve. The resting spores were isolated by centrifugation in a 50 % sucrose gradient, quantified with a haemocytometer (VWR), and adjusted to 1 × 108 resting spores/ml. Total DNA for the standard curve was extracted and serially diluted with sd-H2O at 10-fold intervals down to a dilution of 1 × 102 resting spores/ml. DNA from each dilution in the series was used as a template in the qPCR analysis according to Rennie et al. 2011. The correlation between the amount of pathogen DNA detected and the soil pH was determined for the Czech and Polish samples.
Results
In total, 14 isolates from the Czech Republic and 16 isolates from Poland were evaluated for pathotype classification. Within clubroot populations, the disease severity ranged from low (IDs ≤ 10 %), through moderate (IDs 10–40 %) to high (IDs ≥ 40 %); in most infested fields, however, the severity was low to moderate.
Populations from the Czech Republic
The universally susceptible genotype, ECD 05, was highly susceptible to all of the populations tested (Table 3). Similarly, populations 1–6 caused very high levels of disease on the fodder rape (B. napus var. napus) hosts ECD 07-ECD 09, with IDs ranging from 10 % ± 5 % to 100 % ± 0 % (Table 3). Most of the other B. napus hosts were also highly susceptible; however, the rutabaga ‘Laurentian’ had low levels of infection. The spring oilseed rape ‘Brutor’ developed IDs ranging from 19 % ± 7 % to 100 % ± 0 % in response to inoculation. The reaction of ECD 10 varied widely, with IDs ranging from 0 % ± 0 % to 100 % ± 0 %. The reactions of the B. oleracea cultivars tested were similar to those of the B. napus genotypes (Table 3). Inoculation of ECD 11 and ECD 12 resulted in the development of IDs from as low as 2 % ± 1 % to as high as 90 % ± 6 %. On ECD 15, IDs ranged from 0 % ± 0 % to 52 % ± 2 %, indicating that this genotype was moderately resistant to P. brassicae. High ID values were observed on ECD 13, which were higher than for ECD 11, 12 or 15, and ranged from a low of 32 % ± 1 % to a high of 100 % ± 0 %. Thus, this host appeared to be susceptible to most of the P. brassicae populations tested. Similar ID values were observed on ECD 14; on this cabbage, IDs ranged from 3 % ± 2 % to 99 % ± 1 %. The overall trend showed that the most severe symptoms were associated with populations 1–6 from the same region in the north of the Czech Republic.
The number of pathotypes amongst the P. brassicae populations collected in the Czech Republic varied depending on the classification system and plant reaction threshold used (Tables 1 and 2). Using the threshold of 50 % and classification of Williams (1966), there were six pathotypes: 3, 4 and 9 (1 population each, 7.14 % of the tested populations), 2 (14.28 %), 6 (21.44 %) and 7 (42.86 %). The differentials of Somé et al. (1996) distinguished four pathotypes: P1, P3, P4 and P5, with P3 being predominant (71.4 %). The highest diversity of pathotype classification was obtained with the ECD set, identifying eight pathotypes: 16/14/15 (28.6 % of the tested populations), 16/14/13 (14.3 %), 16/14/31 (14.3 %), 16/2/14 (14.3 %) and also one population (7.1 %) each of following pathotypes: 16/8/4, 16/18/15, 16/14/12 and 16/2/15 (Table 2). Using the threshold of 25 % and the classification of Williams (1966), there were five pathotypes: 1, 2, 4 and 6 (2 populations each, 14.28 % of tested populations) and 7 (42.86 %). According to the system of Somé et al. (1996), the pathotype classifications were: P1, P2, P3, P4 and P5, with P2 being predominant (42.86 %). The highest pathotype diversity with the 25 % threshold was reached with the ECD set, which distinguished 10 pathotypes: 16/15/15 (21.42 %), 16/14/31 and 16/15/31 (14.3 % each) and one population (7.14 %) each of following pathotypes: 16/14/4, 16/14/14, 16/31/12, 16/22/31, 16/22/30, 16/31/15 and 16/3/15 (Table 1). Infested soil samples were tested for the presence of the pathogen by conventional PCR and spore loads were measured by qPCR analysis (Table 5). Two runs of qPCR were performed. Soil samples 2–5, 7–9, 11, 12 and were 14 tested positive in conventional PCR, while samples 1, 6, 13 were negative. The results from qPCR analysis showed spore concentrations that ranged from 3 × 104 spores/g soil in sample 9 (Pohledy) to 4.9 × 106 spores/g soil in sample 4 (Modlibohov). Most of the other samples contained around 1 × 105 spores/g soil. The amount of inoculum in samples 1, 8 and 13 was below the detection level (1 × 103 spores/g soil) (Table 5).
Populations from Poland
The results of conventional PCR and qPCR analysis were comparable. The results of both qPCR runs were consistent, with the exception of sample PL14 and, to some extent, sample PL2, where the spore concentration detected in two runs differed by two orders of magnitude. The highest number of resting spores detected was 8 × 107 spores/g soil (Tables 3, 4 and 5). The number of pathotypes identified in varied depending on the differential system and threshold used (Tables 1 and 2). Using the 50 % threshold and classification by Williams (1966), eight pathotypes was be identified, including pathotypes 6 and 7 (4 populations each, 25 % of the total populations tested), pathotype 4 (18.75 %) and 3, 9, 10, 12, 16 (1 population each, 6.25 %). Classification with the system of Somé et al. (1996) revealed only two pathotypes: P1 and P3 in comparable quantities, with slightly more of P3 (56 %). As with the Czech populations, the highest pathotype diversity in Poland was found using the ECD set, with the following nine pathotypes identified: 16/14/15 (18.75 % of the tested populations in Poland), 16/31/8 and 16/14/12 (12.5 % each), and five populations (6.25 %) of each of the following pathotypes: 16/7/28, 16/31/29, 16/14/30, 16/31/14, 16/2/14 and 16/14/15 (Table 2). Using the threshold of 25 % and classification system of Williams (1966), five pathotypes could be defined: pathotypes 3 and 6 (one population each, 6.25 %), pathotype 9 (12.5 %), pathotype 4 (31.25 %), and pathotype 7 (43.75 %). The differentials of Somé et al. (1996) distinguished two pathotypes, P1 and P3 (56 %). The highest number of pathotypes was identified by the ECD using the 25 % threshold. There were nine pathotypes: 17/31/31 (25 % of all tested populations), 16/14/15 (18.75 %), 16/31/8 and 16/14/31 (two populations each, 12.5 %) and one population (6.25 %) of each of following pathotypes: 16/7/31, 16/31/31, 16/31/14, 16/6/31, 16/14/30 (Table 1).
The differentials ECD 02 - ECD 04 were resistant to all tested Polish populations of P. brassicae (Table 2b). In contrast, the genotype ECD 05 was moderately or highly susceptible to all of the Polish populations. The ID values for the pathogen populations ranged from 62 % ± 4 % to 64 % ± 2 % for PL13 and PL8, respectively, to ID = 100 % ± 0 % for PL5. In the case of the populations PL2 and PL10, the reaction of ECD 05 was intermediate (ID 46 % ± 1 %). Four of the populations from Poland (PL4, PL8, PL10 and PL16) caused intermediate reactions on ECD 14 and ‘Brutor’, while the remainder were highly virulent. The other ECD genotypes, as well as the cultivar ‘Laurentian’, exhibited a wide variety of responses to inoculation with P. brassicae populations collected in Poland. Some of these isolates caused severe clubroot symptoms, whereas others caused no or very small galls on the roots (Table 4).
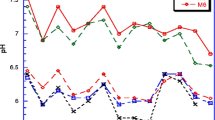
The correlation between soil pH and the concentration of P. brassicae resting spores in the soil samples collected in Poland was weakly negative (−0.495). No spores were detected in three out of four soils of pH equal to or < 6.8. However, in one case, a soil of pH 6.9 contained a substantial amount of spores (4.7 × 106 resting spores/g soil).
Discussion
The results confirm the hypothesis that the P. brassicae pathotype structure in the Czech Republic and Poland differs greatly. The number of pathotypes in the populations of P. brassicae collected in both countries varied depending on the host differential system and the threshold value used to distinguish susceptible vs. resistant host reactions. It can be partly explained by different origin of the isolates, obtained from varieties differing with disease resistance and other characters, as well as different environmental conditions (Russell 2013). When the threshold of ID 25 % (Somé et al. 1996) was used along with the classification of Williams (1966), there were five pathotypes found in both countries, but only three of these (4, 6 and 7) were common to both. Pathotype 7 was predominant and in both Poland and the Czech Republic, representing around 43–44 % of the studied populations. The differences between the countries were even greater when the pathotype classification was based on the threshold of ID < 50 % with CI of 95 %, as described by LeBoldus et al. (2012). In this case, there were eight pathotypes in Poland and six in the Czech Republic, with only five in common to both countries. In this case, pathotypes 6 and 7 both constituted 25 % of the pathogen populations from Poland, whereas in the Czech Republic pathotype 7 was more common (43 %) than pathotype 6 (29 %).
The change of the threshold from ID 25 % to 50 % had a direct effect on pathotype classification; pathotype 1, based on the method described by Somé et al. (1996), was re-classified to 3 or 9, pathotype 4 was reclassified to 7, 10 or 12 and the pathotype 9 was changed to 16. The most common reclassification (occurring six times) was from the pathotype 7 to 6, at 25 % and 50 % threshold values, respectively. Collectively, a total of nine pathotypes were found in the Czech Republic and Poland using Williams’ (1966) classification, with pathotype 7 representing 33 % of all isolates tested from both countries, followed by pathotype 6 (23 %), pathotype 4 (17 %) and other less common pathotypes. This greatly differs from the situation reported in Canada, where the main pathotype found on oilseed rape (canola) was pathotype 3 (Cao et al. 2009), which in Poland and the Czech Republic was very rare. Nonetheless, some of the other pathotypes were common to Canada, the Czech Republic and Poland, including pathotypes 2, 4 and 6 (Cao et al. 2009). In Canada, pathotype 6 was associated only with cruciferous vegetables. In Poland and the Czech Republic, however, it is likely that the pathotypes found on oilseed rape are similar to those of the vegetable brassicas. In Poland, a high incidence of pathotype 7 was reported earlier by Robak (1991) in B. oleracea, in addition to pathotypes 2 and 4. The same situation has also been observed in the Czech Republic, where pathotype 7 was found most frequently in cruciferous vegetables (Rod 1994).
Only two pathotypes were identified in Poland based on the classification system of Somé et al. (1996), regardless of the threshold used to distinguish susceptible or resistant reactions to clubroot. However, in the Czech Republic the variability of pathotypes as assessed on this system was higher. Based on a threshold of 25 % ID, five pathotypes (P1–P5) were found, though changing the threshold to 50 % resulted in the reclassification of pathotype P2 to P3 (6 times) and P1 to P3 (1 time). Using the evaluation system of Somé et al. (1996), 67 % of the populations of P. brassicae in these two countries were classified as P3, whilst 27 % were P1 and a small percentage belonged to P2, P4 or P5. This is again is different from the Canadian situation, where pathotype P2 is dominant (Cao et al. 2009). According to preliminary results from a survey in Germany, pathotypes P1 and P3 are predominant there as well (Somé et al. 1996). In Germany, Lueders et al. (2011) noted the identification mainly of P1 in more northerly collections of P. brassicae and P3 in southern collections, which was similar to the findings in Poland and the Czech Republic.
A heterogeneous mix of pathotypes was identified with the ECD set. Again, the final classification depended somewhat on the threshold value used to distinguish resistant vs. susceptible reactions. Changing the threshold from 25 to 50 % resulted in the reclassification of 77 % of the studied isolates of P. brassicae (23 out of 30). For ID 25 %, there were nine and 10 pathotypes in Poland and the Czech Republic, respectively, with only one (16/14/31) common to both countries. Based on an ID < 50 % with the 95 % CI not exceeding 50 % (LeBoldus et al. 2012), 15 pathotypes in total were identified, with four in common: 16/2/14, 16/14/12, 16/14/15 and 16/31/8. The most common pathotype was 16/14/15 and it constituted over 23 % of the whole population. In Canada, pathotype ECD 16/15/12 was predominant on oilseed rape, but this pathotype was not found in the Czech Republic or Poland. In Poland, two isolates from distant locations (PL4 and PL10), both assigned as pathotype 16/31/8, showed unusually weak reactions (46 ± 1 %) on ECD 05. This accession was introduced originally by Buczacki et al. (1975) in the ECD differential set, due to its high susceptibility to all P. brassicae populations studied at that time. By now, ECD 05 served as susceptible control. One may speculate that the current fast expansion of P. brassicae on oilseed rape gives rise to the development of ‘new variants’ of this pathogen, which are not yet the new pathotypes according to the current evaluation systems, but they surely differ from the pathogen variants described previously. In 1970s the pathogen was found only or mainly on vegetable brassicas, which grow in some areas of the country, on more fertile soils and under different agrotechnical treatments.
It is likely that an even greater diversity of pathotypes would have been identified if single-spore isolates, rather than populations, had been examined. In a study several years ago, the pathotype classification of single-spore isolates of P. brassicae was compared with that of populations of the pathogen, and a greater diversity of pathotypes was found when the single-spore isolates were examined (Somé et al. 1996). Similarly, Xue et al. (2008) reported that more pathotypes were showed when single-spore isolates were evaluated rather than mother populations, suggesting that more rare pathotypes may have been masked in heterogeneous mixes of resting spores. The distinct pathotype composition in the main oilseed growing areas of east-central Europe and Canada has important implications for clubroot management, since resistance sources will have to be evaluated against the most common pathotypes in each region, and will have to be rotated to prevent shifts in the pathogen population.
The correlation between lower soil pH and higher P. brassicae infestation of the soil was relatively weak in Poland and non-existent in the Czech Republic. The average pH of the soils from the Czech Republic was, however, 6.7, the lowest pH value was 6.4 and the pH above 7.0 was observed in two soils only. Therefore, such lack of correlation was to be expected.
In Alberta, Canada, a weak negative correlation was found between these parameters (Gossen et al. 2013). In contrast, in previous studies strong negative correlations were found between P. brassicae and soil pH (Webster and Dixon 1991; Dixon 2009). The absence of a strong correlation in the present study, however, suggests that other factors, such as the length of rotation and amount of soil moisture could have a greater impact on clubroot development than pH.
In the current study, the highest P. brassicae resting spore concentration was detected in a Polish soil sample, collected near Tuczno, where the number of spores exceeded 8 × 107 spores/g dry soil. In the Czech Republic, the highest concentration was 4 × 106 spores/g dry soil, found in a sample from Modlibohov. In most cases, the results from independent determinations of resting spore concentrations in soil samples by qPCR analysis were consistent; they were at least in the same order of magnitude or closer. In a few cases (e.g., CZ10 and PL14) there was a greater difference between independent runs of the same sample, which might reflect the heterogeneous distribution of resting spores in the soil or perhaps artifacts such as sample processing. The results from the conventional PCR and qPCR analysis were generally consistent, but there also were a few cases in which a sample testing negative by conventional PCR still had detectable levels of inoculum. This likely reflects the fact that the qPCR method is more robust and has greater sensitivity. Similar results were reported by Wallenhammar et al. (2012), where the detection limit in soil samples corresponded to 500 resting spores/g soil.
The results of this study, which revealed a diverse pathotype composition in P. brassicae populations from oilseed rape fields in the Czech Republic and Poland, as well as high levels of resting spore inoculum in many of the fields, indicate that the management of clubroot will be challenging in east-central Europe, and will require active screening for host resistance along with practices to reduce soil spore loads. The final classification of the pathotypes depends to some extent on the threshold selected to differentiate between resistant and susceptible host reactions, making it difficult to compare the results between different studies, unless a joint evaluation system is agreed upon and implemented.
References
Buczacki, S. T., Toxopeus, H., Mattusch, P., Johnston, T. D., Dixon, G. R., & Hobolth, L. A. (1975). Study of physiologic specialization in Plasmodiophora brassicae: proposals for attempted rationalization through an international approach. Transaction of British Mycological Society, 65, 295–303.
Cao, T., Tewari, J. P., & Strelkov, S. E. (2007). Molecular detection of Plasmodiophora brassicae, causal agent of clubroot of crucifers, in plant and soil. Plant Disease, 91, 80–87.
Cao, T., Manolii, V. P., Hwang, S. F., Howard, R. J., & Strelkov, S. E. (2009). Virulence and spread of Plasmodiophora brassicae [clubroot] in Alberta, Canada. Canadian Journal of Plant Pathology, 31, 321–329.
Castlebury, L. A., Maddox, J. V., & Glawe, D. A. (1994). A technique for the extraction and purification of viable Plasmodiophora brassicae resting spores from host root tissue. Mycologia, 86, 458–460.
Chytilova, V., & Dusek, K. (2007). Metodika testováni odolnosti brukvovitých plodin k nádorovitosti. Metodika pro praxi (in Czech). Praha: Výzkumný ústav rostlinné výroby.
Czech Statistical Office (2014). http://csugeo.i-server.cz/csu/2015edicniplan.nsf/p/270141-15. Accessed 30 July 2015.
Diederichsen, E., Frauen, M., Linders, E. G. A., Hatakeyama, K., & Hirari, M. (2009). Status and perspectives of clubroot resistance breeding in crucifer crops. Journal of Plant Growth Regulation, 28, 265–281.
Dixon, G. R. (2009). The occurrence and economic impact of Plasmodiophora brassicae and clubroot disease. Journal of Plant Growth Regulation, 28, 194–202.
Dmochowska, H. (2014). Statistical yearbook of agriculture and rural areas. Warsaw: Central Statistical Office (in Polish).
Gossen, B. D., Kasinathan, H., Tsao, R., Manolii, V. P., Strelkov, S. E., Hwang, S. F., & McDonald, M. R. (2013). Interaction of pH and temperature affect infection and symptom development of Plasmodiophora brassicae in canola. Canadian Journal of Plant Pathology, 35, 294–303.
Horiuchi, S., & Hori, M. (1980). A simple greenhouse technique for obtaining high levels of clubroot incidence. Bulletin of the Chugoku National Agricultural Experiment Station Series E (Environment Division), 17, 33–55.
Jedryczka, M., Korbas, M., Jajor, E., Danielewicz, J., & Kaczmarek, J. (2013). The occurrence of Plasmodiophora brassicae in agricultural soils in Wielkopolska region, in 2011–2012 [in Polish with English abstract]. Progress in Plant Protection/Postepy w Ochronie Roslin, 53, 774–778.
Jedryczka, M., Kasprzyk, I., Korbas, M., Jajor, E., & Kaczmarek, J. (2014). Infestation of Polish agricultural soils by Plasmodiophora brassicae along the Polish-Ukrainian border. Journal of Plant Protection Research, 54, 238–241.
Karling, J. S. (1968). The Plasmodiophorales. New York: Hafner Publishing Company.
Kazda, J., Ricarova, V., Prokinova, E., Grimova, L., & Baranyk, P. (2013). Nadorovitost korenu brukvovitych (původce Plasmodiophora brassicae) ohrozuje ozimou repku v Ceske republice (in Czech). Uroda, 6, 28–32.
Kopecky, P., Dolezalova, I., Duchoslav, M., & Dusek, K. (2012). Variability in resistance to clubroot in European cauliflower cultivars. Plant Protection Science, 48, 156–161.
Korbas, M., Jajor, E., & Budka, A. (2009). Clubroot (Plasmodiophora brassicae) – a threat for oilseed rape. Journal of Plant Protection Research, 49, 446–451.
Korbas, M., Jajor, E., Kaczmarek, J., Perek, A., & Jedryczka, M. (2014). Infestation of Polish agricultural soils by Plasmodiophora brassicae on the Polish-Belarussian border in Podlasie province. Integrated Control in Oilseed Crops Bulletin, 104, 167–171.
Kuginuki, Y., Yoshikawa, H., & Hirai, M. (1999). Variation in virulence of Plasmodiophora brassicae in Japan tested with clubroot- resistant cultivars of Chinese cabbage (Brassica rapa L. ssp. pekinesis). European Journal of Plant Pathology, 105, 327–332.
LeBoldus, J. M., Manolii, V. P., Turkington, T. K., & Strelkov, S. E. (2012). Adaptation to Brassica host genotypes by a single-spore isolate and population of Plasmodiophora brassicae (clubroot). Plant Disease, 96, 833–838.
Lueders, W., Abel, S., Friedt, W., Kopahnke, D., & Ordon, F. (2011). Auftreten von Plasmodiophora brassicae als Erreger der Kohlhernie im Winterrapsanbau in Europa sowie Identifizierung, Charakterisierung und molekulare Kartierung neuer Kohlhernieresistenzgene aus genetischen Ressourcen. Julius-Kühn-Archiv, 430, 40–43.
Rennie, D. C., Manolii, V. P., Cao, T., Hwang, S. F., Howard, R. J., & Strelkov, S. E. (2011). Direct evidence of surface infestation of seeds and tubers by Plasmodiophora brassicae and quantification of spore load. Plant Pathology, 60, 811–819.
Robak, J. (1991). Variability of the pathotypes of Plasmodiophora brassicae Wor. present in Poland and their pathogenicity to the cultivars and breeding lines of Brassica oleracea [in Polish]. Habilitation monograph. Skierniewice: Research Institute of Vegetable Crops.
Rod, J. (1994). The occurrence and distribution of pathotypes of Plasmodiophora brassicae Wor. in the Czech Republic and Slovakia. Ochrana Rostlin, 30, 171–182.
Russell, G. E. (2013). Plant breeding for pest and disease resistance. Studies in the agricultural and food sciences. Burlington: Elsevier Science.
Somé, A., Manzanares, M. J., Laurens, F., Baron, F., Thomas, G., & Ouxel, F. R. (1996). Variation for virulence of Brassica napus L., amongst Plasmodiophora brassicae collections from France and derived single-spore isolates. Plant Pathology, 45, 432–439.
Strelkov, S. E., Tewari, J. P., & Smith-Degenhardt, E. (2006). Characterization of Plasmodiophora brassicae populations from Alberta, Canada. Canadian Journal of Plant Pathology, 28, 467–474.
Tewari, J. P., Strelkov, S. E., Orchard, D., Hartman, M., Lange, R. M., & Turkington, T. K. (2005). Identification of clubroot of crucifers on canola (Brassica napus) in Alberta. Canadian Journal of Plant Pathology, 27, 143–144.
Wallenhammar, A. C. (1999). Monitoring and control of Plasmodiophora brassicae in spring oilseed brassica crops. Uppsala: Swedish University of Agricultural Sciences.
Wallenhammar, A. C., Almquist, C., Soderstrom, M., & Jonsson, A. (2012). In-field distribution of Plasmodiophora brassicae measured using quantitative real-time PCR. Plant Pathology, 6, 16–28.
Webster, M. A., & Dixon, G. R. (1991). Calcium, pH and inoculum concentration as factors limiting root hair colonization by Plasmodiophora brassicae Wor. Mycological Research, 95, 64–73.
Williams, P. H. (1966). A system for the determination of races of Plasmodiophora brassicae that infect cabbage and rutabaga. Phytopathology, 56, 624–626.
Xue, S., Cao, T., Howard, R. J., Hwang, S. F., & Strelkov, S. E. (2008). Isolation and variation in virulence of single-spore isolates of Plasmodiophora brassicae from Canada. Plant Disease, 92, 456–462.
Acknowledgments
The experimental work was funded by the National Research Centre in Poland (project N N310 298439), and by grant No. QJ1310227 from the Ministry of Agriculture of the Czech Republic and grant No. 20142031 from the Czech University of Life Sciences, Prague. We thank to the Union of Oilseed Growers and Processors, who provided soil samples from different regions of the Czech Republic. We also would like to thank the lab members in our institutes who provided technical assistance, and Dr. William Truman (Institute of Plant Genetics, Polish Academy of Sciences) for proof-reading this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Veronika Řičařová and Joanna Kaczmarek to be regarded as joint first authors of this paper
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Řičařová, V., Kaczmarek, J., Strelkov, S.E. et al. Pathotypes of Plasmodiophora brassicae causing damage to oilseed rape in the Czech Republic and Poland. Eur J Plant Pathol 145, 559–572 (2016). https://doi.org/10.1007/s10658-016-0939-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-016-0939-1