Abstract
X inactivation, the transcriptional silencing of one of the two X chromosomes in female mammals, achieves dosage compensation of X-linked genes relative to XY males. In eutherian mammals X inactivation is regulated by the X-inactive specific transcript (Xist), a cis-acting non-coding RNA that triggers silencing of the chromosome from which it is transcribed. Marsupial mammals also undergo X inactivation but the mechanism is relatively poorly understood. We set out to analyse the X chromosome in Monodelphis domestica and Didelphis virginiana, focusing on characterizing the interval defined by the Chic1 and Slc16a2 genes that in eutherians flank the Xist locus. The synteny of this region is retained on chicken chromosome 4 where other loci belonging to the evolutionarily ancient stratum of the human X chromosome, the so-called X conserved region (XCR), are also located. We show that in both M. domestica and D. virginiana an evolutionary breakpoint has separated the Chic1 and Slc16a2 loci. Detailed analysis of opossum genomic sequences revealed linkage of Chic1 with the Lnx3 gene, recently proposed to be the evolutionary precursor of Xist, and Fip1, the evolutionary precursor of Tsx, a gene located immediately downstream of Xist in eutherians. We discuss these findings in relation to the evolution of Xist and X inactivation in mammals.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
X inactivation is a process whereby one of the two X chromosomes in female mammals is transcriptionally silenced early in embryogenesis, equalizing the dosage of X chromosome gene expression in males (XY) and females (XX) (Lyon 1961). In eutherian (placental) mammals, X inactivation is governed from a single cis-acting locus on the X chromosome, originally termed the X inactivation centre (Xic), and now identified as the X-inactive specific transcript gene (Xist). The Xist locus produces a large non-coding RNA that coats the X chromosome from which it is transcribed, triggering chromosome-wide silencing (for a recent review see Chow et al. 2005).
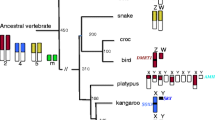
Despite recent advances in our understanding of the molecular mechanisms governing X inactivation, little is known about how the X inactivation process has arisen and developed during the evolution of mammals. Marsupials belong to a branch of mammals that diverged from eutherians about 180 million years ago (Woodburne et al. 2003). The marsupial X chromosome may therefore carry some features of the ancestral mammalian X chromosome. The euchromatic part of the marsupial X consists of only 3% of the total genomic euchromatin (versus 5% in eutherians), and corresponds to the long arm and the pericentric region of the human X (Graves 1996). As in eutherian mammals, dosage compensation of X-linked genes is achieved by X-chromosome inactivation. However, in marsupials X inactivation is paternally imprinted in all tissues (Sharman 1971). Moreover, chromosome silencing is incomplete, and tissue-specific (Cooper et al. 1993). Paternal imprinting of X inactivation resembles X inactivation in extraembryonic tissues of (some) eutherian mammals, and for this reason it has been supposed that marsupial X-chromosome inactivation may reflect an ancestral mammalian X-inactivation system and as such could provide insight into the evolution of X inactivation (discussed by Cooper et al. 1993). However, the molecular mechanisms of marsupial X inactivation remain relatively poorly characterized and a homologue of Xist has not been identified in studies to date (Graves & Westerman 2002, Koina et al. 2005).
As a step towards identifying a marsupial homologue of the Xist gene we set out to perform comparative X-chromosome mapping and X-linked sequence analysis of two species of American marsupials: South American opossum Monodelphis domestica and North American opossum Didelphis virginiana, focusing on the organization of the protein coding genes Chic1 and Slc16a2 that flank Xist in all eutherian mammals studied (Chureau et al. 2002).
A recent study has found that Chic1 and Slc16a2 are linked on chicken chromosome 4 and that they flank a group of protein-coding genes Fip1l2, Lnx3, Rasl11c, Uspl, and Wave4 that, based on sequence comparison, appear to represent the precursors of the eutherian Tsx, Xist, Enox, Ftx, and Cnbp2 genes, comprising the X inactivation centre (Duret et al. 2006). It was hypothesized that the eutherian Xic genes evolved due to pseudogenization of the cognate protein-coding genes. Moreover, opossum genes orthologous to the Lnx3 and Rasl11c of chicken were mapped on the X chromosome of M. domestica, and cloning of a cDNA encoding the M. domestica Lnx3 protein suggested that the noncoding Xist RNA has not evolved in marsupials (Duret et al. 2006). In this study we demonstrate that the Chic1 and Slc16a2 syntenic region is divided in the American marsupial species M. domestica and D. virginiana. We further show that in M. domestica the Lnx3 gene is associated with Chic1 and not with Slc16a2. These results are discussed in the context of understanding the evolution of the X chromosome and X inactivation.
Materials and methods
Cell cultures
The M. domestica long-term female fibroblast cell line FMDL (Nesterova et al. 1997) was maintained in a 1:1 mix of Dulbecco's modified Eagle medium and F12 medium. The D. virginiana long-term female kidney epithelium line OK (ATCC number CRL-1840) was maintained in minimal essential medium alpha. The media was supplemented with 10% fetal calf serum, 1 mM l-glutamine, 50 units/ml penicillin, and 50 µg/ml streptomycin. The media and supplements were purchased from Invitrogen. All cells were grown at 37°C in an atmosphere containing 5% CO2.
M. domestica and D. virginiana genomic BAC libraries
Three opossum genomic BAC libraries, VMRC-6 of M. domestica and OM and LBNL3 of D. virginiana, have been used in this study. High-density filters of VMRC-6, constructed at the Virginia Mason Research Center, and LBNL3, produced at Lawrence Berkeley National Laboratory, were available from CHORI BACPAC Resources (http://bacpac.chori.org/protocols.htm). The OM D. virginiana BAC library has been described previously (Evans et al. 2005).
Probes and screening of opossum BAC libraries
Primers for the Chic1, Slc16a2, Hprt, and Pgk1 genes were designed to the most conserved protein-coding regions found in the alignment of the corresponding gene sequences of mouse, human and, where available, marsupials. Primers for G6pd, Rbmx, and Sox3 were selected to unique sequences present in M. domestica trace database (http://trace.ensembl.org). These sequences are located immediately upstream or downstream of the genes. Probes for all genes were generated by PCR using M. domestica cDNA or the genomic DNA isolated from FMDL cells. The primer pairs, PCR product size, and templates used are listed in Table 1. For blot hybridization experiments the probes were labelled with [α32P]dCTP (Amersham) using a random primer labelling kit (Roche) according to the manufacturer's instructions. The high-density filters for the BAC libraries were screened in ChurchYGilbert hybridization buffer (0.5 M NaHPO4, pH 7.2; 7% SDS) at 65°C overnight. The filters were washed at 65°C in 2× SSC, three times in 2× SSC and 0.1% SDS, and exposed using Kodak XOMAT X-ray films for 6 h up to 1 week at −80°C.
Preparation of metaphase spreads and mechanically stretched chromosomes
Metaphase spreads were prepared according to standard methods. Colchicine-arrested metaphases were collected in culture medium and swelled in 0.2% KCl, 0.2% sodium citrate hypotonic solution for 12–18 min at room temperature. Cells were fixed in methanol:acetic acid (3:1), dropped onto a microscope slide and air-dried.
The mechanically stretched chromosomes were prepared as described (Haaf & Ward 1994). An aliquot of 103–106 mitotically active cells were harvested, washed in phosphate-buffered saline (PBS), and swelled in a hypotonic solution (10 mM HEPES, 30 mM glycerol, 1.0 mM CaCl2, and 0.8 mM MgCl2) for 10 min. Samples (0.5 ml) of the hypotonic cell suspension were centrifuged (Cytospin 2, Shandon) onto clean glass slides at 800 rpm for 4 min and fixed in 70% ethanol at room temperature for 30 min.
Preparation of microdissection probes of opossum X chromosomes
To generate a probe, 15 whole metaphase X chromosomes were dissected using an inverted microscope Axiovert 10 (Zeiss) with micromanipulator MR (Zeiss). The collected X chromosome copies were transferred in 40 nl of buffer solution with a siliconized micropipette tip for proteinase K treatment and than amplified by DOP-PCR withMW6primer (Rubtsov et al. 2000). The microdissected amplified DNA was labelled with biotin 16-dUTP (Roche) or digoxigenin-11-dUTP (Roche) in 20 additional PCR cycles.
Fluorescent in-situ hybridization (FISH)
BAC DNA was labelled by biotin or digoxigenin nick translation kit (Roche). FISH was performed as described in detail (Fantes et al. 1995). Each labelled probe (100 ng) was dissolved in the hybridization solution containing 50% formamide, 10% dextran sulphate, 2× SSC (1× SSC, 0.15MNaCl, 0.015 M sodium citrate (pH 7.0), 0.25 mg/ml opossum Cot-1 DNA (produced from FMDL or OK cells), and 1 mg/ml salmon sperm DNA. Probes were denatured in the hybridization solution for 5min at 75°C and competed with Cot1 DNA for 15 min at 37°C. After an overnight hybridization at 37°C, the preparations were washed with 50% form-amide, 2× SSC three times for 5 min at 42°C, with 2× SSC three times 5 min at 42°C, and with 0.1× SSC once for 5 min at 60°C (when the BAC probes of M. domestica were hybridized to chromosomes of D. virginiana, the last wash at 60°C was omitted). Biotinylated probes were detected with fluorescein-avidin/anti-avidin system (Vector Laboratories) and digoxigenin-labelled DNA was visualized with rhodamine antidigoxigenin/Texas red conjugated antibody system (Vector Laboratories). The chromosomes were counterstained with 4′,6-diamino-2-phenylindol in antifade (Vectashild) and then visualized using a Leitz microscope.
Chromosome walking approach and contig generation
BAC clones positive for Chic1- and Slc16a2-specific probes were sequenced at their insert ends using T7 and SP6 vector primers. Each BAC-end sequence was masked for repeats using RepeatMasker (Smith et al. RepeatMasker Open-3.0 1996–2004, http://www.repeatmasker.org), and a primer pair for the unique region was designed (Table 2). PCR products obtained were tested by Southern blot hybridization with EcoRI-digested opossum genomic DNA to ensure they produce a unique hybridization pattern and were then used as probes for further BAC-library screening. Each newly isolated clone was tested by FISH to ensure that it had the same localization on the opossum X chromosome as the original BAC clone from which end sequence was derived.
The relative order of the BAC clones obtained by chromosome walking was determined by blot hybridization using probes to BAC ends and various exons encoding the sequences of Chic1 and Slc16a2 genes. BAC overlaps thus allowed us to assemble contigs encompassing the genes Slc16a2 and Chic1 of M. domestica and D. virginiana, in conjunction with the sequences obtained from M. domestica genome sequencing project and other available data (AC190120, deposited to the EMBL database).
Sequencing and comparative genomic analysis of BAC clones
BAC clones OM 191G11, OM 158F12, OM 068C1, LBNL3 266N1, and VMRC-6 529L22 were sequenced at the Wellcome Trust Sanger Institute (Cambridge, UK). Sequences are available at the EMBL bank under accession numbers CR385029, CR387999, CR387990, CU075922, and CR753179, respectively.
The comparative genomic analysis was performed using BLAST software (Altschul et al. 1990; http://www.ncbi.nlm.nih.gov/) to search for homologous sequences; RepeatMasker (Smit et al. RepeatMasker Open-3.0.1996–2004 〈http://www.repeatmasker.org〉) to search for interspersed repeats; and CLUSTALX (Jeanmougin et al. 1998) to align two or more sequences. The genomic analysis of large genomic loci was performed using PipMaker software (Schwartz et al. 2000, http://bio.cse.psu.edu) and software and data available on servers (http://genome.ucsc.edu/ and http://www.ensembl.org/). Pairwise comparison of nucleotide and amino acid sequences was carried out with FASTA (Pearson & Lipman 1988).
Results
Comparative gene mapping
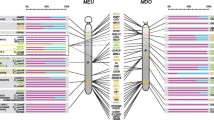
Three BAC libraries, VMRC6 of M. domestica and OM and LBNL3 of D. virginiana, were screened with M. domesteca probes for the X-linked genes Sox3, Rbmx, Chic1, Slc16a2, Pgk1, G6pd, and Hprt. The presence of the cognate genes in the positively hybridizing M. domestica and D. virginiana clones was confirmed by partial sequencing of the BAC clones using universal primers to the BAC ends and also gene-specific primers. The names of the BAC clones and genes identified are listed in Table 3. Some of these have been assigned to the X chromosome of M. domestica and D. virginiana previously, but their exact order had not been established (Samollow et al. 1987, VandeBerg et al. 1987, Nesterova et al. 1997). Using the isolated BAC clones as probes, the gene order in both opossum species was determined by DNA FISH on conventional and mechanically stretched metaphase chromosomes (Figure 1). The small acrocentric X of M. domestica displayed the following gene arrangement: G6pd, and Chic1 were located near the centromere; Hprt, Rbmx, and Sox3 in the middle of the chromosome arm; and Slc16a2 and Pgk1 close to the telomere. Thus, Chic1 and Slc16a2, which are closely linked and flank the Xist gene in all eutherian species studied, are separated on the M. domestica X chromosome. This eutherian linkage group is also split on the D. virginiana X chromosome with Chic1 located on the short arm and Slc16a2 at the telomeric region of the long arm. The order of the other genes was also rearranged on the D. virginiana X (see Figures 1 and 2), indicating that multiple chromosomal inversions have occurred in the evolution of marsupial X chromosomes.
Comparative mapping of M. domestica and D. virginiana X chromosomes. Assignment of the G6pd, Chic1, Hprt, Rbmx, Sox3, Slc16a2 and Pgk1 genes to M. domestica (A-E) and D. virginiana (F-J) X chromosomes. Slc16a2 probe is double labelled to produce yellow overlapping signal in panels B, F, G and H. G-banded ideograms of M. domestica and D. virginiana X chromosomes are shown alongside. Centromere position is indicated by arrowhead
Schematic showing comparative localization of G6pd, Cdx4, Chic1, Fip1, Lnx3, Hprt, Rbmx, Sox3, Slc16a2 and Pgk1 genes on chicken (Gallus gallus) chromosome 4 and on the X chromosome of M. domestica, D. virginiana and Homo sapiens. Only the long arm (Xq) is shown for H. sapiens X chromosome. Note that TSX is H. sapiens orthologue of Fip1 (both shown in grey). Relative position of centromere is indicated by black oval
Organization of opossum X chromosome determined using microdissection probes
To gain a more detailed understanding of the organization of M. domestica and D. virginiana X chromosomes, DNA probes were prepared by microdissection of the whole metaphase X chromosome of each species. The microdissection probes for M. domestica (MD) and D. virginiana (DV) produced a single intense signal along the entire X chromosome on the metaphase spreads of the corresponding opossum species (Figure 3A, C). Hybridization of the DV probe to M. domestica chromosomes and the MD probe to the D. virginiana chromosomes revealed that these probes hybridize predominantly to euchromatic regions of the alternative species, and are almost completely excluded from the heterochromatic regions (Figure 3B, D). Euchromatin is present as a single block on the M. domestica X chromosome and as three separate blocks in D. virginiana. The overall extent of euchromatin identified on the X chromosomes with the microdissection probes is equivalent in both opossum species, whereas the heterochromatic blocks are enlarged on D. virginiana X, increasing the overall size of this chromosome. Small euchromatic regions in D. virginiana, which are located in the middle of the short arm and on the tip of the long arm, may have arisen from a single ancestral block as a result of pericentric and peritelomeric inversions, respectively (Figure 3E). The region involved in pericentric inversion lies above Hprt and includes Chic1 and G6pd genes. The breakage point of the peritelomeric inversion that transferred the euchromatic region to the tip of the long X-chromosome arm of D. virginiana is located distal to Slc16a2 and Pgk1, as these genes are not involved in the rearrangement.
Delineation of the homologous regions of the opossum X chromosomes by microdissection probes. A, C: Hybridization of M. domestica (MD) and D. virginiana (DV) microdissection probes to the X chromosomes of corresponding species. B, D: Cross-species hybridization of the DV probe to M. domestica X chromosome and MD probe to the D. virginiana X chromosome. Ideograms of M. domestica and D. virginiana X chromosomes are shown alongside. E: Reconstruction of the opossum X chromosome evolution
Identification of contigs surrounding the Chic1 and Slc16a2 genes in M. domestica and D. virginiana
To investigate the organization of Chic1 and Slc16a2 relative to other genes linked to Xist in eutherians we assembled long-range contigs using chromosome walking, identifying and sequencing overlapping BAC clones and additionally integrating recently obtained sequence data from the M. domestica sequencing project (Figure 4 and see Materials and methods for full details).
Genomic contigs surrounding Chic1 (A) and Slc16a2 (B) genes in M. domestica and D. virginiana. Solid lines represent BAC clones isolated in this study and their relative order. The name of each BAC clone is shown alongside. AC190120 sequence is obtained from EMBL bank. Asterisks on the lines show location of the probes used for ordering the BAC clones. Numbers above the asterisks correspond to the primer pair numbers from Table 2. Primer pairs for probes 3 and 9 correspond to Chic1 and Slc16a2 specific primers listed in Table 1. Circled numbers indicate the BAC clones used to design the primer pairs to generate probes for chromosome walking analysis. Lines with diamonds represent the available sequences. Solid lines with diamonds correspond to the sequences determined in this study. Dotted lines with diamonds indicate the sequences obtained from the M. domestica genome sequencing project. The bold dashed line indicates a region where the nucleotide sequence is not completely determined. Open boxes show the position of the genes identified, and arrows above indicate the predicted direction of transcription.
Sequences of opossum protein-coding genes, including promoters, exons, and 3′- and 5′-untranslated regions, compare unambiguously with the corresponding genes of human and mouse. Thus the putative exonYintron structure of Chic1, Slc16a2, Rnf12, and Kaa2022 genes in opossum is identical to the patterns for human and mouse genes. Nucleotide sequence homology of coding regions is approximately 60% whilst at the amino acid level homology is approximately 70%. The analogous characteristics when comparing the sequences of M. domestica and D. virginiana are approximately 85% and 95%, respectively, and for mouse and human sequences, approximately 80% and 90%. Homology between introns of eutherians and opossum is absent. Similarly, the intergenic regions of eutherians and opossum are as a rule not homologous. Nonetheless, in certain cases highly homologous regions are detected in unique intergenic sequences.
Organization of genes linked to Slc16a2
Analysis of the Slc16a2 contig demonstrated the presence of two genes, Rnf12 and Kiaa2022, both located downstream of Slc16a2 (Figure 4B). The relative order of these genes in opossum corresponds to the ancestral arrangement that is characteristic of chicken and human, and differs from the pattern of mouse, whose genome contains microinversions in this region. The sequences of these Slc16a2 contigs display a higher interspecies similarity, amounting on average to 80%. Sequences located directly upstream of the Slc16a2 promoter available in the contig of M. domestica contain dispersed repeats and species-specific unique sequences lacking similarity to any known genes.
Organization of genes linked to Chic1
Within the Chic1 contig two copies of the Cdx4 gene were found in both opossum species. The orthologous region in the chicken genome also contains two Cdx genes designated Cdx4 and Cdxb. Cdxb has not been identified in the orthologous region of eutherians. The homology of exons of M. domestica and D. virginiana Cdx4 genes and Chic1 gene amounts on the average to 90%; the homology of introns to 80%. The average level of similarity in the intergenic regions is 70%. Upstream of Cdx4b in M. domestica we have identified homologies to Fzr1, Kdr and Arhgef 9 genes, also linked to Chic1 in chicken, but not in eutherian lineages.
The sequence adjacent to the 3′ end of the Chic1 gene, available for analysis in D. virginiana and M. domestica, comprises 24 and 82 kb, respectively. In mouse the Tsx gene and a part of the 30 exons of Xist are located within 75 kb downstream of Chic1 (Chureau et al. 2002). A detailed analysis of this region in opossum reveals that, starting from approximately 20 kb downstream of Chic1, both species display a fragmentary homology to Fip1 gene sequence, the precursor of Tsx located in the orthologous region on chicken chromosome 4 (Duret et al. 2006 and Figure 2). The homology to the first and second exons ismost distinctly detectable, whereas other exons and introns have diverged more pronouncedly due to mutations and insertions of mobile elements. Presumably this gene in opossum is already non-functional. In the last 7 kb of the M. domestica contig that were added from AC190120 sequence deposited at EMBL, we detected two exons which displayed a 100% similarity to the cDNA sequence of the M. domestica Lnx3 gene, which in eutherians represents the gene from which Xist is proposed to have arisen by pseudogenization (Duret et al. 2006).
Taken together, analysis of the Slc16a1 and Chic1 contig data reveal the location of an evolutionary breakpoint in marsupials that separates genes that are closely linked and flank Xist in eutherians (Figure 2). The organization of linked genes in marsupials more closely resembles that found in chicken, compared with the eutherian Xic, further supporting the conclusion that a direct homologue of Xist has not evolved in the marsupial lineage.
Discussion
In this study we demonstrate that, despite the considerable variability in the size and morphology of M. domestica and D. virginiana X chromosomes, both species have similar euchromatin/genic content. The differences in X chromosome morphology result from a pericentric inversion in D. virginiana, which has transferred a small euchromatin region over the centromere, resulting in formation of an X chromosome short arm. Enlargement of the D. virginiana X chromosome is probably attributable to amplification of species-specific repeats forming the pericentromeric and peritelomeric blocks of heterochromatin.
Mapping of X-linked genes in opossum and comparison with gene order in the ancient human XCR and chicken chromosome 4 indicates that opossum X chromosomes have undergone numerous rearrangements during evolution (see Figure 2). One of these rearrangements has separated the eutherian Xic region defined by the flanking genes Chic1 and Slc16a2. These genes are tightly linked in chicken and in all eutherian mammals analysed. D. virginiana carries an additional inversion as compared with M. domestica, which transferred Cdx4 and Chic1 to the short arm of X chromosome, such that these genes are separated by the centromere and a block of pericentromeric heterochromatin from the larger part of X euchromatin that includes the Slc16a2, Pgk1, G6pd, and Hprt genes.
When analysing available sequences surrounding the Slc16a2 gene, we have not identified any homology to Xist or other sequences in the Xic domain. The sequences downstream of opossum Chic1, however, display a homology to Fip1 and Lnx3 genes, present in the orthologous locus on chicken chromosome 4. Thus, Lnx3 and Rasl11c genes, mapped earlier on the X chromosome of opossums (Duret et al. 2006), are directly linked to the Chic1 gene. The opossum Fip1 gene has diverged considerably and is not functional, whereas Lnx3 produces mRNA, has a native open reading frame (Duret et al. 2006), and, presumably, functions as a protein-coding gene, not as an untranslated nuclear RNA similar to Xist. Overall these findings are in agreement with the conclusion reached by Duret et al. (2006), suggesting that, despite having an X inactivation system, marsupials do not utilize a direct homologue of Xist.
If marsupials have no direct homologue of Xist, how is X inactivation regulated? One idea, discussed previously, is that the paternal X retains a repressive chromatin structure established during meiotic sex chromosome inactivation in spermatogenesis (Cooper et al. 1993). Whilst feasible, a difficulty with this hypothesis is that packaging of DNA in sperm, at least in other mammals, involves replacement of core chromatin proteins with protamines. With this in mind it is unclear how a repressive chromatin configuration could be transmitted into the zygote unless replacement is incomplete. An alternative explanation is that an Xist-like regulator, i.e. a cis-acting non-coding RNA, has evolved independently in the marsupial lineage. Studies on imprinting clusters in eutherians suggests that the co-ordinate regulation of chromosome domains may in some cases result from imprinted expression of a cis-acting non-coding RNA (Delaval & Feil 2004), indicating that Xist is not unique in this regard. The availability of the M. domestica genome sequence opens up the possibility to carry out a systematic search for candidate sequences.
References
Altschul SF, Gish W, Miller W, Meyers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215: 403-10.
Chow JC, Yen Z, Ziesche SM, Brown CJ (2005) Silencing of the mammalian X chromosome. Annu Rev Genomics Hum Genet 6: 69-2.
Chureau C, Prissette M, Bourdet A et al. (2002) Comparative sequence analysis of the X-inactivation center region in mouse, human and bovine. Genome Res 12: 894-08.
Cooper DW, Johnston PG, Watson JM, Graves JAM (1993) X-inactivation in marsupials and monotremes. Dev Biol 4: 117-28.
Delaval K, Feil R (2004) Epigenetic regulation of mammalian genomic imprinting. Curr Opin Genet Dev 14: 188-95.
Duret L, Chureau C, Samain S, Weissenbach J, Avner P (2006) The Xist RNA gene evolved in eutherians by pseudogenization of a protein-coding gene. Science 312: 1653-655.
Evans HK, Weidman JR, Cowley DO, Jirtle RL (2005) Comparative phylogenetic analysis of blcap/nnat reveals eutherian-specific imprinted gene. Mol Biol Evol 22: 1740-1748.
Fantes JA, Oghene K, Boyle S et al. (1995) A high resolution integrated physical, cytogenetic and genetic map of human chromosome 11 from the distal region of p13 to the proximal part of p15.1. Genomics 25: 447-61.
Graves JAM (1996) Mammals that break the rules: genetics of marsupials and monotremes. Annu Rev Genet 30: 233-60.
Graves JAM, Westerman M (2002) Marsupial genetics and genomics. Trends Genet 18: 517-21.
Haaf T, Ward DC (1994) Structural analysis of alpha-satellite DNA and centromere proteins using extended chromatin and chromosomes. Hum Mol Genet 3: 697-09.
Jeanmougin F, Thompson JD, Gouy M, Higgins DG, Gibson TJ (1998) Multiple sequence alignment with Clustal X. Trends Biochem Sci 23: 403-405.
Koina E, Wakefield MJ, Walcher C et al. (2005) Isolation, X location and activity of the marsupial homologue of SLC16A2, an XIST-flanking gene in eutherian mammals. Chromosome Res 13: 687-98.
Lyon MF (1961) Gene action in the X-chromosome of the mouse (Mus musculus L.). Nature 190: 372-73.
Nesterova TB, Isaenko AA, Matveeva NM et al. (1997) Novel strategies for eutherian × marsupial somatic cell hybrids: mapping the genome of Monodelphis domestica. Cytogenet Cell Genet 76: 115-22.
Pearson WR, Lipman DJ (1988) Improved tools for biological sequence comparison. Proc Natl Acad Sci 85: 2444-448.
Rubtsov NB, Karamisheva TV, Astakhova NM, Liehr T, Claussen U, Zhdanova NS (2000) Zoo-FISH with region-specific paints for mink chromosome 5q: delineation of inter- and intrachromosomal rearrangements in human, pig, and fox. Cytogenet Cell Genet 90: 268-70.
Samollow PB, Ford AL, VandeBerg JL (1987) X-linked gene expression in the Virginia opossum: differences between the paternally derived Gpd and Pgk-A loci. Genetics 115: 185-95.
Schwartz S, Zhang Z, Frazer KA et al. (2000) PipMaker–a web server for aligning two genomic DNA sequences. Genome Res 10: 577-86.
Sharman GB (1971) Late DNA replication in paternally derived X chromosome of female kangaroos. Nature 230: 231-32.
Smit AFA, Hubley R, Green P. RepeatMasker at http://www.repeatmasker.org.
VandeBerg JL, Robinson ES, Samollow PB, Jonhston PG (1987) X linked gene expression and X chromosome inactivation: marsupials, mouse and man compared. In Markert CL, ed. Isozymes: Current Topics in Biological and Medical Research. New York: Alan R. Liss Inc. vol.15: 225–253.
Woodburne MO, Rich TH, Springer MS (2003) The evolution of tribospheny and the antiquity of mammalian clades. Mol Phylogenet Evol 28: 360-85.
Acknowledgement
This work was supported by funding from the Wellcome Trust, UK (an International Development Award, 067065/Z/02/Z, for A.I.S.), from the Russian Foundation for Fundamental Research (grants 06-04-48337 and 05-04-48230a) and the Russian Academy of Science Foundation Program for Molecular and Cell Biology. We also thank the Sequencing Division at the Wellcome Trust Sanger Institute.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License ( https://creativecommons.org/licenses/by-nc/2.0 ), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Shevchenko, A.I., Zakharova, I.S., Elisaphenko, E.A. et al. Genes flanking Xist in mouse and human are separated on the X chromosome in American marsupials. Chromosome Res 15, 127–136 (2007). https://doi.org/10.1007/s10577-006-1115-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-006-1115-9