Abstract
The high transmission and mortality rates associated with SARS-CoV-2 have led to tragic consequences worldwide. Large-scale whole-genome sequencing of the SARS-CoV-2 genome since its identification in late 2019 has identified many sequence changes and the emergence of novel strains, each described by co-segregation of a particular set of sequence variations. Variants designated G, alpha (B.1.1.7), beta (B.1.351), gamma (P.1), and delta (B.1.617.2) are important lineages that emerged sequentially and are considered variants of concern. A notable feature of the last four, each of which ultimately evolved from clade G, is the large number (≥ 20) of co-segregating sequence variations associated with them. Several variations are in the spike gene, and some variations are shared among or between strains. Meanwhile, observation of recurrent infections with the same or different SARS-CoV-2 lineages has raised concerns about the duration of the immune responses induced by the initial infection or the vaccine that was administered. While the alpha strain is sensitive to immune responses induced by earlier strains, the beta, gamma, and delta strains can escape antibody neutralization. Apart from random replication errors, intra-host RNA editing, chronic infections, and recombination are processes that may promote the accumulation of sequence changes in the SARS-CoV-2 genome. The known contribution of recombination to coronavirus evolution and recent data pertaining to SARS-CoV-2 suggest that recombination may be particularly important. Continued surveillance of the SARS-CoV-2 genome is imperative.
Similar content being viewed by others
Introduction
Late in December 2019, a number of patients with pneumonia of unknown cause were observed in Wuhan, China. Deep metagenomic sequencing of bronchoalveolar lavage fluid samples of the first patients resulted in the identification of a new SARS-like coronavirus [1, 2]. The novel virus was designated as SARS-CoV-2 by the International Committee on Taxonomy of Viruses, and the disease related to this virus was named coronavirus disease 19 (COVID-19) by WHO [3]. Prior to the discovery of SARS-CoV-2, six human-pathogenic coronaviruses (hCoVs) – NL63, 229E, OC43, HKU1, SARS-CoV, and MERS-CoV – had been identified [4]. While the first four were relatively benign, SARS-CoV (severe acute respiratory syndrome coronavirus) and MERS-CoV (Middle East respiratory syndrome coronavirus) were highly pathogenic [4, 5]. SARS-CoV first emerged in southern China and caused an epidemic that lasted from November of 2002 to July of 2003; over 8,000 infected cases and 774 deaths in more than 25 countries were reported during this period [6,7,8]. MERS-CoV appeared in Saudi Arabia in 2012, and by 2020, approximately 2,500 cases of MERS-CoV infection and over 850 deaths had been confirmed in 27 countries [8, 9]. The new virus, SARS-CoV-2, which contains a linear positive-strand RNA genome of about 30,000 nucleotides, is associated with mild to severe clinical symptoms and sometimes death [1, 10]. The higher person-to-person transmission rate of SARS-CoV-2 as compared with the other hCoVs has led to rapid spread of this virus, and on March 11, 2020, the World Health Organization (WHO) declared this spread to be a global pandemic [11]. Various lines of evidence support a possible contribution of recombination events to tropism and adaptation of coronaviruses to new host species and to the emergence and diversity of human pathogenic coronaviruses, including SARS-CoV-2 [12,13,14,15,16,17,18,19]. Recombination may be a normal feature of CoV replication, and its role in the emergence of new strains may be quite significant [20].
Emergence and expansion of SARS-CoV-2 clades
Very shortly after the sequencing of SARS-CoV-2 genomes isolated from the first infected individuals from China, thousands of whole-genome sequences of virus isolates from across the world were submitted to public databases such as GISAID (the Global Initiative on Sharing All Influenza Data; GISAID https://www.gisaid.org/). Moreover, other bioinformatics platforms including Nextstrain (https://nextstrain.org/ncov/global) were developed that continuously monitor the submitted sequences and enable real-time tracking and visualization of sequence variations. These developments and other analyses have allowed close monitoring of the evolution of the SARS-CoV-2 genome. Analysis of various SARS-CoV-2 genome sequence data sets has led to an estimated evolutionary rate of approximately 1.1 × 10-3 substitutions/site/year [21]. Although rapid, the mutation rate is lower than that of some other RNA viruses, and this may partly be due to presence of a 3'-to-5' exoribonuclease (ExoN)-encoding gene in the SARS-CoV-2 genome whose protein product may correct some of the errors that occur during replication [22]. In a survey of 2,790 complete and high-coverage sequences available on April 2, 2020, in GISAID, 13 distinct haplotype groups (H1-H13) and their associated daughter haplotypes were described. Each haplotype was defined by the co-segregation of two or more Tag nucleotide sequence variations in a significant number of available genome sequences [23]. Some haplotypes, after initial emergence, spread to various regions of the world, while others remained localized or were eliminated. For example, the haplotype group H2, which contained the sequence variations 8782C>T and 28144T>C, first appeared in the Far East but soon spread to other continents. This haplotype group had earlier been recognized as the S lineage by others [24]. On the other hand, a haplotype (H13) with a large deletion (27848_28229del), after its appearance and initial local expansion in a country in the Far East, essentially disappeared with the passage of time [23, 25]. Differences in the history of sequence variations at least partly reflect their contribution to viral fitness. This is well exemplified by the haplotype H1, defined by co-segregation of four variations: 241C>T, 3037C>T, 14408C>T, and 23403A>G. This haplotype is named clade G in GISAID and 20A in Nextstrain. 23403A>G causes substitution of aspartic acid at position 614 of the spike glycoprotein by glycine (p.D614G) [26]. The p.D614G variation had a rare global presence in March 2020 but became a dominant variation that was present in more than 74% of available sequences by June 2020 [26, 27]. Analysis of approximately 50,000 complete and high-coverage sequences collected between August 1 and November 15, 2020, showed that 99.7 percent of sequences had the four defining sequence variations of clade G (https://cov.lanl.gov/content/index) [28]. It has been reported that this variation confers a competitive advantage because of enhancement of viral replication, viral load, and infectivity [26, 27, 29,30,31]. The SARS-CoV-2 spike protein has two functional subunits, S1 and S2, which play crucial roles, respectively, in the attachment of the virus to the host-cell receptor (angiotensin-converting enzyme 2; ACE2) through its receptor-binding domain (RBD) and mediation of fusion of viral and host membranes, leading to entry of the virus into the host cell [32, 33]. At the molecular level, p.D614G induces an open conformation in RBD, which facilitates virus entry [34, 35]. Some subclades, including GR, GH, and GV, have evolved from the G clade. Three adjacent nucleotide variations (28881_28883GGG>AAC in N) define GR (H1a), one (25563G>T in orf3a) defines GH (H1b), and seven (445T>C in nsp1, 6286C>T in nsp3, 21255G>C in 2ʹ‐O‐ribose methyltransferase, 22227C>T in S, 26801C>G in M, 28932C>T in N, and 29645G>T in orf10) define GV (H1r) (https://www.gisaid.org/references/statements-clarifications/clade-and-lineage-nomenclature-aids-in-genomic-epidemiology-of-active-hcov-19-viruses/) [28]. The three-nucleotide variation in subclade GR causes the amino acid substitutions p.R203K and p.G204R in the nucleocapsid protein, which is encoded by the N gene. The wild-type N protein binds the virus genome and establishes a condensed organization that is crucial for virion assembly and protection of the viral genome against host RNA sensors and, consequently, immune surveillance [36, 37]. Formation of the RNA-protein complex relies on a liquid–liquid phase separation (LLPS) of the N protein [37, 38]. It has been suggested that p.R203K and p.G204R affect this physiochemical process in a manner that causes an increased tendency of the N protein to undergo LLPS and greater inhibition of host antiviral responses [37]. The 25563G>T variation in GH causes p.Gln57His in the protein encoded by orf3a, which is a putative viral ion channel [39]. However, cryo-electron microscopy and bioinformatics studies on the ORF3a structure have predicted that p.Gln57His does not affect the permeability properties of the ion channel [40, 41]. The 22227C>T variation of the GV subclade causes p.A222V in the N-terminal domain (NTD) of the spike protein, but this variation was reported not to inhibit binding of monoclonal antibodies specific for the spike NTD [42]. It is noteworthy that new sub-clades (lineages) are continually being identified (Pango lineages; https://cov-lineages.org/index.html) [43].
The issue of SARS-CoV-2 re-infection
Soon after the beginning of the COVID-19 outbreak, various companies and governments initiated programs aimed at the development of anti-SARS-CoV-2 vaccines. It is believed that vaccination of a substantial proportion of the world population can limit the outbreak by ultimately reducing the number of circulating viruses [44]. In a study in which symptomatic patients infected very early in the pandemic were followed up, it was found that spike RBD- and nucleocapsid (N)-recognizing IgG antibodies that could bind and neutralize viruses were present at high positivity rates and at high titers for more than six months after recovery. The data were suggestive of durable humoral immunity and long-lasting protection [45]. On the other hand, multiple reports of SARS-CoV-2 re-infection in individuals who had only recently recovered from an earlier infection have created concerns for scientists and health officials. These reports raise questions about the duration of immunity and the protection conferred by anti-COVID-19 vaccines/antibodies [46,47,48,49]. It is hoped that repeated booster vaccine shots may induce long-term immunity and increase vaccine effectiveness [50, 51].
Emergence of SARS-CoV-2 variants of concern with large numbers of sequence variations
A SARS-CoV-2 variant of interest (VOI) is a virus strain associated with an increase in public-health-related parameters such as transmissibility, pathogenicity, severity of clinical presentation, therapeutic escape, and antigenicity (https://www.who.int/en/activities/tracking-SARS-CoV-2-variants/). A SARS-CoV-2 variant of concern (VOC) is a strain associated with more-drastic changes in these parameters. As of the end of July 2021, four VOIs (eta [η], iota [ι], kappa [κ], and lambda [λ]) and four VOCs (alpha [α], beta [β], gamma [γ], and delta [δ] were recognized by WHO [https://www.who.int/en/activities/tracking-SARS-CoV-2-variants/]. Although various SARS-CoV-2 strain nomenclatures have been used, WHO has proposed that the letters of the Greek alphabet be used as labels for global SARS-CoV-2 variants of interest and variants of concern alongside the scientific nomenclature in communications about the variants to the public (https://www.who.int/en/activities/tracking-SARS-CoV-2-variants/). The objective was to introduce easy-to-pronounce and non-stigmatizing labels. As the scientific community is also familiar with the more facile Greek letter nomenclature of the VOCs, these (sometimes in conjunction with the scientific names) will hereafter be used for the strains discussed.
In mid-December 2020, a new lineage of the GR clade with an unusual cluster of 17 additional non-synonymous, nonsense, and deletion mutations was identified in the United Kingdom (UK). It was designated as VOC lineage B.1.1.7 (lineage alpha). Almost half of the new mutations were located in the spike protein, including the mutations Δ69-70, p.N501Y, and p.P681H, which affect key epitopes of the protein. The possible effects of these mutations are discussed below. No lineage that resembled this strain had been reported previously [52,53,54]. Based on data available at GISAID on June 13, 2021, GV was the major clade circulating in the UK in November 2020; this dominance was taken over by the alpha strain by the second half of December 2020, indicative of a rapid and remarkable shift in the virus profile of the UK [28]. The alpha strain was reported in approximately 90 countries/territories by the end of December. An unprecedentedly rapid global spread of this strain was observed by the beginning of March 2021. Soon after identification of the alpha strain, other potentially dangerous lineages with large numbers of mutations were observed in South Africa (B.1.351; beta), Brazil (B.1.1.28.1/P.1; gamma), and India (B.1.617.2; delta) [55,56,57,58]. Based on strain designations by GISAID on June 13, 2021, more than 93% of samples collected in the United Kingdom in April 2021 had alpha strain sequences, whereas among samples collected between May 1 and June 7, 2021, only 38% had the alpha strain sequence and 60% had the delta strain sequence. This suggests a marked rapid displacement of the alpha strain by the delta strain. Rapid expansion of VOCs suggested that they may have properties that confer a fitness advantage. The emergence of VOCs raised questions about their origins, and their rapid expansion raised questions and concerns regarding their infectivity and transmission rate, and more importantly, the effectiveness of the vaccines being developed in targeting the novel strains.
Concerns pertaining to the effectiveness of current vaccines against VOC lineages
Recent empirical and population-based findings suggest that the alpha strain has a significant transmission advantage over earlier strains, causes a higher virus load, and is associated with a higher risk of infection for individuals under 20 years old. A substantial correlation between the strain and disease severity or mortality rate was not observed [59,60,61]. Antibody sensitivity experiments indicate that viruses of the G clade (the earliest strains shown to have the p.D614G variation) and the alpha strain viruses are highly susceptible to neutralization by antibodies present in sera or nasal swabs from individuals who had recovered from infection with viruses circulating before the emergence of these strains, or from individuals who had been vaccinated with existing vaccines [62,63,64]. However, there is substantial evidence that antibodies in the sera of convalescent individuals who had been infected in early 2020 and antibodies induced by some vaccines have decreased neutralization potential against the beta, gamma, and delta VOCs [51, 62,63,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79].
The alpha, beta, and gamma SARS-CoV-2 strains all share the mutation that causes the p.N501Y change in the RBD domain of the spike protein. In fact, position 501 is within the region of the RBD (receptor-binding motif; RBM) that makes contact with the host ACE2 receptor [80]. It has been shown that the p.N501Y change plays a critical role in promoting the increased infectivity and transmissibility of the alpha strain. Increased infectivity may be a consequence of a gain in replication fitness conferred by the amino acid change, and increased transmissibility may be a consequence of considerable strengthening of the spike/cell receptor interaction [81, 82]. The beta and gamma strains also share the p.E484K-causing mutation in the same domain. Additionally, the lysine at position 417 of the earliest sequenced SARS-CoV-2 strains is changed to asparagine and threonine, respectively, in the beta, and gamma strains [55, 58]. It is believed that mutated amino acids at positions 417, 484, and 501 result in a tighter interaction between the spike protein and the ACE2 host cell receptor, probably by induction of conformational changes. The p.E484K mutation may also cause a favorable change in the charge of the flexible loop region of the spike RBD [83]. Reduced neutralization of the beta and gamma strains by antibodies in the sera from convalescent patients may be partly or largely attributable to steric clashes and charge switches at antibody-binding sites caused by the p.E484K mutation in these strains [75, 78, 83,84,85,86,87,88]. The alpha (p.P681H) and delta (p.P681R) strains both contain a variation at position 681 of the spike protein. The mutations that affect p.681 are believed to be associated with enhanced furin cleavage at the suboptimal S1/S2 junction site during virus and host-cell membrane fusion, which may ultimately result in increased transmissibility of the virus [89,90,91]. The delta strain contains a unique p.L452R variation in the RBD domain of the spike protein. Antigenicity experiments have shown that this mutation is linked to a marked resistance to some neutralizing monoclonal antibodies [92]. The Δ69-70 deletion (p.69-70delHV) in the spike protein that is observed only in the alpha lineage is proposed to contribute to increased infectivity, possibly by promotion of spike protein cleavage [93]. The apparent biological and molecular consequences of some of the potentially important sequence variations observed in SARS-CoV-2 VOCs are summarized in Table 1.
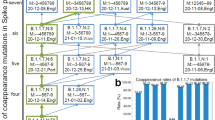
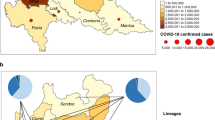
Figure 1 shows the defining genome sequence changes of the four VOCs described above and the effects of these changes on the amino acid sequences of the encoded proteins. The distribution of the amino acid changes in the domains of the spike protein is shown in Figure 2. Shared variations between any two or among three or four of the VOCs are also shown in the two figures. These figures are based on analysis of 50 alpha, beta, gamma, and delta SARS-CoV-2 genome sequences from the UK, South Africa, Brazil, and India, respectively, that were designated as the respective strain in GISAID. GISAID, in addition to serving as a database for reported SARS-CoV-2 genome sequences, itself designates the lineage associated with each of the sequences (https://www.gisaid.org/). The sequences chosen for the analysis included the first and most recently (as of June 21, 2021) reported sequence associated with the lineage, and other sequences fairly equally distributed with respect to date of sample collection in the interim (Supplementary Tables S1-S4). Nucleotide sequence variations in the chosen sequences were identified by alignment to the reference genome sequence of the Wuhan‐Hu‐1 isolate (NC_045512.2) as described previously [23]. Haplotype network and phylogenetic analyses of all the sequences were also performed as described previously [23]. Figure 3 shows haplotype networks based on whole-genome sequences of the VOCs, and Figure 4 shows the networks based only on the S gene sequences. Phylogenic trees were also constructed based on whole-genome and S gene sequences (available upon request). As expected, the sequences pertaining to each of the four VOCs cluster together in the networks and in the phylogenetic trees. As compared to the whole-genome sequence network, there is less branching at each of the foci in the S gene network; this is because there is less variation among the 50 S gene sequences of each of the VOCs than among the 50 whole-genome sequences of each of the VOCs. Figures 1 and 2 show the variations in each of the strains that were present in at least 80% of the 50 selected sequences. There were 32 (Supplementary Table S1), 21 (Supplementary Table S2), 35 (Supplementary Table S3), and 20 (Supplementary Table S4) sequence changes among the alpha, beta, gamma, and delta genome sequences, respectively, that were present in at least 40 of the 50 sequences (the 80% threshold). All of the sequence changes described for each VOC were usually, but not always, present together. All of the sequence changes pertaining to the alpha, beta, gamma, and delta VOCs that are described in Figure 1 were present together in 50 (100%), 37 (74%), 34 (68%), and 38 (76%) of the 50 sequences of the respective VOC genomes. These data suggest that the set of variations described for the alpha strain indeed describes a robust haplotype; the haplotypes that are described for each of the remaining three VOCs may be less robust. Four sequence variations that define clade G (major haplotype group H1) were shared among all four VOCs, suggesting that the VOCs are derived from this major clade. Furthermore, the alpha and gamma VOCs contain the subclade GR (H1a sub-haplotype)-defining variations (28881_28883GGG>AAC), and the beta VOC has the subclade GH (H1b sub-haplotype)-defining variation (25563G>T). These observations are consistent with these three VOCs having evolved from GR or GH subclades or having been products of recombination events in which one of the participating genomes was GR or GH. The haplotype network based on whole-genome sequences also suggests a common origin (presumably GR) for the alpha and gamma VOCs (Fig. 3). A mutation (11288-11296deltctggtttt) in nsp6 that causes a three amino-acid-deletion (p.106-108delSGF) in the encoded protein was shared among three VOCs (alpha, beta, and gamma strains). The nsp6 (non-structural protein 6) protein of coronaviruses has been suggested to affect autophagy and possibly thus promote viral infection [96, 97]. Additionally, it has been shown that the SARS-CoV-2 nsp6 affects the host cell immune response by suppression of interferon production [96]. To the best of our knowledge, the effect of the p.106-108delSGF mutation on these nsp6 functions has not been studied.
Defining genome sequence variations of SARS-CoV-2 variants of concern (VOCs). A schematic representation of the SARS-CoV-2 genome is shown at the top of the figure. Nucleotide sequence variations pertaining to the alpha (B.1.1.7), beta (B.1.351), gamma (P.1), and delta (B.1.617.2) lineages (relative to Wuhan-Hu-1 sequence at https://www.ncbi.nlm.nih.gov/nuccore/NC_045512.2) and effects on amino acid sequences of the encoded proteins are shown. The defining sequence variations were selected as described in the text. Green, red, and blue dots indicate synonymous, non-synonymous, and nonsense mutations, respectively. Black dashes represent indels. 5' UTR, 5'- untranslated region; nsp1-16, non-structural proteins 1-16; orf1ab, open reading frame 1ab; S, spike; orf3a, open reading frame 3a; E, envelope; M, membrane; orf6, open reading frame 6; orf7a, open reading frame 7a; orf7b, open reading frame 7b; orf8, open reading frame 8; N, nucleocapsid; orf10, open reading frame 10; 3' UTR, 3'- untranslated region. ***, variations present in all four VOCs; **, variations present in three VOCs; *, variations present in two VOCs
Effects of defining genome sequence variations of SARS-CoV-2 variants of concern (VOCs) on the domains of the spike protein. A schematic representation of the domains of the spike protein is shown on top of the figure. Amino acids affected by the defining genome sequence variations of the alpha (B.1.1.7), beta (B.1.351), gamma (P.1), and delta (B.1.617.2) lineages are indicated, and their corresponding genome positions are shown in parentheses. S1, S1 subunit: S2, S2 subunit; S1/S2, S1 and S2 interface – spike cleavage site; SP, signal peptide; NTD, N-terminal domain; RBD, receptor-binding domain; RBM, receptor-binding motif; HR1, heptad repeat 1; HR2, heptad repeat 2; TM, transmembrane domain, CP, cytoplasmic domain. Green, red, and blue dots indicate synonymous, non-synonymous, and nonsense mutations, respectively. Black dashes represent deletions. ***, variations present in all four VOCs; **, variations present in three VOCs; *, variations present in two VOCs
Haplotype network of 200 whole-genome sequences of VOC. The nodes represent the various whole-genome sequences. Because of space limitations, the samples are abbreviated as alpha1, alpha2, etc., and the corresponding GISAID IDs are presented in Supplementary Table S5. The number of crossbeams on the connecting lines between nodes represents the number of sequence differences between the respective sequences.
Haplotype network of 200 S gene sequences of VOCs. The nodes represent the various S gene sequences. Because of space limitations, the samples are abbreviated as alpha1, alpha2, etc., and the corresponding GISAID IDs are presented in Supplementary Table S5. The number of crossbeams on the connecting lines between nodes represents the number of sequence differences between the respective sequences.
With regard to vaccine efficacy, a promising finding was increased effectiveness of second-dose vaccine shots against the alpha and delta strains. This emphasizes the importance of booster vaccinations [51]. Also, it has been reported that a G614 pseudovirus was more susceptible to neutralization by sera from earlier infected convalescent individuals than viruses with the D614-encoding genotype. The molecular explanation for this may be the RBD “up” conformation induced by p.D614G. In any case, the finding that G614 is not an escape variation implies that the global dominance of G614 over D614 is not expected to affect the effectiveness of the SARS-CoV-2 vaccines that have been developed [35].
Potential sources of changes in the SARS-CoV-2 genome
Genome replication infidelity [98], intra-host viral evolution in prolonged infections, which predominantly occurs in immunocompromised individuals [54, 99,100,101,102], host RNA-editing systems [103], and within-host recombination events occurring subsequently to co- or super-infection with different circulating strains [104] are considered the potential sources of SARS-CoV-2 genome sequence variability. Random mutations account for the majority of genomic variations [103, 105]. Although most immunocompromised persons effectively clear SARS-CoV-2 infection, accelerated viral evolution in persistently infected immunocompromised individuals has been reported [54, 103, 106]. In a publication that reported findings on a carefully monitored patient, it was shown that a cluster of mutations accumulated rapidly, sometimes within a span of approximately 50 days [99]. These new mutations disproportionately changed amino acids in the spike protein and in its receptor-binding domain. Definitive empirical evidence for involvement of cellular RNA editing processes in promoting sequence changes in the SARS-CoV-2 genome is lacking. However, the observation of an enrichment of C-to-U transitions among the reported single-nucleotide variations in the genome and the known activities of enzymes of the editing process (including the APOBEC cytidine deaminases and ADAR adenosine deaminases) that promote such changes suggest that RNA editing may contribute to the variability of the SARS-CoV-2 genome [103, 107,108,109].
Presently, recombination may be the best candidate competitor of genome replication errors as the driving force for emergence of new SARS-CoV-2 strains. Recombination occurs with many RNA viruses, and the contribution of recombination to the evolution of betacoronaviruses is well established [4, 19, 110,111,112]. Recognition of recombination between SARS-CoV-2 viruses requires co-infection with viruses with distinct sequences. Co-infection with different SARS-CoV-2 strains has been documented [113,114,115]. With increased frequencies of infections worldwide, at least partly due to the emergence of strains with higher transmissibility, it is expected that the frequency of co-infections will also rise. Since the new strains have distinct combinations of variations, there is also a greater possibility of co-infection with strains with different combinations of sequence variations and subsequent recombination events that would produce novel genome sequences. To the best of our knowledge, the first report of recombination between SARS-CoV-2 strains was published in August of 2020 [104]. In a few subsequent studies, recombination events between SARS-CoV-2 strains were surmised on the basis of defining markers of major clades or sequence variations in locally circulating strains [26, 105, 115,116,117]. It is not unlikely that some recombinant genomes have not been recognized because of technical issues in sequence analysis [105]. In light of the importance of recombination vis a vis the emergence of novel virus strains with a large number of mutations, efforts must be made to resolve these impediments [54, 106]. Although strains with large numbers of variations may have evolved by accumulation of sequential mutations and subsequent expansion due to stochastic events or selection, an alternative scenario is that they are products of one or more recombination events between virus strains that each contained a subset of the variations. It is to be noted that coinfection and recombination are most likely to occur between locally or globally dominant strains [26, 115]. In a recent study in which SARS-CoV-2 genome sequences in various cities of a single country were analyzed using non-conventional protocols, notably high frequencies of co-infection as well as sequences that were probable products of recombination events between locally or globally dominant strains were indeed observed [115]. For example, one putative recombinant genome simultaneously had GR and GH marker sequence variations. Selective advantages may have contributed to the dominant frequency status of some strains, including some VOCs, and recombination between any two of these strains may produce a recombinant strain that has the combined advantages associated with both of the parental strains. This renders the recognition of such possibly evolving strains even more important. The proposed role of recombination in the emergence of new strains is reminiscent of the origin of SARS-CoV-2 itself [15, 16].
Conclusion
Since the first identification of SARS-CoV-2, numerous variations in the genomes of evolved viruses have been identified. It is expected that the majority of evolved variations have neutral effects and that those with highly deleterious consequences for the virus will be rapidly eliminated. Mutations that confer an advantage to the virus, such as p.D614G, tend to be retained and can lead to a notable shift in the spectrum of the local or worldwide virus population in a short time period. The presence of a significant proportion of the retained frequent variations in the spike S1 subunit, and especially in the RBD domain, of the variants of concern and the proposed pathogenic consequences associated with these sequence variations provide strong evidence of the critical importance of S1 in terms of infectivity and transmissibility as well as immune escape. Some reports emphasize the biased concentration of recombination events in the spike coding gene of SARS-CoV-2 [19, 26]. Given the essential role of the spike protein in the infection process, this protein or its main domains have been targeted in developing immunization protocols. In the face of resistance of some variants to antibody neutralization, the robust cross-reactive responses elicited by the South African strain against other virus strains are somewhat reassuring. This observation suggests that vaccine development strategies should take into consideration features of the spike proteins of the VOC strains [118]. The continuous tracking of novel and potentially clinically important sequence variations and recombinant and/or highly mutated strains is of immense importance in light of public health, disease control, and design of new preventive immunization strategies.
References
Wu F, Zhao S, Yu B et al (2020) A new coronavirus associated with human respiratory disease in China. Nature 579:265–269. https://doi.org/10.1038/s41586-020-2008-3
Zhou P, Lou YX, Wang XG et al (2020) A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579:270–273. https://doi.org/10.1038/s41586-020-2012-7
Gorbalenya AE, Baker SC, Baric RS et al (2020) The species Severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat Microbiol 5:536–544. https://doi.org/10.1038/s41564-020-0695-z
Su S, Wong G, Shi W et al (2016) Epidemiology, genetic recombination, and pathogenesis of coronaviruses. Trends Microbiol 24:490–502. https://doi.org/10.1016/j.tim.2016.03.003
Forni D, Cagliani R, Clerici M, Sironi M (2017) Molecular evolution of human coronavirus genomes. Trends Microbiol 25:35–48. https://doi.org/10.1016/j.tim.2016.09.001
Ksiazek TG, Erdman D, Goldsmith CS et al (2003) A novel coronavirus associated with severe acute respiratory syndrome. N Engl J Med 348:1953–1966. https://doi.org/10.1056/nejmoa030781
Peiris JSM, Yuen KY, Osterhaus ADME, Stöhr K (2003) The severe acute respiratory syndrome. N Engl J Med 349:2431–2441. https://doi.org/10.1056/NEJMra032498
De Wit E, Van Doremalen N, Falzarano D, Munster VJ (2016) SARS and MERS: recent insights into emerging coronaviruses. Nat Rev Microbiol 14:523–534. https://doi.org/10.1038/nrmicro.2016.81
Memish ZA, Perlman S, Van Kerkhove MD, Zumla A (2020) Middle East respiratory syndrome. Lancet 395:1063–1077. https://doi.org/10.1016/S0140-6736(19)33221-0
Guan W, Ni Z, Hu Y et al (2020) Clinical characteristics of Coronavirus Disease 2019 in China. N Engl J Med 382:1708–1720. https://doi.org/10.1056/nejmoa2002032
World Health Organization (2020) WHO Director-General’s opening remarks at the media briefing on COVID-19-11 Mar 2020
Hon C-C, Lam T-Y, Shi Z-L et al (2008) Evidence of the recombinant origin of a bat severe acute respiratory syndrome (SARS)-like coronavirus and its implications on the direct ancestor of SARS coronavirus. J Virol 82:1819–1826. https://doi.org/10.1128/jvi.01926-07
Wang Y, Liu D, Shi W et al (2015) Origin and possible genetic recombination of the middle east respiratory syndrome coronavirus from the first imported case in china: phylogenetics and coalescence analysis. MBio. https://doi.org/10.1128/mBio.01280-15
Zhang Z, Shen L, Gu X (2016) Evolutionary dynamics of MERS-CoV: potential recombination. Posit Sel Transm Sci Rep 6:25049. https://doi.org/10.1038/srep25049
Li X, Giorg EE, Marichannegowda MH et al (2020) Emergence of SARS-CoV-2 through recombination and strong purifying selection. Sci Adv 6:eabb9153. https://doi.org/10.1126/sciadv.abb9153
Zhu Z, Meng K, Meng G (2020) Genomic recombination events may reveal the evolution of coronavirus and the origin of SARS-CoV-2. Sci Rep 10:1–10. https://doi.org/10.1038/s41598-020-78703-6
Forni D, Cagliani R, Sironi M (2020) Recombination and positive selection differentially shaped the diversity of betacoronavirus subgenera. Viruses 12:1313. https://doi.org/10.3390/v12111313
Patiño-Galindo JÁ, Filip I, Rabadan R (2021) Global patterns of recombination across human viruses. Mol Biol Evol 38:2520–2531. https://doi.org/10.1093/molbev/msab046
Bobay LM, O’Donnell AC, Ochman H (2020) Recombination events are concentrated in the spike protein region of Betacoronaviruses. PLoS Genet 16:e1009272. https://doi.org/10.1371/JOURNAL.PGEN.1009272
Gribble J, Stevens LJ, Agostini ML et al (2021) The coronavirus proofreading exoribonuclease mediates extensive viral recombination. PLoS Pathog 17:e1009226. https://doi.org/10.1371/journal.ppat.1009226
Duchene S, Featherstone L, Haritopoulou-Sinanidou M et al (2020) Temporal signal and the phylodynamic threshold of SARS-CoV-2. Virus Evol 6:veaa061. https://doi.org/10.1093/ve/veaa061
Yan L, Yang Y, Li M et al (2021) Coupling of N7-methyltransferase and 3′-5′ exoribonuclease with SARS-CoV-2 polymerase reveals mechanisms for capping and proofreading. Cell 184:3474-3485.e11. https://doi.org/10.1016/j.cell.2021.05.033
Safari I, InanlooRahatloo K, Elahi E (2021) Evolution of SARS-CoV-2 genome from December 2019 to late March 2020: emerged haplotypes and informative Tag nucleotide variations. J Med Virol 93:2010–2020. https://doi.org/10.1002/jmv.26553
Tang X, Wu C, Li X et al (2020) On the origin and continuing evolution of SARS-CoV-2. Natl Sci Rev 7:1012–1023. https://doi.org/10.1093/nsr/nwaa036
Su YCF, Anderson DE, Young BE et al (2020) Discovery and genomic characterization of a 382-nucleotide deletion in ORF7B and orf8 during the early evolution of SARS-CoV-2. MBio 11:1–9. https://doi.org/10.1128/mBio.01610-20
Korber B, Fischer WM, Gnanakaran S et al (2020) Tracking changes in SARS-CoV-2 spike: evidence that D614G increases infectivity of the COVID-19 virus. Cell 182:812-827.e19. https://doi.org/10.1016/j.cell.2020.06.043
Plante JA, Liu Y, Liu J et al (2021) Spike mutation D614G alters SARS-CoV-2 fitness. Nature 592:116–121. https://doi.org/10.1038/s41586-020-2895-3
Safari I, InanlooRahatloo K, Elahi E (2021) World-wide tracking of major SARS-CoV-2 genome haplotypes in sequences of June 1 to November 15, 2020 and discovery of rapid expansion of a new haplotype. J Med Virol 93:3251–3256. https://doi.org/10.1002/jmv.26802
Yurkovetskiy L, Wang X, Pascal KE et al (2020) Structural and functional analysis of the D614G SARS-CoV-2 spike protein variant. Cell 183:739-751.e8. https://doi.org/10.1016/j.cell.2020.09.032
Volz E, Hill V, McCrone JT et al (2021) Evaluating the effects of SARS-CoV-2 spike mutation D614G on transmissibility and pathogenicity. Cell 184:64–75
Lorenzo-Redondo R, Nam HH, Roberts SC et al (2020) A clade of SARS-CoV-2 viruses associated with lower viral loads in patient upper airways. EBioMedicine 62:103112. https://doi.org/10.1016/j.ebiom.2020.103112
Tai W, He L, Zhang X et al (2020) Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell Mol Immunol 17:613–620. https://doi.org/10.1038/s41423-020-0400-4
Hu B, Guo H, Zhou P, Shi Z-L (2020) Characteristics of SARS-CoV-2 and COVID-19. Nat Rev Microbiol 19:141–154
Gobeil SMC, Janowska K, McDowell S et al (2021) D614G mutation alters SARS-CoV-2 spike conformation and enhances protease cleavage at the S1/S2 junction. Cell Rep 34:108630. https://doi.org/10.1016/j.celrep.2020.108630
Weissman D, Alameh MG, de Silva T et al (2021) D614G spike mutation increases SARS CoV-2 susceptibility to neutralization. Cell Host Microbe 29:23-31.e4. https://doi.org/10.1016/j.chom.2020.11.012
Zúñiga S, Sola I, Moreno JL et al (2007) Coronavirus nucleocapsid protein is an RNA chaperone. Virology 357:215–227. https://doi.org/10.1016/j.virol.2006.07.046
Zhao M, Yu Y, Sun LM et al (2021) GCG inhibits SARS-CoV-2 replication by disrupting the liquid phase condensation of its nucleocapsid protein. Nat Commun 12:1–14. https://doi.org/10.1038/s41467-021-22297-8
Chen H, Cui Y, Han X et al (2020) Liquid–liquid phase separation by SARS-CoV-2 nucleocapsid protein and RNA. Cell Res 30:1143–1145. https://doi.org/10.1038/s41422-020-00408-2
Lu W, Zheng BJ, Xu K et al (2006) Severe acute respiratory syndrome-associated coronavirus 3a protein forms an ion channel and modulates virus release. Proc Natl Acad Sci USA 103:12540–12545. https://doi.org/10.1073/pnas.0605402103
Kern DM, Sorum B, Mali SS et al (2021) Cryo-EM structure of SARS-CoV-2 ORF3a in lipid nanodiscs. Nat Struct Mol Biol 28:573–582. https://doi.org/10.1038/s41594-021-00619-0
Bianchi M, Borsetti A, Ciccozzi M, Pascarella S (2021) SARS-Cov-2 ORF3a: mutability and function. Int J Biol Macromol 170:820–826. https://doi.org/10.1016/j.ijbiomac.2020.12.142
McCallum M, De Marco A, Lempp FA et al (2021) N-terminal domain antigenic mapping reveals a site of vulnerability for SARS-CoV-2. Cell 184:2332-2347.e16. https://doi.org/10.1016/j.cell.2021.03.028
Rambaut A, Holmes EC, O’Toole Á et al (2021) Addendum: a dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat Microbiol 6:415. https://doi.org/10.1038/s41564-021-00872-5
Fontanet A, Cauchemez S (2020) COVID-19 herd immunity: where are we? Nat Rev Immunol 20:583–584
Wu J, Liang B, Chen C et al (2021) SARS-CoV-2 infection induces sustained humoral immune responses in convalescent patients following symptomatic COVID-19. Nat Commun 12:1–9. https://doi.org/10.1038/s41467-021-22034-1
Kang H, Wang Y, Tong Z, Liu X (2020) Retest positive for SARS-CoV-2 RNA of “recovered” patients with COVID-19: Persistence, sampling issues, or re-infection? J Med Virol 92:2263–2265. https://doi.org/10.1002/jmv.26114
To KK-W, Hung IF-N, Ip JD et al (2020) Coronavirus Disease 2019 (COVID-19) re-infection by a phylogenetically distinct severe acute respiratory syndrome Coronavirus 2 strain confirmed by whole genome sequencing. Clin Infect Dis. https://doi.org/10.1093/cid/ciaa1275
Brouqui P, Colson P, Melenotte C et al (2021) COVID-19 re-infection. Eur J Clin Invest 51:e13537. https://doi.org/10.1111/eci.13537
Bongiovanni M (2021) COVID-19 reinfection in a healthcare worker. J Med Virol 93:4058–4059. https://doi.org/10.1002/jmv.26565
Ledford H (2020) COVID-19 reinfection: three questions scientists are asking. Nature 585:168–169
Lopez Bernal J, Andrews N, Gower C et al (2021) Effectiveness of Covid-19 vaccines against the B. 1.617. 2 (delta) variant. N Engl J Med 385:585–594. https://doi.org/10.1056/NEJMoa2108891
Wise J (2020) Covid-19: New coronavirus variant is identified in UK. BMJ 371:m4857
Mahase E (2020) Covid-19: What have we learnt about the new variant in the UK? BMJ 371:m4944
Rambaut A, Loman N, Pybus O et al (2020) Preliminary genomic characterisation of an emergent SARS-CoV-2 lineage in the UK defined by a novel set of spike mutations—Virological. Virological.org, pp 1–9
Tegally H, Wilkinson E, Giovanetti M et al (2021) Detection of a SARS-CoV-2 variant of concern in South Africa. Nature 592:438–443. https://doi.org/10.1038/s41586-021-03402-9
Voloch CM, da Silva FR, de Almeida LGP et al (2021) Genomic characterization of a novel SARS-CoV-2 lineage from Rio de Janeiro, Brazil. J Virol. https://doi.org/10.1128/jvi.00119-21
Singh J, Rahman SA, Ehtesham NZ et al (2021) SARS-CoV-2 variants of concern are emerging in India. Nat Med 27:1131–1133. https://doi.org/10.1038/s41591-021-01397-4
Faria NR, Mellan TA, Whittaker C et al (2021) Genomics and epidemiology of the P. 1 SARS-CoV-2 lineage in Manaus, Brazil. Science 372:815–821
Volz E, Mishra S, Chand M, Barrett JC, Johnson R, Geidelberg L, Hinsley WR, Laydon DJ, Dabrera G, O’Toole ÁAR (2021) Assessing transmissibility of SARS-CoV-2 lineage B.1.1.7 in England enhanced reader.pdf. Nature 593:266–269. https://doi.org/10.1038/s41586-021-03470-x
Frampton D, Rampling T, Cross A et al (2021) Genomic characteristics and clinical effect of the emergent SARS-CoV-2 B.1.1.7 lineage in London, UK: a whole-genome sequencing and hospital-based cohort study. Lancet Infect Dis 3099:00170–00175. https://doi.org/10.1016/S1473-3099(21)00170-5
Davies NG, Abbott S, Barnard RC et al (2021) Estimated transmissibility and impact of SARS-CoV-2 lineage B.1.1.7 in England. Science 3055:eabg3055. https://doi.org/10.1126/science.abg3055
Planas D, Bruel T, Grzelak L et al (2021) Sensitivity of infectious SARS-CoV-2 B. 1.1. 7 and B. 1.351 variants to neutralizing antibodies. Nat Med 27:917–924. https://doi.org/10.1038/s41591-021-01318-5
Shen X, Tang H, McDanal C et al (2021) SARS-CoV-2 variant B.1.1.7 is susceptible to neutralizing antibodies elicited by ancestral spike vaccines. Cell Host Microbe 29:529-539.e3. https://doi.org/10.1016/j.chom.2021.03.002
Muik A, Wallisch AK, Sänger B et al (2021) Neutralization of SARS-CoV-2 lineage B.1.1.7 pseudovirus by BNT162b2 vaccine–elicited human sera. Science 371:1152–1153. https://doi.org/10.1126/science.abg6105
Wall EC, Wu M, Harvey R et al (2021) Neutralising antibody activity against SARS-CoV-2 VOCs B.1.617.2 and B.1.351 by BNT162b2 vaccination. Lancet 397:2331–2333. https://doi.org/10.1016/S0140-6736(21)01290-3
Yadav PD, Sapkal G, Ella R et al (2021) Neutralization against B. 1.351 and B. 1.617. 2 with sera of COVID-19 recovered cases and vaccinees of BBV152. bioRxiv. https://doi.org/10.1101/2021.06.05.447177
Edara VV, Norwood C, Floyd K et al (2021) Infection-and vaccine-induced antibody binding and neutralization of the B. 1.351 SARS-CoV-2 variant. Cell Host Microbe 29:516–521
Rees-Spear C, Muir L, Griffith SA et al (2021) The effect of spike mutations on SARS-CoV-2 neutralization. Cell Rep 34:108890. https://doi.org/10.1016/j.celrep.2021.108890
Shen X, Tang H, Pajon R et al (2021) Neutralization of SARS-CoV-2 variants B.1.429 and B.1.351. N Engl J Med 384:2352–2354. https://doi.org/10.1056/nejmc2103740
Wibmer CK, Ayres F, Hermanus T et al (2021) SARS-CoV-2 501Y. V2 escapes neutralization by South African COVID-19 donor plasma. Nat Med 27:622–625
Cele S, Gazy I, Jackson L et al (2021) Escape of SARS-CoV-2 501Y. V2 from neutralization by convalescent plasma. Nature 593:142–146
Abu-Raddad LJ, Chemaitelly H, Butt AA (2021) Effectiveness of the BNT162b2 Covid-19 Vaccine against the B.1.1.7 and B.1.351 Variants. N Engl J Med 385:187–189. https://doi.org/10.1056/nejmc2104974
Dejnirattisai W, Zhou D, Supasa P et al (2021) Antibody evasion by the P. 1 strain of SARS-CoV-2. Cell 184:2939–2954
Wang Z, Schmidt F, Weisblum Y et al (2021) mRNA vaccine-elicited antibodies to SARS-CoV-2 and circulating variants. Nature 592:616–622. https://doi.org/10.1038/s41586-021-03324-6
Zhou D, Dejnirattisai W, Supasa P et al (2021) Evidence of escape of SARS-CoV-2 variant B.1.351 from natural and vaccine-induced sera. Cell 184:2348-2361.e6. https://doi.org/10.1016/j.cell.2021.02.037
Hoffmann M, Arora P, Groß R et al (2021) SARS-CoV-2 variants B.1.351 and P.1 escape from neutralizing antibodies. Cell 184:2384-2393.e12. https://doi.org/10.1016/j.cell.2021.03.036
Li Q, Nie J, Wu J et al (2021) SARS-CoV-2 501Y.V2 variants lack higher infectivity but do have immune escape. Cell 184:2362-2371.e9. https://doi.org/10.1016/j.cell.2021.02.042
Wang P, Nair MS, Liu L et al (2021) Antibody resistance of SARS-CoV-2 variants B. 1.351 and B. 1.1. 7. Nature 593:130–135. https://doi.org/10.1038/s41586-021-03398-2
Liu J, Liu Y, Xia H et al (2021) BNT162b2-elicited neutralization of B.1.61.7 and other SARS-CoV-2 variants. Nature. https://doi.org/10.1038/s41586-021-03693-y
Shang J, Ye G, Shi K et al (2020) Structural basis of receptor recognition by SARS-CoV-2. Nature 581:221–224. https://doi.org/10.1038/s41586-020-2179-y
Liu Y, Liu J, Plante KS et al (2021) The N501Y spike substitution enhances SARS-CoV-2 transmission. BioRxiv. https://doi.org/10.1101/2021.03.08.434499
Tian F, Tong B, Sun L et al (2021) Mutation N501Y in RBD of spike protein strengthens the interaction between COVID-19 and its receptor ACE2. BioRxiv. https://doi.org/10.1101/2021.02.14.431117
Nelson G, Buzko O, Spilman PR et al (2021) Molecular dynamic simulation reveals E484K mutation enhances spike RBD-ACE2 affinity and the combination of E484K, K417N and N501Y mutations (501Y. V2 variant) induces conformational change greater than N501Y mutant alone, potentially resulting in an esca. BioRxiv. https://doi.org/10.1101/2021.01.13.426558
Weisblum Y, Schmidt F, Zhang F et al (2020) Escape from neutralizing antibodies 1 by SARS-CoV-2 spike protein variants. Elife 9:1. https://doi.org/10.7554/eLife.61312
Greaney AJ, Loes AN, Crawford KHD et al (2021) Comprehensive mapping of mutations in the SARS-CoV-2 receptor-binding domain that affect recognition by polyclonal human plasma antibodies. Cell Host Microbe 29:463-476.e6. https://doi.org/10.1016/j.chom.2021.02.003
Jangra S, Ye C, Rathnasinghe R et al (2021) SARS-CoV-2 spike E484K mutation reduces antibody neutralisation. Lancet Microbe 2:e283–e284. https://doi.org/10.1016/S2666-5247(21)00068-9
Liu Z, VanBlargan LA, Bloyet L-M et al (2021) Identification of SARS-CoV-2 spike mutations that attenuate monoclonal and serum antibody neutralization. Cell Host Microbe 29:477–488
Andreano E, Piccini G, Licastro D et al (2020) SARS-CoV-2 escape in vitro from a highly neutralizing COVID-19 convalescent plasma. bioRxiv. https://doi.org/10.1101/2020.12.28.424451
Brown JC, Goldhill DH, Zhou J et al (2021) Increased transmission of SARS-CoV-2 lineage B.1.1.7 (VOC 2020212/01) is not accounted for by a replicative advantage in primary airway cells or antibody escape. bioRxiv. https://doi.org/10.1101/2021.02.24.432576
Lubinski B, Tang T, Daniel S et al (2021) Functional evaluation of proteolytic activation for the SARS-CoV-2 variant B. 1.1.7: role of the P681H mutation. bioRxiv. https://doi.org/10.1101/2021.04.06.438731
Peacock TP, Sheppard CM, Brown JC et al (2021) The SARS-CoV-2 variants associated with infections in India, B. 1.617, show enhanced spike cleavage by furin. bioRxiv. https://doi.org/10.1101/2021.05.28.446163
Li Q, Wu J, Nie J et al (2020) The impact of mutations in SARS-CoV-2 spike on viral infectivity and antigenicity. Cell 182:1284–1294
Kemp BA, Papa SA, Datir G et al (2021) Recurrent emergence of SARS-CoV-2 spike deletion H69/V70 and its role in the variant of concern lineage B.1.1.7 Genomics UK (COG-UK) Consortium. Cell Rep 35:109292
McCarthy KR, Rennick LJ, Nambulli S et al (2021) Recurrent deletions in the SARS-CoV-2 spike glycoprotein drive antibody escape. Science 371:1139–1142. https://doi.org/10.1126/science.abf6950
Lan J, Ge J, Yu J et al (2020) Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581:215–220. https://doi.org/10.1038/s41586-020-2180-5
Cottam EM, Whelband MC, Wileman T (2014) Coronavirus NSP6 restricts autophagosome expansion. Autophagy 10:1426–1441. https://doi.org/10.4161/auto.29309
Benvenuto D, Angeletti S, Giovanetti M et al (2020) Evolutionary analysis of SARS-CoV-2: how mutation of Non-Structural Protein 6 (NSP6) could affect viral autophagy. J Infect 81:e24–e27. https://doi.org/10.1016/j.jinf.2020.03.058
Grubaugh ND, Petrone ME, Holmes EC (2020) We shouldn’t worry when a virus mutates during disease outbreaks. Nat Microbiol 5:529–530
Choi B, Choudhary MC, Regan J et al (2020) Persistence and evolution of SARS-CoV-2 in an immunocompromised host. N Engl J Med 383:2291–2293
Baang JH, Smith C, Mirabelli C et al (2021) Prolonged severe acute respiratory syndrome Coronavirus 2 replication in an immunocompromised patient. J Infect Dis 223:23–27
Voloch CM, Ronaldo da Silva F, de Almeida LGP et al (2020) Intra-host evolution during SARS-CoV-2 persistent infection. medRxiv. https://doi.org/10.1101/2020.11.13.20231217
Kemp SA, Collier DA, Datir RP et al (2021) SARS-CoV-2 evolution during treatment of chronic infection. Nature 592:277–282. https://doi.org/10.1038/s41586-021-03291-y
van Dorp L, Richard D, Tan CCS et al (2020) No evidence for increased transmissibility from recurrent mutations in SARS-CoV-2. Nat Commun 11:1–8. https://doi.org/10.1038/s41467-020-19818-2
Yi H (2020) 2019 Novel coronavirus is undergoing active recombination. Clin Infect Dis 71:884–887. https://doi.org/10.1093/cid/ciaa219
Varabyou A, Pockrandt C, Salzberg SL, Pertea M (2021) Rapid detection of inter-clade recombination in SARS-CoV-2 with Bolotie. Genetics 218:iyab074. https://doi.org/10.1093/genetics/iyab074
Jackson B, Rambaut A, Pybus O et al (2021) Recombinant SARS-CoV-2 genomes involving lineage B.1.1.7 in the UK—Virological. In: Virological. https://virological.org/t/recombinant-sars-cov-2-genomes-involving-lineage-b-1-1-7-in-the-uk/658
Sharma S, Patnaik SK, Thomas Taggart R et al (2015) APOBEC3A cytidine deaminase induces RNA editing in monocytes and macrophages. Nat Commun 6:1–15. https://doi.org/10.1038/ncomms7881
Lamers MM, van den Hoogen BG, Haagmans BL (2019) ADAR1: “Editor-in-Chief” of cytoplasmic innate immunity. Front Immunol 10:1763. https://doi.org/10.3389/fimmu.2019.01763
Hamilton CE, Papavasiliou FN, Rosenberg BR (2010) Diverse functions for DNA and RNA editing in the immune system. RNA Biol 7:220–228. https://doi.org/10.4161/rna.7.2.11344
Lai MMC, Cavanagh D (1997) The molecular biology of coronaviruses. Adv Virus Res 48:1–100
Simon-Loriere E, Holmes EC (2011) Why do RNA viruses recombine? Nat Rev Microbiol 9:617–626. https://doi.org/10.1038/nrmicro2614
Graham RL, Baric RS (2010) Recombination, reservoirs, and the modular spike: mechanisms of coronavirus cross-species transmission. J Virol 84:3134–3146. https://doi.org/10.1128/jvi.01394-09
Francisco RS, Benites LF, Lamarca AP et al (2021) Pervasive transmission of E484K and emergence of VUI-NP13L with evidence of SARS-CoV-2 co-infection events by two different lineages in Rio Grande do Sul. Brazil. Virus Res 296:198345. https://doi.org/10.1016/j.virusres.2021.198345
Pedro N, Silva CN, Magalhães AC et al (2021) Dynamics of a dual sars-cov-2 lineage co-infection on a prolonged viral shedding COVID-19 case: Insights into clinical severity and disease duration. Microorganisms 9:1–10. https://doi.org/10.3390/microorganisms9020300
Taghizadeh P, Salehi S, Heshmati A et al (2021) Study on SARS-CoV-2 strains in Iran reveals potential contribution of co-infection with and recombination between different strains to the emergence of new strains. Virology 562:63–73. https://doi.org/10.1016/j.virol.2021.06.004
VanInsberghe D, Neish AS, Lowen AC, Koelle K (2020) Identification of SARS-CoV-2 recombinant genomes. BioRxiv. https://doi.org/10.1101/2020.08.05.238386
Ignatieva A, Hein J, Jenkins PA (2021) Evidence of ongoing recombination in SARS-CoV-2 through genealogical reconstruction. BioRxiv. https://doi.org/10.1101/2021.01.21.427579
Moyo-Gwete T, Madzivhandila M, Makhado Z et al (2021) Cross-reactive neutralizing antibody responses elicited by SARS-CoV-2 501Y.V2 (B.1.351). N Engl J Med 384:2161–2163. https://doi.org/10.1056/NEJMc2104192
Funding
This research was supported by a grant from the National Institute for Genetic Engineering and Biotechnology, Tehran, Iran.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There is no conflict of interest.
Data availability statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Additional information
Handling Editor: William G Dundon.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Safari, I., Elahi, E. Evolution of the SARS-CoV-2 genome and emergence of variants of concern. Arch Virol 167, 293–305 (2022). https://doi.org/10.1007/s00705-021-05295-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05295-5