Abstract
Porcine reproductive and respiratory syndrome (PRRS) is one of the most economically important infectious diseases of swine worldwide. Immunization with an attenuated vaccine is considered an effective method for reducing the economic losses resulting from porcine reproductive and respiratory syndrome virus (PRRSV) infection. Several studies have shown that PRRSV can be attenuated by passage in Marc-145 cells, but it is still not clear whether this attenuation influences the immunogenicity of PRRSV and what the mechanism of attenuation is. In order to study the mechanism of attenuation and immunogenicity of highly pathogenic (HP) PRRSV, the HP-PRRSV strain XH-GD was serially 122 times passaged in Marc-145 cells. Genomic sequence comparisons were made at selected passages. To explore the differences in pathogenicity and immunogenicity at different passages, three passages (P5, P62 and P122) were selected for an animal challenge experiment, which showed that passage in Marc-145 cells resulted in attenuation of the virus. After 122 passages, 35 amino acid changes were observed in the structural proteins and non-structural proteins. The animal challenge experiment showed that pathogenicity decreased with increasing passage number. The N antibody level and specific neutralizing (SN) antibody titers also decreased with increasing passage number in the late stage of the animal experiment. This study indicated that the virulence of XH-GD was decreased by passage in Marc-145 cells and that overattenuation might influence the immunogenicity of virus. These results might contribute to our understanding of the mechanism of attenuation.
Similar content being viewed by others
Introduction
Porcine reproductive and respiratory syndrome (PRRS) is caused by porcine reproductive and respiratory syndrome virus (PRRSV), which is a member of the family Arteriviridae, order Nidovirales [2, 27]. PRRSV is an enveloped, positive-strand RNA virus, and its genome is approximately 15 kb in length. PRRSV has two different genotypes: the European type (type 1) and the American type (type 2) [6]. The genome of PRRSV has at least nine overlapping open reading frames (ORF1-7) which encode fourteen non-structural proteins (Nsp1α, Nsp1β, Nsp2-6, Nsp7α, Nsp7β, Nsp8-12) and eight structural proteins (GP1, GP2, E, GP3, GP4, GP5, M and N) [7]. In 2011, a new ORF was discovered and named ORF5a [10]. Recently, Nsp2 was shown to be a virion-associated structural protein [11].
PRRS was first reported in North America in 1987 [34] and has since become prevalent worldwide [16]. It is one of the most economically important infectious diseases for the swine industry and is characterized by respiratory disorder in piglets and abortion in pregnant sows [32]. PRRS has caused a loss of almost 560 million dollars per year, or $1.5 million dollars every day, to the U.S. economy [24]. In 1996, PRRSV was found in China and since then it has become prevalent throughout China. In 2006, highly pathogenic (HP)-PRRSV emerged in China and caused disease characterized by high fever and high mortality in piglets of all ages [18]. An obvious feature of the newly emerged strains was a discontinuous deletion of 30 amino acids. This virus has spread rapidly and has become the most prevalent strain in China [9].
The use of attenuated vaccines is considered an effective method for reducing losses from PRRS, and many attenuated vaccines are available on the market [25], such as MLV, VR2332, JXA1, and HuN4-F11 [15, 20]. The purpose of this study was to identify mutations selected during serial passage that could be associated with attenuation, changes in the immunogenicity of the virus, or adaption of the virus to cells.
Materials and methods
Virus and cells
Marc-145 cells are derivatives of the African green monkey kidney cell line MA-104. Marc-145 cells were cultured in Dulbecco’s modified Eagle medium (DMEM) containing 10 % fetal bovine serum (FBS; Gibco) and streptomycin at 37 °C in a 5 % CO2 atmosphere.
The PRRSV strain XH-GD was isolated from Guangdong province in 2007 [35], where the disease situation was complex [3, 28]. It is one of the epidemic HP-PRRSV strains in Guangdong, and it exhibits 99.1 % nucleotide sequence identity to strain JXA1. The recently isolated viruses in Guangdong also share high similarity with strain XH-GD [30, 35]. The strain was continuously passaged 122 times in Marc-145 cells and plaque purified at every tenth passage [40]. The 5th (P5), 62nd (P62), and 122nd (P122) passages of XH-GD were used for the animal infection experiment.
RT-PCR and sequencing
The complete genome sequences of the selected passages (P2, P12, P22, P32, P42, P52, P62, P72, P82, P92, P102, P112, P122) were sequenced and analyzed as reported previously [39]. RNA was extracted from infected cells, using RNAfast200 according to manufacturer’s directions (Fastagen, Shanghai FASTAGEN Biotech Co, Ltd, China). Five μl of cDNA, which was generated from 5 µg of RNA using reverse transcriptase (M-MLV, Takara), was used as the template in the subsequent PCR. The genome was divided into 14 segments, and the sense and antisense primers used for each segment and the PCR protocol were in accordance with a previous report [39]. PCR products were sent to Invitrogen (Shanghai, China) for sequencing. Sanger sequencing techniques were used. The Clustal W method of the Lasergene software (version 7.1.0) (DNASTAR, Inc., Madison, WI) was used for sequence comparison and analysis [1].
Animals
Twenty 24-days-old piglets were used for the virus infection experiment. The piglets were provided by Qingyuan Pig Farm in Guangdong province. The piglets were tested and shown to be free of PRRSV, CSFV (classical swine fever virus, PRV (porcine pseudorabies virus) (IDEXX Labs Inc. USA), PCV2 (porcine circovirus type 2) (Biochek, The Netherlands), and PPV (porcine parvovirus) (LSI, France) by ELISA.
Animal experiment
Twenty piglets were randomly placed into four independent pigpens with five piglets in each group. The piglets in groups 1, 2 and 3 were inoculated with 2 ml of P5 (group 1), P62 (group 2), and P122 (group 3), respectively, of XH-GD virus, which was injected intratracheally at a concentration of 2 × 104.8TCID50/mL. The pigs in group 4 were inoculated with 2 ml of PBS as negative controls. Clinical signs in the experimental animals were observed, and the rectal temperature was recorded daily. Blood samples were collected at 1, 4, 7, 10, 14, 17, 20, 24, 28 days post-inoculation (dpi), and a commercial ELISA for detection of IgG directed against the N protein of PRRSV was used for detection of PRRSV antigens and antibodies. Anatomic pathological changes were observed when the piglets were euthanized at 28 dpi. The lungs were collected, tissue sections were prepared, and histological examination was carried out.
Viremia detection by RT-PCR
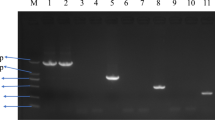
To detect whether the piglets had viremia, blood samples collected at 1, 4, 7, 10, 14, 17, 20, 24, 28 dpi were tested by RT-PCR. Total RNA was isolated as described above, and the PCR was carried out as reported previously [39]. The primers ORF5-F (5’-GTTTAGCCTTCTTTTGCC-3’) and ORF5-R (5’-TATATCATCACTGGCGTGTAGG-3’) were used for amplification of the ORF5 fragment, which was 731 bp in length [36].
The histological examination of the lung
At necropsy, tissue samples from lungs of the study animals were collected for histopathology analysis. Lung samples were placed in 10 % neutral buffered formalin and stained with hematoxylin and eosin (H&E) The sections were examined by light microscopy [17].
Neutralization test
Twofold serial dilutions of sera (20, 2−1, 2−2, 2−3, 2−4 and 2−5) were made with DMEM. About 200 TCID50 of XH-GD (50 µl) was incubated with treated serum (50 µl) or DMEM (as a control) at 37 °C for 1h, and the mixtures were then transferred to 96-wells plate containing Marc-145 cells. Each well was examined for cytopathic effect (CPE) after 48 h. TCID50 was calculated according to the method of Reed and Muench [30].
Statistical analysis
All experiments were repeated independently at least three times, and the data were analyzed using GraphPad Prism (version 5.0) software. Survival curves were draw using SPSS (version 21.0). An independent t-test was used for variation analysis.
Ethics statement
In order to ensure the environment security, repeatability and reliability of the animal experiments, the piglets were raised in a negative-pressure animal house at South China Agriculture University. The experiments were approved by the institute and carried out in strict accordance with animal ethics guidelines and approved protocols.
Results
Amino acid and nucleotide changes during passage of XH-GD
The XH-GD strain was serial passaged 122 times, and thirteen different viruses were selected and sequenced. Except for NSP1a, NSP5, NSP6, NSP8 and NSP12, mutations were observed in all of the genes (Table 1). In ORF1a, there were 14 predicted amino acid changes: one in NSP4, three each in NSP3 and NSP7, and six in NSP2. In ORF1b, there were four changes: one each in NSP10 and NSP11 and two in NSP9. ORF2a, ORF2b, ORF4, ORF5 and ORF6 all contained two changes: ORF3 had seven changes. Only one mutation was observed in ORF5a. Among the 36 amino acid mutations, 50 % (18/36) occurred in non-structural proteins, and the rest 50 % (18/36) were located in structural proteins.
Nucleotide and amino acid analysis of different viruses
The mutation rates were different for the different structural proteins or dissimilar non-structural proteins. NSP2 and GP3 had the highest mutation frequency in their nucleotide and the amino acid sequences during passage. The mutational relationship between genes and the amino acids is shown in Table 2 [20, 31]. The mutations in GP2, E, GP3 and GP4 were shown previously to be present in other attenuated viruses [38]. Further investigation is needed to determine whether these mutations lead to a decline in virulence.
Clinical signs and pathological examination of piglets infected with different virus passages
The deaths of the animals in each group were recorded. Two piglets (no. 1 and no. 5) in group 1 died (Fig. 1A, Supplementary Documents). Total mortality was 40 % (Fig. 1B, Supplementary Documents). All piglets inoculated with P62 or P122 survived throughout the experiment (Supplementary Documents). These results clearly show that the virulence of the virus was reduced after serial passage in Marc-145 cells (Table 3).
The clinical symptoms in animals of group 1 inoculated with XH-GD P5 included depression, loss of appetite, inertia and sluggishness at 3 dpi, and the piglets showed typical signs of PRRS, such as coughing, sneezing, constipation, erubescence and weight loss at 7 dpi. At 14 dpi, the piglets began to shiver. Then, two piglets died at 20 (no. 5) and 22 (no. 1) dpi. Pigs in group 2 (inoculated with P62) showed depression and anorexia at 4 dpi, and a slight cough and individual constipation were observed at 7 dpi. All of these piglets recovered at 15 dpi; those in group 3 (inoculated with P122) had only a slight cough and then recovered at 8 dpi. There were no obvious differences in the activity and reaction of the pigs when compared with those of group 4 (negative control group). The group 4 pigs did not show any obvious clinical signs (Table 3). The average body temperature (Fig. 1) of the pigs in group 1 was 40 °C at 4 dpi, continuing until the end of the experiment. There were 12 days on which the average temperature was higher than 40.5 °C. In group 2, the average temperature of the piglets was above 40.0 °C and lasted for 8 days. Their average temperature was also lower than that of group 1. The average temperature of group 3 increased slightly. The temperature exceeded 40.0 °C at 11 dpi and 14 dpi. The average temperature of the control group was around 39.6 °C, which is within the normal range.
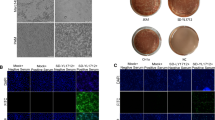
The surviving piglets were euthanized at 28 dpi, and pathological changes were observed (Fig. 2). The piglets in group 1 (P5) showed hemorrhage, congestion, interstitial broadening, extravasated blood at inguinal lymph nodes, and lymphocyte infiltration – all features of interstitial pneumonia. The major lesion of piglets in group 2 (P62) was hemorrhage in lung and inguinal lymph nodes. Only slight interstitial pneumonia was observed, but lymphocytic infiltration of skin persisted. There were no obvious lesions in group 3 (P122) or in the control group.
The pathological changes in the lung at 28 dpi (×200). A. The group infected with the 5th-passage virus showed severe hemorrhage and congestion. B. The group infected with the 62nd-passage virus showed slight hemorrhage and interstitial pneumonia. C. No evident lesions were found in the group infected with the 122nd-passage virus. D. The negative control had no observed pathological change
Serological response differences in swine infected with parental and highly passaged XH-GD strains
Serum specimens were collected at 1, 4, 7, 10,14, 17, 20, 24, and 28 dpi, and RT-PCR was used for detection of the viral nucleic acid. Viremia was detected at 4 dpi in group 1. The whole group was positive from 7 dpi until the end, and the piglets infected with P62 were viremic from 14 dpi to 20 dpi. Only two pigs had viremia at 28 dpi, and those in group 3 showed only slight viremia and became negative at 24 dpi (Table 4).
All serum samples were tested by ELISA. The results are shown in Fig. 3. The antibody levels in the three groups increased at 10 dpi. Furthermore, except for group 3, the antibody level increased continuously in the other groups, and the antibody level also varied among the groups. The antibody levels in group 3 (P122) were always lower than those in group 1 (P5) and group 2 (P62) and were significantly lower than those in group 1 at 24 dpi and 28 dpi and group 2 at 28 dpi. We also found that there was no obvious relationship between viremia and the N antibody level (data not shown).
Antibody responses of piglets infected with different passages of the XH-GD strain. The mean value is shown for the different groups. Samples with S/P ≥ 0.4 were considered PRRSV antibody positive. Differences were deemed to be statistically significant if the P-value was below 0.05. *, P < 0.05; **, P < 0.01; ***, P < 0.001. a The dashed line at y = 0.4 is the value of the standard, which was used as the cutoff value for positivity. b *, P < 0.05; **, P < 0.01; ***, P < 0.001
Since it was reported previously that PRRSV specific neutralizing (SN) antibody is produced 3 weeks after inoculation [21], we collected serum samples from 21 dpi to 28 dpi for neutralization tests (Fig. 4), which showed that SN antibody induced by P5 was detected at 24 dpi. SN antibody induced by P62 and P122 was not detected until 28 dpi, and the higher virus passage, the lower the SN titer. There was a significant difference in the SN antibody level between P5 and P122 at 28 dpi, indicating that P5 may induce a superior virus neutralizing antibody response. However, the SN titers were low in all groups at 28 dpi.
Discussion
PRRS has been recognized as one of the most serious swine diseases associated with significant economic losses in China. Furthermore, unprecedented large-scale outbreaks of a previously unknown ‘high-fever’ disease, HP-PRRS, swept across the whole country in 2006 [26]. Different from the typical PRRS, HP-PRRS was more severe and characterized by high fever, high morbidity and high mortality in all ages of piglets. Therefore, it is extremely urgent to restrict the spread of HP-PRRS. It is clear that the live attenuated vaccines are the most effective approach for controlling the clinical symptoms of this disease, but the mechanism by which the virus can be attenuated is still unclear [29].
In this study, XH-GD was continuously passaged in Marc-145 cells. As reported previously, in the process of passaging, the virus titer increased from 104.6 TCID50/ml to 106.5 TCID50/ml [31]. This indicates that the virus adapts gradually to Marc-145 cells. In our animal experiment, we found significant differences in the levels of antibodies against the N protein and in the SN in a later stage of the experiment. This may be due to decreased replication of the virus in vivo after passaging, leading to fewer viral particles, less viral antigen, and less stimulation of the host immune system. This suggests that overpassaging of PRRSV on Marc-145 cells affects the immunogenicity of this virus. In this study, HP-PRRSV XH-GD was attenuated by passaging in Marc-145 cells. During the passaging, the Nsp4, Nsp6, Nsp12 and N sequences were absolutely conserved, suggesting that these proteins might not be involved in adaptation of the virus to Marc-145 cells.
The NSPs of PRRSV play a critical role in the replication and maturation of the virus. In order to identify mutations in NSPs that could play a critical role in attenuation, comparisons were made among different attenuated strains; however, no such mutations were found. Recently, researchers found that the NSP9 and NSP10 proteins contribute to the virulence of PRRSV [17]. NSP9, which is the core protein of the polymerase complex, is one of the most variable proteins according to amino acid mutation analysis. Most of the mutations in this protein were located in the region from aa 135 to 420. Partial superposition with members of the RNA-directed RNA polymerase (RdRp) superfamily has been observed [36], but it needs to be investigated whether those mutations lead to attenuation.
The minor structural proteins, including GP2, E, GP3 and GP4, can connect with each other to form a complex that is important for virus infectivity and also helps to induce an immune response [5]. A comparison with other attenuated viruses showed the occurrence of mutations at aa 9 of GP2a, especially in attenuated viruses derived from HP-PRRSV, which contains an Asp residue at this position.
The E protein of PRRSV is vital for infection. It has been speculated that the E protein acts as an ion channel protein and is present in the viral envelope [13]. The E protein may play a role in the shell of the virus and entry of viral genes into the cell [14]. In a comparison with other viruses, we found that six attenuated viruses had similar mutations at the ninth position of E, most of which were D9N. This might be used as a marker to distinguish vaccine strains from naturally circulating viruses and it might also be involved in the adaptation of the virus to Marc-145 cells (Table 5).
GP3 is a glycosylated protein with seven glycosylation sites [22]. It has high antigenicity [8]. Antibodies against GP3 participate in the process of virus neutralization. Some consistent mutations in GP3 at positions 79, 85 and 165 were found in JXA1/JXAP170 and XH-GDP5/XH-GDP122. Most noteworthy is the fact that amino acids 85 and 165 of GP3 were both changed after the virus was attenuated and therefore may have a relationship to overattenuation and the decrease in the antibody level [33].
The GP4 is a type I membrane protein [4]. It also can combine with CD163 and induce the host to produce SN antibody [19]. At position 43 of GP4, most attenuated have a mutation from Asp to another amino acid, but XH-GD/XH-GDP122 did not have this mutation.
GP5 is a major structural protein [23] that is very important for infection and packaging of the viral genome [37]. There have been many reports showing that an antibody against GP5 can neutralize the virus and reduce the damage caused by PRRS [12]. It is noteworthy that a Q196R mutation in GP5 is commonly found in other vaccine strains. Whether this mutation reduces the virulence of the virus needs further investigation.
To sum up, the virulence of the XH-GD strain was attenuated by serial passage. The immunogenicity of the virus decreased when it became overattenuated, which may result in the failure of vaccination. We speculate that aa 9 of E, 43 of GP4, and 196 of GP5 may be involved in attenuation and that aa 79, 85, and 165 of GP3 may re involved in overattenuation. Further investigation is needed to determine whether this strain generates adequate immunity against field strains or whether these mutations lead to attenuation or overattenuation. To develop an ideal vaccine against PRRSV, safety and stability will be the main factors that we should take into consideration. Therefore, much work remains to be done in the future.
References
Burland TG (2000) DNASTAR’s Lasergene sequence analysis software. Methods Mol Biol 132:71–91
Cavanagh D (1997) Nidovirales: a new order comprising Coronaviridae and Arteriviridae. Arch Virol 142:629–633
Chen J, Fu X, Chen Y, He S, Zheng Y, Cao Z, Yu W, Zhou H, Su S, Zhang G (2014) Identification of four genotypes of H3N2 swine influenza virus in pigs from southern China. Arch Virol 159:2705–2709
Costers S, Lefebvre DJ, Van Doorsselaere J, Vanhee M, Delputte PL, Nauwynck HJ (2010) GP4 of porcine reproductive and respiratory syndrome virus contains a neutralizing epitope that is susceptible to immunoselection in vitro. Arch Virol 155:371–378
Das PB, Dinh PX, Ansari IH, de Lima M, Osorio FA, Pattnaik AK (2010) The minor envelope glycoproteins GP2a and GP4 of porcine reproductive and respiratory syndrome virus interact with the receptor CD163. J Virol 84:1731–1740
de Lima M, Pattnaik AK, Flores EF, Osorio FA (2006) Serologic marker candidates identified among B-cell linear epitopes of Nsp2 and structural proteins of a North American strain of porcine reproductive and respiratory syndrome virus. Virology 353:410–421
Fang Y, Snijder EJ (2010) The PRRSV replicase: exploring the multifunctionality of an intriguing set of nonstructural proteins. Virus Res 154:61–76
Gonin P, Mardassi H, Gagnon CA, Massie B, Dea S (1998) A nonstructural and antigenic glycoprotein is encoded by ORF3 of the IAF-Klop strain of porcine reproductive and respiratory syndrome virus. Arch Virol 143:1927–1940
Guo XK, Zhang Q, Gao L, Li N, Chen XX, Feng WH (2013) Increasing expression of microRNA 181 inhibits porcine reproductive and respiratory syndrome virus replication and has implications for controlling virus infection. J Virol 87:1159–1171
Johnson CR, Griggs TF, Gnanandarajah J, Murtaugh MP (2011) Novel structural protein in porcine reproductive and respiratory syndrome virus encoded by an alternative ORF5 present in all arteriviruses. J Gen Virol 92:1107–1116
Kappes MA, Miller CL, Faaberg KS (2013) Highly divergent strains of porcine reproductive and respiratory syndrome virus incorporate multiple isoforms of nonstructural protein 2 into virions. J Virol 87:13456–13465
Karniychuk UU, Nauwynck HJ (2013) Pathogenesis and prevention of placental and transplacental porcine reproductive and respiratory syndrome virus infection. Vet Res 44:95
Lee C, Yoo D (2006) The small envelope protein of porcine reproductive and respiratory syndrome virus possesses ion channel protein-like properties. Virology 355:30–43
Lee C, Hodgins D, Calvert JG, Welch SK, Jolie R, Yoo D (2006) Mutations within the nuclear localization signal of the porcine reproductive and respiratory syndrome virus nucleocapsid protein attenuate virus replication. Virology 346:238–250
Leng X, Li Z, Xia M, Li X, Wang F, Wang W, Zhang X, Wu H (2012) Mutations in the genome of the highly pathogenic porcine reproductive and respiratory syndrome virus potentially related to attenuation. Vet Microbiol 157:50–60
Li Y, Wang X, Bo K, Wang X, Tang B, Yang B, Jiang W, Jiang P (2007) Emergence of a highly pathogenic porcine reproductive and respiratory syndrome virus in the Mid-Eastern region of China. Vet J 174:577–584
Li Y, Zhou L, Zhang J, Ge X, Zhou R, Zheng H, Geng G, Guo X, Yang H (2014) Nsp9 and Nsp10 contribute to the fatal virulence of highly pathogenic porcine reproductive and respiratory syndrome virus emerging in China. Plos Pathog 10:e1004216
Liu JK, Wei CH, Yang XY, Hou XL, Dai AL, Li XH, Wei MK, Pan XZ (2013) Genetic diversity and evolutionary characterization of Chinese porcine reproductive and respiratory syndrome viruses based on NSP2 and ORF5. Arch Virol 158:1811–1816
Lopez OJ, Osorio FA (2004) Role of neutralizing antibodies in PRRSV protective immunity. Vet Immunol Immunopathol 102:155–163
Lu W, Sun B, Mo J, Zeng X, Zhang G, Wang L, Zhou Q, Zhu L, Li Z, Xie Q, Bi Y, Ma J (2014) Attenuation and immunogenicity of a live high pathogenic PRRSV vaccine candidate with a 32-amino acid deletion in the nsp2 protein. J Immunol Res 2014:810523
Meier WA, Galeota J, Osorio FA, Husmann RJ, Schnitzlein WM, Zuckermann FA (2003) Gradual development of the interferon-gamma response of swine to porcine reproductive and respiratory syndrome virus infection or vaccination. Virology 309:18–31
Meulenberg JJ, Petersen-den BA, de Kluyver EP, Moormann RJ, Schaaper WM, Wensvoort G (1995) Characterization of structural proteins of Lelystad virus. Adv Exp Med Biol 380:271–276
Nam HM, Chae KS, Song YJ, Lee NH, Lee JB, Park SY, Song CS, Seo KH, Kang SM, Kim MC, Choi IS (2013) Immune responses in mice vaccinated with virus-like particles composed of the GP5 and M proteins of porcine reproductive and respiratory syndrome virus. Arch Virol 158:1275–1285
Neumann EJ, Kliebenstein JB, Johnson CD, Mabry JW, Bush EJ, Seitzinger AH, Green AL, Zimmerman JJ (2005) Assessment of the economic impact of porcine reproductive and respiratory syndrome on swine production in the United States. J Am Vet Med Assoc 227:385–392
Nilubol D, Tripipat T, Hoonsuwan T, Tipsombatboon P, Piriyapongsa J (2014) Dynamics and evolution of porcine reproductive and respiratory syndrome virus (PRRSV) ORF5 following modified live PRRSV vaccination in a PRRSV-infected herd. Arch Virol 159:17–27
Olanratmanee EO, Nuntawan NAS, Thanawongnuwech R, Kunavongkrit A, Tummaruk P (2013) Reproductive parameters following a PRRS outbreak where a whole-herd PRRS MLV vaccination strategy was instituted post-outbreak. Trop Anim Health Prod 45:1099–1106
Shi Y, Hu Z, Xiong Z, Zhou Y, Jin X, Gu C, Hu X, Cheng G, Song N, Zhang W (2013) Analysis of molecular variation of porcine reproductive and respiratory syndrome virus in Central China from 2006 to 2012. Arch Virol 158:717–721
Su S, Bi Y, Wong G, Gray GC, Gao GF, Li S (2015) The epidemiology, evolution and recent outbreaks of avian influenza viruses in China: a review. J Virol
Tian ZJ, An TQ, Zhou YJ, Peng JM, Hu SP, Wei TC, Jiang YF, Xiao Y, Tong GZ (2009) An attenuated live vaccine based on highly pathogenic porcine reproductive and respiratory syndrome virus (HP-PRRSV) protects piglets against HP-PRRS. Vet Microbiol 138:34–40
Wang R, Xiao Y, Opriessnig T, Ding Y, Yu Y, Nan Y, Ma Z, Halbur PG, Zhang YJ (2013) Enhancing neutralizing antibody production by an interferon-inducing porcine reproductive and respiratory syndrome virus strain. Vaccine 31:5537–5543
Wei Y, Li S, Huang L, Tang Q, Liu J, Liu D, Wang Y, Wu H, Liu C (2013) Experimental infection and comparative genomic analysis of a highly pathogenic PRRSV-HBR strain at different passage levels. Vet Microbiol 166:337–346
Wensvoort G (1993) Lelystad virus and the porcine epidemic abortion and respiratory syndrome. Vet Res 24:117–124
Wieringa R, de Vries AA, Raamsman MJ, Rottier PJ (2002) Characterization of two new structural glycoproteins, GP(3) and GP(4), of equine arteritis virus. J Virol 76:10829–10840
Wootton S, Yoo D, Rogan D (2000) Full-length sequence of a Canadian porcine reproductive and respiratory syndrome virus (PRRSV) isolate. Arch Virol 145:2297–2323
Xie J, Zhu W, Chen Y, Wei C, Zhou P, Zhang M, Huang Z, Sun L, Su S, Zhang G (2013) Molecular epidemiology of PRRSV in South China from 2007 to 2011 based on the genetic analysis of ORF5. Microb Pathog 63:30–36
Xie J, Zhou H, Cui J, Chen Y, Zhang M, Deng S, Zhou P, Su S, Zhang G (2014) Inhibition of porcine reproductive and respiratory syndrome virus by specific siRNA targeting Nsp9 gene. Infect Genet Evol 28:64–70
Xue C, Wang W, Liu Q, Miao Z, Liu K, Shen H, Lv L, Li X, Chen X, Cao Y (2014) Chimeric influenza-virus-like particles containing the porcine reproductive and respiratory syndrome virus GP5 protein and the influenza virus HA and M1 proteins. Arch Virol 159:3043–3051
Yu X, Chen N, Deng X, Cao Z, Han W, Hu D, Wu J, Zhang S, Wang B, Gu X, Tian K (2013) Genomic sequencing reveals mutations potentially related to the overattenuation of a highly pathogenic porcine reproductive and respiratory syndrome virus. Clin Vaccine Immunol 20:613–619
Zhang M, Xie J, Sun L, Cao Z, Gu H, Deng S, Chen Y, Cao Z, Tang F, Su S, Zhang G (2013) Phylogenetic analysis and molecular characteristics of 17 porcine reproductive and respiratory syndrome virus isolates in Southern China from 2010 to 2011. Microb Pathog 65:67–72
Zhang X, Song Z, He J, Yen HL, Li J, Zhu Z, Tian D, Wang W, Xu L, Guan W, Liu Y, Wang S, Shi B, Zhang W, Qin B, Cai J, Wan Y, Xu C, Ren X, Chen H, Liu L, Yang Y, Zhou X, Zhou W, Xu J, Zhang X, Peiris M, Hu Y, Yuan Z (2014) Drug susceptibility profile and pathogenicity of H7N9 influenza virus (Anhui1 lineage) with R292K substitution. Emerg Microbes Infect 3:e78
Acknowledgments
This project was supported in part by the National Natural Science Foundation of China (grant no. 31272564), the Joint Fund of NSFC Guangdong of China (U0931003), and the Modern Agro-Industry Technology Research System (CARS-36).
Author information
Authors and Affiliations
Corresponding author
Additional information
Y. Chen, S. He and L. Sun contributed equally.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, Y., He, S., Sun, L. et al. Genetic variation, pathogenicity, and immunogenicity of highly pathogenic porcine reproductive and respiratory syndrome virus strain XH-GD at different passage levels. Arch Virol 161, 77–86 (2016). https://doi.org/10.1007/s00705-015-2597-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-015-2597-6