Abstract
Cellulose microfibrils are a key component of plant cell walls, which in turn compose most of our renewable biomaterials. Consequently, there is considerable interest in understanding how cellulose microfibrils are made in living cells by the plant cellulose synthesis complex (CSC). This remarkable multi-subunit complex contains cellulose synthase (CESA) proteins, and it is often called a rosette due to its six-lobed shape. Each CSC moves within the plasma membrane as it spins a strong cellulose microfibril in its wake. To accomplish this biological manufacturing process, the CESAs harvest an activated sugar substrate from the cytoplasm for use in the polymerization of glucan chains. An elongating glucan is simultaneously translocated across the plasma membrane by each CESA, where the group of chains emanating from one CSC co-crystallizes into a cellulose microfibril that becomes part of the assembling cell wall. Here we review major advances in understanding CESA and CSC structure/function relationships since 2013, when ground-breaking insights about the structure of cellulose synthases in bacteria and plants were published. We additionally discuss: (a) the relationship of CSC substructure to the size of the fundamental cellulose fibril; (b) an evolutionary perspective on the driving force behind the existence of hetero-oligomeric CSCs that currently appear to dominate in land plants; and (c) how cellulose properties may be regulated by CESA and CSC activity. We also pose major questions that still remain in this rapidly changing and exciting research field.
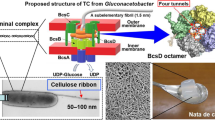
Graphical abstract



Similar content being viewed by others
Abbreviations
- BcsA:
-
Cellulose synthase in bacteria
- BcsB:
-
Non-catalytic partner protein for BscA
- CSC:
-
Cellulose synthesis complex
- CESA:
-
Cellulose synthase in plants
- CNE:
-
Constructive neutral evolution
- CSR:
-
Class-specific region of CESA
- DP:
-
Degree of polymerization
- FF-TEM:
-
Freeze fracture transmission electron microscopy
- P-CR:
-
Plant-conserved region of CESA
- PCW:
-
Primary cell wall
- PCW CSC:
-
Primary cell wall cellulose synthesis complex
- SCW:
-
Secondary cell wall
- SCW CSC:
-
Secondary cell wall cellulose synthesis complex
- SSNMR:
-
Solid-state nuclear magnetic resonance
- TMH:
-
Transmembrane helix
- UDP-glucose:
-
Uridine diphosphate-alpha-d-glucose
References
Arioli T et al (1998) Molecular analysis of cellulose biosynthesis in Arabidopsis. Science 279:717–720
Atalla RH, Vanderhart DL (1984) Native cellulose: a composite of two distinct crystalline forms. Science 223:283–285
Atanassov II, Pittman JK, Turner SR (2009) Elucidating the mechanisms of assembly and subunit interaction of the cellulose synthase complex of Arabidopsis secondary cell walls. J Biol Chem 284:3833–3841
Bashline L, Lei L, Li S, Gu Y (2014) Cell wall, cytoskeleton, and cell expansion in higher plants. Mol Plant 7:586–600. https://doi.org/10.1093/mp/ssu018
Betancur L, Singh B, Rapp RA, Wendel JF, Marks MD, Roberts AW, Haigler CH (2011) Phylogenetically distinct cellulose synthase genes support secondary wall thickening in arabidopsis shoot trichomes and cotton fiber. J Integr Plant Biol 52:205–220
Bowling AJ, Brown RM Jr (2008) The cytoplasmic domain of the cellulose-synthesizing complex in vascular plants. Protoplasma 233:115–127
Brown DM, Zeef LA, Ellis J, Goodacre R, Turner SR (2005) Identification of novel genes in Arabidopsis involved in secondary cell wall formation using expression profiling and reverse genetics. Plant Cell 17:2281–2295
Carpita NC (2011) How plants make cellulose and other (1->4)-b-d-glycans. Plant Physiol 155:171–184
Carroll A, Specht CD (2011) Understanding plant cellulose synthases through a comprehensive investigation of the cellulose synthase family sequences Front. Plant Sci 2:5. https://doi.org/10.3389/fpls.2011.00005
Chen S, Ehrhardt DW, Somerville CR (2010) Mutations of cellulose synthase (CESA1) phosphorylation sites modulate anisotropic cell expansion and bidirectional mobility of cellulose synthase. Proc Natl Acad Sci USA 107:17188–17193
Cho SH et al (2017) Synthesis and self-assembly of cellulose microfibrils from reconstituted cellulose synthase. Plant Physiol 175:146–156. https://doi.org/10.1104/pp.17.00619
Conant GC, Wolfe KH (2008) Turning a hobby into a job: how duplicated genes find new functions. Nat Rev Genet 9:938–950
Cosgrove DJ (2014) Re-constructing our models of cellulose and primary cell wall assembly. Curr Opin Plant Biol 22:122–131. https://doi.org/10.1016/j.pbi.2014.11.001
Delmer DP (1987) Cellulose biosynthesis. Ann Rev Plant Physiol 38:259–290
Delmer DP (1999) Cellulose biosynthesis: exciting times for a difficult field of study. Annu Rev Plant Physiol Plant Mol Biol 50:245–276
Desprez T et al (2007) Organization of cellulose synthase complexes involved in primary cell wall synthesis in Arabidopsis thaliana. Proc Natl Acad Sci USA 104:15572–15577
Diotallevi F, Mulder B (2007) The cellulose synthase complex: a polymerization driven supramolecular motor. Biophys J 92:2666–2673
Donaldson L (2007) Cellulose microfibril aggregates and their size variation with cell wall type. Wood Sci Technol 41:443–460
Doolittle WF (2012) Evolutionary biology: a ratchet for protein complexity. Nature 481:270–271
Dupree R, Simmons TJ, Mortimer JC, Patel D, Iuga D, Brown SP, Dupree P (2015) Probing the molecular architecture of Arabidopsis thaliana secondary cell walls using two- and three-dimensional (13)C solid state nuclear magnetic resonance spectroscopy. Biochemistry 54:2335–2345. https://doi.org/10.1021/bi501552k
Emons AMC (1991) Role of particle rosettes and terminal globules in cellulose synthesis. In: Haigler CH, Weimer PJ (eds) Biosynthesis and biodegradation of cellulose. Marcel Dekker, New York, pp 71–98
Endler A, Persson S (2011) Cellulose synthases and synthesis in Arabidopsis. Mol Plant 4:199–211. https://doi.org/10.1093/mp/ssq079
Fang L, Catchmark JM (2014) Structure characterization of native cellulose during dehydration and rehydration. Cellulose 21:3951–3963
Fernandes AN et al (2011) Nanostructure of cellulose microfibrils in spruce wood. Proc Natl Acad Sci USA 108:E1195–E1203. https://doi.org/10.1073/pnas.1108942108
Finnigan GC, Hanson-Smith V, Stevens TH, Thornton JW (2012) Evolution of increased complexity in a molecular machine. Nature 481:360–364
Foston M et al (2011) Chemical, ultrastructural and supramolecular analysis of tension wood in Populus tremula x alba as a model substrate for reduced recalcitrance. Energy Environ Sci 4:4962–4971
Fujita M et al (2013) The anisotropy1 D604N mutation in the Arabidopsis cellulose synthase1 catalytic domain reduces cell wall crystallinity and the velocity of cellulose synthase complexes. Plant Physiol 162:74–85. https://doi.org/10.1104/pp.112.211565
Giddings TH Jr, Brower DL, Staehelin LA (1980) Visualization of particle complexes in the plasma membrane of Micrasterias denticulata associated with the formation of cellulose fibrils in primary and secondary cell walls. J Cell Biol 84:327–339
Gonneau M, Desprez T, Guillot A, Vernhettes S, Hofte H (2014) Catalytic subunit stoichiometry within the cellulose synthase complex. Plant Physiol 166:1709–1712. https://doi.org/10.1104/pp.114.250159
Griffiths JS et al (2015) Unidirectional movement of cellulose synthase complexes in Arabidopsis seed coat epidermal cells deposit cellulose involved in mucilage extrusion, adherence, and ray formation. Plant Physiol 168:502–520. https://doi.org/10.1104/pp.15.00478
Guerriero G, Fugelstad J, Bulone V (2010) What do we really know about cellulose biosynthesis in higher plants? J Integr Plant Biol 52:161–175
Haigler CH, Grimson MJ, Gervais J, Le Moigne N, Hofte H, Monasse B, Navard P (2014) Molecular modeling and imaging of initial stages of cellulose fibril assembly: evidence for a disordered intermediate stage. PLoS ONE 9:e93981. https://doi.org/10.1371/journal.pone.0093981
Haigler CH, Davis J, Slabaugh E, Kubicki JD (2016) Biosynthesis and assembly of cellulose. In: Rose JK (ed) Molecular cell biology of the growth and differentiation of plant cells. CRC Press, Boca Raton, pp 120–138. https://doi.org/10.1201/b20316
Hamann T, Osborne E, Youngs HL, Misson J, Nussaume L, Somerville C (2004) Global expression analysis of CESA and CSL genes in Arabidopsis. Cellulose 11:279–286
Harholt J, Moestrup Ø, Ulvskov P (2016) Why plants were terrestrial from the beginning. Trends Plant Sci 21:96–101
Harris D, Stork J, Debolt S (2009) Genetic modification in cellulose-synthase reduces crystallinity and improves biochemical conversion to fermentable sugar. Glob Change Biol 1:51–61
Harris DM et al (2012) Cellulose microfibril crystallinity is reduced by mutating C-terminal transmembrane region residues CESA1A903V and CESA3T942I of cellulose synthase. Proc Natl Acad Sci USA 109:4098–4103
Herth W (1983) Arrays of plasma-membrane “rosettes” involved in cellulose microfibril formation of Spirogyra. Planta 159:347–356
Herth W (1985) Plant cell wall formation. In: Robards AW (ed) Botanical microscopy. Oxford University Press, Oxford, pp 285–310
Hill JL Jr, Hammudi MB, Tien M (2014) The Arabidopsis cellulose synthase complex: a proposed hexamer of CESA trimers in an equimolar stoichiometry. Plant Cell 26:4834–4842. https://doi.org/10.1105/tpc.114.131193
Hill JL Jr, Hill AN, Roberts AW, Haigler CH, Tien M (2018a) Domain swaps of Arabidopsis secondary wall cellulose synthases to elucidate their class specificity. Plant Direct 2:e00061
Hill JL Jr, Josephs C, Barnes WJ, Anderson CT, Tien M (2018b) Longevity in vivo of primary cell wall cellulose synthases. Plant Mol Biol 96:279–289. https://doi.org/10.1007/s11103-017-0695-4
Ivakov A et al (2017) Cellulose synthesis and cell expansion are regulated by different mechanisms in growing Arabidopsis hypocotyls. Plant Cell 29:1305–1315. https://doi.org/10.1105/tpc.16.00782
Jarvis MC (2013) Cellulose biosynthesis: counting the chains. Plant Physiol 163:1485–1486. https://doi.org/10.1104/pp.113.231092
Jarvis MC (2018) Structure of native cellulose microfibrils, the starting point for nanocellulose manufacture. Phil Trans R Soc A. https://doi.org/10.1098/rsta.2017.0045
Jokipii-Lukkari S, Sundell D, Nilsson O, Hvidsten TR, Street NR, Tuominen H (2017) NorWood: a gene expression resource for evo-devo studies of conifer wood development. New Phytol 216:482–494. https://doi.org/10.1111/nph.14458
Jones DM, Murray CM, Ketelaar KJ, Thomas JJ, Villalobos JA, Wallace IS (2016) The emerging role of protein phosphorylation as a critical regulatory mechanism controlling cellulose biosynthesis Front. Plant Sci 7:684. https://doi.org/10.3389/fpls.2016.00684
Kimura S, Laosinchai W, Itoh T, Cui X, Linder CR, Brown RM Jr (1999) Immunogold labeling of rosette terminal cellulose-synthesizing complexes in the vascular plant Vigna angularis. Plant Cell 11:2075–2085
Knott BC, Crowley MF, Himmel ME, Zimmer J, Beckham GT (2016) Simulations of cellulose translocation in the bacterial cellulose synthase suggest a regulatory mechanism for the dimeric structure of cellulose. Chem Sci 7:3108–3116. https://doi.org/10.1039/C5SC04558D
Kubicki JD, Yang H, Sawada D, O’Neill H, Oehme D, Cosgrove D (2018) The shape of native plant cellulose microfibrils. Sci Rep 8:13983. https://doi.org/10.1038/s41598-018-32211-w
Kumar M, Turner S (2015) Plant cellulose synthesis: CESA proteins crossing kingdoms. Phytochemistry 112:91–99. https://doi.org/10.1016/j.phytochem.2014.07.009
Kumar M et al (2009) An update on the nomenclature for the cellulose synthase genes in Populus. Trends Plant Sci 14:248–254
Kumar M, Atanassov I, Turner S (2016) Functional analysis of cellulose synthase CESA protein class-specificity. Plant Physiol 173:970–983
Kumar M, Mishra L, Carr P, Pilling M, Gardner P, Mansfield SD, Turner S (2018) Exploiting Cellulose Synthase (CESA) class specificity to probe cellulose microfibril biosynthesis. Plant Physiol 177:151–167. https://doi.org/10.1104/pp.18.00263
Lang D et al (2018) The Physcomitrella patens chromosome-scale assembly reveals moss genome structure and evolution. Plant J 93:515–533. https://doi.org/10.1111/tpj.13801
Lee CM, Kafle K, Belias DW, Park YB, Glick RE, Haigler CH, Kim SH (2015) Comprehensive analysis of cellulose content, crystallinity, and lateral packing in Gossypium hirsutum and Gossypium barbadense cotton fibers using sum frequency generation, infrared and Raman spectroscopy, and X-ray diffraction. Cellulose 22:971–989
Lei L et al (2014) The jiaoyao1 mutant Is an allele of korrigan1 that abolishes endoglucanase activity and affects the organization of both cellulose microfibrils and microtubules in Arabidopsis. Plant Cell 26:2601–2616. https://doi.org/10.1105/tpc.114.126193
Li X (2017) Characterization of cellulose synthesis complexes and Physcomitrella patens. University of Rhode Island, Kingston
Li Y, Qi B (2017) Progress toward Understanding protein S-acylation: prospective in plants. Front Plant Sci 8:346. https://doi.org/10.3389/fpls.2017.00346
Li S, Bashline L, Lei L, Gu Y (2014) Cellulose synthesis and its regulation. The Arabidopsis book. Am Soc Plant Biol 12:e0169. https://doi.org/10.1199/tab.0169
Li S et al (2016) Cellulose synthase complexes act in a concerted fashion to synthesize highly aggregated cellulose in secondary cell walls of plants. Proc Natl Acad Sci USA 113:11348–11353. https://doi.org/10.1073/pnas.1613273113
Liu Z, Persson S, Sanchez-Rodriguez C (2015) At the border: the plasma membrane-cell wall continuum. J Exp Bot 66:1553–1563. https://doi.org/10.1093/jxb/erv019
Liu D et al (2017) Imaging cellulose synthase motility during primary cell wall synthesis in the grass Brachypodium distachyon. Sci Rep 7:15111. https://doi.org/10.1038/s41598-017-14988-4
Martinez-Sanz M, Pettolino F, Flanagan B, Gidley MJ, Gilbert EP (2017) Structure of cellulose microfibrils in mature cotton fibres. Carbohydr Polym 175:450–463. https://doi.org/10.1016/j.carbpol.2017.07.090
McFarlane HE, Doring A, Persson S (2014) The cell biology of cellulose synthesis. Annu Rev Plant Biol 65:69–94. https://doi.org/10.1146/annurev-arplant-050213-040240
McNamara JT, Morgan JL, Zimmer J (2015) A molecular description of cellulose biosynthesis. Annu Rev Biochem 84:895–921. https://doi.org/10.1146/annurev-biochem-060614-033930
Meents MJ, Watanabe Y, Samuels AL (2018) The cell biology of secondary cell wall biosynthesis. Annu Bot 121:1107–1125. https://doi.org/10.1093/aob/mcy005
Mølhøj M, Ulvskov P, Dal Degan F (2001) Characterization of a functional soluble form of a Brassica napus membrane-anchored endo-1,4-β-glucanase heterologously expressed in Pichia pastoris. Plant Physiol 127:674–684
Morgan JLW, Strumillo J, Zimmer J (2013) Crystallographic snapshot of cellulose synthesis and membrane translocation. Nature 493:181-U192. https://doi.org/10.1038/Nature11744
Morris JL et al (2018) The timescale of early land plant evolution. Proc Natl Acad Sci USA 115:E2274–E2283. https://doi.org/10.1073/pnas.1719588115
Mueller SC, Brown RM Jr (1980) Evidence for an intramembrane component associated with a cellulose microfibril-synthesizing complex in higher plants. J Cell Biol 84:315–326
Mühlethaler K (1967) Ultrastructure and formation of plant cell walls. Annu Rev Plant Physiol 18:1–24
Mutwil M, Debolt S, Persson S (2008) Cellulose synthesis: a complex complex. Curr Opin Plant Biol 11:252–257
Newman RH, Hill SJ, Harris PJ (2013) Wide-angle x-ray scattering and solid-state nuclear magnetic resonance data combined to test models for cellulose microfibrils in mung bean cell walls. Plant Physiol 163:1558–1567. https://doi.org/10.1104/pp.113.228262
Nixon BT, Mansouri K, Singh A, Du J, Davis JK, Lee J-G, Slabaugh E, Vandavasi VG, O’Neill H, Roberts EM, Roberts AW, Yingling YG, Haigler CH (2016) Comparative structural and computational analysis supports eighteen cellulose synthases in the plant cellulose synthesis complex Sci Rep 6:28696
Norris JH et al (2017) Functional specialization of cellulose synthase isoforms in a moss shows parallels with seed plants. Plant Physiol 175:210–222
Oehme D, Doblin MS, Wagner J, Bacic A, Downton MT, Gidley MJ (2015a) Gaining insight into cell wall cellulose macrofibril organisation by simulating microfibril adsorption. Cellulose 22:3501–3520
Oehme DP, Downton MT, Doblin MS, Wagner J, Gidley MJ, Bacic A (2015b) Unique aspects of the structure and dynamics of elementary Ibeta cellulose microfibrils revealed by computational simulations. Plant Physiol 168:3–17. https://doi.org/10.1104/pp.114.254664
Okuda K, Brown RM Jr (1992) A new putative cellulose-synthesizing complex of Coleochaete scutata. Protoplasma 168:51–63
Olek AT et al (2014) The structure of the catalytic domain of a plant cellulose synthase and its assembly into dimers. Plant Cell 26:2996–3009. https://doi.org/10.1105/tpc.114.126862
Omadjela O, Narahari A, Strumillo J, Melida H, Mazur O, Bulone V, Zimmer J (2013) BcsA and BcsB form the catalytically active core of bacterial cellulose synthase sufficient for in vitro cellulose synthesis. Proc Natl Acad Sci USA 110:17856–17861. https://doi.org/10.1073/pnas.1314063110
Paredez AR, Somerville CR, Ehrhardt DW (2006) Visualization of cellulose synthase demonstrates functional association with microtubules. Science 312:1491–1495
Park YB, Cosgrove DJ (2015) Xyloglucan and its interactions with other components of the growing cell wall. Plant Cell Physiol 56:180–194. https://doi.org/10.1093/pcp/pcu204
Pear JR, Kawagoe Y, Schreckengost WE, Delmer DP, Stalker DM (1996) Higher plants contain homologs of the bacterial celA genes encoding the catalytic subunit of cellulose synthase. Proc Natl Acad Sci USA 93:12637–12642
Persson S et al (2007) Genetic evidence for three unique components in primary cell-wall cellulose synthase complexes in Arabidopsis. Proc Natl Acad Sci USA 104:15566–15571
Purushotham P, Cho SH, Díaz-Moreno SM, Kumar M, Nixon BT, Bulone V, Zimmer J (2016) A single heterologously expressed plant cellulose synthase isoform is sufficient for cellulose microfibril formation in vitro. Proc Natl Acad Sci USA 113:11360–11365
Ren R et al (2018) Widespread whole genome duplications contribute to genome complexity and species diversity in angiosperms. Mol Plant 11:414–428. https://doi.org/10.1016/j.molp.2018.01.002
Roberts AW, Bushoven JT (2007) The cellulose synthase (CESA) gene superfamily of the moss Physcomitrella patens. Plant Mol Biol 63:207–219
Roberts AW, Roberts EM, Haigler CH (2012) Moss cell walls: structure and biosynthesis. Front Plant Sci 3:166
Ruprecht C, Mutwil M, Saxe F, Eder M, Nikoloski Z, Persson S (2011) Large-scale co-expression approach to dissect secondary cell wall formation across plant species Front. Plant Sci 2:23. https://doi.org/10.3389/fpls.2011.00023
Rushton PS, Olek AT, Makowski L, Badger J, Steussy CN, Carpita NC, Stauffacher CV (2017) Rice Cellulose SynthaseA8 plant-conserved region is a coiled-coil at the catalytic core entrance. Plant Physiol 173:482–494. https://doi.org/10.1104/pp.16.00739
Sahoo DK, Stork J, DeBolt S, Maiti IB (2013) Manipulating cellulose biosynthesis by expression of mutant Arabidopsis proM24:CESA3(ixr1-2) gene in transgenic tobacco. Plant Biotechnol J 11:362–372. https://doi.org/10.1111/pbi.12024
Saxena IM, Lin FC, Brown RM Jr (1990) Cloning and sequencing of the cellulose synthase catalytic subunit gene of Acetobacter xylinum. Plant Mol Biol 15:673–683
Scavuzzo-Duggan TR et al (2018) Cellulose synthase ‘class specific regions’ are intrinsically disordered and functionally undifferentiated. J Integr Plant Biol 60:481–497. https://doi.org/10.1111/jipb.12637
Scheible W-R, Eshed R, Richmond T, Delmer D, Somerville C (2001) Modifications of cellulose synthase confer resistance to isoxaben and thiazolidinone herbicides in Arabidopsis Ixr1 mutants. Proc Natl Acad Sci USA 98:10079–10084
Schneider B, Herth W (1986) Distribution of plasma membrane rosettes and kinetics of cellulose formation in xylem development of higher plants. Protoplasma 131:142–152
Schneider R, Hanak T, Persson S, Voigt CA (2016) Cellulose and callose synthesis and organization in focus: What’s new? Curr Opin Plant Biol 34:9–16. https://doi.org/10.1016/j.pbi.2016.07.007
Schneider R et al (2017) Two complementary mechanisms underpin cell wall patterning during xylem vessel development. Plant Cell 29:2433–2449. https://doi.org/10.1105/tpc.17.00309
Sethaphong L, Haigler CH, Kubicki JD, Zimmer J, Bonetta D, DeBolt S, Yingling YG (2013) Tertiary model of a plant cellulose synthase. Proc Natl Acad Sci USA 110:7512–7517. https://doi.org/10.1073/pnas.1301027110
Sethaphong L, Davis JK, Slabaugh E, Singh A, Haigler CH, Yingling YG (2016) Prediction of the structures of the plant-specific regions of vascular plant cellulose synthases and correlated functional analysis. Cellulose 23:145–161
Shim I, Law R, Kileeg Z, Stronghill P, Northey JGB, Strap JL, Bonetta D (2018) Alleles causing resistance to isoxaben and flupoxam highlight the significance of transmembrane domains for CESA protein function. Front Plant Sci 9:1152
Slabaugh E, Davis JK, Haigler CH, Yingling YG, Zimmer J (2014) Cellulose synthases: new insights from crystallography and modeling. Trends Plant Sci 19:99–106. https://doi.org/10.1016/j.tplants.2013.09.009
Song D, Shen J, Li L (2010) Characterization of cellulose synthase complexes in Populus xylem differentiation. New Phytol 187:777–790
Speicher TL, Li PZ, Wallace IS (2018) Phosphoregulation of the plant cellulose synthase complex and cellulose synthase-like proteins. Plants. https://doi.org/10.3390/plants7030052
Taylor JB (1957) The water solubilities and heats of solution of short chain cellulosic oligosaccharides. Trans Faraday Soc 53:1198–1203
Taylor NG (2007) Identification of cellulose synthase AtCesA7 (IRX3) in vivo phosphorylation sites—a potential role in regulating protein degradation. Plant Mol Biol 64:161–171
Taylor NG, Howells RM, Huttly AK, Vickers K, Turner SR (2003) Interactions among three distinct CesA proteins essential for cellulose synthesis. Proc Natl Acad Sci USA 100:1450–1455
Taylor NG, Gardiner JC, Whiteman R, Turner SR (2004) Cellulose synthesis in the Arabidopsis secondary cell wall. Cellulose 11:329–338
Thomas LH, Forsyth VT, Martel A, Grillo I, Altaner CM, Jarvis MC (2014) Structure and spacing of cellulose microfibrils in woody cell walls of dicots. Cellulose 21:3887–3895
Timpa JD, Triplett BA (1993) Analysis of cell-wall polymers during cotton fiber development. Planta 189:101–108
Tsekos I (1999) The sites of cellulose synthesis in algae: diversity and evolution of cellulose-synthesizing enzyme complexes. J Phycol 35:635–655
Turner S, Kumar M (2018) Cellulose synthase complex organization and cellulose microfibril structure. Phil Trans R Soc A. https://doi.org/10.1098/rsta.2017.0048
Turner SR, Somerville CR (1997) Collapsed xylem phenotype of Arabidopsis identifies mutants deficient in cellulose deposition in the secondary cell wall. Plant Cell 9:689–701
Umemura M, Yuguchi Y, Hirotsu T (2004) Interaction between cellooligosaccharides in aqueous solution from molecular dynamics simulation: comparison of cellotetraose, cellopentaose, and cellohexaose. J Phys Chem A 108:7063–7070
Vain T et al (2014) The cellulase KORRIGAN is part of the cellulose synthase complex. Plant Physiol 165:1521–1532. https://doi.org/10.1104/pp.114.241216
Vandavasi VG et al (2016) A structural study of CESA1 catalytic domain of Arabidopsis cellulose synthesis complex: evidence for CESA trimers. Plant Physiol 170:123–135. https://doi.org/10.1104/pp.15.01356
Vergara CE, Carpita NC (2001) b-d-glycan synthases and the CesA gene family: lessons to be learned from the mixed-linkage (1→3), (1→4)b-d-glucan synthase. Plant Mol Biol 47:145–160
Wang T, Hong M (2016) Solid-state NMR investigations of cellulose structure and interactions with matrix polysaccharides in plant primary cell walls. J Exp Bot 67:503–514. https://doi.org/10.1093/jxb/erv416
Wang J, Howles PA, Cork AH, Birch RJ, Williamson RE (2006) Chimeric proteins suggest that the catalytic and/or C-terminal domains give CesA1 and CesA3 access to their specific sites in the cellulose synthase of primary walls. Plant Physiol 142:685–695
Wang T, Park YB, Cosgrove DJ, Hong M (2015) Cellulose-pectin spatial contacts are inherent to never-dried arabidopsis primary cell walls: evidence from solid-state nuclear magnetic resonance. Plant Physiol 168:871–884. https://doi.org/10.1104/pp.15.00665
Wang T, Yang H, Kubicki JD, Hong M (2016) Cellulose structural polymorphism in plant primary cell walls investigated by high-field 2D solid-state NMR spectroscopy and density functional theory calculations. Biomacromol 17:2210–2222. https://doi.org/10.1021/acs.biomac.6b00441
Watanabe Y et al (2015) Visualization of cellulose synthases in Arabidopsis secondary cell walls. Science 350:198–203. https://doi.org/10.1126/science.aac7446
Watanabe Y et al (2018) Cellulose synthase complexes display distinct dynamic behaviors during xylem transdifferentiation. Proc Natl Acad Sci USA 115:E6366–E6374. https://doi.org/10.1073/pnas.1802113115
Wickett NJ et al (2014) Phylotranscriptomic analysis of the origin and early diversification of land plants. Proc Natl Acad Sci USA 111:E4859–E4868. https://doi.org/10.1073/pnas.1323926111
Willison JHM (1976) Plasmodesmata: a freeze-fracture view. Can J Bot 54:2842–2847
Wong HC et al (1990) Genetic organization of the cellulose synthase operon in Acetobacter xylinum. Proc Natl Acad Sci USA 87:8130–8134
Woodley M, Mulvihill A, Fujita M, Wasteneys GO (2018) Exploring microtubule-dependent cellulose-synthase-complex movement with high precision. Part Track Plants. https://doi.org/10.3390/plants7030053
Xi W, Song D, Sun J, Shen J, Li L (2017) Formation of wood secondary cell wall may involve two type cellulose synthase complexes in Populus. Plant Mol Biol 93:419–429. https://doi.org/10.1007/s11103-016-0570-8
Yang H, Zimmer J, Yingling YG, Kubicki JD (2015) How cellulose elongates—A QM/MM study of the molecular mechanism of cellulose polymerization in bacterial CESA. J Phys Chem B 119:6525–6535. https://doi.org/10.1021/acs.jpcb.5b01433
Yin Y, Huang J, Xu Y (2009) The cellulose synthase superfamily in fully sequenced plants and algae. BMC Plant Biol 9:99
Yin Y, Johns MA, Cao H, Rupani M (2014) A survey of plant and algal genomes and transcriptomes reveals new insights into the evolution and function of the cellulose synthase superfamily. BMC Genom 15:260. https://doi.org/10.1186/1471-2164-15-260
Yoon HS, Hackett JD, Ciniglia C, Pinto G, Bhattacharya D (2004) A molecular timeline for the origin of photosynthetic eukaryotes. Mol Biol Evol 21:809–818
Yu L, Sun J, Li L (2013) PtrCel9A6, an endo-1,4-β-glucanase, is required for cell wall formation during xylem differentiation in populus. Mol Plant 6:1904–1917. https://doi.org/10.1093/mp/sst104
Zhang T, Mahgsoudy-Louyeh S, Tittmann B, Cosgrove DJ (2014) Visualization of the nanoscale pattern of recently-deposited cellulose microfibrils and matrix materials in never-dried primary walls of the onion epidermis. Cellulose 21:853–862
Zhang T, Zheng Y, Cosgrove DJ (2016) Spatial organization of cellulose microfibrils and matrix polysaccharides in primary plant cell walls as imaged by multichannel atomic force microscopy. Plant J 85:179–192. https://doi.org/10.1111/tpj.13102
Zhang X et al (2018) Cellulose synthase stoichiometry in aspen differs from Arabidopsis and Norway spruce. Plant Physiol 177:1096–1107. https://doi.org/10.1104/pp.18.00394
Acknowledgments
Under the Creative Commons License (http://creativecommons.org/licenses/by/4.0/), the graphical abstract includes a low resolution subset of the images originally published by the authors and their coworkers in Fig. 2 of Scientific Reports (Nixon et al. 2016). The writing of this review was supported by the US Department of Energy, Office of Science, Basic Energy Sciences as part of The Center for LignoCellulose Structure and Formation, an Energy Frontier Research Center under Award # DE-SC0001090.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Haigler, C.H., Roberts, A.W. Structure/function relationships in the rosette cellulose synthesis complex illuminated by an evolutionary perspective. Cellulose 26, 227–247 (2019). https://doi.org/10.1007/s10570-018-2157-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10570-018-2157-9