Abstract
The genus Methylobacter is considered an important and often dominant group of aerobic methane-oxidizing bacteria in many oxic ecosystems, where members of this genus contribute to the reduction of CH4 emissions. Metagenomic studies of the upper oxic layers of geothermal soils of the Favara Grande, Pantelleria, Italy, revealed the presence of various methane-oxidizing bacteria, and resulted in a near complete metagenome assembled genome (MAG) of an aerobic methanotroph, which was classified as a Methylobacter species. In this study, the Methylobacter sp. B2 MAG was used to investigate its metabolic potential and phylogenetic affiliation. The MAG has a size of 4,086,539 bp, consists of 134 contigs and 3955 genes were found, of which 3902 were protein coding genes. All genes for CH4 oxidation to CO2 were detected, including pmoCAB encoding particulate methane monooxygenase (pMMO) and xoxF encoding a methanol dehydrogenase. No gene encoding a formaldehyde dehydrogenase was present and the formaldehyde to formate conversion follows the tetrahydromethanopterin (H4MPT) pathway. “Ca. Methylobacter favarea” B2 uses the Ribulose-Mono-Phosphate (RuMP) pathway for carbon fixation. Analysis of the MAG indicates that Na+/H+ antiporters and the urease system might be important in the maintenance of pH homeostasis of this strain to cope with acidic conditions. So far, thermoacidophilic Methylobacter species have not been isolated, however this study indicates that members of the genus Methylobacter can be found in distinct ecosystems and their presence is not restricted to freshwater or marine sediments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Volcanic and geothermal areas are hostile environments characterized by low pH, high temperature, geothermal gas emissions, and low O2 concentrations. One of the emitted geothermal gases is CH4, a potent greenhouse gas. Multiple studies have shown that aerobic methanotrophs may be important in reducing the emissions of geothermally produced CH4 (D'Alessandro et al. 2009; Etiope and Klusman 2002).
Phylogenetically, aerobic methanotrophs belong to the phyla Alphaproteobacteria, Gammaproteobacteria or Verrucomicrobia (Op den Camp et al. 2009). These methanotrophs are microorganisms that conserve energy by oxidizing CH4 to CO2, while using O2 as terminal electron acceptor (Hanson and Hanson 1996). The first step of the methane oxidation pathway involves the conversion of methane into methanol catalysed by a soluble or membrane-bound methane monooxygenase (Hanson and Hanson 1996). In the following steps, methanol is converted into CO2 via formaldehyde and formate using either lanthanide-dependent XoxF-type methanol dehydrogenase or a calcium-dependent MxaF-type methanol dehydrogenase (Keltjens et al. 2014). Carbon fixation occurs via the Ribulose-Mono-Phosphate (RuMP) pathway, the Serine Pathway or the Calvin–Benson–Bassham (CBB) cycle (Chistoserdova 2011; Khadem et al. 2011; Murrell 1992; Rasigraf et al. 2014; Sharp et al. 2014).
Previous studies using analysis of 16S rRNA genes or the diagnostic pmoA gene revealed the presence of methanotrophs in volcanic areas (Gagliano et al. 2014, 2016; Niemann et al. 2006). Thermoacidophilic methanotrophs from geothermal areas have been isolated and resulted in the first pure cultures of methanotrophic members of the phylum Verrucomicrobia (Dunfield et al. 2007; Erikstad et al. 2019; Islam et al. 2008; Pol et al. 2007; van Teeseling et al. 2014).
A recent metagenomic analysis of the volcanic soils of the Favara Grande, the main geothermal active area of Pantelleria Island, Italy, indicated the presence of a unique methanotrophic community, composed of Verrucomicrobia and Gammaproteobacteria (Picone et al. 2020). Different metagenome assembled genomes (MAGs) were retrieved. One of these MAGs was nearly complete and phylogenetic analysis showed that it represents the genome of a novel Methylobacter species. Typically, Methylobacter species are found in freshwater sediments and wetland soils, where they account for a large fraction of aerobic methanotrophs (Smith et al. 2018). Thermoacidophilic Methylobacter species have, so far, not been isolated. In this study, we determined the phylogenetic position of this Methylobacter, and analysed the encoded metabolic potential.
Materials and methods
Sampling location and DNA isolation
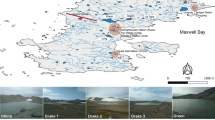
Samples were collected at Favara Grande, Pantelleria, Italy 2017 (FAV1, 36° 50′ 80″ N; 11° 57′ 170″ E) and (FAV2, 36° 50′ 77″ N; 11°57′ 160″ E) during a field campaign in June 2017 (Picone et al. 2020). Soil samples (1–10, 10–15 and 12–20 cm depth) were taken using a core sampler (diameter 1.5 cm), stored in sterile 50 mL tubes and kept at 4 °C until DNA was extracted. In situ pH values were 4–4.5 with temperatures from 60 to 67 °C. Two different DNA extraction methods were used, namely Fast DNA Spin kit for soil (MP Biomedicals, Santa Ana, California), according to manufacturer’s instructions, and the CTAB method (Allen et al. 2006). DNA extraction was only successful from the FAV2 sampling site and the reads from the different depths and different extraction methods were combined for assembly and binning. For more detail see Picone et al. (2020).
Genome sequencing, assembly and binning
The metagenome was sequenced on the Illumina sequencing platform. For library preparation the Nextera XT kit (Illumina, San Diego, California) was used according to the manufacturer’s instructions. Enzymatic tagmentation was performed starting with 1 ng of DNA, followed by incorporation of the indexed adapters and amplification of the library. After purification of the amplified library using AMPure XP beads (Beckman Coulter, Indianapolis), libraries were checked for quality and size distribution using the Agilent 2100 Bioanalyzer and the High sensitivity DNA kit. Quantitation of the library was performed by Qubit using the Qubit dsDNA HS Assay Kit (Thermo Fisher Scientific, Waltham, Massachusetts). The libraries were pooled, denatured and sequenced with the Illumina Miseq sequence machine (San Diego, California). Paired end sequencing of 2 × 300 base pairs was performed using the MiSeq Reagent Kit v3 (Illumina, San Diego, California) according the manufacturers protocol.
Reads were trimmed using BBDuk (BBMap), assembled by MEGAHIT v1.0.3 (Li et al. 2015) and binned using an in-house pipeline, using different binning algorithms, including BinSanity (Graham et al. 2017), COCACOLA (Lu et al. 2017), CONCOCT (Alneberg et al. 2014), MaxBin 2.0 (Wu et al. 2016), and MetaBAT 2 (Kang et al. 2019). DAS Tool 1.0 was used for consensus binning (Sieber et al. 2018) and CheckM was used to assess the MAG quality (Parks et al. 2015). The average nucleotide identity using BLAST (ANIb) is calculated using JSpeciesWS software with standard settings (Richter et al. 2016). An up-to-date Bacterial Core Gene (UBCG) phylogenetic tree was constructed using RAxML (Stamatakis 2014) on CIPRES Science Gateway V. 3.3 platform (Miller et al. 2012). PROKKA and the MicroScope platform were used to automatically annotate the draft genome (Seemann 2014; Vallenet et al. 2013) and genomic features were manually checked.
Results and discussion
The Favara Grande is the main geothermal gas-emitting area on Pantelleria Island, Italy. The soil in this region is acidic, of high temperature and exposed to geothermal gas emission (D'Alessandro et al. 2009). It is devoid of any plant growth. At the FAV2 site the following physicochemical parameters were observed: temperatures 60–67 °C, pH 4–4.5, CH4 1000–18,000 ppm and H2 125–8400 ppm (see also Picone et al. 2020). CH4 and H2 concentrations were lowest close to soil surface, indicating active consumption of these gases. Metagenomic analysis of FAV2 soil samples revealed the presence of a diverse community of methanotrophs including those belonging to the phyla Verrucomicrobia and Gammaproteobacteria, at 6–11% and 2.5–3% relative abundance, respectively. No alphaproteobacterial methanotroph was detected (Picone et al. 2020) (Fig. 1). One of these FAV2 methanotrophs was classified as a Methylobacter species. The Methylobacter sp. B2 MAG was chosen for detailed analysis to achieve a better understanding of its metabolic potential and its relevance in the carbon cycle of geothermal soils. The MAG had a size of 4,086,539 bp, consisted of 134 contigs and had a GC content of 47.2%. CheckM analysis revealed that the completeness of this MAG was 99.1% with only 0.4% contamination. A total of 3955 genes could be identified, of which 3902 were protein coding genes and 53 were RNA genes. Functions could be assigned to 2164 protein coding genes (Table 1). Moreover, 88.4% of the predicted genes were assigned into Clusters of Orthologous Groups and these COG functional categories are compiled in Table 2.
The relative abundance of the different binned MAGs (completeness > 95%) at the different depths of the geothermal soil of the Favara Grande, Pantelleria Island, Italy. The relative abundance of Methylobacter sp. B2 (MAG2) is 0.49%, 0.71% and 1.17% in the top layer, at 10–15 cm depth and at 15–20 cm depth, respectively
Phylogeny
The draft genome of Methylobacter sp. B2 contained one 16S rRNA gene. This gene is located at the end of a contig and the rest of the ribosomal RNA operon could not be detected. This often happens during binning due to high conservation of the rRNA gene sequences. We were able to assemble a contig with the full rRNA operon using all reads from the metagenome dataset and the contig with the 16S rRNA gene in MAG as a seed (Supplementary Material). Phylogenetic analysis of the 16S rRNA gene revealed that this gene clusters together with Methylobacter species (Supplementary Fig. S1, Kumar et al. 2016; Tamura 1992). The closest cultivated relative is Methylobacter psychrophilus Z-0021, showing a 16S rRNA gene identity of only 96.5%. Using a species boundary of 98.2%, this 16S rRNA identity indicated that the B2 MAG represented a novel species within the genus Methylobacter. Besides the 16S rRNA gene, pmoA is considered as molecular marker gene for defining methanotrophic taxa (Knief 2015). Phylogenetic analysis revealed of the pmoA gene in this MAG did not cluster with other Methylobacter species (Supplementary Fig. S2, Saitou and Nei 1987; Zuckerkandl and Pauling 1965). This is not uncommon as for Methylobacter species, single-gene phylogenies resulted in inconsistencies, since this genus is assumed to be polyphyletic (i.e. having more than one common ancestor) (Orata et al. 2018). To circumvent this one-gene-polyphyletic classification problem, an UBCG (Up-to-date bacterial core gene set) phylogenetic tree was constructed. Rather than a single gene, this tree is based on 92 bacterial core genes (Na et al. 2018). The UBCG phylogenetic analysis showed that Methylobacter sp. B2 clusters within the Methylobacter genus (Fig. 2). Furthermore, average nucleotide identity (ANI) calculations gave values well below the threshold for species delimitation (95–96%), demonstrating that this MAG indeed represents a novel species within the genus Methylobacter (Table 3) (Chun et al. 2018), for which we propose the name “Candidatus Methylobacter favarea” B2.
Up-to-date Bacterial Core Gene (UBCG) phylogenetic tree of MAG2 and members of the family Methylomonaceae, all with a genome completeness of > 90%. Hyphomicrobium denitrificans was used to root the tree, but removed from the tree for clarity. The tree was constructed using RAxmL. Bootstrap analysis was carried using 100 replications and percentage bootstrap values > 95% are indicated by black circles
Analysis of the encoded metabolic potential
Methanotrophy
The first step in the methane oxidation pathway is the conversion of methane into methanol catalyzed by a membrane-bound or soluble methane monooxygenase. Analysis of the draft genome (see Fig. 3 for a schematic representation) revelaed one pmoCAB gene cluster, encoding for the particulate membrane-bound, methane monooxygenase (pMMO). Typically, but not in all methanotrophs, the pmoCAB gene cluster is complemented with a pmoD gene. Recent studies show that pmoD is essential for pMMO activity, and probably involved in copper incorporation (Fisher et al. 2018). In draft genome of “Ca. Methylobacter favarea” B2, a homolog of the pmoD gene (METHB2_v1_630010) was present, but the gene is not located near the pmoCAB gene cluster. Genes encoding for the soluble methane monooxygenase (sMMO) were not detected.
Cell metabolism of “Ca. Methylobacter favarea” B2. Pmo particulate methane monoxygenase, XoxF XoxF-type methanol dehydrogenase, FDH formate dehydrogenase, NifDHK nitrogenase, Nirk dissimilatory nitrite reductase, NorB nitric oxide reductase, HoxHY hydrogenase. Enzyme complexes of the electron transport chain are labeled by Roman numerals
“Ca. Methylobacter favarea” B2 catalyses the second step of the methane oxidation pathway, the conversion of methanol into formaldehyde, using a rare earth element (REE) and pyrroloquinoline quinone (PQQ) dependent XoxF-type methanol dehydrogenase (MDH) (Keltjens et al. 2014; Pol et al. 2014). In the genome, XoxF (METHB2_v1_30111), the catalytic MDH protein and XoxJ (METHB2_v1_30112), involved in protein binding for stability or electron transfer (Versantvoort et al. 2019), were encoded. The xoxG gene, which encodes for the cytochrome cL, that accepts electrons from MDH (Keltjens et al. 2014), could not be detected within the xoxF/xoxJ gene cluster. However, XoxF-MDH systems do not follow a general organizational pattern at genomic level (Keltjens et al. 2014) and this MAG contains several mono-heme cytochromes with the heme-binding motif CxxCH and the conserved methionine typical for XoxG proteins (METHB2_v1_70040, METHB2_v1_200048, METHB2_v1_230021, METHB2_v1_350009, METHB2_v1_350010 and METHB2_v1_510007). These cytochromes could serve as electron acceptor for XoxF (Versantvoort et al. 2019). During methanol oxidation, electrons are transferred to the cytochrome via the cofactor PQQ and all genes for the synthesis of PQQ were present in this MAG. No genes encoding for the calcium-dependent MxaF-MDH could be identified.
XoxF-MDH requires lanthanides as metal cofactor (Pol et al. 2014) but the mechanism of their uptake in bacterial cells in not completely clarified. Recently, the lanthanide binding protein Lanmodulin was identified and postulated to function as cargo protein for lanthanides from a lanthanide transporter to MDH (Cotruvo et al. 2018). Lanmodulin seems not to be encoded in the MAG. Other studies indicated that TonB-dependent transporters might be involved in REE transport over the outer membrane, which are encoded by tonB or cirA genes (Ochsner et al. 2019; Roszczenko-Jasińska et al. 2020). The draft genome of “Ca. Methylobacter favarea” B2 contains several tonB and cirA genes (METHB2_v1_20123, METHB2_v1_30066, METHB2_v1_130008, METHB2_v1_200030 and METHB2_v1_300037) whereby METHB2_v1_20123 shows the highest similarity (33% based on amino acid composition) with the putative REE transporter of Methylorubrum extorquens PA1. None of these transporters are located near the methanol dehydrogenase genes as in Methylorubrum extorquens PA1 (Ochsner et al. 2019).
In the next step of the CH4 oxidation pathway, formaldehyde is oxidized to formate. Formaldehyde is a key metabolite, since it is used for carbon assimilation by proteobacterial methanotrophs. Furthermore, formaldehyde oxidation is considered to be essential, since intracellular concentrations of toxic formaldehyde should remain low (Chistoserdova 2011). However, this MAG does not seem to contain a gene encoding for formaldehyde dehydrogenase, which prevents direct formaldehyde to formate conversion. In vitro studies showed that XoxF-MDH from M. extorquens AM1 and M. fumariolicum SolV can use formaldehyde as substrate leading to the hypothesis that XoxF-type MDHs oxidize methanol into formate (Pol et al. 2014; Good et al. 2018). In contrast, in vivo, the XoxF-type MDH of M. extorquens AM1 was shown to produce formaldehyde, which can be used for carbon assimilation (Good et al. 2018).
In M. extorquens AM1 the lanthanide-dependent ethanol dehydrogenase ExaF was involved in formaldehyde oxidation (Good et al. 2018). As there was no ethanol dehydrogenase nor a MxaF-MDH found in the draft genome of “Ca. Methylobacter favarea” B2 to oxidize formaldehyde, indicating that this strain possibly uses a different pathway for formaldehyde oxidation. The most widespread formaldehyde conversion pathway is the glutathione-linked formaldehyde oxidation pathway. The first step in this pathway is the reaction of formaldehyde with glutathione. The glutathione-dependent formaldehyde dehydrogenase enzyme accelerates this spontaneous reaction (Goenrich et al. 2002). This enzyme could be detected in the “Ca. Methylobacter favarea” B2 MAG (METHB2_v1_3500004). In the following two steps, formate is formed by glutathione-dependent formaldehyde dehydrogenase and S-formylglutathione hydrolase, however none of these enzymes could be detected within “Ca. Methylobacter favarea” B2.
Alternatively, formaldehyde can be converted to formate by the tetrahydromethanopterin (H4MPT) pathway or the methylene-tetrahydrofolate (methylene-H4F) pathway. The genes for both pathways were detected in draft genome of “Ca. Methylobacter favarea” B2, suggesting that this strain can use both pathways, as described for Methylococcus capsulatus Bath (Chistoserdova et al. 2005). Proteomics studies revealed that the H4MPT-pathway is used for formaldehyde to formate conversion in M. capsulatus Bath (Kao et al. 2004). Whether “Ca. Methylobacter favarea” B2 uses the H4MPT-pathway or the methylene-H4F-pathway remains uncertain. However, the genes for the synthesis of the cofactors methanofuran (MFR) and tetrahydromethanopterin (THPMT) were lacking, whereas the genes needed for the synthesis of tetrahydrofolate (THF) were present. This suggests that the H4MPT-pathway can only be used whenever the cofactors are supplied by other members of the microbial community.
Formate dehydrogenase catalyses the final step in the methane oxidation pathway, namely the conversion of formate into CO2. Typically, methanotrophs encode for multiple formate dehydrogenases (Chistoserdova 2011; Flynn et al., 2016). “Ca. Methylobacter favarea” B2 encodes for two different formate dehydrogenases, a NAD-dependent FDH encoded by a single gene (METHB2_v1_310032) and a molybdopterin containing formate dehydrogenase encoded by the three genes fdhABC (METHB2_v1_500021, METHB2_v1_500022 and METHB2_v1_500023).
Energy conservation and respiration
“Ca. Methylobacter favarea” B2 uses O2 as terminal electron acceptor. The NADH:ubiquinone reductase genes (complex I, nuoABCDEFGHIJKLMN) were found in the MAG, together with genes encoding the succinate dehydrogenase (complex II), cytochrome bc1 (complex III) and cytochrome-c-oxidase (complex IV) complexes. The proton motive force generated by the respiratory chain can be used by the ATP-generating ATPase (complex V).
Carbon fixation
“Ca. Methylobacter favarea” B2 uses the ribulose-mono-phosphate (RuMP) pathway for carbon fixation, as do other Methylobacter species (Flynn et al. 2016). Interestingly, a nearly complete serine pathway could also be detected in the genome of “Ca. Methylobacter favarea” B2. Usually, the serine pathway is used by alphaproteobacterial methanotrophs as carbon fixation pathway and gammaproteobacterial methanotrophs only contain parts of the serine pathway. Typically, gammaproteobacterial methanotrophs lack the enzyme phosphoenolpyruvate carboxylase (Chistoserdova 2011), however, the “Ca. Methylobacter favarea” B2 MAG encodes for every enzyme in the serine pathway except the serine-glyoxylate aminotransferase. In contrast to Verrucomicrobial methanotrophs (Khadem et al. 2011), the Calvin–Benson–Bassham cycle cannot be used for carbon fixation in “Ca. Methylobacter favarea” B2, since RuBisCO genes are lacking.
Alternative substrates
All genes for the glycolysis and gluconeogenesis were found and the TCA cycle genes were present. Genes of the glyoxylate shunt were partially detected, the gene encoding for isocitrate lyase is found in the draft genome, but a gene encoding for malate synthase could not be detected. The pentose phosphate pathway was present. Furthermore, transporters for a variety of organic molecules could be predicted. This indicates that “Ca. Methylobacter favarea” B2 could benefit from a mixotrophic lifestyle or survival modus. Furthermore, “Ca. Methylobacter favarea” B2 seemed to be able to store a variety of carbon compounds, including glycogen, polyhydroxybutyrate and polyphosphates.
The “Ca. Methylobacter favarea” B2 MAG contained an oxygen tolerant NAD-coupled hydrogenase, belonging to group 3d bidirectional hydrogenases (Greening et al. 2016). H2 is an abundant electron donor in the FAV2 soils (D'Alessandro et al. 2009; Gagliano et al. 2016; Picone et al. 2020) and simultaneous CH4 and H2 oxidation is reported for different methanotrophs (Chen and Yoch 1987; Hanczar et al. 2002; Mohammadi et al. 2017; Carere et al. 2017). However, it is unlikely that “Ca. Methylobacter favarea” B2 can grow as ‘Knallgas’ bacterium, since it requires formaldehyde for carbon fixation. A mechanism for CO2 fixation is not detected in this MAG.
Nitrogen
Ammonia can be used as nitrogen source and is assimilated using either the glutamate dehydrogenase (METHB2_v1_670013) or the glutamine synthetase (METHB2_v1_100031) and glutamate synthase (METHB2_v1_410017). Nitrate and nitrite could also serve as nitrogen source. The activity of the assimilatory nitrate reductase NasA (METHB2_v1_40058) and nitrite reductase NirBD (METHB2_v1_40055, METHB2_v1_40054) would results in the production of ammonium. Genes encoding urease (METHB2_v1_20024, METHB2_v1_20025 and METHB2_v1_20026) and nitrogenase (nifHDK, METHB2_v1_180032, METHB2_v1_180033 and METHB2_v1_180034) were present, indicating that urea and N2 could also be used as nitrogen source in this severely N-limited ecosystem. The genes encoding the nitrite reductases NirK (METHB2_v1_00392), NirS (METHB2_v1_02231) and nitric oxide reductase NorBC (METHB2_v1_01347, METHB2_v1_01348) were found, suggesting that “Ca. Methylobacter favarea” B2 may be capable of partial denitrification.
pH homeostasis
In order for “Ca. Methylobacter favarea” B2 to thrive in an acid environment, it is important to control the intracellular pH. The maintenance of pH homeostasis is a result of restriction of proton permeation, internal consumption of protons and enhancement of proton pumps (Guan and Liu 2020). ATPases can pump out electrons and release acid stress, but this requires ATP (Liu et al. 2015a). “Ca. Methylobacter favarea” B2 encodes for different Na+/H+ antiporters (METHB2_v1_30076, METHB2_v1_70105, METHB2_v1_150016 and METHB2_v1_840011), which might be important for proton exchange as well (Slonczewski et al. 2009).
There are different mechanisms on intracellular proton consumption, which generates alkaline products. Several microorganisms use an amino-acid tolerance system to decrease the intracellular pH. However, both the arginine deaminase (ADI) system and the glutamate-dependent acid tolerance system (Liu et al. 2015b; Reeve and Reid 2016; Shabayek and Spellerberg 2017) could not be detected in the draft genome of “Ca. Methylobacter favarea” B2 Instead, “Ca. Methylobacter favarea” B2 could use the urease system for proton consumption. Since the genome encodes a urease, this enzyme can transform urea into ammonia and CO2 at the expense of a proton, whereby it regulates the internal pH (Miller and Maier 2014). Interestingly, the urease genes are widespread amongst the geothermal microorganisms of the Favara Grande (Picone et al. 2020), indicating that this urease system might be an important mechanisms in pH homeostasis within geothermal microorganisms.
Ecological role
Typically, Methylobacter species are found in freshwater oxic sediments, terrestrial habitats and marine ecosystems, where they account for a large fraction of aerobic methanotrophy (Hao et al., 2020; Khatri et al. 2019; Smith et al. 2018). Thermoacidophilic Methylobacter species have, so far, not been isolated. Thermophilic Gammaproteobacteria are found within the family Methylothermaceae and the genus Methylocaldum (Houghton et al. 2019) and not within the family Methylomonaceae. Previously, 16S rRNA gene amplicon sequencing and metagenomic sequencing revealed that Methylobacter sp. are abundant methanotrophs in the geothermal soils of the Favara Grande (Gagliano et al. 2016). Other geothermal soil microbial communities, such as the one in the Solfatara Crater near Naples, Italy, did not show the presence of Methylobacter species (Crognale et al., 2018), indicating that we still have to learn more about the metabolic diversity of this important group of methanotrophs.
Availability of data and materials
DNA sequences (raw sequence reads and MAGs) have been deposited in NCBI BioProject database with accession number PRJEB36447. The draft genome is available at NCBI under Accession No. GCA_902806695.
References
Allen GC, Flores-Vergara MA, Krasynanski S, Kumar S, Thompson WF (2006) A modified protocol for rapid DNA isolation from plant tissues using cetyltrimethylammonium bromide. Nat Protoc 1:2320–2325. https://doi.org/10.1038/nprot.2006.384
Alneberg J, Bjarnason BS, De Bruijn I, Schirmer M, Quick J, Ijaz UZ, Lahti L, Loman NJ, Andersson AF, Quince C (2014) Binning metagenomic contigs by coverage and composition. Nat Methods 11:1144–1146. https://doi.org/10.1038/nmeth.3103
Carere CR, Hards K, Houghton KM, Power JF, Mcdonald B, Collet C, Gapes DJ, Sparling R, Boyd ES, Cook GM, Greening C, Stott MB (2017) Mixotrophy drives niche expansion of verrucomicrobial methanotrophs. ISME J 11:2599–2610. https://doi.org/10.1038/ismej.2017.112
Chen YP, Yoch DC (1987) Regulation of two nickel-requiring (inducible and constitutive) hydrogenases and their coupling to nitrogenase in Methylosinus trichosporium OB3b. J Bacteriol 169:4778–4783. https://doi.org/10.1128/jb.169.10.4778-4783.1987
Chistoserdova L (2011) Modularity of methylotrophy, revisited. Environ Microbiol 13:2603–2622. https://doi.org/10.1111/j.1462-2920.2011.02464.x
Chistoserdova L, Vorholt JA, Lidstrom ME (2005) A genomic view of methane oxidation by aerobic bacteria and anaerobic archaea. Genome Biol 6:208. https://doi.org/10.1186/gb-2005-6-2-208
Chun J, Oren A, Ventosa A, Christensen H, Arahal DR, Da Costa MS, Rooney AP, Yi H, Xu XW, De Meyer S, Trujillo ME (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68:461–466. https://doi.org/10.1099/ijsem.0.002516
Cotruvo JA Jr, Featherston ER, Mattocks JA, Ho JV, Laremore TN (2018) Lanmodulin: a highly selective lanthanide-binding protein from a lanthanide-utilizing bacterium. J Am Chem Soc 140:15056–15061. https://doi.org/10.1021/jacs.8b09842
Crognale S, Venturi S, Tassi F, Rossetti S, Rashed H, Cabassi J, Capecchiacci F, Nisi B, Vaselli O, Morrison HG, Sogin ML, Fazi S (2018) Microbiome profiling in extremely acidic soils affected by hydrothermal fluids: the case of the Solfatara Crater (Campi Flegrei, southern Italy). FEMS Microbiol Ecol 94:fiy190. https://doi.org/10.1093/femsec/fiy190
D’Alessandro W, Bellomo S, Brusca L, Fiebig J, Longo M, Martelli M, Pecoraino G, Salerno F (2009) Hydrothermal methane fluxes from the soil at Pantelleria island (Italy). J Volcanol Geotherm Res 187:147–157. https://doi.org/10.1016/j.jvolgeores.2009.08.018
Dunfield PF, Yuryev A, Senin P, Smirnova AV, Stott MB, Hou S, Ly B, Saw JH, Zhou Z, Ren Y, Wang J, Mountain BW, Crowe MA, Weatherby TM, Bodelier PLE, Liesack W, Feng L, Wang L, Alam M (2007) Methane oxidation by an extremely acidophilic bacterium of the phylum Verrucomicrobia. Nature 450:879–882. https://doi.org/10.1038/nature06411
Erikstad H-A, Ceballos RM, Smestad NB, Birkeland N-K (2019) Global biogeographic distribution patterns of thermoacidophilic Verrucomicrobia methanotrophs suggest allopatric evolution. Front Microbiol 10:1129. https://doi.org/10.3389/fmicb.2019.01129
Etiope G, Klusman RW (2002) Geologic emissions of methane to the atmosphere. Chemosphere 49:777–789. https://doi.org/10.1016/S0045-6535(02)00380-6
Fisher OS, Kenney GE, Ross MO, Ro SY, Lemma BE, Batelu S, Thomas PM, Sosnowski VC, Dehart CJ, Kelleher NL, Stemmler TL, Hoffman BM, Rosenzweig AC (2018) Characterization of a long overlooked copper protein from methane- and ammonia-oxidizing bacteria. Nat Commun 9:4276. https://doi.org/10.1038/s41467-018-06681-5
Flynn JD, Hirayama H, Sakai Y, Dunfield PF, Klotz MG, Knief C, Op den Camp HJM, Jetten MSM, Khmelenina VN, Trotsenko YA, Murrell JC, Semrau JD, Svenning MM, Stein LY, Kyrpides N, Shapiro N, Woyke T, Bringel F, Vuilleumier S, Dispirito AA, Kalyuzhnaya MG (2016) Draft genome sequences of gammaproteobacterial methanotrophs isolated from marine ecosystems. Genome Announc 4:e01629. https://doi.org/10.1128/genomeA.01629-15
Gagliano AL, D’Alessandro W, Tagliavia M, Parello F, Quatrini P (2014) Methanotrophic activity and diversity of methanotrophs in volcanic geothermal soils at Pantelleria (Italy). Biogeosciences 11:5865–5875. https://doi.org/10.5194/bg-11-5865-2014
Gagliano AL, Tagliavia M, D’Alessandro W, Franzetti A, Parello F, Quatrini P (2016) So close, so different: geothermal flux shapes divergent soil microbial communities at neighbouring sites. Geobiology 14:150–162. https://doi.org/10.1111/gbi.12167
Goenrich M, Bartoschek S, Hagemeier CH, Griesinger C, Vorholt JA (2002) A Glutathione-dependent Formaldehyde-activating Enzyme (Gfa) from Paracoccus denitrificans Detected and Purified via Two-dimensional Proton Exchange NMR Spectroscopy. J Biol Chem 277(5):3069–3072. https://doi.org/10.1074/jbc.C100579200
Good NM, Walser ON, Moore RS, Suriano CJ, Huff AF, Martínez-Gómez NC (2018) Investigation of lanthanide-dependent methylotrophy uncovers complementary roles for alcohol dehydrogenase enzymes. bioRxiv 329011. https://doi.org/10.1101/329011
Graham ED, Heidelberg JF, Tully BJ (2017) BinSanity: unsupervised clustering of environmental microbial assemblies using coverage and affinity propagation. PeerJ 5:e3035. https://doi.org/10.7717/peerj.3035
Greening C, Biswas A, Carere CR, Jackson CJ, Taylor MC, Stott MB, Cook GM, Morales SE (2016) Genomic and metagenomic surveys of hydrogenase distribution indicate H2 is a widely utilised energy source for microbial growth and survival. ISME J 10:761–777. https://doi.org/10.1038/ismej.2015.153
Guan N, Liu L (2020) Microbial response to acid stress: mechanisms and applications. Appl Microbiol Biotechnol 104:51–65. https://doi.org/10.1007/s00253-019-10226-1
Hanczar T, Csaki R, Bodrossy L, Murrell JC, Kovacs KL (2002) Detection and localization of two hydrogenases in Methylococcus capsulatus (Bath) and their potential role in methane metabolism. Arch Microbiol 177:167–172. https://doi.org/10.1007/s00203-001-0372-4
Hanson RS, Hanson TE (1996) Methanotrophic bacteria. Microbiol Rev 60:439–447
Hao Q, Liu F, Zhang Y, Wang O, Xiao L (2020) Methylobacter accounts for strong aerobic methane oxidation in the Yellow River Delta with characteristics of a methane sink during the dry season. Sci Total Environ 704:135383. https://doi.org/10.1016/j.scitotenv.2019.135383
Houghton KM, Carere CR, Stott MB, McDonald IR (2019) Thermophilic methanotrophs: in hot pursuit. FEMS Microbiol Ecol 95:fiz125. https://doi.org/10.1093/femsec/fiz125
Islam T, Jensen S, Reigstad LJ, Larsen O, Birkeland N-K (2008) Methane oxidation at 55 degrees C and pH 2 by a thermoacidophilic bacterium belonging to the Verrucomicrobia phylum. Proc Natl Acad Sci USA 105:300–304. https://doi.org/10.1073/pnas.0704162105
Kang DD, Li F, Kirton E, Thomas A, Egan R, An H, Wang Z (2019) MetaBAT 2: an adaptive binning algorithm for robust and efficient genome reconstruction from metagenome assemblies. PeerJ 7:e7359. https://doi.org/10.7717/peerj.7359
Kao WC, Chen YR, Yi EC, Lee H, Tian Q, Wu KM, Tsai SF, Yu SS, Chen YJ, Aebersold R, Chan SI (2004) Quantitative proteomic analysis of metabolic regulation by copper ions in Methylococcus capsulatus (Bath). J Biol Chem 279:51554–51560. https://doi.org/10.1074/jbc.M408013200
Keltjens JT, Pol A, Reimann J, Op den Camp HJM (2014) PQQ-dependent methanol dehydrogenases: rare-earth elements make a difference. Appl Microbiol Biotechnol 98:6163–6183. https://doi.org/10.1007/s00253-014-5766-8
Khadem AF, Pol A, Wieczorek A, Mohammadi SS, Francoijs K-J, Stunnenberg HG, Jetten MSM, Op den Camp HJM (2011) Autotrophic methanotrophy in Verrucomicrobia: Methylacidiphilum fumariolicum SolV uses the Calvin–Benson–Bassham cycle for carbon dioxide fixation. J Bacteriol 193:4438–4446. https://doi.org/10.1128/jb.00407-11
Khatri K, Mohite JA, Pandit PS, Bahulikar R, Rahalkar MC (2019) Description of ‘Ca. Methylobacter oryzae’ KRF1, a novel species from the environmentally important Methylobacter clade 2. Antonie Van Leeuwenhoek 113:729–735. https://doi.org/10.1007/s10482-019-01369-2
Knief C (2015) Diversity and habitat preferences of cultivated and uncultivated aerobic methanotrophic bacteria evaluated based on pmoA as molecular marker. Front Microbiol 6:1346–1346. https://doi.org/10.3389/fmicb.2015.01346
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Li D, Liu CM, Luo R, Sadakane K, Lam TW (2015) MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31:1674–1676. https://doi.org/10.1093/bioinformatics/btv033
Liu Y, Lv C, Xu Q, Li S, Huang H, Ouyang P (2015a) Enhanced acid tolerance of Rhizopus oryzae during fumaric acid production. Bioprocess Biosystems Eng 38:323–328. https://doi.org/10.1007/s00449-014-1272-8
Liu Y, Tang H, Lin Z, Xu P (2015b) Mechanisms of acid tolerance in bacteria and prospects in biotechnology and bioremediation. Biotechnol Adv 33:1484–1492. https://doi.org/10.1016/j.biotechadv.2015.06.001
Lu YY, Chen T, Fuhrman JA, Sun F (2017) COCACOLA: binning metagenomic contigs using sequence COmposition, read CoverAge, CO-alignment and paired-end read LinkAge. Bioinformatics 33:791–798. https://doi.org/10.1093/bioinformatics/btw290
Miller EF, Maier RJ (2014) Ammonium metabolism enzymes aid Helicobacter pylori acid resistance. J Bacteriol 196:3074–3081. https://doi.org/10.1128/jb.01423-13
Miller MA, Pfeiffer W, Schwartz T (2012) The CIPRES science gateway: enabling high-impact science for phylogenetics researchers with limited resources. In: XSEDE '12: proceedings of the 1st conference of the extreme science and engineering discovery environment: bridging from the extreme to the campus and beyond, vol 39, pp 1–8. https://doi.org/10.1145/2335755.2335836
Mohammadi S, Pol A, van Alen TA, Jetten MSM, Op den Camp HJM (2017) Methylacidiphilum fumariolicum SolV, a thermoacidophilic ‘Knallgas’ methanotroph with both an oxygen-senstitive and—insensitive hydrogenase. ISME J 11:945–958. https://doi.org/10.1038/ismej.2016.171
Murrell JC (1992) Genetics and molecular biology of methanotrophs. FEMS Microbiol Rev 8:233–248. https://doi.org/10.1111/j.1574-6968.1992.tb04990.x
Na SI, Kim YO, Yoon SH, Ha SM, Baek I, Chun J (2018) UBCG: up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J Microbiol 56:280–285. https://doi.org/10.1007/s12275-018-8014-6
Niemann H, Lösekann T, De Beer D, Elvert M, Nadalig T, Knittel K, Amann R, Sauter EJ, Schlüter M, Klages M, Foucher JP, Boetius A (2006) Novel microbial communities of the Haakon Mosby mud volcano and their role as a methane sink. Nature 443:854–858. https://doi.org/10.1038/nature05227
Ochsner AM, Hemmerle L, Vonderach T, Nüssli R, Bortfeld-Miller M, Hattendorf B, Vorholt JA (2019) Use of rare-earth elements in the phyllosphere colonizer Methylobacterium extorquens PA1. Mol Microbiol 111:1152–1166. https://doi.org/10.1111/mmi.14208
Op den Camp HJM, Islam T, Stott MB, Harhangi HR, Hynes A, Schouten S, Jetten MSM, Birkeland N-K, Pol A, Dunfield PF (2009) Environmental, genomic and taxonomic perspectives on methanotrophic Verrucomicrobia. Environ Microbiol Rep 1:293–306. https://doi.org/10.1111/j.1758-2229.2009.00022.x
Orata FD, Meier-Kolthoff JP, Sauvageau D, Stein LY (2018) Phylogenomic analysis of the gammaproteobacterial methanotrophs (order Methylococcales) calls for the reclassification of members at the genus and species levels. Front Microbiol 9:3162. https://doi.org/10.3389/fmicb.2018.03162
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW (2015) CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res 25:1043–1055. https://doi.org/10.1101/gr.186072.114
Picone N, Hogendoorn C, Cremers G, Poghosyan L, Pol A, van Alen TA, Gagliano AL, D’Alessandro W, Quatrini P, Jetten MSM, Op den Camp HJM, Berben T (2020) Geothermal gases shape the microbial community of the volcanic soil of Pantelleria, Italy. mSystems 5:e00517-e520. https://doi.org/10.1128/mSystems.00517-20
Pol A, Heijmans K, Harhangi HR, Tedesco D, Jetten MSM, Op den Camp HJM (2007) Methanotrophy below pH 1 by a new Verrucomicrobia species. Nature 450:874–878. https://doi.org/10.1038/nature06222
Pol A, Barends TRM, Dietl A, Khadem AF, Eygensteyn J, Jetten MSM, Op den Camp HJM (2014) Rare earth metals are essential for methanotrophic life in volcanic mudpots. Environ Microbiol 16:255–264. https://doi.org/10.1111/1462-2920.12249
Rasigraf O, Kool DM, Jetten MS, Sinninghe Damste JS, Ettwig KF (2014) Autotrophic carbon dioxide fixation via the Calvin–Benson–Bassham cycle by the denitrifying methanotroph “Candidatus Methylomirabilis oxyfera". Appl Environ Microbiol 80:2451–2460. https://doi.org/10.1128/aem.04199-13
Reeve BWP, Reid SJ (2016) Glutamate and histidine improve both solvent yields and the acid tolerance response of Clostridium beijerinckii NCP 260. J Appl Microbiol 120:1271–1281. https://doi.org/10.1111/jam.13067
Richter M, Rossello-Mora R, Oliver Glockner F, Peplies J (2016) JSpeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics 32:929–931. https://doi.org/10.1093/bioinformatics/btv681
Roszczenko-Jasińska P, Vu HN, Subuyuj GA, Crisostomo RV, Cai J, Lien NF, Clippard EJ, Ayala EM, Ngo RT, Yarza F, Wingett JP, Raghuraman C, Hoeber CA, Martinez-Gomez NC, Skovran E (2020) Gene products and processes contributing to lanthanide homeostasis and methanol metabolism in Methylorubrum extorquens AM1. Sci Rep 10:12663. https://doi.org/10.1038/s41598-020-69401-4
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069. https://doi.org/10.1093/bioinformatics/btu153
Shabayek S, Spellerberg B (2017) Acid stress response mechanisms of group B Streptococci. Front Cell Infect Microbiol 7:395. https://doi.org/10.3389/fcimb.2017.00395
Sharp CE, Smirnova AV, Graham JM, Stott MB, Khadka R, Moore TR, Grasby SE, Strack M, Dunfield PF (2014) Distribution and diversity of Verrucomicrobia methanotrophs in geothermal and acidic environments. Environ Microbiol 16:1867–1878. https://doi.org/10.1111/1462-2920.12454
Sieber CMK, Probst AJ, Sharrar A, Thomas BC, Hess M, Tringe SG, Banfield JF (2018) Recovery of genomes from metagenomes via a dereplication, aggregation and scoring strategy. Nat Microbiol 3:836–843. https://doi.org/10.1038/s41564-018-0171
Slonczewski JL, Fujisawa M, Dopson M, Krulwich TA (2009) Cytoplasmic pH measurement and homeostasis in Bacteria and Archaea. Adv Microb Physiol 55(1–79):317. https://doi.org/10.1016/S0065-2911(09)05501-5
Smith GJ, Angle JC, Solden LM, Borton MA, Morin TH, Daly RA, Johnston MD, Stefanik KC, Wolfe R, Gil B, Wrighton KC (2018) Members of the genus Methylobacter are inferred to account for the majority of aerobic methane oxidation in oxic soils from a freshwater wetland. mBio 9:e00815–e00818. https://doi.org/10.1128/mBio.00815-18
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Tamura K (1992) Estimation of the number of nucleotide substitutions when there are strong transition-transversion and G+C-content biases. Mol Biol Evol 9:678–687. https://doi.org/10.1093/oxfordjournals.molbev.a040752
Vallenet D, Belda E, Calteau A, Cruveiller S, Engelen S, Lajus A, Le Fevre F, Longin C, Mornico D, Roche D, Rouy Z, Salvignol G, Scarpelli C, Thil Smith AA, Weiman M, Medigue C (2013) MicroScope—an integrated microbial resource for the curation and comparative analysis of genomic and metabolic data. Nucl Acids Res 41:D636–D647. https://doi.org/10.1093/nar/gks1194
van Teeseling MCF, Pol A, Harhangi HR, van der Zwart S, Jetten MSM, Op den Camp HJM, Van Niftrik L (2014) Expanding the verrucomicrobial methanotrophic world: description of three novel species of Methylacidimicrobium gen. nov. Appl Environ Microbiol 80:6782–6791. https://doi.org/10.1128/aem.01838-14
Versantvoort W, Pol A, Daumann LJ, Larrabee JA, Strayer AH, Jetten MSM, van Niftrik L, Reimann J, Op den Camp HJM (2019) Characterization of a novel cytochrome cGJ as the electron acceptor of XoxF-MDH in the thermoacidophilic methanotroph Methylacidiphilum fumariolicum SolV. Biochim Biophys Acta 1867:595–603. https://doi.org/10.1016/j.bbapap.2019.04.001
Wu YW, Simmons BA, Singer SW (2016) MaxBin 2.0: an automated binning algorithm to recover genomes from multiple metagenomic datasets. Bioinformatics 32:605–607. https://doi.org/10.1093/bioinformatics/btv638
Zuckerkandl E, Pauling L (1965) Molecules as documents of evolutionary history. J Theor Biol 8:357–366. https://doi.org/10.1016/0022-5193(65)90083-4
Acknowledgements
CH, NP and HOdC were supported by the European Research Council (ERC Advanced Grant Project VOLCANO 669371), LP was supported by the Netherlands Earth System Science Centre (NESSC 024002001) and MSMJ was supported by the European Research Council (ERC Advanced Grant Project Eco_MoM 339880) and The Soehngen Institute of Anaerobic Microbiology (SIAM 024002002).
Funding
The research was supported by the European Research Council (ERC Advanced Grant Project VOLCANO 669371 and ERC Advanced Grant project Eco_MoM 339880), by the Netherlands Earth System Science Centre (NESSC 024002001) and by The Soehngen Institute of Anaerobic Microbiology (SIAM 024002002).
Author information
Authors and Affiliations
Contributions
CH, NP, AP, MJ, and HOdC designed the projects and experiments. CH, NP, AP, AG, WD’A, PQ, and HOdC sampled the geothermal soils. CH and NP performed DNA extraction. TA sequenced the genome. JF and TA reconstructed the genome and FH, SV, LP, CH, NP and HOdC annotated the genome. CH, FH, SV and HOdC wrote the manuscript. All authors contributed to revision of the manuscript, and read and approved the submitted version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Ethical standards
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hogendoorn, C., Picone, N., van Hout, F. et al. Draft genome of a novel methanotrophic Methylobacter sp. from the volcanic soils of Pantelleria Island. Antonie van Leeuwenhoek 114, 313–324 (2021). https://doi.org/10.1007/s10482-021-01525-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-021-01525-7