Abstract
Background
RNA editing is catalyzed by adenosine deaminases acting on RNA (ADARs). ADAR2 is the main enzyme responsible for recoding editing in humans. Adenosine-to-inosine (A-to-I) editing at the Q/R site is reported to be decreased in gliomas; however, the expression of ADAR2 mRNA was not greatly affected.
Methods
We determined ADAR2 mRNA expression in human glioblastoma cell lines and in normal human glial cells by real-time RT-PCR. We also determined ADAR2 mRNA expression in 44 glioma tissues and normal white matter. After identifying an alternative splicing variant (ASV) of ADAR2 in gliomas, we performed sequencing. We then classified glioblastomas based on the presence (+) or absence (–) of the ASV to determine the correlations between ASV + and malignant features of glioblastomas, such as invasion, peritumoral brain edema, and survival time.
Results
There were no significant differences in ADAR2 mRNA expression among human glioblastoma cell lines or in gliomas compared with normal white matter (all p > 0.05). The ASV, which contained a 47-nucleotide insertion in the ADAR2 mRNA transcript, was detected in the U251 and BT325 cell lines, and in some glioma tissues. The expression rate of ASV differed among gliomas of different grades. ASV + glioblastomas were more malignant than ASV – glioblastomas.
Conclusions
ADAR2 is a family of enzymes in which ASVs result in differences in enzymatic activity. The ADAR2 ASV may be correlated with the invasiveness of gliomas. Identification of the mechanistic characterization of ADAR2 ASV may have future potential for individualized molecular targeted-therapy for glioma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gliomas are the most common primary malignant brain tumor in humans. They can be typed as low-grade astrocytomas (LGA), oligodendrogliomas (OG), anaplastic astrocytomas (AA), and glioblastomas multiforme (GBM), among other subtypes. These tumors are often invasive, making surgical resection difficult. Despite the best treatment regimen currently available, which consists of surgery, radiotherapy, and systematic chemotherapy, the prognosis of patients with gliomas is poor. The 5-year survival of patients with GBM is less than 3 % [4, 32].
The widespread RNA–DNA variations found in human brain tissues indicate that RNA sequences are not identical to their corresponding DNA sequences [19]. Most RNA–DNA differences are caused by RNA editing, a process that generates many different mRNAs from the same gene after the post-transcriptional events [2, 3, 8, 23]. The potential impact of RNA editing on the etiology or progression of human diseases has now been realized. Deficient or hyperactive adenosine-to-inosine (A-to-I) RNA editing is associated with several human diseases including epilepsy, malignant brain cancer, amyotrophic lateral sclerosis, immunological disorders, and depression [13, 21, 31]. The analysis of RNA editing in various cancers, including brain, prostate, lung, kidney, bladder, colorectal, and testicular tumors, is expected to provide new diagnostic and prognostic markers [6, 7, 11, 12, 26, 28].
In mammals, RNA editing by site-selective adenosine deamination regulates the key functional properties of neurotransmitter receptors in the central nervous system [2]. Many RNA editing sites have now been reported [3, 8, 9, 21]. Almost all editing of the glutamate receptor (GluR) subunit B (GluR-B) occurs at one position, the Q/R-site, in a process that is essential for normal receptor function [4, 16, 34]. A general decrease in RNA editing activity has been observed in malignant gliomas in the brain. The Q/R site of GluR-B is frequently underedited in malignant gliomas compared with control tissues [5, 22, 32]. Alu sequences within MED13 transcripts are also underedited in brain tumors [26]. RNA editing is performed by adenosine deaminases acting on RNA (ADARs), of which ADAR2 is the main enzyme responsible for recoding editing in the brain [24, 31]. However, the ADAR2 mRNA expression levels in gliomas were inconsistent in three previous reports. Furthermore, there was no consistent correlation between editing efficiency and ADAR2 mRNA expression level in those studies [5, 22, 26, 32]. Therefore, the regulation of the RNA editing activity of ADAR2 in vivo is still largely unknown.
In this study, we determined ADAR2 mRNA expression levels in glioma-derived cell lines and in glioma tissues. We have presented evidence for an alternative splicing variant (ASV) of ADAR2 in gliomas. Furthermore, we explored the correlation between ADAR2 ASV expression and the invasiveness of GBM. We demonstrate that (1) ADAR2 mRNA levels were not altered in glioma-derived cell lines or in glioma tissues; (2) self-editing of ADAR2 pre-mRNA generates an ADAR2 ASV in glioma-derived cell lines and glioma tissues; and (3) the ADAR2 ASV may be correlated with the invasiveness of gliomas.
Materials and methods
Clinical specimens and cell lines
Glioma tissues and normal white matter tissue samples were obtained from the China–Japan Union Hospital and the First Hospital of Jilin University, Changchun, China between January 2007 and June 2008. Tumor tissue samples were taken from 44 patients with a primary glioma (age range at the time of diagnosis, 17–55 years). None of the patients previously received radiotherapy or chemotherapy. Tumor tissue samples were dissected from both the core and the edge of the tumors. Diagnosis was defined according to the guidelines of the World Health Organization, and was obtained by two glioma neuropathologists who performed independent immunohistochemical analyses of all of glioma samples. Tumor tissues included 12 cases of GBM, eight cases of AA, 10 cases of LGA, and six cases of OG. As normal controls, we used six non-tumoral white matter samples obtained from six anonymized patients (age range at the time of diagnosis, 21–54 years) undergoing focal brain resection during acute internal decompression operation for severe traumatic brain injury such as brain contusions [5, 32]. As normal controls, we choose white matter because tumor cells of gliomas originate from glial cells which are particularly abundant in white matter [17]. We obtained IRB approval prior to obtaining the normal white matter samples from the six patients [15]. Tumor and control tissues were frozen immediately after their removal at surgery and were kept at −70 °C until further use. The number of samples tested in each assay differed because of the limited amount of RNA that was available for each sample. The study obtained IRB approval regarding the use of human samples for experimental studies from the ethics committees of China–Japan Union Hospital and the First Hospital of Jilin University, China.
An astrocytoma cell line (SHG44) and two human GBM cell lines (U251 and BT325) were obtained from the Cell Center of the Chinese Academic Medical College (Beijing, China). Glioma cells were routinely cultured in Dulbecco’s modified Eagle’s medium (GIBCO®; Life Technologies, Inc., Gaithersburg, MD, USA) at 37 °C under 5 % CO2. Normal human astrocytes (NHA) (#CC-2565, Lot 80982; Lonza, Walkersville, MD, USA) were grown in a humidified incubator at 37 °C under 5 % CO2.
RNA extraction and real-time RT-PCR
Total RNA was extracted from the clinical samples and cultured cells using Trizol (Invitrogen, Life Technologies, Carlsbad, CA, USA) according to the manufacturer’s instructions. Quantitative real-time PCR of ADAR2 cDNA was performed using an Mx3000P real-time PCR instrument (Agilent, Santa Clara, CA, USA) with SYBR green I fluorescence, as follows. The reaction mixture consisted of 12.5 μL of 2× SYBR, 0.5 μL each of the forward (5′-GTATTTTGCCATGGA TATAGAAGATG-3′; sense, exon 1, 10 μmol L−1) and reverse primers (5′-GTACTGGGATCCAGGCTTGATCTCATTCAGCTG-3′; antisense, exon 2, 10 μmol L−1), and 2.0 μL of cDNA 2.0 μL, in a final reaction volume of 25 μL with DNase- and RNase-free water. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH; assay ID Hs9999905_m1; Applied Biosystems, Foster City, CA, USA) was used as a reference gene. The PCR conditions consisted of 10 min at 95 °C, followed by 45 cycles of 95 °C for 20 s, 60 °C for 30 s, and 72 °C for 45 s. Real-time PCR was performed using serial dilutions of each cDNA template (1×, 1:4, 1:16, 1:64, and 1:256). The cycle threshold (Ct value) was defined as the number of cycles required to reach the fluorescence threshold. As a representative example, an amplification plot was generated for ADAR2 in which fluorescence intensity was plotted against cycle number. The comparative Ct method was used to quantify transcript number, and the normalized Ct value (2–ΔΔCt) was used to estimate the level of ADAR2 gene expression. The odds ratio and 95 % confidence interval were calculated automatically by UNPHASED software (MRC Biostatistics Unit, Cambridge, UK). Real-time assays were performed in triplicate from two independent RT reactions.
RT-PCR
cDNA was synthesized using SuperScript II reverse transcriptase (Invitrogen) and random hexamer primers. To analyze human ADAR2 alternative splicing, we performed PCR reactions using forward (5′-GTATTTTGCCATGGATATAGAAGATG-3′; sense, exon 1) and reverse primers (5′-GTACTGGGATCCAGGCTTGATCTCATTCAGCTG-3′; antisense, exon 2), which amplified both splice variants from cDNA.
Calculation of peritumoral brain edema (PTBE)
PTBE contributes to the morbidity and mortality of brain tumors. The development of PTBE is influenced by many factors. PTBE in brain tumors might be a prognostic indicator of patient survival [25]. The extent of PTBE and tumor size were measured by magnetic resonance imaging (MRI) [25]. The tumor volume was measured on enhanced T1 weighted images and the volume of edema surrounding the tumors was measured on T2 weighted images [33]. The edema/tumor volume ratio was defined as the edema index (EI), and was used as an indicator of PTBE [30]. EI was calculated as PTBE area/tumor area. Thus, if EI = 1, there is no PTBE [14]. An EI of ≤3 was defined as weak PTBE (+), an EI of >3 to <7 was defined as moderate PTBE (++), and an EI of ≥7 was defined as strong PTBE (+++) [33].
Statistical analysis
SPSS for Windows (version 14.0; SPSS, Inc., Chicago, IL, USA) was used to perform the Mann–Whitney U test to examine the difference in ADAR2 expression between patients and controls. Survival rates were estimated by the Kaplan–Meier method and overall survival was compared between patients with (+) and without (–) an ADAR2 ASV using the log-rank test. Statistical significance (p < 0.05) was determined by descriptive statistics.
Results
ADAR2 expression in human glioblastoma cell lines
Quantitative real-time RT-PCR revealed no marked differences in ADAR2 mRNA expression between NHA, SHG44, U251, and BT325 cells (Fig. 1).
Relative ADAR2 expression levels in human glioma cell lines. Real-time RT-PCR for ADAR2 was performed using a human astrocytoma cell line (SHG44), human glioblastoma cell lines (U251 and BT325), and normal human astrocytes (NHA) as a control. Error bars indicate the standard deviation of triplicate experiments
ADAR2 mRNA expression in human gliomas
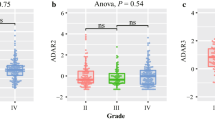
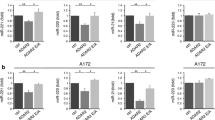
We performed semi-quantitative RT-PCR using 44 glioma samples, and performed quantitative real-time RT-PCR using six normal white matter tissue samples and 36 glioma samples (12 GBM, eight AA, 10 LGA, and six OG). We found that ADAR2 mRNA expression was not significantly different between the gliomas and normal white matter tissues (all p > 0.05; Fig. 2a).
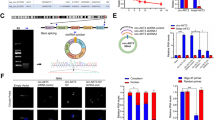
Expression of ADAR2 mRNA in human glioblastomas and analysis of ADAR2 mRNA alternatively splicing variants (ASV). a Expression of ADAR2 mRNA in human glioblastomas. Real-time PCR was performed to quantify the mRNA expression levels of ADAR2 in normal tissues and in different grades and types of brain tumor. LGA, low-grade astrocytoma; AA, anaplastic astrocytoma; GBM, glioblastoma multiforme; OG, oligodendroglioma. Quantitative real-time PCR showed that the expression of ADAR2 mRNA was not significantly different among the tumor tissues. b Analysis of ADAR2 ASVs in glioma-derived cell lines and glioblastomas by RT-PCR. The migration position of the PCR reaction products corresponding to the transcripts with and without the 47-nucleotide insert are indicated. c Analysis of ADAR2 ASVs in glioma tissues. The ADAR2 ASV was detected in 1/10 LGAs, 1/6 OGs, 1/8 AAs, and 3/12 GBMs, corresponding to rate of self-editing-induced ASVs of 10 %, 16.7 %, 12.5 %, and 25 %, respectively
Expression of ADAR2 ASVs in glioma cell lines
RT-PCR amplification of total RNA isolated from the NHA and SHG44 cells presented a single electrophoretic band. RT-PCR amplification of total RNA isolated from the glioma-derived cell line U251 and BT325 revealed two distinct electrophoretic bands (Fig. 2b). Sequence analysis of both isolated ADAR2 cDNAs indicated that these two isoforms were produced by alternative splicing. The longer isoforms contained a 47-nucleotide sequence (GATCCTGCAACGAAGGCGTTGTAAGTTACTCTTTCTGGGCACCACAG) inserted after the first 28 nucleotides of the coding region. This ASV is identical to the one that was previously identified in a human ADAR2 transcript containing a 47-nucleotide insertion at the analogous position [18, 20, 27].
Expression of ADAR2 ASVs in glioma tissues
In glioma samples, we detected ADAR2 ASV mRNA which was identical to that found in the U251 and BT325 cell lines. The ASV was not detected in any of the normal human brain tissue samples. The ADAR2 ASV was expressed in 1/10 LGAs, 1/6 OGs, 1/8 AAs, and 3/12 GBMs, corresponding to rates of self-editing induced ASVs of 10 %, 16.7 %, 12.5 %, and 25 %, respectively. The expression of the ASV was highest in GBMs, the most aggressive type of gliomas. The level of ASV expression was correlated with tumor grade (Fig. 2c).
ADAR2 ASV expression is correlated with the extent of PTBE in patients with GBM
Patients with GBM were classified into two groups according to the presence (+; n = 3) or absence (–; n = 9) of the ADAR2 ASV. For the three ADAR2 ASV + patients, weak, moderate, and strong EI scores were observed in 0 (0 %), 0 (0 %), and three (100 %) patients, respectively. By contrast, the EI scores were classified as weak, moderate, and strong in seven (78 %), two (22 %), and 0 (0 %) in the ADAR2 ASV – patients, respectively (Table 1). The EI scores were significantly greater in the ADAR2 ASV + patients than in the ADAR2 ASV – patients (p < 0.01).
ADAR2 ASV expression and the invasiveness of GBM
To analyze the association between ADAR2 ASV expression and invasiveness of GBM, we used MRI to examine the extent of tumor invasion in the GBM patients. For the three ADAR2 ASV + patients, the tumor had invaded two lobes in one patient and three lobes in two patients. By contrast, among the nine ADAR2 ASV – patients, the tumor had invaded one lobe in four patients, two lobes in four patients, and three lobes in one patient. Therefore, the tumors in the ADAR2 ASV + patients were more aggressive than those in the ADAR2 ASV – patients (p < 0.01; Fig. 3a).
Magnetic resonance imaging (MRI) of glioblastomas and overall survival analysis of patients with GBM. a MRI of glioblastomas. A Patient with ADAR2 ASV– GBM. The lesion has invaded the temporal and frontal lobes. B–D Patients with ADAR2 ASV + GBM. B the lesion has invaded the temporal and frontal lobes. C The lesion has invaded the temporal, parietal, and occipital lobes. D The lesion has invaded the temporal, frontal, and parietal lobes. A1 axial T1-weighted image; A2, D2 sagittal T1-weighted images; A3 coronal contrast-enhanced T1-weighted image; B1, C1, D1 axial T2-weighted images; B2, C2 sagittal contrast-enhanced T1-weighted image; B3, C3, D3 coronal contrast-enhanced T1-weighted images. b Overall survival analysis of patients with GBM. The median survival time was significantly shorter in ADAR2 ASV + patients compared with ADAR2 ASV – patients (log-rank test, p < 0.0001)
Overall survival in patients with GBM
The median survival time was 4.53 months for the three ADAR2 ASV + patients compared with 9.98 months for the ADAR2 ASV – patients. The difference in median survival time between the two groups of patients was statistically significant (p < 0.0001; Fig. 3b).
Discussion
RNA editing is a common phenomenon in eukaryotic cells that leads to post-transcriptional base changes in mRNA. In mammals, a growing number of genes have been identified that undergo a type of RNA editing characterized by site-selective A-to-I modification [2, 3, 8]. The best-studied A-to-I editing substrates are the brain-specific transcripts coding the GluR. For GluR-B, the Q/R site controls the Ca2+ permeability of the ion channel [16, 34]. ADAR2 is the main enzyme responsible for recoding editing in the brain. It is predominantly expressed in neurons, and is weakly expressed in gliocytes [17]. Mammalian RNA editing catalyzed by ADAR1 and ADAR2 plays pivotal roles in the brain by controlling functional modifications of neurotransmitter receptors and ion channels [31]. The ADAR proteins and RNA editing targets have been studied in many human diseases associated with RNA editing, including various cancers [6, 7, 11, 12, 26, 28].
Gliomas are the most common malignant tumors in the central nervous system, and are almost always fatal. The molecular basis for malignant progression in gliomas involves several collaborative processes. Thus far, several studies have shown that a general state of underediting exists in gliomas in some RNA editing sites, including the Q/R site, the 5HT2C serotonin receptor site, and the Glu/Arg site. For regulated normal cells, these findings suggest a key role of ADAR2 editing activity in controlling the growth of gliomas [5, 22, 26, 32] because this enzyme seems to prevent cell proliferation by the Akt pathway [16, 34] and/or modulates the cell cycle [10]. It was also reported that the Q/R site of GluR-B in gliomas is underedited compared with that in normal tissue. Underediting at this position might be due to decreased ADAR2 expression or downregulation of ADAR2 enzymatic activity.
The recent findings regarding ADAR2 mRNA expression in gliomas were somewhat inconsistent [5, 22, 26]. The first report, by Maas et al., showed that the ADAR2 mRNA expression was not significantly altered in one OG and seven GBM samples, although the Q/R site of GluR-B was underedited compared with that in control tissues [22]. Paz et al. observed a prominent reduction in ADAR2 mRNA levels in four types of brain tumors (GBM, LGA, AA, and OG) in an analysis of 18 brain tumors; however, their control was normal brain tissue containing neurons, which show high ADAR2 expression [26]. Cenci et al. analyzed 14 astrocytoma tissue samples from 10 children, and found no significant differences in ADAR2 mRNA levels between tumor tissue and normal white matter [5].
In our study, we examined the expression of ADAR2 mRNA in human glioblastoma cell lines and in NHA, and found no differences in its expression among these cell types. We also determined the expression of ADAR2 mRNA in various gliomas of different grades, and found that it was not significantly different between gliomas and normal white matter. Based on the results of previous studies and our study, we consider that the expression of ADAR2 mRNA is not significantly altered in gliomas. Thus, we speculate that underediting at the Q/R site in gliomas is not simply due to decreased ADAR2 expression.
The human ADAR2 gene comprises 14 exons [23, 29], and several ASVs have been identified [1, 18, 20, 23, 27, 29]. For example, one involving self-editing of ADAR2 pre-mRNA creates a 3′-prime splice site within intron 1 leading to the insertion of a 47-nucleotide sequence into the intronic sequence. Another ASV includes an exon located 18 kb upstream of the previously annotated first coding exon, and extends the open reading frame of ADAR2 by 49 amino acids [20]. In mammals, alternative splicing occurs within the catalytic domain, the RNA-binding domain, and at the carboxy terminus. Of these ASVs, some have no effect on editing activity whereas others decrease the editing activity of ADAR2. Therefore, the characterization of ADAR2 transcription and alternative splicing is essential to understand the regulation of RNA editing.
The regulation of RNA editing activity in vivo remains unknown: the amount of total ADAR2 mRNA is one factor regulating RNA editing, but the level of total ADAR2 does not necessarily represent the editing activity, and alternative splicing may be another factor affecting RNA editing. Alternative splicing events have been demonstrated to affect the catalytic activity of ADAR2. Rueter et al. [27] demonstrated experimentally that the addition of 47 nucleotides to the 5’ end of exon2 occurs due to RNA editing within intron 1 and decreases ADAR2 activity in vivo because of the inefficient initiation of translation from Met25. The results of Rueter et al. are cited by several subsequent studies [18, 20]. In the present study, we identified an ASV containing a 47-nucleotide insertion by RT-PCR and sequencing. This ASV is identical to that identified by Rueter et al., which has lower RNA editing activity. We also detected that the expression rate of ASV is higher in glioma than in normal white matter. On this basis, we speculate that the expression of a less-active ASV of ADAR2 may correlate with downregulation of A-to-I editing in gliomas.
In our study, we found an ADAR2 ASV in two glioma-derived cell lines, U251 and BT325, and in human glioma tissues. The ASV was detected in 10 %, 16.7 %, 12.5 %, and 25 % of LGA, OG, AA, and GBM tissues, respectively. The mRNA sequence of the ADAR2 ASV was identical to that originally reported by Rueter et al. [27] and by many other authors since then. The splicing variant is generated via a proximal 3′ acceptor site, at which 47 nucleotides are added to the ADAR2 coding region, changing the reading frame of the mature ADAR2 transcript. Insertion of a 47-nucleotide cassette into the coding region shifts the translation start to an inefficient downstream methionine. In vitro and tissue culture model systems have indicated that the ASV downregulates the catalytic activity of the human ADAR2 protein isoforms without affecting the substrate. This finding is consistent with the assumption that downregulation of A-to-I editing in gliomas may be due to the expression of a less-active ASV of ADAR2.
We found that the positive rate of the self-editing-induced ASV increased significantly with increasing malignancy of the gliomas, being greatest in GBMs, the most malignant type of glioma. To determine whether ADAR2 ASV expression was correlated with the malignant features of gliomas, we divided the patients with GBM into two groups according to expression of the ASV, and compared PTBE, invasiveness, and median survival time between the two groups. These analyses revealed that ADAR2 ASV + patients had more severe PTBE, tumor invasion into more brain lobes, and had a shorter median survival time compared with ADAR2 ASV – patients.
Conclusions
Our data indicate that (1) ADAR2 mRNA levels were not altered in glioma-derived cell lines or in glioma tissues; (2) self-editing of ADAR2 pre-mRNA generates an ADAR2 ASV in glioma-derived cell lines and glioma tissues; and (3) expression of ADAR2 ASV may be correlated to the malignancy of gliomas. Identification of the mechanistic characterization of ADAR2 ASV may be helpful in the future development of individualized molecular-targeted therapy for glioma.
Abbreviations
- A-to-I:
-
adenosine-to-inosine
- AA:
-
anaplastic astrocytomas
- ADAR:
-
adenosine deaminases acting on RNA
- ASV:
-
alternative splicing variant
- Ct:
-
cycle threshold
- EI:
-
edema index
- GAPDH:
-
glyceraldehyde 3-phosphate dehydrogenase
- GBM:
-
glioblastomas multiforme
- GluR-B:
-
glutamate receptor subunit B
- LGA:
-
low-grade astrocytomas
- MRI:
-
magnetic resonance imaging
- NHA:
-
normal human astrocytes
- OG:
-
oligodendrogliomas
- PTBE:
-
peritumoral brain edema
- RT-PCR:
-
reverse-transcription–polymerase chain reaction
References
Agranat L, Sperling J, Sperling R (2010) A novel tissue-specific alternatively spliced form of the A-to-I RNA editing enzyme ADAR2. RNA Biol 7:253–262
Athanasiadis A, Rich A, Maas S (2004) Widespread A-to-I RNA editing of Alu-containing mRNAs in the human transcriptome. PLoS Biol 2:e391
Bahn JH, Lee JH, Li G, Greer C, Peng GD, Xiao XS (2012) Accurate identification of A-to-I RNA editing in human by transcriptome sequencing. Genome Research 22:142–150
Buonerba C, Di Lorenzo G, Marinelli A, Federico P, Palmieri G, Imbimbo M, Conti P, Peluso G, De Placido S, Sampson JH (2010) A comprehensive outlook on intra- cerebral therapy of malignant gliomas. Crit Rev Oncol Hematol 80:54–68
Cenci C, Barzotti R, Galeano F, Corbelli S, Rota R, Massimi L, Di Rocco C, O’Connell MA, Gallo A (2008) Downregulation of RNA editing in pediatric astrocytomas: ADAR2 editing activity inhibits cell migration and proliferation. J Biol Chem 283:7251–7260
Chin L, Gray JW (2008) Translating insights from the cancer genome into clinical practice. Nature 452:553–563
Dominissini D, Moshitch-Moshkovitz S, Amariglio N, Rechavi G (2011) Adenosine-to-inosine RNA editing meets cancer. Carcinogenesis 32:1569–1577
Eisenberg E, Li JB, Levanon EY (2010) Sequence based identification of RNA editing sites. RNA Biol 7:248–252
Galeano F, Leroy A, Rossetti C, Gromova I, Gautier P, Keegan LP, Massimi L, Di Rocco C, O’Connell MA, Gallo A (2010) Human BLCAP transcript: new editing events in normal and cancerous tissues. Int J Cancer 127:127–137
Galeano F, Rossetti C, Tomaselli S, Cifaldi L, Lezzerini M, Pezzullo M, Boldrini R, Massimi L, Rocco CD, Locatelli F, Gallo A (2013) ADAR2-editing activity inhibits glioblastoma growth through the modulation of the CDC14B/Skp2/p21/p27 axis. Oncogene 32:998–1009
Galeano F, Tomaselli S, Locatelli F, Gallo A (2012) A-to-I RNA editing: The “ADAR” side of human cancer. Semin Cell Dev Biol 23:244–250
Gallo A, Galardi S (2008) A-to-I RNA editing and cancer: from pathology to basic science. RNA Biol 5:135–139
Gallo A, Locatelli F (2012) ADARs: allies or enemies? The importance of A-to-I RNA editing in human disease: from cancer to HIV-1. Biol Rev Camb Philos Soc 87:95–110
Gurkanlar D, Er U, Sanli M, Ozkan M, Sekerci Z (2005) Peritumoral brain edema in intracranial meningiomas. J Clin Neurosci 12:750–753
Hoelzinger DB, Mariani L, Weis J, Woyke T, Berens TJ, McDonough WS, Sloan A, Coons SW, Berens ME (2005) Gene expression profile of glioblastoma multiforme invasive phenotype points to new therapeutic targets. Neoplasia 7:7–16
Ishiuchi S, Yoshida Y, Sugawara K, Aihara M, Ohtani T, Watanabe T, Saito N, Tsuzuki K, Okado H, Miwa A, Nakazato Y, Ozawa S (2007) Ca 2+ permeable AMPA receptors regulate growth of human glioblastoma via Akt activation. J Neurosci 27:7987–8001
Jacobs MM, Fogg RL, Emeson RB, Stanwood GD (2009) ADAR1 and ADAR2 expression and editing activity during forebrain development. Dev Neurosci 31:223–237
Kawahara Y, Ito K, Ito M, Tsuji S, Kwak S (2005) Novel splice variants of human ADAR2 mRNA: Skipping of the exon encoding the dsRNA-binding domains, and multiple C-terminal splice sites. Gene 22:193–201
Li M, Wang IX, Li Y, Bruzel A, Richards AL, Toung JM, Cheung VG (2011) Widespread RNA and DNA Sequence Differences in the Human Transcriptome. Science 333:53–58
Maas S, Gommans WM (2009) Novel exon of mammalian ADAR2 extends open reading frame. PLoS One 4:e4225
Maas S, Kawahara Y, Tamburro KM, Nishikura K (2006) A-to-I RNA editing and human disease. RNA Biol 3:1–9
Maas S, Patt S, Schrey M, Rich A (2001) Underediting of glutamate receptor GluR-B mRNA in malignant gliomas. Proc Natl Acad Sci USA 98:14687–14692
Mittaz L, Scott HS, Rossier C, Seeburg PH, Higuchi M, Antonarakis SE (1997) Cloning of a human RNA editing deaminase (ADARB1) of glutamate receptors that maps to chromosome 21q22.3. Genomics 41:210–217
Nishikura K (2010) Functions and regulation of RNA editing by ADAR2 deaminases. Annu Rev Biochem 79:321–349
Park MW, Kim CH, Cheong JH, Bak KH, Kim JM, Oh SJ (2006) Occludin expression in brain tumors and its relevance to peritumoral edema and survival. Cancer Res Treat 38:139–143
Paz N, Levanon EY, Amariglio N, Heimberger AB, Ram Z, Constantini S, Barbash ZS, Adamsky K, Safran M, Hirschberg A, Krupsky M, Ben-Dov I, Cazacu S, Mikkelsen T, Brodie C, Eisenberg E, Rechavi G (2007) Altered adenosine-to-inosine RNA editing in human cancer. Genome Res 17:1586–1595
Rueter SM, Dawson TR, Emeson RB (1999) Regulation of alternative splicing by RNA editing. Nature 399:75–80
Skarda J, Amariglio N, Rechavi G (2009) RNA editing in human cancer: review. APMIS 117:551–557
Slavov D, Gardiner K (2002) Phylogenetic comparison of the pre-mRNA adenosine deaminase ADAR2 genes and transcripts: conservation and diversity in editing site sequence and alternative splicing patterns. Gene 299:83–94
Tamiya T, Ono Y, Matsumoto K, Ohmoto T (2001) Peritumoral brain edema in intracranial meningiomas: effects of radiological and histological factors. Neurosurgery 49:1046–1051
Tariq A, Jantsch MF (2012) Transcript diversification in the nervous system: A to I RNA editing in CNS function and disease development. Front Neurosci 6:99
Tomaselli S, Galeano F, Massimi L, Rocco CD, Lauriola L, Mastronuzzi A, Locatelli F, Gallo A (2013) ADAR2 editing activity in newly diagnosed versus relapsed pediatric high-grade astrocytomas. BMC Cancer 13:255
Wang P, Ni RY, Chen MN, Mou KJ, Mao Q, Liu YH (2011) Expression of aquaporin-4 in human supratentorial meningiomas with peritumoral brain edema and correlation of VEGF with edema formation. Genet Mol Res 10:2165–2171
Wright A, Vissel B (2012) The essential role of AMPA receptor GluA2 subunit RNA editing in the normal and diseased brain. Front Mol Neurosci 5:34
Acknowledgments
This work was supported by grants from the National Science Foundation of China (no. 30672159), the New Century Excellent Talents in the Universities of China program (no. 060306), and the Graduate Innovation Fund of Jilin University (no. 20121124).
Conflicts of Interest
None.
Author information
Authors and Affiliations
Corresponding author
Additional information
Jun Wei and Zhaohui Li are the first co-authors
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Wei, J., Li, Z., Du, C. et al. Abnormal expression of an ADAR2 alternative splicing variant in gliomas downregulates adenosine-to-inosine RNA editing. Acta Neurochir 156, 1135–1142 (2014). https://doi.org/10.1007/s00701-014-2004-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00701-014-2004-1