Abstract
Background
CDH1 germline mutations lead to hereditary diffuse gastric carcinomas. However, it is unclear whether genetic variations in the CDH1 promoter affect the progression of sporadic gastric carcinomas (SGCs).
Methods
SGC patients in two independent cohorts with follow-up data were enrolled. The CDH1 genotypes, including the − 73A > C polymorphism (rs28372783), were determined by PCR sequencing. The CDH1 promoter activity was determined using reporter assays. SNAIL bound to CDH1 alleles was determined by chromatin immunoprecipitation primer extension PCR. CDH1 DNA methylation was determined by bisulfite-based PCR analyses.
Results
Kaplan–Meier analyses showed that the overall survival (OS) of the − 73C/C patients was significantly longer than that of the − 73A/C or − 73A/A patients in a Chinese cohort [n = 526; hazard ratio 0.68 (95% CI 0.47–1.00)], which was validated in an independent Korea cohort [n = 215; hazard ratio 0.49 (95% CI 0.26–0.94)]. Moreover, the transcription activity of the − 73C alleles was significantly higher than that of the − 73A alleles in vitro and in vivo. The ratio of SNAIL recruited to the promoter regions of the − 73C and − 73A alleles was 1:10, indicating a strong influence of this polymorphism on the recruitment of SNAIL to the flanking E-box. The prevalence of DNA methylation of the CpG island and shore within the promoter of the − 73C allele was much less than that of the − 73A allele in both gastric tissues and cancer cell lines.
Conclusion
The − 73A > C variation may lead to differences in the overall survival of SGC patients and allele-specific repressions of CDH1.
Similar content being viewed by others
Background
E-cadherin, an epithelial adhesion molecule encoded by the CDH1 gene, participates in the establishment and maintenance of intercellular adhesion, cell polarity, and tissue architecture. It is well recognized that the loss of E-cadherin expression promotes cancer metastasis through the epithelial–mesenchymal transition (EMT) mechanism. CDH1 germline mutations have been found in patients from hereditary diffuse gastric cancer (HDGC) families [1, 2]. The single nucleotide polymorphisms (SNPs) − 160C > A (rs16260) and − 347del > A (rs5030625) in the CDH1 promoter contribute to the susceptibility of sporadic gastric carcinomas (SGCs) [3, 4]. The − 160C > A and − 73A > C (rs28372783) SNPs affect CDH1 transcription suppression [5,6,7,8]. However, it is unknown whether SNPs in the CDH1 promoter affect the progression of SGCs. In addition, decreased CDH1 expression through promoter DNA methylation is frequently detected in many cancers and is associated with poor prognosis [9,10,11,12,13,14,15,16]. However, it is not clear if the methylation status of the CDH1 CpG islands is affected by genetic factors.
The − 73C allele is a common CDH1 allele in East Asians, with an allele frequency of 12.0% in the Japanese population (NCBI Sub SNP ss35074623) and 15.5% in the Chinese population [3]. In our previous study, we did not observe a significant correlation between this polymorphism and a susceptibility to SGCs [3]. We subsequently carried out a long-term follow-up and report here, for the first time, that the overall survival (OS) of SGC patients with the − 73C/C genotype of CDH1 is much longer than those with other genotypes of CDH1 in two independent cohorts. The possible mechanisms accounting for this allele-specific difference were also extensively studied.
Methods
Subjects and CDH1 genotyping
572 SGC patients surgically treated at the Peking University Cancer Hospital and Institute (2000–2006) were enrolled in our previous CDH1 SNP case–control study in 2008 [3], and were followed up for at least five years. An additional 215 SGC patients from the Seoul National University Hospital in 2004 were recruited into the present study as a validation cohort. The 2003 TNM-6 stage system was used for to classify SGC [17]. Patients without survival data or those who died within 2 months after surgery were excluded from the study.
Genotypes of the CDH1 gene in Korean patients were determined through the direct sequencing of a 119 bp PCR product amplified with the primer set (forward, 5′-cgcgtctatg cgaggccgggt-3′; reverse, 5′-accgcccccc gtaccgctga-3′) under thermal cycling conditions (95 °C for 5 min, [95 °C 30 s, 58 °C for 30 s, 72 °C for 30 s] × 35 cycles, followed by 72 °C for 10 min) using the genomic DNA extracted from formalin-fixed paraffin-embedded (FFPE) surgical margin (SM) tissue specimens as templates.
Cell lines, culture, and authentication
The cancer cell lines Caski, SiHa, HeLa, and MCF7 (kindly provided by Prof. Yang Ke at the Peking University Cancer Hospital) were cultured in DMEM medium supplemented with 10% fetal bovine serum (FBS, Gibco). The MKN74 cell line (kindly provided by Prof. Yasuhito Yuasa at the Tokyo Medical and Dental University) was cultured in RPMI1640 medium with 10% FBS. AGS (kindly provided by Dr. Chengchao Shou at the Peking University Cancer Hospital) and PC-3 (purchased from Cell Line Bank, Chinese Academy of Medical Sciences) were cultured in F12 medium with 10% FBS. All of these cell lines were maintained at 37 °C in humidified air with 5% CO2. These cell lines were tested and authenticated by the Beijing GENEWIZ Biotechnology Company before they were used in this study. STR patterns were analyzed using the Promega GenePrint10 System Kit and were matched with the American Type Culture Collection (ATCC) [18].
Quantitative RT-PCR
Total RNA was extracted from cell lines and frozen gastric tissues using TRIZol Reagent (Invitrogen) and was then reverse transcribed with the EasyScript First-Strand cDNA Synthesis SuperMix (Transgen Biotech Co. Beijing). The RT-PCR primers for amplifying CDH1 mRNA (151-bp) were 5′-gaacgcattg ccacatacac-3′ and 5′-gaattcgggc ttgttgtcat-3′. GAPDH (226-bp), and Alu repeat transcripts were used as reference genes in the regular RT-PCR (35 cycles) and quantitative RT-PCR, respectively [19].
Western blot
E-cadherin protein was analyzed by western blot as previously described [8]. The MYC-tag antibody (AB9132, Abcam) was used to detect the expression level of SNAIL. GAPDH or ACTIN was used as an internal reference.
Dual-luciferase reporter assay
The CDH1 promoter of the pGL3-A and pGL3-C reporter plasmids (pGL-A and pGL-C; − 108 nt ~ + 31 nt) were the same as described previously (Fig. S1 in the Electronic supplementary material, ESM) [8]. These reporter plasmids were co-transfected into the Caski and MKN74 cell lines using the Roche X-tremeGENE HP DNA Transfection Reagent (Cat. 06366236001) with the pcDNA3.1-Snail-myc (85 nt ~ 879 nt in Snail mRNA NM_005985) or its control vector (kindly provided by Prof. Zhiqian Zhang, Peking University Cancer Hospital) and the pRL-SV40 vector containing the Renilla luciferase gene. Luciferase activities were measured with a Dual-Luciferase Reporter Assay System (Promega, USA) and a luminometer (Molecular Devices, LMAXII Microplate Reader), and were normalized against the activity of the Renilla luciferase. The pGL3-Basic vector was used as a negative control. Cells in three parallel wells were tested for each treatment. Independent experiments were performed in triplicate. Statistics were performed using an unpaired two-tailed Student’s t test.
Chromatin immunoprecipitation (ChIP) primer extension assay
The Caski and MKN74 cell lines were transfected with pcDNA3.1-Snail-myc (or empty vector) and an equal amount of the pGL-A and pGL-C reporter vectors for 48 h. These cells were then collected and used in the ChIP assay as previously described [20, 21]. Briefly, the goat anti-MYC-tag (AB9132, Abcam) antibody was used to precipitate the overexpressed SNAIL protein and the bound DNA. The 119-bp endogenous CDH1 promoter fragment covering the − 73SNP site was amplified with HiFi DNA polymerase (Beijing Transgen Biotech) as described above, and was purified using the Tiangen DNA Purification Kit for subsequent primer extension analyses. The extension primer (5′-ctgattggct gtggccggca ggtga-3′) matching the 5′-flanking region of the − 73SNP was extended at 96 °C for 1 min, followed by 50 cycles (96 °C 10 s, 55 °C 20 s, 65 °C 40 s) of PCR. The 20 μl of reaction mix consisted of 50 ng of purified PCR product, 0.5 μl of Thermo Sequenase, 2 μl of 10× buffer, 15 pmol of extension primer, and 50 μmol/l ddATP and ddCTP. The − 73C and − 73A extended products were separated using DHPLC at 80 °C (oligo detection model) [22]. The peak-height ratio of the − 73A product to the − 73C product was calculated and was adjusted for the ratio of the input sample. The PCR products of the pGL-A and pGL-C reporter vectors were used as − 73C and − 73A positive controls, respectively. All of the assays were run at least in triplicate.
Analysis of the methylation status of the CDH1 CpG islands
Genomic DNA was isolated via phenol–chloroform extraction from the gastric tissues and cell lines and modified with 5 mol/l sodium bisulfite at 50 °C overnight [23].
All SGC tissue samples from − 73C/C patients (n = 14), and about four times that number of SGC samples from − 73A/A patients (n = 54) and from − 73A/C patients (n = 52), as characterized in our published study [3], were selected as representative samples for comparing CDH1 methylation levels. The methylation status of the CDH1 promoter was analyzed using the reported methylation-specific PCR (MSP) method (Fig. S1) [24] and a denatured high-performance liquid chromatography assay (DHPLC) [25, 26]. Briefly, a CpG-free primer set (forward primer, 5′-gattttagta attttaggtt agagggt-3′; reverse primer, 5′-actccaaaaa cccataacta acc-3′) was used to amplify both the methylated and unmethylated 329-bp DNA templates in the CDH1 CpG island in a HotStart touch-down PCR reaction (Fig. S1 in the ESM). The PCR conditions were as follows: 95 °C for 15 min to activate the Taq DNA polymerase (Qiagen GmbH, Hilden, Germany); then 15 touch-down cycles at 95 °C for 30 s, 58 °C for 30 s (−1 °C/cycle), and 72 °C for 30 s; followed by 25 regular cycles of 95 °C for 30 s, 43 °C for 30 s, and 72 °C for 30 s; and a final extension at 72 °C for 10 min. The methylated and unmethylated CDH1 PCR products were separated and quantified using a DHPLC set (WAVE™ DNA Fragment Analysis System) coupled to a high-sensitivity fluorescence (FL) detector (excitation at 450 nm, emission at 520 nm) at the partial denaturation temperature of 55.6 °C. The proportion of methylated CDH1 was calculated as follows: proportion of methylated CDH1 (%) = [peak area of methylated CDH1]/[combined peak area of methylated and unmethylated CDH1] × 100. The 329-bp bisulfite-PCR products were also used for clone sequencing.
To determine the methylation status of the CpG island 5′-shore within the CDH1 promoter, the 443-bp and 519-bp fragments were amplified from the sense- and antisense-strands of the CDH1 promoter with two CpG-free universal primer sets (S-forward primer, 5′-ttttagttat tagagaggtt gg-3′; S-reverse primer, 5′-taactacaac caaataaacc c-3′; AS-forward primer, 5′-gttttaaggg tttatggttg gt-3′; AS-reverse primer, 5′-accactacac tccaacttaa ataaa-3′) and were clone-sequenced, respectively (Fig. S1 in the ESM).
Statistical analysis
Differences in the methylation-positive rates were analyzed by the chi-square test, while differences in the proportion of methylated CDH1 and the relative mRNA levels were analyzed by the nonparametric Kruskal–Wallis test. The univariate survival rate was estimated by the Kaplan–Meier method, and the multivariate survival was analyzed using Cox proportional hazard regression models. All of the analyses were carried out using the SPSS 16.0 statistical software package.
Results
Overall survival of the − 73C/C patients was significantly longer than that of the − 73A/A or − 73A/C patients
To study whether SNPs in the CDH1 promoter are associated with the survival of SGC patients, all of the SGC patients (n = 572) enrolled in our previous CDH1 SNP study [3] were followed up for at least 5 years. Postoperative OS data were obtained from 526 patients (92.0%; median OS 34 months) (Table 1). 171 patients (32.5%) were at pTNM stage IV, and 330 patients (62.7%) died during the follow-up. The Kaplan–Meier analyses showed that the OS of the − 73C/C carriers (n = 12) was twofold higher than that of the − 73A/A or − 73A/C carriers (n = 514) [hazard ratio (HR) 0.68 (95% CI 0.47–1.00)]; log rank p = 0.036) (Table 1 and Fig. 1a). Age, location, vessel embolus, and pTNM stage were also significant survival factors. In the multivariate analysis, the − 73C/C genotype was still a significant indicator of good survival after adjusting for age, sex, location, differentiation, pTNM stage, and adjunctive therapy (Table 2). Such a difference in survival was not observed among the patients with different genotypes of − 160C > A and − 347del > A variations (Fig. 1b, c).
The overall survival of SGC patients with various genotypes of the CDH1 gene. Kaplan–Meier survival curves are shown for Chinese and Korean SGC patients with different genotypes of − 73A > C (a and d), − 160C > A (b), and − 374del > A (c) polymorphisms; the significant p values in the log rank test are also labeled
To confirm the above results, an additional 215 Korea SGC patients were used as an independent validation cohort. Fifty-nine patients (27.4%) were at pTNM stage IV, and 79 patients (36.7%) died during the follow-up. The − 73A/A genotype significantly increased the risk of lymph metastasis for SGC patients in the Korean cohort (p < 0.001), but did not increase the risk of metastasis in the Chinese cohort (Table S1 in the ESM). Because only two − 73C/C patients were present in this cohort, we compared the OS of the − 73C/C (n = 2) or − 73A/C (n = 42) patients with that of the − 73A/A patients. As expected, the OS of the − 73C/C or − 73A/C patients was also significantly longer than that of the − 73A/A patients (n = 171) [HR 0.49 (95% CI 0.26–0.94); Fig. 1d]. Together, these results suggest that the CDH1 − 73C allele may be a favorable survival factor for SGC patients.
Transcription activity of the − 73C allele is higher than that of the − 73A allele
To study the molecular mechanisms of the − 73A > C genotypes affecting SGC progression, we compared the transcription level of CDH1 in representative gastric samples from patients with different − 73A > C genotypes. A regular RT-PCR analysis showed that the CDH1 mRNA-positive rate in the − 73C/C gastric samples was significantly higher than in the − 73A/C or − 73A/A gastric samples (SGCs: 6/6 vs. 28/38), especially in SM samples (8/11 vs. 12/38, p = 0.033; Fig. 2a). A quantitative RT-PCR analysis confirmed this. The expression level of the CDH1 mRNA was gradually decreased in the SGCs from the − 73C/C carriers (n = 10) to the − 73A/C carriers (n = 35) and the − 73A/A carriers (n = 88): 18.13 vs. 4.89 vs. 3.69 (median − 73A/A vs. − 73C/C, p = 0.081; Fig. 2b).
The mRNA expression of CDH1 in gastric tissue samples from SGC patients with various − 73A > C genotypes. a CDH1 mRNA level in representative SGC (T) and paired surgical margin (N) samples from patients carrying the − 73A/A, − 73A/C, and − 73C/C genotypes based on regular RT-PCR analysis (35 cycles). The CDH1 mRNA-positive rate in the − 73C/C, − 73A/C, and − 73A/C samples was 100% (6/6), 75.0% (12/16), and 72.7% (16/22), respectively, for SGCs; and 72.7% (8/11), 18.8% (3/16), and 40.9% (9/22), respectively, for surgical margins (8/11 vs. 12/38, p = 0.033). b Relative CDH1 mRNA level (×10−5) in SGC tissues with different genotypes based on quantitative RT-PCR analysis. The median value for each group of samples is indicated
A dual-luciferase reporter assay was then used to investigate whether the − 73A > C variation directly affects the CDH1 promoter activity (Fig. S2 in the ESM). Reporter analysis showed that the promoter activity of the pGL-C vector was significantly higher than that of the pGL-A vector in the human gastric cancer cell lines AGS and MKN74, as well as in three other cancer cell lines—MCF7, PC-3, and Caski—in which the CDH1 CpG islands are not methylated [8]. Together, these results suggest that the CDH1 − 73C alleles are more transcriptionally active than the − 73A alleles in vitro and in vivo.
Reduced recruitment of the transcriptional repressor SNAIL to the − 73C allele compared with the − 73A allele
SNAIL is a strong repressor of CDH1 transcription that directly binds to the E-box sequences in the CDH1 promoter [27, 28]. One of the E-boxes flanks the − 73A > C SNP (Fig. S1 in the ESM). To investigate whether the − 73A > C variation affects the repressive effect of SNAIL on CDH1 transcription, the above reporter assay was also carried out in the gastric cancer cell line MKN74 [carrying the − 73A/A CDH1] with or without epitopic SNAIL overexpression (Fig. 3a). Remarkably, significant inhibition of the reporter activity due to SNAIL overexpression was observed for pGL-A but not for pGL-C (Fig. 3b).
Repression of the promoter activities of the − 73C and − 73A alleles by SNAIL. a Results of western blot analysis for the ectopic SNAIL protein expression. b Comparison of the promoter activities of pGL-A and pGL-C in MKN74 cells with SNAIL overexpression. The decrease in transcription activity is indicated (p < 0.001). c Comparison of the binding of SNAIL to the endogenous − 73A and − 73C alleles in Caski cells; the ratio for the binding of SNAIL to the − 73C and − 73A alleles is indicated
Most importantly, a significant difference in the binding of SNAIL to the endogenous − 73A and − 73C alleles in Caski cells [carrying the − 73A/C CDH1] was observed following chromatin DNA immunoprecipitation with a SNAIL antibody, as determined by the ChIP primer extension DHPLC analysis. The amount of endogenous − 73A allele bound to SNAIL was tenfold (3.53:0.35) higher than the amount of − 73C allele bound to SNAIL (Fig. 3c). These results strongly suggest that the − 73C allele is more refractory to transcriptional repression by SNAIL than the − 73A allele.
Less CDH1 methylation in the − 73C allele than in the − 73A allele
To investigate whether the − 73A > C variation causes long-term effects on CDH1 transcription through epigenetic mechanisms, the prevalence of methylation in the CpG island near the CDH1 transcription start site was studied. DHPLC analysis showed that the CDH1 methylation-positive rate was significantly increased in SGC from 121 representative patients with − 73C/C, − 73A/C, or − 73A/A genotypes (42.9 vs. 50.0 vs. 74.5%, trend test, p = 0.013; Table S2 in the ESM). The proportion of methylated-CDH1 CpG islands in the CDH1 methylation-positive SGCs was also increased (p = 0.019). Furthermore, the prevalence of CDH1 methylation in the − 73A/C SGCs was significantly lower than that in the − 73A/A SGCs. Subtle CDH1 methylation differences were also observed among the SGC samples with different − 160C > A or − 347del > A genotypes. These results indicate that the CpG island that contains the − 73C variation is more resistant to methylation than its − 73A counterpart in SGC tissues.
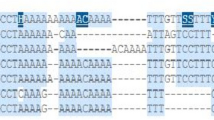
Bisulfite sequencing further confirmed the difference in methylation between the − 73C and − 73A alleles in the CDH1 promoter region in three CDH1 methylation-positive samples (#0163N, #0379N, and #0375T) in the DHPLC analysis (Fig. 4a). The cluster of extensively methylated CpGs in the promoter region was only detected in the − 73A alleles, not in the − 73C alleles (the regions highlighted in red in Fig. 4b). No cluster of methylated CpGs was observed in the clones from the CDH1 methylation-negative control sample (#0385N).
The methylation status of the CDH1 CpG islands in representative SGC samples. a DHPLC chromatograms for the methylated-CDH1 (M) and unmethylated-CDH1 (U) PCR products (329 bp). The cancer cell lines MCF7 and HeLa were used as CDH1 methylation-negative and -positive controls. b Results of clone sequencing for the 329-bp PCR products. Each line represents one clone. The black dots represent methylated CpG sites. The allelic information at the − 160 and − 73 SNPs in each clone is labeled. The fragments highlighted in red are the − 73C allele-specific promoter regions in which methylated CpG clusters were not observed
In addition, we mined the publicly available databases and found that four CpG sites within the CpG island 5′-shore region may be seeding-methylation sites for the CDH1 CpG islands in both human cell lines and tissues (Figs. S3 and S4 in the ESM). Notably, we observed a − 73A allele-specific 5′-shore methylation in both of the available cancer cell lines [carrying − 73A/C CDH1] and the gastric cancer tissues by deep clone sequencing. The proportion of the − 73A clones containing at least three methylated CpG sites in the 5′-shore region was significantly higher than that of the − 73C clones in two CDH1-expressing cell lines: 14/14 vs. 0/20 for the Caski cells, and 19/22 vs. 10/28 for the Siha cells (Fig. 5; p < 0.001). The proportion of shore-methylated clones among the − 73A alleles was also higher than that among the − 73C clones in two representative gastric SM samples: 12/129 vs. 1/70 (p = 0.036) for sample #0187N (Fig. S5 in the ESM) and 12/145 vs. 1/52 (p = 0.190) for sample #0379 N (Fig. S6 in the ESM). Collectively, all of the above results indicate that the CDH1 promoter is marked by the − 73A allele-specific methylation.
Comparison of the methylation status of the CpG sites in the promoter CpG island and the 5′-shore regions in two CDH1-expressing and − 73A/C genotype cancer cell lines. a Results of the RT-PCR analysis of CDH1 mRNA; cDNA from HeLa cells was used as the negative control. b Results of the bisulfite sequencing of the 519-bp fragment around the CDH1 transcription start site. The proportion of the − 73A clones that contained at least three methylated CpGs in the shore region was much higher than the corresponding proportion of the − 73C clones [14/14 vs. 0/20 for the Caski cell line and 19/22 vs. 10/28 for the Siha cell line (p < 0.001)]. Deep-red dots represent methylated-CpG sites, green dots indicate − 73A or − 347A, pink dots indicate − 73C or − 160C, blue dots indicate − 347del, highlighted clones are shore-unmethylated clones
Discussion
Studies of whether these polymorphisms contribute to CDH1 methylation and SGC prognosis are rare. In the present study, we report, for the first time, that the − 73C allele, especially the − 73C/C genotype, of CDH1 is a significant, independent, indicator of good survival for Chinese SGC patients. Similar results were also observed in a Korea validation cohort. Compared to the − 73A allele, the − 73C allele has high transcriptional activity, a low sensitivity to SNAIL repression, and a low probability of DNA methylation.
It is well recognized that epigenetic alterations that result from crosstalk between the genome and environmental factors play important roles in cancer development. The Genotype-Tissue Expression (GTEx) project demonstrated that local genetic variation affected gene expression levels for the majority of genes [29,30,31]. However, whether the epigenetic alteration mediates the effect of genetic variation on gene expression and whether the epigenetic alteration has a genetic foundation remain hot topics for epigenetic studies. Corso et al. reported promoter hypermethylation in an HDGC family with a CDH1 P373L germline missense mutation [32]. Yamada et al. reported the absence of monoallelic promoter hypermethylation of CDH1 in the gastric tissues from three SGC patients with lost CDH1 expression [7]. Here, we report that the CpG island and 5′-shore regions within the CDH1 promoter are marked by the − 73A allele-specific methylation in both gastric tissues and cancer cell lines. More binding of the repressor SNAIL to the − 73A alleles than the − 73C alleles may contribute to the − 73A allele-specific methylation.
The − 73A variant is highly linked with the − 160A and − 347A variants in the human genome [3]. Li et al. reported that the − 160A allele decreased the promoter activity by 68% compared with the − 160C allele, and Shin et al. reported that the − 347A allele decreased the promoter activity by tenfold compared with the − 347del allele using the luciferase reporter assay [5, 33]. Borges et al. reported a higher methylation frequency in the promoter region in the − 160A CDH1 allele than in the − 160C allele in gastric tissues of SGC patients [34]. In the present study, we found a slight increase in CDH1 promoter methylation in SGC from patients carrying the − 160C and − 347del alleles compared to those carrying the − 160A and − 347A alleles, but this increase is not statistically significant. In addition, a relationship between the − 160C > A or − 347del > A polymorphism and overall survival of SGC patients was not found. Upon analyzing the results of the deep bisulfite clone sequencing for a representative sample carrying the − 347del/del and − 160C/C CDH1 (#0187N; Fig. S6 in the ESM), the − 73A allele-specific methylation was still observed in the shore region, indicating that it is the − 73A variant, not the − 160A or the − 347A variant, that contributes to allele-specific CDH1 promoter methylation. This may also account for the result that it is − 73A > C, not− 160C > A or − 347del > A, that is significantly associated with patient survival. Our results support the notion that reconstituting E-cadherin expression or targeting molecules downstream holds promise in cancer therapies [35].
Interestingly, the − 73A/A genotype was a statistically significant risk factor for lymph metastasis of SGC among Korean patients, but not among Chinese patients. A significant survival difference between − 73A/A patients and − 73A/C and C/C patients was also observed in the Korean cohort, but not in the Chinese cohort. Generally, the population heterogeneity in South Korea is much smaller than that in mainland China. The SGC risk factors and genetic backgrounds may also be somewhat different for these two countries. The factors that contribute to the difference in − 73A > C sensitivity between the SGC patients in these two cohorts should be studied further.
Conclusions
The present study indicates that the − 73C allele of the CDH1 gene may be a favorable genetic survival factor for SGC patients. Compared to the − 73A allele, the − 73C allele has a higher promoter activity, a decreased repressor binding affinity, and a lower DNA methylation sensitivity, which may contribute to the longer survival of SGC patients. CDH1 and molecules downstream may serve as potential molecular cancer therapy targets.
References
van Roy F, Berx G. The cell–cell adhesion molecule E-cadherin. Cell Mol Life Sci. 2008;65:3756–8.
Peek R, Reddy KR. Genetic mutations identified for hereditary diffuse gastric cancer. Gastroenterology. 2007;133:379–80.
Zhang B, Pan K, Liu Z, Zhou J, Gu L, Ji J, et al. Genetic polymorphisms of the E-cadherin promoter and risk of sporadic gastric carcinoma in Chinese populations. Cancer Epidemiol Biomark Prev. 2008;17:2402–8.
Li YL, Tian Z, Zhang JB, Fu BY. CDH1 promoter polymorphism and stomach cancer susceptibility. Mol Biol Rep. 2012;39:1283–6.
Li LC, Chui RM, Sasaki M, Nakajima K, Perinchery G, Au HC, et al. A single nucleotide polymorphism in the E-cadherin gene promoter alters transcriptional activities. Cancer Res. 2000;60:873–6.
Kuraoka K, Oue N, Yokozaki H, Kitadai Y, Ito R, Nakayama H, et al. Correlation of a single nucleotide polymorphism in the E-cadherin gene promoter with tumorigenesis and progression of gastric carcinoma in Japan. Int J Oncol. 2003;23:421–7.
Yamada H, Shinmura K, Goto M, Iwaizumi M, Konno H, Kataoka H, et al. Absence of germline mono-allelic promoter hypermethylation of the CDH1 gene in gastric cancer patients. Mol Cancer. 2009;8:63.
B-z Zhang, D-j Deng. Effects of methylation status of CpG islands on results of luciferase reporter assay. Zhonghua Yufang Yixue Zazhi. 2009;43:601–6.
Grady W, Willis J, Guilford P, Dunbier A, Toro T, Lynch H, et al. Methylation of the CDH1 promoter as the second genetic hit in hereditary diffuse gastric cancer. Nat Genet. 2000;26:16–7.
Corso G, Carvalho J, Marrelli D, Vindigni C, Carvalho B, Seruca R, et al. Somatic mutations and deletions of the E-cadherin gene predict poor survival of patients with gastric cancer. J Clin Oncol. 2013;31:868–75.
Yu QM, Wang XB, Luo J, Wang S, Fang XH, Yu JL, et al. CDH1 methylation in preoperative peritoneal washes is an independent prognostic factor for gastric cancer. J Surg Oncol. 2012;106:765–71.
Yamada S, Nomoto S, Fujii T, Takeda S, Kanazumi N, Sugimoto H, et al. Frequent promoter methylation of M-cadherin in hepatocellular carcinoma is associated with poor prognosis. Anticancer Res. 2007;27:2269–74.
Marsit CJ, Posner MR, McClean MD, Kelsey KT. Hypermethylation of E-cadherin is an independent predictor of improved survival in head and neck squamous cell carcinoma. Cancer. 2008;113:1566–71.
Graziano F, Arduini F, Ruzzo A, Bearzi I, Humar B, More H, et al. Prognostic analysis of E-cadherin gene promoter hypermethylation in patients with surgically resected, node-positive, diffuse gastric cancer. Clin Cancer Res. 2004;10:2784–9.
Yi TZ, Guo J, Zhou L, Chen X, Mi RR, Qu QX, et al. Prognostic value of E-cadherin expression and CDH1 promoter methylation in patients with endometrial carcinoma. Cancer Invest. 2011;29:86–92.
Shimada S, Mimata A, Sekine M, Mogushi K, Akiyama Y, Fukamachi H, et al. Synergistic tumour suppressor activity of E-cadherin and p53 in a conditional mouse model for metastatic diffuse-type gastric cancer. Gut. 2012;61:344–53.
Sobin L, Wittekind C. TNM classification of malignant tumours. 6th ed. New York: Wiley; 2002.
Liu ZJ, Zhang J, Gao YH, Pei LR, Zhou J, Gu LK, et al. Large-scale characterization of DNA methylation changes in human gastric carcinomas with and without metastasis. Clin Cancer Res. 2014;20:4598–612.
Zheng X, Zhou J, Zhang B, Zhang J, Wilson J, Gu L, et al. Critical evaluation of Cbx7 downregulation in primary colon carcinomas and its clinical significance in Chinese patients. BMC Cancer. 2015;15:1172.
Shang Y, Hu X, DiRenzo J, Lazar MA, Brown M. Cofactor dynamics and sufficiency in estrogen receptor-regulated transcription. Cell. 2000;103:843–52.
Zhang B, Xiang S, Zhong Q, Yin Y, Gu L, Deng D. The p16-specific reactivation and inhibition of cell migration through demethylation of CpG islands by engineered transcription factors. Hum Gene Ther. 2012;23:1071–81.
Bai H, D-J Deng. Analysis of demethylation-related HPV16 reactivation by DHPLC-primer extension assay. Zhonghua Yufang Yixue Zazhi. 2007;41:81–3.
Eads CA, Laird PW. Combined bisulfite restriction analysis (COBRA). Methods Mol Biol. 2002;200:71–85.
Herman JG, Graff JR, Myöhänen S, Nelkin BD, Baylin SB. Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc Natl Acad Sci USA. 1996;93:9821–6.
Deng D, Deng G, Smith MF, Zhou J, Xin H, Powell SM, et al. Simultaneous detection of CpG methylation and single nucleotide polymorphism by denaturing high performance liquid chromatography. Nucleic Acids Res. 2002;30:E13.
Luo DY, Zhang BZ, Lv LB, Xiang SY, Liu YH, Ji JF, et al. Methylation of CpG islands of p16 associated with progression of primary gastric carcinomas. Lab Invest. 2006;86:591–8.
Batlle E, Sancho E, Francí C, Domínguez D, Monfar M, Baulida J, et al. The transcription factor snail is a repressor of E-cadherin gene expression in epithelial tumour cells. Nat Cell Biol. 2000;2:84–9.
Cano A, Pérez-Moreno MA, Rodrigo I, Locascio A, Blanco MJ, del Barrio MG, et al. The transcription factor snail controls epithelial–mesenchymal transitions by repressing E-cadherin expression. Nat Cell Biol. 2000;2:76–83.
Li X, Kim Y, Tsang EK, Davis JR, Damani FN, Chiang C, et al. The impact of rare variation on gene expression across tissues. Nature. 2017;550:239–43.
GTEx Consortium. Genetic effects on gene expression across human tissues. Nature. 2017;550:204–13.
Tukiainen T, Villani AC, Yen A, Rivas MA, Marshall JL, Satija R, et al. Landscape of X chromosome inactivation across human tissues. Nature. 2017;550:244–8.
Corso G, Roviello F, Paredes J, Pedrazzani C, Novais M, Correia J, et al. Characterization of the P373L E-cadherin germline missense mutation and implication for clinical management. Eur J Surg Oncol. 2007;33:1061–7.
Shin Y, Kim IJ, Kang HC, Park JH, Park HR, Park HW, et al. The E-cadherin −347G- > GA promoter polymorphism and its effect on transcriptional regulation. Carcinogenesis. 2004;25:895–9.
BoN Borges, EaS Santos, Bastos CE, Pinto LC, Anselmo NP, Quaresma JA, et al. Promoter polymorphisms and methylation of E-cadherin (CDH1) and KIT in gastric cancer patients from northern Brazil. Anticancer Res. 2010;30:2225–33.
Carneiro P, Figueiredo J, Bordeira-Carriço R, Fernandes MS, Carvalho J, Oliveira C, et al. Therapeutic targets associated to E-cadherin dysfunction in gastric cancer. Expert Opin Ther Targets. 2013;17:1187–201.
Acknowledgements
This work is supported by funding from the National Natural Science Foundation of China (nos. 81572762 and 31261140372) and the National Basic Research Program of China (no. 2015CB553902) to B. Zhang, WH Kim, and D. Deng. We thank Dr. Huidong Shi at the GRU Cancer Center at Georgia Reagents University, Augusta (USA) for language editing.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors disclose no potential conflicts of interest.
Ethical approval
All procedures followed were in accordance with the ethical standards of the responsible committee on human experimentation (institutional and national) and with the Helsinki Declaration of 1964 and later versions.
Informed consent
Informed consent or a substitute for it was obtained from all patients before they were included in the study unless the local institution review board permitted a waiver.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10120_2017_778_MOESM1_ESM.tif
Locations of the SNPs, CpG sites, and various PCR amplicons within the CDH1 CpG islands, 5’-shore, and the Alu element. TSS transcription start site (+1-nt), E-boxes; SNAIL binding sites (TIFF 149 kb)
10120_2017_778_MOESM2_ESM.tif
Comparison of the promoter activity of the –73A allele with that of the –73C allele in various human cancer cell lines. a The methylation status of the CDH1 promoter detected by the 115-bp methylation-specific PCR. b and c Levels of protein and mRNA of CDH1 detected by western blot and RT-PCR, respectively; cDNA from HeLa cells was used as the CDH1-expression-negative control. d Comparison of the promoter activity of the –73C allele with the –73A allele in various cell lines. * p<0.001 (TIFF 245 kb)
10120_2017_778_MOESM3_ESM.tif
Graphic view of the methylation status of the CpG island and the 5′-shore regions around the transcription start site of the CDH1 gene in various human cell lines. The methylation datasets were detected using reduced representative bisulfite sequencing. This image was modified from the image for the CDH1 gene visualized with the UCSC Genome Browser (Human hg19) found at the web site http://genome.ucsc.edu. Each red bar represents a methylated CpG site; the yellow bar represents a partially methylated CpG site; the green bar represents a CpG site that is not methylated. (TIFF 222 kb)
10120_2017_778_MOESM4_ESM.tif
Graphical view of the methylation status of the CpG island and shore regions around the transcription start site of the CDH1 gene in various adult human tissues/cells. The methylation datasets were detected using human whole genome bisulfite sequencing. This image was modified from the image for the CDH1 gene visualized with the WashU EpiGenome Browser (Human hg19) found at the web site http://epigenomegateway.wustl.edu/browser/. The height of each blue bar represents the combined read depth for the corresponding methylated CpG site; the height of each gray bar represents the combined read depth of the corresponding CpG. (TIFF 504 kb)
10120_2017_778_MOESM5_ESM.tif
Comparison of the methylation statuses of the CpG sites in the promoter CpG island and the shore regions in the representative gastric tissue sample #0187N from a patient carrying the –347del/del and –160C/C genotype of CDH1. The proportion of the –73A clones that were underlined clones containing at least three methylated CpGs in the shore region was much higher than the corresponding proportion of the –73C clones [12/129 vs. 1/70 (p=0.036)]. Red dots represent methylated CpG sites. (TIFF 2122 kb)
10120_2017_778_MOESM6_ESM.tif
Comparison of the methylation statuses of the CpG sites in the promoter CpG island and the shore regions in the representative gastric tissue sample #0379N. The proportion of the –73A clones that were underlined clones containing at least three methylated CpGs in the shore region was much higher than the corresponding proportion of the –73C clones [12/145 vs. 1/52 (p=0.190)]; the proportions were 2/21 vs. 11/176 in the −347A and –347del clones. Red dots represent methylated CpG sites. (TIFF 2117 kb)
Rights and permissions
About this article
Cite this article
Zhang, B., Zhou, J., Liu, Z. et al. Clinical and biological significance of a − 73A > C variation in the CDH1 promoter of patients with sporadic gastric carcinoma. Gastric Cancer 21, 606–616 (2018). https://doi.org/10.1007/s10120-017-0778-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10120-017-0778-6