Abstract
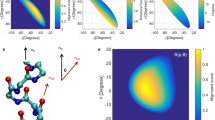
We have developed a geometrical approach to quantify differences in the stereochemistry of α-helical and turning regions in four iron proteins. Two spatial signatures are used to analyze residue coordinate data for each protein; and a third is employed to analyze amino-acid molecular volume data. The residue-by-residue analysis of the results, taken together with the finding that two major factors stabilize an α-helix (minimization of side-chain steric interference and intrachain H-bonding), lead to the conclusion that certain residues are preferentially selected for α-helix formation. In the sequential, de novo synthesis of a turning region, residues are preferentially selected such that the overall molecular volume profile (representing purely repulsive, excluded-volume effects) spans a small range Δ of values (Δ = 39.1 Å3) relative to the total range that could be spanned (Δ = 167.7 Å3). It follows that excluded-volume effects are of enormous importance for residues in helical regions as well as those in adjacent turning regions. Once steric effects are taken into account, down-range attractive interactions between residues come into play in the formation of α-helical regions. The geometry of α-helices can be accommodated by conformational changes in less-structured turning regions of a polypeptide, thereby producing a globally optimized (native) protein structure.








Similar content being viewed by others
References
Ramachandran GN, Ramakrishnan C, Sasisekharan V (1963) Stereochemistry of polypeptide chain configurations. J Mol Biol 7:95–99
Ramachandran GN, Sasisekharan V (1968) Conformation of polypeptides and proteins. Adv Protein Chem 23:283–438
Saven JG, Wolynes PG (1996) Local conformational signals and the statistical thermodynamics of collapsed helical proteins. J Mol Biol 257:199–216
Onuchic JN, Luthey-Schulten Z, Wolynes PG (1997) Theory of protein folding: the energy landscape perspective. Annu Rev Phys Chem 48:545–600
Wolynes PG (2015) Evolution, energy landscapes and the paradoxes of protein folding. Biochimie 119:218–230
Lin X, Schafer NP, Lu W, Jin S, Chen M, Onuchic JN, Wolynes PG (2019) Forging tools for refining predicted protein structures. Proc Natl Acad Sci USA 116(19):9400–9409
Kozak JJ, Gray HB, Wittung-Stafshede P (2018) Geometrical description of protein structural motifs. J Phys Chem B 122:11289–11294
Faraone-Mennella J, Tezca FA, Gray HB, Winkler JR (2006) Stability and folding kinetics of structurally characterized cytochrome c-b562. Biochemistry 45:10504–10511
Lee JC, Engman KC, Tezcan FA, Gray HB, Winkler JR (2002) Structural feature of cytochrome c’ folding intermediates revealed by fluorescence energy transfer kinetics. Proc Natl Acad Sci USA 99:14778–14782
Isogai Y, Imamura H, Nakae S, Sumi T, Takahashi KI, Nakagawa T, Tsuneshige A, Shirai T (2018) Tracing whale myoglobin evolution by resurrecting ancient proteins. Sci Rep 8:16883
Sugimoto H, Makino M, Sawai H, Kawada N, Yoshizato K, Shiro Y (2004) Structural basis of human cytoglobin for ligand binding. J Mol Biol 339:873–885
Cohn EJ, Edsall JT (1943) In proteins, amino acids and peptides. Rheinhold Publishing Corporation, New York, pp 155–176 and 370–381
Zamyatin AA (1972) Protein volume in solution. Prog Biophys Mol Biol 24:107–123
Perkins SJ (1986) Protein volumes and hydration effects. The calculations of partial specific volumes, neutron scattering matchpoints and 280-nm absorption coefficients for protein and glycoproteins from amino acid sequences. Eur J Biochem 157:169–180
Kozak JJ, Piasecki J, Szymczak P (2016) Distribution function approach to the stability of fluids. Adv Chem Phys 161:359–394
Dill KA, MacCallum JL (2012) The protein-folding problem 50 years on. Science 338:1042–1046
Dobson CM (2004) Principles of protein folding, misfolding and aggregation. Semin Cell Dev Biol 15:3–16
Leopold PE, Montal M, Onuchic JN (1992) Protein folding funnels: a kinetic approach to the sequence-structure relationship. Proc Natl Acad Sci USA 89:8721–8725
Wolynes P, Luthey-Schulten Z, Onuchic J (1996) Fast-folding experiments and the topography of protein folding energy landscapes. Chem Biol 3:425–432
Acknowledgments
Work at Caltech was supported by the National Institutes of Health (DK-019038 to HBG).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendices
Appendices
Appendix 1
Appendix 2
Rights and permissions
About this article
Cite this article
Kozak, J.J., Gray, H.B. Stereochemistry of residues in turning regions of helical proteins. J Biol Inorg Chem 24, 879–888 (2019). https://doi.org/10.1007/s00775-019-01696-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-019-01696-9