Abstract
Main conclusion
Genome-wide identification, classification, expression analyses, and functional characterization of GRAS genes in oil crop, Brassica napus, indicate their importance in root development and stress response.
Abstract
GRAS proteins are a plant-specific transcription factor gene family involved in tissues development and stress response. We classified 87 putative GRAS genes in the Brassica napus genome (BnGRASs) into 13 subfamilies by phylogenetic analysis. The C-terminal GRAS domains of Brassica napus (B. napus) proteins were less conserved among subfamilies, but were conserved within each subfamily. A series of analyses revealed that 89.7% of the BnGRASs did not have intron insertions, and 24 specific-motifs were found at the N-terminal. A highly conserved microRNA 171 (miRNA171) target was observed specifically in the HAM subfamily across land plants. A total of 868 pairs of interaction proteins were predicted, the primary of which were transcription factors involved in transcriptional regulation and signal transduction. Integrated comparative analysis of GRAS genes across 26 species of algae, mosses, ferns, gymnosperms, and angiosperms revealed that this gene family originated in early mosses and was classified into 19 subfamilies, 14 of which may have originated prior to bryophyte evolution. RNA-Seq analysis demonstrated that most BnGRASs were widely expressed in different tissues/organs at different stages in B. napus, and 24 BnGRASs were highly/specifically expressed in roots. Results from a qRT-PCR analysis suggested that two BnGRASs belonging to SCR and LISCL subfamilies potentially have important roles in the stress response of roots.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
GRAS proteins are an important family of transcription factors (TF) in plants (Silverstone et al. 1998). Typically, GRAS proteins include 400–700 amino acids, with a variable N-terminal and highly conserved C-terminal containing several motifs with conserved sequence characteristics and order, such as leucine heptad repeat I (LHRI), VHIID, leucine heptad repeat II (LHRII), PFYRE, and SAW (Lee et al. 2008). The N-terminal domains contain numerous molecular recognition features (MoRFs) representing potential protein–protein binding sites that are potentially involved in molecular recognition during plant development (Sun et al. 2011).
GRAS transcription factor functions vary and include microspore and root development (Morohashi et al. 2003), signal transduction (Bolle et al. 2000), stem cell maintenance (Stuurman et al. 2002; Li et al. 2003), and stress response (Fode et al. 2008; Ma et al. 2010). Genome-wide analyses in a range of plants, such as Chinese cabbage (Brassica rapa), tomato (Solanum lycopersicum), and woad (Isatis indigotica) (Song et al. 2014; Huang et al. 2015; Zhang et al. 2016), have generally divided the GRAS gene family into eight subfamilies: DELLA, SCARECROW (SCR), LATERAL SUPPRESSOR (LS), HAIRY MERISTEM (HAM), phytochrome A signal transduction 1 (PAT1), SCL4/7, SHORT-ROOT (SHR), and SCARECROW-LIKE9 (SCL9) (Lee et al. 2008; Guo et al. 2017). However, 13 subfamilies have been identified in Populus trichocarpa (Liu and Widmer 2014) and 16 in Medicago truncatula (Song et al. 2017). Recently, the family was newly classed into 17 subfamilies in angiosperms (Cenci and Rouard 2017). Thus, the classification of this gene family is not yet uniform.
Brassica plants are widely used as vegetables, oilseed, and fodder, and B. napus is one of the most economically important oil crops in China, Europe, and North America. Recently, the genome of B. napus was sequenced (Chalhoub et al. 2014; Sun et al. 2017), but the GRAS gene family has yet to be reported. Investigating this family in B. napus would provide insights into the evolutionary mechanisms of hybridisation (allopolyploid) between B. rapa and Brassica oleracea and a theoretical basis for future studies on the roles of GRAS proteins in B. napus.
In this study, we identified 87 GRAS members in the B. napus genome (BnGRASs) mapped to the 19 B. napus chromosomes. According to the results of the phylogenetic analysis and the gene functions, the angiosperm GRAS gene family is divided into 17 subfamilies and the BnGRASs were found to be distributed in 13 of these. Allopolyploidization was the main reason for the rapid expansion of this gene family in B. napus. The expression profiles of candidate BnGRASs in 50 samples revealed that they were widely expressed in different tissues at different stages in B. napus. Their expression patterns within the same subfamily were found to be generally similar, suggesting functional similarity. Meanwhile, up to 24 BnGRASs were highly or specifically expressed in roots, suggesting important roles for GRAS genes in root development. qRT-PCR analysis of nine screened BnGRASs from LS, LISCL, SCL4/7, and SCR subfamilies under PEG and salt stresses showed that some genes were up- or down-regulated by abiotic stresses. Among which, BnGRAS86 and BnGRAS25 are potential candidates for breeding stress resistant B. napus. The results of our study provide key insights into the evolution, classification, and function of GRAS proteins in B. napus.
Materials and methods
Sequence retrieval
The sequences of 34 Arabidopsis thaliana GRAS proteins (AtGRASs) were retrieved from the TAIR (http://www.arabidopsis.org/). To identify the putative GRAS genes in the B. napus genome, we performed BLASTP searches in the GENOSCOPE database (Darmor–bzh, http://www.genoscope.cns.fr/brassicanapus/) (Chalhoub et al. 2014) using all of the A. thaliana, Oryza sativa, and Vitis vinifera GRAS protein sequences as queries (Cenci and Rouard 2017). We further manually checked the sequence information of candidate genes according to that of the GRAS genes in another sequenced B. napus genome in NCBI database (ZS11, https://www.ncbi.nlm.nih.gov/genome/annotation_euk/Brassica_napus/101/) (Sun et al. 2017). To verify the reliability of our results, all putative non-redundant sequences were assessed with PROSITE profiling to confirm the candidates possessing the typical GRAS domain (http://prosite.expasy.org/). The information of cDNA sequences and genomic sequences was acquired from the B. napus genome, as well. Similarly, the candidate GRAS genes in B. oleracea (v1.0) were identified by the same method in the BRAD database (http://brassicadb.org/brad/), and the candidates in B. rapa and other species were identified in the Phytozome v12.1 (https://phytozome.jgi.doe.gov/pz/portal.html).
Multiple sequence alignment, detection of orthologous groups, and phylogenetic analysis
Multiple sequence alignment of GRAS domains of candidate proteins was performed using the online MAFFT version 7 software under default parameters (https://mafft.cbrc.jp/alignment/server/) (Katoh and Standley 2013) and then edited by MEGA version 7.0 software (Kumar et al. 2016). Protein sequence characteristics were viewed by WebLogo (http://weblogo.berkeley.edu/logo.cgi).
The manual inference of orthologous groups (OGs) was to confirm the protein belonging to the OG by mutual BLASTp, as described in Cenci and Rouard (2017). To investigate the evolutionary history of the B. napus GRAS gene family, we constructed a neighbour joining (NJ) tree based on the multiple sequence alignment of the GRAS domains using MEGA 7.0. To determine the statistical reliability, we applied bootstrap analysis with 1000 replicates and with the following main parameters: p distance and pairwise deletion. The JJT amino acid substitution model with estimation of the gamma distribution shape parameter (JJT+G) was suggested to be the best evolutionary model, based on the Akaike information criterion (AIC) in MEGA7.0. The maximum-likelihood (ML) trees were constructed using PhyML 3.0 (Guindon et al. 2009) with 100 replicates and the JJT+G model. The tree files were viewed and edited using FigTree v1.3.1 (http://tree.bio.ed.ac.uk/software/figtree/).
Detection of the physical and chemical characteristics and conserved motifs
The physical and chemical characteristics of the candidate GRAS proteins were predicted using the Protparam tool (http://web.expasy.org/) (Bjellqvist et al. 1993). Subcellular localisation prediction was applied by Plant-mPLoc (http://www.csbio.sjtu.edu.cn/bioinf/plantmulti/) (Chou and Shen 2008).
Conserved motifs of GRAS proteins were identified with the MEME 4.11.1 program with default parameters, except for the maximum number of motifs (5) and the maximum width (100). Only motifs with an e value < 1e-20 were kept for further analysis.
Analyses of intron patterns, chromosomal location, miRNA, and the interaction network
The intron insertion sites of GARS genes in B. napus, B. rapa, and B. oleracea were manually analyzed by comparing the DNA and CDS sequences of each gene using MEGA 7.0. The intron insertion information of GRAS genes in A. thaliana was acquired from Phytozome v12.1 (https://phytozome.jgi.doe.gov/pz/portal.html). The intron patterns of B. napus GARS genes were visualised by the Gene Structure Display Server (GSDS) 2.0 (http://gsds.cbi.pku.edu.cn/) (Hu et al. 2015). MapChart software was used to draw the chromosome map of candidate genes. Collinearity analysis of GRAS genes in the A. thaliana, B. oleracea, B. rapa, and B. napus genomes was obtained from the Brassica Database (BRAD), and the R package (Krzywinski et al. 2009) was used to view collinearity. Gene replication event was analyzed by CoGe online software (https://genomevolution.org/CoGe/). Nucleotide substitution rate (Ka/Ks) of duplicate genes was calculated by KaKs_calculator 2.0 using the LWL method (Wang et al. 2010a). Plant small RNA-targeted gene prediction was performed by psRNATarget (http://plantgrn.noble.org/psRNATarget/analysis/). Protein interaction network prediction was performed by STRING (https://string-db.org/cgi/input.pl?sessionId) (Szklarczyk et al. 2017) and visualised with Cytoscape version 3.4.0. The gene ontology (GO) annotation information of interaction proteins was obtained by GO database (http://www.geneontology.org/) (Ashburner et al. 2000).
The expression analyses of GRAS genes in B. rapa, B. oleracea, and B. napus
The RNA-Seq of B. napus variety “Zhongshuang 11” (ZS11) in distinct tissues at different developmental stages (BioProject ID PRJNA358784) and under five hormones induction (IAA, auxin; GA3, gibberellin; 6-BA, cytokinin; ABA, abscisic acid and ACC, ethylene) (unpublished data) was used to analyze the expression profiles of candidate BnGRASs. The data were analyzed by Cluster 3.0 and a heatmap was drawn by Java Treeview software. The RNA-seq data of B. rapa and B. oleracea were obtained from BRAD (http://brassicadb.org/brad/genomeDominanceData.php) and GEO Data Sets (Accession number: GSE42891), and then were used to analyze the expression profiles of candidate BoGRASs and BrGRASs, respectively.
For qRT-PCR analyses, seeds of ZS11 were obtained from the College of Agriculture and Biotechnology and grown in an artificial climatic chamber at 25 °C with a 14:10 photoperiod (day:night). 5-leaf seedlings of ZS11 were used for different treatments. Seedlings were treated in Hoagland liquid medium containing 200 mM NaCl, 15% PEG6000 (to mimic drought stress) and 50 μM ABA for 0, 1, 3, 6, 12, and 24 h, respectively, and then, the root tissues were collected. All materials were frozen in liquid nitrogen immediately and stored at −80 °C for RNA isolation.
Total RNA was extracted using EASYspin total RNA Extraction kit (Biomed, Beijing, China); the quality and concentration of each RNA sample was determined using gel electrophoresis and a NanoDrop 2000 spectrophotometer. Only the RNA samples that met the criteria A260/280 ratio = 1.8–2.1 and A260/230 ratio ≥ 2.0 were used for further analyses. The total RNA sample was treated with DNase I (Promega) before use. First-strand cDNA synthesis was performed using an oligo (dT) primer and 2 μg of total RNA in a 20 μl reaction volume, according to the manufacturer’s instructions for the M-MuLV RT kit (Takara, Dalian, China). The fluorescence was measured after the extension step using the CFX Connect™ Real-Time System (Bio-Rad, Chongqing, China) and the SYBR-Green PrimeScript RT-PCR Kit (Takara). The B. napus Actin7 (BnActin7) (GenBank Accession no. AF024716) was used as the reference gene. The primers used in this analysis are listed in Table S1. Three biological replicates were included for each treatment, and each consisted of three technical replicates. Expression levels were calculated as the mean signal intensity across the three replicates, and the relative expression levels were determined using the 2(−ΔΔCt) method (Livak and Schmittgen 2001).
Results
Genes encoding GRAS proteins in the B. napus genome
To identify GRAS encoding genes in the B. napus genome, a preliminary repeated BLASTP search (e values of < 1.0) was performed using the A. thaliana, O. sativa, and V. vinifera GRAS proteins as queries. To ensure the integrity of the B. napus GRAS protein data, we searched the two sequenced B. napus cultivar genomes in GENOSCOPE (Darmor–bzh, http://www.genoscope.cns.fr/brassicanapus/) (Chalhoub et al. 2014) and NCBI database (ZS11, https://www.ncbi.nlm.nih.gov/genome/annotation_euk/Brassica_napus/101/) (Sun et al. 2017), and then, we manually compared and corrected the sequence information of the candidate genes between these two cultivars.
A large number of deduced amino acid sequences containing GRAS or GRAS-like repeats were obtained. Redundant sequences were discarded, and the GRAS domain was verified in the remaining sequences by ExPASy. The sequences possessing incomplete open reading frames (ORFs), especially long deletion in the GRAS domain, were excluded for further analysis. Finally, based on the comparative analyses of the GRAS sequences between ZS11 and Darmor–bzh genomes, we found 92 GRAS candidate genes in Darmor–bzh. Among them, 85 genes encode typical non-redundant GRAS proteins with relative complete ORFs, one gene (BnaC06g03620D) should be split into two proteins, (BnGRAS55.1 and BnGRAS55.1) due to annotated error, and five genes may be pseudogenes and were excluded in our further analysis (Table S2). The candidate proteins were referred to as: BnGRAS1-BnGRAS86 based on their chromosomal order, where the BnaC06g03620D was named as BnGRAS55.1 and BnGRAS55.2 (Table 1). Accordingly, we used the 87 GRAS proteins (BnGRASs) from Darmor–bzh in our further analyses.
The 87 BnGRASs ranged from 44.47 (BnGRAS86) to 83.25 kDa (BnGRAS25) and were located in the nucleus (Table 1). They were primarily dominated by alpha helices and random coils and secondarily by extended strands and beta turns with an average incidence of 44.58, 40.23, 10.64, and 4.54% (Table S3), respectively. All except one had a theoretical PI value < 7 (Table 1). Most had an instability index > 40, indicating that they were unstable and all of them had a hydropathicity value < 0, suggesting that they were hydrophilic (Table S4).
We also identified 46 and 51 sequences encoding GRAS proteins in the genomes of B. rapa (BrGRASs) and B. oleracea (BoGRASs), respectively (Table S5). For the phylogenetic analysis, we retained the GRAS proteins that lacked some sequence at the N-terminus (non-GRAS domain) to ensure the representativeness of the data. This did not affect the phylogenetic tree construction. One GRAS gene was excluded from B. rapa in our data compared with a previous study (Song et al. 2014), due to the updated genome. The 34 GRAS genes in A. thaliana (AtGRASs) and 52 GRAS genes in V. vinifera (VvGRASs) were obtained from the previous reports (Cenci and Rouard 2017; Guo et al. 2017).
Phylogenetic analysis of GRAS gene family in B. napus
To explore the evolution and classification of candidate BnGRASs, the GRAS domains of the 87 BnGRASs, 34 AtGRASs, 46 BrGRASs, 51 BoGRASs, and 49 VvGRASs were aligned using MAFFT online software, and then, were used to construct NJ and ML phylogenetic trees by MEGA 7.0 and PhyML3.0, respectively, based on the alignment.
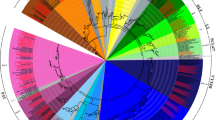
Our results showed that the topologies and bootstrap support values of the NJ and ML trees were highly congruent (Fig. 1 and Fig. S1), and we divided the candidates into 13 subfamilies based on high bootstrap support (Fig. 1). The distributions of candidate BnGRASs among the different subfamilies were as follows: HAM (13), LISCL (19), SHR (3), NSP1 (1), SCL32 (3), PAT (21), DELLA (10), SCR (3), SCL3 (4), DLT (4), SCL4/7 (2), LS (2), and NSP2 (2). To further confirm our classification, we referred to that of Cenci and Rouard (2017), who classified the GRAS proteins of eight representative angiosperms into 29 OGs based on orthologous relationships (Table S5). Our results were consistent with 17 OGs of the 29 OGs, and that 12 OGs representatives were lost in Brassicaceae (Cenci and Rouard 2017). However, homologous genes of subfamilies SCLB, SCLA, RAM1, and RAD1 were not identified in B. rapa, B. oleracea, nor B. napus, and, therefore, may not exist in the Brassica lineage. The number of BnGRASs in each subfamily was generally equal to the number in B. rapa and B. oleracea. However, some BnGRASs were missing from LISCL, HAM, SHR, SCL32, and NSP1 compared to those of B. rapa and B. oleracea, inferring gene loss in B. napus, potentially following the hybridisation process.
Phylogenetic relationships of GRAS proteins. The phylogenetic tree (neighbour joining) was constructed using all of the candidate GRAS proteins in A. thaliana (At: 34), B. rapa (Br: 46), B. oleracea (Bo: 51), B. napus (Bn: 87), and V. vinifera (Vv: 49) which were classified into 17 subfamilies (29 OGs) based on the bootstrap value and topology. The branches sharing the same background colour are in the same subfamily, except for four representatives of OG-SCLA, OG-RAM1, OG-RAD1, and OG-SCLB subfamilies, which did not have homologous genes in Brassica species
Sequence feature analysis of BnGRAS proteins
To analyze the sequence features in GRAS domains, we performed a multiple sequence alignment analysis of the GRAS domains of the 87 BnGRASs, using MAFFT with default parameters. The GRAS domains were located at the C-terminal. The length of which varied among candidates but was similar within each subfamily ranging from 230–400 amino acids. Furthermore, the similarity between all BnGRASs domains was only 35.88%, whereas, within each subfamily, it was 59.68–100% (Fig. S2). Accordingly, the sequence similarity of the GRAS domains was less conserved among the whole gene family, but was highly conserved within each subfamily (Fig. S2 and Fig. S3). Consistent with the previous studies (Liu and Widmer 2014), the GRAS domain comprised five main motifs, namely, LHRI, VHIID, LHRII, PFYRE, and SAW with distanced degree of conservation. Most of the BnGRASs contained the nuclear localisation signal region (NLS) of the LHRI motif at the C-terminal (Figs. 2, S2 and S3). The LHRII motif was relatively conserved, especially the Lx6Lx6L (L=leucine, x=any amino acid) sub-motif in the first half part that was highly conserved in BnGRASs (Fig. 2). The xHxxD (H=histidine, D=aspartic acid) sub-motif in the VHIID motif was involved in transcriptional activation activity (Gallagher and Benfey 2009). Accordingly, we found that the residues of H-162 (99%) and D-165 (100%) in this motif were highly conservative (Fig. S2), implying that they might be important for transcriptional activities. The PFYRE motif was relatively less conserved and contained three sub-motifs (P, FY, and RE) that had a short segment insertion at the N-terminal in the SCL3 subfamily (Fig. S2). The SAW motif comprised four sub-motifs: RVER, W-G, L-W, and SAW (Fig. 2 and Fig. S2), consistent with the previous results (Yoon et al. 2016).
We further identified 24 conserved motifs outside the GRAS domain at the N-terminal of the GRAS proteins in A. thaliana, B. rapa, B. oleracea, and B. napus using the MEME website. In general, the distribution of these motifs varied among subfamilies (Fig. S4). For example, motif 1 was only located in PAT subfamily; motifs 15, 16, and 17 in LISCL subfamily; and motifs 7, 8, and 9 in SCL4/7 subfamily (Table S6). The diversity of motifs between subfamilies may be related to their functional evolution.
Intron/exon structure and miRNA target analysis
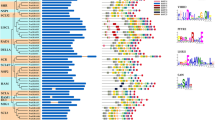
Using the online GSDS 2.0 software, we analyzed the intron insertions of the 87 BnGRASs (Fig. 3). Our results show that most BnGRASs have only one GRAS domain. About 89.7% (78/87) of the genes did not have intron insertion sites along their full-length ORF regions. The last (10.3%) have similar insertion sites in the same branch (Fig. 3), among which nine BnGRASs have one intron insertion in the GRAS domains. Similarly, we analyzed the intron insertions of GRAS genes in A. thaliana, B. oleracea, and B. rapa, and found that 88% (30/34), 89% (41/46), and 86% (44/51) of the candidates had no intron insertion in these three species, respectively (Fig. S5). Moreover, we found that the insertion positions and phases of the introns in the orthologs genes were highly conserved in each clade. For example, all members of the same clade in the SCR subfamily (AtGRAS21, BrGRAS39, BoGRAS29, BnGRAS31, and BnGRAS66) had the same intron insertion pattern (Fig. S5). These results revealed that the intron patterns of the GRAS gene family are conserved within each branch.
Phylogenetic relationship and gene structures of BnGRASs. The neighbour joining (NJ) tree was generated based on the multiple sequence alignment of the 87 BnGRAS proteins. The coloured backgrounds represent the 13 subfamilies that were supported with high predictive value (> 99%). Green boxes indicate the exons, black lines indicate the introns, and pink boxes represent the GRAS domains
MicroRNAs (miRNAs) played an important role in the expression of transcription factor genes in plants (Axtell 2013), and it was reported that GRAS members are regulated by miRNA171 (Ma et al. 2014). Our results showed that eight genes in HAM (BnGRAS6, BnGRAS7, BnGRAS22, BnGRAS39, BnGRAS40, BnGRAS48, BnGRAS51, and BnGRAS80) were unambiguously complementary to the miRNA171 sequence. This result was confirmed by psRNATarget (Table S7). In addition, another 26 miRNAs were also found to bind to 63 BnGRASs with about 90% complementary. These results suggested that miRNAs may play an important role in regulating the expression of BnGRASs.
Chromosome distribution and collinearity of the B. napus GRAS gene family
Based on genome annotation, the distribution of the BnGRASs was drawn. Most of the 87 BnGRASs were mapped on the 19 B. napus chromosomes (Fig. S6), except for one gene in the Unn subgenome (BnGRAS86), four genes in the Ann subgenome (BnGRAS38-BnGRAS41), and ten genes in the Cnn subgenome (BnGRAS75-BnGRAS85), the locations of which were unclear yet. In general, the distribution on each chromosome was uneven. For example, chromosomes An09 had seven genes, while chromosome An02 only had one. In all, the An and Cn subgenomes contain 41 and 45 BnGRASs, respectively, proving that there is not an obvious bias between these two subgenomes. This was similar to the gene number in the ancestors of B. rapa (46 genes) and B. oleracea (51 genes) genomes, respectively.
To explore the amplification mechanism of BnGRASs, we performed collinearity analysis of GRAS genes in the A. thaliana, B. oleracea, B. rapa, and B. napus genomes (Fig. 4) by Brassica Database (BRAD). We further demonstrated that up to 22.99% (16 genes from B. rapa and four genes from B. oleracea) of BnGRASs originated from chromosome doubling; 19.54% genes (12 genes from B. rapa and five genes from B. oleracea) originated from segmental exchange; 32.18% genes (24 genes from B. rapa and four genes from B. oleracea) originated from homologous exchange; 8.05% genes (seven genes) were involved in segmental duplications by GoGe. Only three pairs (BnGRAS32/BnGRAS33, BnGRAS39/BnGRAS40, and BnGRAS67/BnGRAS68) were tandem duplications (4.59%) that occurred in B. napus following hybridisation (Fig. S6; Table S8). We further analyzed the loss of the GRAS gene after polyploidisation events. The results showed that most of the BnGRASs derived from B. rapa were retained in the An subgenome, and a small number of candidate genes were exchanged to the Cn subgenome. However, many BnGRASs derived from B. oleracea were lost in B. napus; accordingly, a relatively small number of genes from B. oleracea were retained in the Cn subgenome (Table S8). Given that B. napus was hybridised by B. rapa and B. oleracea about 7500 years ago, these results suggest that genomic heterogeneous doubling (allopolyploid) was the main driving force for the large expansion of GRAS genes in B. napus genome, which tended to retain the genes from B. rapa.
Collinearity analysis of GRAS genes from A. thaliana (At), B. rapa (Br), B. oleracea (Bo), and B. napus (Bn). The outer circle indicates the chromosome number and the inner circle indicates the locations of the AtGRASs, BoGRASs, BnGRASs, and BrGRASs on each chromosome. The blue lines link two syntenic GRAS genes from A. thaliana, B. oleracea, B. napus, and B. rapa. The red line indicates tandem duplications
To explore the selective pressures acting upon the duplicated genes, we determined the non-synonymous/synonymous substitution ratio (Ka/Ks) for each pair of duplicated genes. Our results showed that the Ka/Ks ratios of the duplicates were commonly less than 1, indicating that these genes were subject to purifying selection (Table S9).
Interaction network analysis of GRAS proteins in B. napus
In the GRAS domain, the LHRI motif was known to mediate protein–protein interactions that are important for plant development. For instance, in rice, a conserved LZ domain in the LHRI motif of SLR1 was able to interact with itself to form a homodimer and inhibit the protein functions (Itoh et al. 2002), while, in M. truncatula, the LHRI motif of NSP2 interacted with NSP1 to form a heterodimer that was required for nodule formation (Hirsch et al. 2009). These results suggested that BnGRASs might function by forming homo- and hetero-protein complexes. Thus, we used the STRING database to construct a protein interaction network for BnGRASs based on the orthology analysis with A. thaliana GRAS proteins.
A total of 149 interaction protein pairs were predicted in A. thaliana (Table S10). The number and the types of interaction proteins for each subfamily were significantly different. Among the 13 subfamilies, SCL4/7 and NSP2 had no interaction proteins; DELLA, PAT, and LISCL had more interaction proteins. The interaction proteins of GRAS proteins were found to be diverse, including transcription factors (37.3%), enzymes (19.6%) involved in signal transduction, and other types of proteins (43.1%) (Table S10), consistent with the previous studies. For example, members of SHR and SCR interacted with JKD and MGP transcription factors to control an oriented cell division in ground tissue stem cell daughters (Welch et al. 2007). RGA1 (DELLA) interacted with GID1 (receptor protein) and PIF3 (transcription factor) to mediate gibberellin acid (GA) signal transduction and activate a light-responsive transcription factor that was required for GA signaling and controls root growth, seed germination, and flower development (Feng et al. 2008). HAM members (SCL27, SCL22, and HAM4) interacted with WUS-like transcription factors (WUS and WOX4) to specify stem cell identity in meristems (such as shoot apical meristem, SAM).
In B. napus, 45 BnGRASs have orthologs in A. thaliana, based on the syntenic analysis data in BRAD and up to 868 interaction proteins of BnGRASs were predicted on the basis of the orthology analysis with A. thaliana GRAS proteins (Fig. 5; Table S10). Accordingly, members of PAT, LISCL, DELLA, and HAM have relatively more interaction proteins than the others. In addition, the types of interaction proteins were generally different among subfamilies. For instance, the interaction proteins were mainly transcription factors in SCR, SHR, LS, and SCL3, whereas, in DLT, the interaction proteins ware mainly other types of proteins. GO enrichment analysis showed that interaction proteins mainly exist in the nucleus (208 genes), plasma membrane (30 genes), and cytoplasm (22 genes), and functions were generally sequence-specific transcription factor activity (124 genes), protein binding (79 genes), and DNA binding (65 genes). Interaction proteins participated in up to 289 biological processes including transcription regulation (124 genes), jasmonic acid mediated signaling pathway (37 genes), and response to chitin (35 genes) (Table S11; Fig. S7).
Interaction network of BnGRAS proteins in B. napus. 868 pairs of interacting proteins for 11 BnGRAS subfamilies, except for SCL4/7 and NSP2 that had no interaction proteins. The red rhombi indicate the BnGRAS proteins in each subfamily; the other coloured diagrams symbolise the four main types to which the interaction proteins in each clade belong
In all, our results show that most of the GRAS proteins tend to form protein complexes, suggesting that it is the major way for GRAS genes to regulate plant growth and development. Although the BnGRASs’ interaction network needs to be verified experimentally, our results provide important theoretical evidence for the molecular mechanisms of BnGRASs.
Expression analysis of BnGRASs at different developmental stages
Gene expression patterns are closely related to function, so we further analyzed the expression levels of BnGRASs in various tissues using the published RNA-seq data (BioProject ID PRJNA358784).
Our results showed that most of the BnGRASs were differentially expressed in different tissues in both vegetative and reproductive organs at different stages in B. napus. Except for BnGRAS68 that had no detectable expression level, most BnGRASs had high expression levels in roots, stems, leaves, flowers, and seed tissues. For example, members of SCR, SHR, and LS were mainly expressed in roots; members of HAM and SCL3 were mainly expressed in reproductive organs; members of PAT, LISCL, DELLA, and SCL4/7 were expressed in different vegetative organisation (Fig. 6). In addition, we found that 21 BnGRASs from PAT (BnGRAS2, BnGRAS34, BnGRAS43, BnGRAS46, BnGRAS56, BnGRAS59, BnGRAS71, and BnGRAS81), DELLA (BnGRAS16, BnGRAS30, BnGRAS61, BnGRAS74, and BnGRAS75), SHR (BnGRAS24 and BnGRAS63), SCL4/7 (BnGRAS21 and BnGRAS85), and LISCL subfamilies (BnGRAS25, BnGRAS3, BnGRAS65, and BnGRAS73) were highly expressed in roots and three BnGRASs from SCR (BnGRAS86) and LS subfamilies (BnGRAS11 and BnGRAS57) were specifically expressed in roots. These results demonstrated that many BnGRASs (24 genes) have a relative higher expression levels in roots, suggesting they may play an important role in root development in B. napus (Fig. 6).
Expression profiles of BnGRASs in 50 different samples during plant development by RNA-seq. Ro root, Ststem, Le leaf, Sp silique pericarp, Sc seed coat, Em embryo, Ao anthocaulus, Se seed, Hy hypocotyl, GS germination seeds, Cap capillament, Pi pistil, Cal calyx, Co cotyledon, Pe petal. The ‘h’, ‘d’, ‘s’, ‘b’ ‘i’, ‘f’ indicate hour, day, seeding, budding, initial flowering, and full-bloom stages, respectively. Colour bar at the top represents log2 (FPKM) expression values: green represents low expression value, black represents mean expression level, and red represents high expression value
We further analyzed the expression patterns of ten pairs of duplication genes in B. napus, and the results showed that the expression patterns of each pair were similar (Fig. S8c and d), indicating that they were functionally redundant. We compared the expression profiles of the homologous GRAS genes in roots, stems, and leaves at the seedling stage of B. rapa and B. oleracea. Our results showed that most of the BnGRASs have similar expression patterns with their homologs in B. rapa and B. oleracea, indicating their functional conservation. For instance, homologs from B. rapa and B. napus in subfamily DELLA were all expressed in roots and stems; homologs from B. rapa, B. oleracea, and B. napus in subfamily HAM have similar expression profiles in roots and stems (Fig. S8a–c), etc. However, we also found that members of three subfamilies, PAT, SCL32, and DLT, have different expression patterns, where members of PAT in B. napus were mainly expressed in leaves, while these in B. rapa and B. oleracea were mainly expressed in roots and stems (Fig. S8a–c). This may indicate functional diversification of duplicates.
Expression analysis of BnGRASs under hormone induction
We analyzed the hormone-induced expression patterns of BnGRASs under ABA, IAA, GA3, ACC, and 6-BA treatments in roots as well, using recently created RNA-seq data. The BnGRASs with expression levels less than one (FPKM < 1) were removed from our analysis (Fig. 7).
Expression profiles of BnGRASs under five hormone induction in B. napus root by RNA-seq. IAA, indoleacetic acid; ACC, 1-aminocyclopropanecarboxylic acid; ABA, abscisic acid; GA3, gibberellin acid 3; 6-BA, cytokinin. The BnGRASs with no or weak expression levels (FPKM < 1) were removed from the figure. Colour bar at the top represents log2 (FPKM) expression values: olive colour represents low expression value, white represents mean expression level, and red represents high expression value
We found that most of BnGRASs were not obviously induced by hormone treatments, while a few BnGRASs from PAT, LISCL, SHR, SCR, DELLA, and HAM were obviously up- or down-regulated. The genes in the same branch generally have the same expression pattern within each subfamily. Such as members of SCR subfamily (BnGRAS31 and BnGRAS66) were up-regulated by 6-BA induction; members of DELLA (BnGRAS10 and BnGRAS54), SCL32 (BnGRAS60), SHR (BnGRAS24 and BnGRAS49) LISCL (BnGRAS3 and BnGRAS36), PAT (BnGRAS46 and BnGRAS41), and SCL3 (BnGRAS13) subfamilies were down-regulated by ACC and GA3; members of PAT (BnGRAS27, BnGRAS2, BnGRAS14, and BnGRAS20) were up-regulated by GA; members of PAT (BnGRAS14 and BnGRAS15) subfamily were up-regulated by ACC (Fig. 7).
Overall, our results showed that the GRAS gene family has a wide expression pattern in B. napus, and some BnGRASs were obviously induced by hormone inductions in roots. Up to 24 BnGRASs may be involved in B. napus root development, especially these in SCR and LS.
Expression analysis of BnGRASs under abiotic stresses by qRT-PCR
As mentioned above, members of SCL4/7, LISCL, LS, and SCR subfamilies were highly or specifically expressed in roots (Fig. 6), suggesting their possible roles in root. Abiotic stress is a key global issue that reduces crop yield, especially drought and excess salt. The root resistance of plants plays important roles in response to drought and salt stress. Therefore, to examine possible roles of BnGRASs under abiotic stress response in B. napus roots, we selected nine genes, which were highly or specificity expression in roots from SCL4/7 (BnGRAS21 and BnGRAS85), LISCL (BnGRAS25 and BnGRAS65), LS (BnGRAS11 and BnGRAS57), and SCR (BnGRAS31, BnGRAS66, and BnGRAS86) subfamilies to analyze their expression profiles in root under PEG, salt stresses, and ABA induction by qRT-PCR.
As shown in Fig. 8a and c, the expressions of six candidate genes (BnGRAS11, BnGRAS57, BnGRAS31, BnGRAS66, BnGRAS85, and BnGRAS21) were down-regulated and three genes (BnGRAS86, BnGRAS25, and BnGRAS65) were up-regulated under NaCl stress and ABA induction. The expressions of four genes from SCR (BnGRAS31, BnGRAS66, and BnGRAS86) and LISCL (BnGRAS25) were up-regulated under PEG stress (Fig. 8b), suggesting that they may be candidates for stress-resistance functions in B. napus root systems. Another five genes from LS (BnGRAS11 and BnGRAS57), SCL4/7 (BnGRAS85 and BnGRAS21), and LISCL (BnGRAS65) were not evident under PEG stress (Fig. 8b), suggesting that they may be involved in other biological processes. In addition, we found that two genes in SCR (BnGRAS86) and LISCL (BnGRAS25) were up-regulated under NaCl and PEG stresses and ABA induction (Fig. 8a–c), indicating that they may respond to drought stress through the ABA signaling pathway in B. napus roots. Conversely, the two genes from SCR (BnGRAS31 and BnGRAS66) subfamily were up-regulated under PEG stresses and down-regulated under NaCl stresses and ABA induction (Fig. 8a–c), suggesting that these genes may respond to drought resistance through the ABA independent pathway.
In all, our results showed that the BnGRAS86 and BnGRAS25 obviously respond to drought and salt stress, suggesting that they may be good genes for analyzing the mechanisms of GRAS gene regulation of B. napus abiotic stress response.
Origin, evolution, and diversification of GRAS gene family in the plant kingdom
There is yet to be a standard classification for the GRAS gene family, e.g., Bolle (2004) divided the rice and A. thaliana GRAS genes into eight subfamilies, although some genes were not well defined. In tomato (Niu et al. 2017), P. trichocarpa (Liu and Widmer 2014), and Gossypium hirsutum L. (Zhang et al. 2018), the family was divided into 10, 13, and 14 subfamilies, respectively, with some genes individually divided into a group due to species specificity. Based on eight angiosperm species, Cenci and Rouard (2017) divided the family into 17 subfamilies. This may be attributed to the relatively small sample sizes of previous studies. Thus, we further analyzed the distribution of GRAS genes in 26 plant genomes, including five green algae (Chlamydomonas reinhardtii, Dunaliella salina, Micromonas pusilla, Ostreococcus lucimarinus, and Klebsormidium nitens), two mosses (Marchantia polymorpha and Physcomitrella patens), one fern (Selaginella moellendorffii), one gymnosperm (Pinus abies), and 17 angiosperms. Among them, GRAS genes in nine species (D. salina, M. pusilla, K. nitens, M. polymorpha, Amborella trichopoda, Glycine max, B. rapa, B. oleracea, and B. napus) were first identified in the present study, whereas the remainder were acquired from other reports (Lee et al. 2008; Liu and Widmer 2014; Wang et al. 2016; Cenci and Rouard 2017; Guo et al. 2017; Niu et al. 2017; Wang et al. 2018; Zhang et al. 2018) (Fig. 9; Table S5).
In all, we obtained 1388 GRAS candidate genes in 21 land plants. Consistent with the previous reports (Song et al. 2017; Wang et al. 2018; Zhang et al. 2018), no GRAS members were identified in algae, but they were in a moss (M. polymorpha). It was previous reported that GRAS proteins were present in bacteria and horizontally transferred from bacteria to plants close to the origin of land plants (Zhang et al. 2012). Thus, this gene family may have originated from the horizontal transfer process from bacteria to early land plants ~ 443 millions of years ago (MA) (Steemans et al. 2009). To systematically understand the evolution of GRAS in land plants, 1265 of the 1388 candidates with relatively complete ORF regions were applied for phylogenetic analysis (NJ and ML tree), 98 candidates with large sequence deletions in the GRAS domains were not included in the phylogenetic analysis due to technical reason (no common sites for the sequence pairs) with classifications defined based on sequence homologous, while 17 genes with no typical GRAS character (e.g., the GRAS domain) by ExPASy analysis were excluded from this study (Table S5; Fig. S9 and S10). To ensure the accuracy of our classification, we considered the current classification and gene functions of different species (Liu and Widmer 2014; Cenci and Rouard 2017; Niu et al. 2017; Song et al. 2017). As an SCLB member in M. truncatula was recently characterised and named MIG1 (Heck et al. 2016), we renamed SCLB subfamily to MIG1.
Based on the NJ and ML phylogenetic trees, we divided the candidate GRAS genes from the 21 species into 19 subfamilies (Fig. S9 and S10). This number is larger than that reported previously (Liu and Widmer 2014; Niu et al. 2017). However, our result is highly consistent with the classification of the OGs and subfamilies for the GRAS gene family in eight angiosperms by Cenci and Rouard (2017). In addition, we identified the 29 OGs in all of the angiosperms investigated in this study (Fig. 9 and Table S5), and the 17 subfamilies defined by Cenci and Rouard (2017), supporting that they were conserved in higher plants. As mentioned above, the amino acid sequence (Fig. S2), expression patterns (Fig. 6), and functions (Table S12) showed subfamily-specific conservation across these subfamilies, providing additional support for our classification. However, some GRAS proteins from lower plants and G. hirsutum L. and P. trichocarpa did not have orthologous relationships with the 29 well-known OGs and were clustered into two different clades in the phylogenetic analysis, and, thus, were defined as PSG and PG subfamilies, respectively. Consistent with the results of Zhang et al. (2018), PSG only includes moss and fern genes, and PG only contains G. hirsutum L. and P. trichocarpa genes, and, thus, may be species- and/or lineage-specific. In the previous reports, members of the PG subfamily were classified as the SHR (Zhang et al. 2018) or DELLA (Liu and Widmer 2014) subfamilies. However, their protein motifs, intron patterns, and expression patterns were different from those of SHR or DELLA subfamilies. Most members of the PG subfamily contain the zinc finger and ubiquitin-like protease at the C-terminal instead (Liu and Widmer 2014; Zhang et al. 2018). Thus, it may be species- and/or lineage-specific as well.
Of the 19 subfamilies, PSG is found in lower plants but not higher plants; SCLA appeared in the early ancestral angiosperm A. trichopoda about ∼ 130 MA (Crane et al. 1995); RAD1 and NSP2 included fern and angiosperm members and may have originated in a tracheophyte; PG included P. trichocarpa and G. hirsutum L. genes and may be species-specific; the last 14 subfamilies (PAT, LISCL, DELLA, SHR, SCR, SCL3, SCL32, DLT, HAM, LS, SCL4/7, RAM1, MIG1, and NSP1) existed in moss (Fig. 9), suggesting that they might have evolved earlier, after the divergence of land plants, and were retained in land plants. In addition, some subfamilies were lost in specific species, such as, RAM1, RAD1, and SCLA were lost in gymnosperm and Brassicaceae plants and MIG1 were lost in Brassicaceae plants.
In summary, our results support that the GRAS gene family in land plants should be classified into 19 subfamilies, with 17 of them being highly conserved across angiosperms. There may exist some species- and lineage-specific subfamilies, implying different expansion and loss of some subfamilies in some land plants.
Discussion
Functional conservation and diversity of GRAS genes
The functions of some GRAS genes have been previously studied, mainly focusing on tissue/organ development, stress-resistance processes, signal transduction, and symbiotic processes with fungi. Functional characterisation of GRAS proteins has indicated the conserved function of putative orthologues in each subfamily and/or subclade. Therefore, we summarised the functions of plant GRAS genes (Table S12).
As plant-specific transcription factors, GRAS genes are well known to be involved in various processes of tissue or organ development. In SCR subfamily, A. thalianaSCR gene is required for asymmetric cell division that is responsible for ground tissue formation in roots and shoots (Wysocka-Diller et al. 2000). The homologs in rice and B. napus, OsSCR, and BnGRASs (BnGRAS31 and BnGRAS68) also have similar expression patterns and functions (Gao et al. 2004). In LS subfamily, LAS gene in A. thaliana has genetic regulation of axillary meristem formation (Greb et al. 2003); similarly, the rice OsMOCI, S. lycopersicum Ls, and B. napus homologous genes have similar functions (Li et al. 2003; Yang et al. 2011). In SHR subfamily, SHR protein usually interacts with SCR and SCL23 proteins, which is essential for bundle sheath cell-fate specification (Cui et al. 2014). In HAM subfamily, A. thalianaHAM1, HAM2, HAM3, and HAM4 genes functional redundancy, which regulated stem tips, root tips, and bud meristems (Engstrom et al. 2011). In Petunia hybrid, the PhHAM gene also plays a key role in shoot meristem and stem cell maintenance (Stuurman et al. 2002). The LISCL subfamily members have a wide range of tissue expression profiles (Fig. 6); in Lilium longiflorum, LISCL is involved in transcriptional regulation during microsporogenesis within the lily anther (Morohashi et al. 2003).
GRAS genes are also considered to act as key regulators of responses to hormone induction and abiotic stress (cold, hot, drought, and salt). For example, DELLA members play an important role in GA biosynthesis and negatively regulate GA signaling pathways in A. thaliana (Silverstone et al. 1998). In A. thaliana, DELLA subfamily members (AtRGL3, AtRGA, and AtGAI) respond to GA and were related to tissue development in seeds, roots, stems, and leaves (Tyler et al. 2004). It has been reported that RGL2 is inhibited by GA3 induction to inhibit seed germination (Lee et al. 2002). In our results, AtRGL2 homologous genes (BnGRAS10 and BnGRAS55) were down-regulated under GA3 induction, as well (Fig. 6). The SCL3 subfamily member (SCL3) acted as a positive regulator of GA signaling, SCL3 and DELLA subfamily members interact, which maintains GA homeostasis and controls GA-mediated growth and development (Zhang et al. 2011). In DLT subfamily, OsDLT gene was found to be involved in negative feedback regulation of brassinosteroid (BR) biosynthesis (Tong et al. 2009). The SCL4/7 subfamily members appear to be involved in plant stress tolerance. In A. thaliana, AtSCL7 gene was up-regulated, while AtSCL4 gene was down-regulated under drought and salt stress conditions (Kilian et al. 2007). Overexpressing poplar PeSCL7 gene in A. thaliana could enhance stress tolerance (Ma et al. 2010). The LISCL subfamily member AtSCL14 gene could activate stress-inducible promoters and regulate the expression of stress-responsive genes, such as drought stress (Fode et al. 2008).
In addition, the GRAS gene is also involved in signal transduction pathways and symbiotic relationships between plants and microorganisms. For example, in PAT subfamily, AtPAT1 and SCL13 genes are involved in phytochrome A and B signal transduction (Bolle et al. 2000). In NSP1 and NSP2 subfamilies, both of NSP1 and NSP2 genes fulfil dual regulatory functions involved in nod factor from rhizobial bacteria signal transduction as well as strigolactone (SL) biosynthesis in non-symbiotic conditions in M. truncatula and rice (Liu et al. 2011). In RAD1 and RAM1 subfamilies, RAD1 and PAM1 genes were involved in mycorrhizal signaling and promote mycorrhizal colonisation by regulating cutin biosynthesis (Gobbato et al. 2012; Xue et al. 2015). In the MIG1 subfamily, symbiotic fungi control plant root cortex development through the MIG1 homolog genes (Heck et al. 2016). It is clear that different subfamily members have different functions, but the same subfamily orthologous genes function, similarly, indicating the diversity and conservation of GRAS gene function.
The conservatism of miRNA171 regulating GRAS genes across land plants
MiRNAs bind to specific sequences as a negative regulator; this can be deployed for a wide variety of functions (Axtell 2013). A previous analysis of the origin and evolution of the miRNA family found that some originated from bryophytes or angiosperms (Cuperus et al. 2011), suggesting that they may be closely related to the origin of the gene family and/or the lineage-specific subfamily.
It was reported that miRNA171 was involved in the regulation of shoot branching by regulating the transcription of GRAS genes (Wang et al. 2010b). This mechanism is identified in many species, for instance, A. thaliana (Wang et al. 2010b), rice (Fan et al. 2015), and grapevine (Sun et al. 2016). Our results showed that HAM comprised two clades; the first clade (OG-HAM-II) contained eight genes (BnGRAS6, BnGRAS7, BnGRAS22, BnGRAS39, BnGRAS40, BnGRAS48, BnGRAS51, and BnGRAS80) which were highly expressed in the stem (Fig. 6) and were unambiguously complementary to the miRNA171 sequence (Table S7), suggesting that these genes may be the target genes of miRNA171. The second clade (OG-HAM-I) contains five genes (BnGRAS1, BnGRAS42, BnGRAS62, BnGRAS17, and BnGRAS58) which were mainly expressed in flowers (Fig. 6). Interestingly, these genes were not the targets of miRNA171. These results indicate that the function of the HAM genes was differentiated during evolution, but was conserved within clades.
In plants, only a minority of annotated miRNA gene families are conserved across all plants and most are species-specific (Cuperus et al. 2011). In addition, miRNAs often have a defined set of miRNA targets that they co-evolved with. miRNA171 is considered to have originated from moss (P. patens) and then expanded in angiosperms (Nozawa et al. 2012). According to our results, the GRAS gene family appeared in early land plants. HAM was one of the most ancient subfamilies, members of which were identified in moss. Accordingly, we found that the target sites of miRNA171 existed in the HAM homologs in moss (Sphagnum fallax), P. radiate, basal angiosperms (A. trichopoda), dicots (A. thaliana), and monocots (O. sativa), except for ferns due to incomplete information on gene annotation (Table S13). The target sequences were highly conserved across different plant species. These results suggest that the mechanism with which miRNA171 regulates HAM homologs was formed in ancestral species to meet the needs of shoot tip branch development. This result further confirmed that species traits and gene regulation mechanisms were coevolutionary. In conclusion, BnGRASs may be regulated by miRNA and especially the miRNA171. The role of HAM-like genes regulating branching through miRNA171 was conserved in land plant.
Conclusions
In this study, 87 GRAS genes (BnGRASs) were identified in the B. napus genome and were further divided into 13 subfamilies. Most BnGRASs were found to have no intron insertion and the GRAS domains were less conservative across different subfamilies, but are conserved within the same subfamily. GRAS genes were only distributed in land plants and were divided into 19 subfamilies, 14 of which may have originated before bryophyte differentiation. Genome-wide duplication was its main amplification mechanism in land plants. Eight BnGRASs in HAM may be regulated by miRNA171 and this mechanism was conserved throughout land plants. The expression patterns of BnGRASs were commonly diversified and up to 24 genes may be related to root development. Among them, two genes (BnGRAS86 and BnGRAS25) may response to PEG and NaCl stress by the ABA signal pathway. In summary, our results provide valuable data for further exploring the functions of candidate BnGRASs, especially their roles in root development and stress response.
Author contributions statement
HD contributed to the conception of the study. PCG, HD, and JY drafted and revised the manuscript. PCG, HD, JW, JY, MMW, FR, and MML contributed to data analysis. PCG, HD, YZK, YWW, PFL, and JNL conceived of and designed the experiments. All authors reviewed and approved the final manuscript.
Abbreviations
- BRAD:
-
Brassica database
- FPKM:
-
Fragments per kilobase of transcript per million mapped reads
- OGs:
-
Orthologous groups
- PEG:
-
Polyethylene glycol
References
Ashburner M, Ball CA, Blake JA (2000) Gene ontology: tool for the unification of biology. The gene ontology consortium. Nat Genet 25:25–29. https://doi.org/10.1038/75556
Axtell MJ (2013) Classification and comparison of small RNAs from plants. Annu Rev Plant Biol 64:137–159. https://doi.org/10.1146/annurev-arplant-050312-120043
Bjellqvist B, Hughes GJ, Pasquali C, Paquet N, Ravier F, Sanchez JC, Frutiger S, Hochstrasser D (1993) The focusing positions of polypeptides in immobilized pH gradients can be predicted from their amino acid sequences. Electrophoresis 14:1023–1031. https://doi.org/10.1002/elps.11501401163
Bolle C (2004) The role of GRAS proteins in plant signal transduction and development. Planta 218:683–692. https://doi.org/10.1007/s00425-004-1203-z
Bolle C, Koncz C, Chua NH (2000) PAT1, a new member of the GRAS family, is involved in phytochrome a signal transduction. Genes Dev 14:1269–1278. https://doi.org/10.1101/gad.14.10.1269
Cenci A, Rouard M (2017) Evolutionary analyses of GRAS transcription factors in angiosperms. Front Plant Sci 8:273. https://doi.org/10.3389/fpls.2017.00273
Chalhoub B, Denoeud F, Liu S et al (2014) Plant genetics. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 345:950–953. https://doi.org/10.1126/science.1253435
Chou KC, Shen HB (2008) Cell-PLoc: a package of web-servers for predicting subcellular localization of proteins in various organisms. Nat Protoc 3:153–162. https://doi.org/10.1038/nprot.2007.494
Crane PR, Friis EM, Pedersen KR (1995) The origin and early diversification of angiosperms. Nature 374:27–33. https://doi.org/10.1038/374027a0
Cui H, Kong D, Liu X, Hao Y (2014) SCARECROW, SCR-LIKE 23 and SHORT-ROOT control bundle sheath cell fate and function in Arabidopsis thaliana. Plant J 78:319–327. https://doi.org/10.1111/tpj.12470
Cuperus JT, Fahlgren N, Carrington JC (2011) Evolution and functional diversification of miRNA genes. Plant Cell 23:431–442. https://doi.org/10.1105/tpc.110.082784
Engstrom EM, Andersen CM, Gumulak-Smith J, Hu J, Orlova E, Sozzani R, Bowman JL (2011) Arabidopsis homologs of the petunia hairy meristem gene are required for maintenance of shoot and root indeterminacy. Plant Physiol 155:735–750. https://doi.org/10.1104/pp.110.168757
Fan T, Li X, Yang W, Xia K, Ouyang J, Zhang M (2015) Rice osa-miR171c mediates phase change from vegetative to reproductive development and shoot apical meristem maintenance by repressing four OsHAM transcription factors. PLoS One 10:e0125833. https://doi.org/10.1371/journal.pone.0125833
Feng S, Martinez C, Gusmaroli G, Wang Y, Zhou J, Wang F, Chen L, Yu L, Iglesias-Pedraz JM, Kircher S, Schäfer E, Fu X, Fan LM, Deng XW (2008) Coordinated regulation of Arabidopsis thaliana development by light and gibberellins. Nature 451:475–479. https://doi.org/10.1038/nature06448
Fode B, Siemsen T, Thurow C, Weigel R, Gatz C (2008) The Arabidopsis GRAS protein SCL14 interacts with class II TGA transcription factors and is essential for the activation of stress inducible promoters. Plant Cell 20:3122–3135. https://doi.org/10.1105/tpc.108.058974
Gallagher KL, Benfey PN (2009) Both the conserved GRAS domain and nuclear localization are required for SHORT-ROOT movement. Plant J 57:785–797. https://doi.org/10.1111/j.1365-313X.2008.03735.x
Gao MJ, Parkin I, Lydiate D, Hannoufa A (2004) An auxin-responsive SCARECROW-like transcriptional activator interacts with histone deacetylase. Plant Mol Biol 55:417–431. https://doi.org/10.1007/s11103-004-0892-9
Gobbato E, Marsh JF, Vernié T, Wang E, Maillet F, Kim J, Miller JB, Sun J, Bano SA, Ratet P (2012) A GRAS-type transcription factor with a specific function in mycorrhizal signaling. Curr Biol 22:2236–2241. https://doi.org/10.1016/j.cub.2012.09.044
Greb T, Clarenz O, Schafer E, Muller D, Herrero R, Schmitz G, Theres K (2003) Molecular analysis of the LATERAL SUPPRESSOR gene in Arabidopsis reveals a conserved control mechanism for axillary meristem formation. Genes Dev 17:1175–1187. https://doi.org/10.1101/gad.260703
Guindon S, Delsuc F, Dufayard JF, Gascuel O (2009) Estimating maximum likelihood phylogenies with PhyML. Methods Mol Biol 537:113–137. https://doi.org/10.1007/978-1-59745-251-9_6
Guo Y, Wu H, Li X, Li Q, Zhao X, Duan X, An Y, Lv W, An H (2017) Identification and expression of GRAS family genes in maize (Zea mays L.). PLoS One 12:e0185418. https://doi.org/10.1371/journal.pone.0185418
Heck C, Kuhn H, Heidt S, Walter S, Rieger N, Requena N (2016) Symbiotic fungi control plant root cortex development through the novel GRAS transcription factor MIG1. Curr Biol 26:2770–2778. https://doi.org/10.1016/j.cub.2016.07.059
Hirsch S, Kim J, Muñoz A, Heckmann AB, Downie JA, Oldroyd GE (2009) GRAS proteins form a DNA binding complex to induce gene expression during nodulation signaling in Medicago truncatula. Plant Cell 21:545–557. https://doi.org/10.1105/tpc.108.064501
Hu B, Jin J, Guo AY, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31:1296–1297. https://doi.org/10.1093/bioinformatics/btu817
Huang W, Xian Z, Kang X, Tang N, Li Z (2015) Genome-wide identification, phylogeny and expression analysis of GRAS gene family in tomato. BMC Plant Biol 15:209. https://doi.org/10.1186/s12870-015-0590-6
Itoh H, Ueguchi-Tanaka M, Sato Y, Ashikari M, Matsuoka M (2002) The gibberellin signaling pathway is regulated by the appearance and disappearance of SLENDER RICE1 in nuclei. Plant Cell 14:57–70. https://doi.org/10.1105/tpc.010319
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kilian J, Whitehead D, Horak J, Wanke D, Weinl S, Batistic O, D’Angelo C, Bornberg-Bauer E, Kudla J, Harter K (2007) The AtGenExpress global stress expression data set: protocols, evaluation and model data analysis of UV-B light, drought and cold stress responses. Plant J 50:347–363. https://doi.org/10.1111/j.1365-313X.2007.03052.x
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19:1639–1645. https://doi.org/10.1101/gr.092759.109
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Lee S, Cheng H, King KE, Wang W, He Y, Hussain A, Lo J, Harberd NP, Peng J (2002) Gibberellin regulates Arabidopsis seed germination via RGL2, a GAI/RGA-like gene whose expression is up-regulated following imbibition. Genes Dev 16:646–658. https://doi.org/10.1101/gad.969002
Lee MH, Kim B, Song SK, Heo JO, Yu NI, Lee SA, Kim M, Kim DG, Sohn SO, Lim CE, Chang KS, Lee MM, Lim J (2008) Large-scale analysis of the GRAS gene family in Arabidopsis thaliana. Plant Mol Biol 67:659–670. https://doi.org/10.1007/s11103-008-9345-1
Li X, Qian Q, Fu Z, Wang Y, Xiong G, Zeng D, Wang X, Liu X, Teng S, Hiroshi F, Yuan M, Luo D, Han B, Li J (2003) Control of tillering in rice. Nature 422:618–621. https://doi.org/10.1038/nature01518
Liu XY, Widmer A (2014) Genome-wide comparative analysis of the GRAS gene family in Populus, Arabidopsis and rice. Plant Mol Biol Report 32:1129–1145. https://doi.org/10.1007/s11105-014-0721-5
Liu W, Kohlen W, Lillo A, Op den Camp R, Ivanov S, Hartog M, Limpens E, Jamil M, Smaczniak C, Kaufmann K (2011) Strigolactone biosynthesis in Medicago truncatula and rice requires the symbiotic GRAS-type transcription factors NSP1 and NSP2. Plant Cell 23:3853–3865. https://doi.org/10.1105/tpc.111.089771
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Ma HS, Liang D, Shuai P, Xia XL, Yin WL (2010) The salt- and drought-inducible poplar GRAS protein SCL7 confers salt and drought tolerance in Arabidopsis thaliana. J Exp Bot 61:4011–4019. https://doi.org/10.1093/jxb/erq217
Ma Z, Hu X, Cai W, Huang W, Zhou X, Luo Q, Yang H, Wang J, Huang J (2014) Arabidopsis miR171-targeted scarecrow-like proteins bind to GT cis-elements and mediate gibberellin-regulated chlorophyll biosynthesis under light conditions. PLoS Genet 10:e1004519. https://doi.org/10.1371/journal.pgen.1004519
Morohashi K, Minami M, Takase H, Hotta Y, Hiratsuka K (2003) Isolation and characterization of a novel GRAS gene that regulates meiosis-associated gene expression. J Biol Chem 278:20865–20873. https://doi.org/10.1074/jbc.M301712200
Niu Y, Zhao T, Xu X, Li J (2017) Genome-wide identification and characterization of GRAS transcription factors in tomato (Solanum lycopersicum). Peer J 5:e3955. https://doi.org/10.7717/peerj.3955
Nozawa M, Miura S, Nei M (2012) Origins and evolution of microRNA genes in plant species. Genome Biol Evol 4:230–239. https://doi.org/10.1093/gbe/evs002
Silverstone AL, Ciampaglio CN, Sun T (1998) The Arabidopsis RGA gene encodes a transcriptional regulator repressing the gibberellin signal transduction pathway. Plant Cell 10:155–169. https://doi.org/10.1105/tpc.10.2.155
Song XM, Liu TK, Duan WK, Ma QH, Ren J, Wang Z, Li Y, Hou XL (2014) Genome-wide analysis of the GRAS gene family in Chinese cabbage (Brassica rapa ssp. pekinensis). Genomics 103:135–146. https://doi.org/10.1016/j.ygeno.2013.12.004
Song L, Tao L, Cui H, Ling L, Guo C (2017) Genome-wide identification and expression analysis of the GRAS family proteins in Medicago truncatula. Acta Physiol Plant 39:93. https://doi.org/10.1007/s11738-017-2393-x
Steemans P, Hérissé AL, Melvin J, Miller MA, Paris F, Verniers J, Wellman CH (2009) Origin and radiation of the earliest vascular land plants. Science 324:353. https://doi.org/10.1126/science.1169659
Stuurman J, Jäggi F, Kuhlemeier C (2002) Shoot meristem maintenance is controlled by a GRAS-gene mediated signal from differentiating cells. Genes Dev 16:2213–2218. https://doi.org/10.1101/gad.230702
Sun X, Xue B, Jones WT, Rikkerink E, Dunker AK, Uversky VN (2011) A functionally required unfoldome from the plant kingdom: intrinsically disordered N-terminal domains of GRAS proteins are involved in molecular recognition during plant development. Plant Mol Biol 77:205–223. https://doi.org/10.1007/s11103-011-9803-z
Sun X, Xie Z, Zhang C, Mu Q, Wu W, Wang B, Fang J (2016) A characterization of grapevine of GRAS domain transcription factor gene family. Funct Integr Genome 16:347–363. https://doi.org/10.1007/s10142-016-0479-y
Sun F, Fan G, Hu Q et al (2017) The high-quality genome of Brassica napus cultivar ‘ZS11’ reveals the introgression history in semi-winter morphotype. Plant J 92:452–468. https://doi.org/10.1111/tpj.13669
Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Bork P (2017) The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res 45:D362–D368. https://doi.org/10.1093/nar/gkw937
Tong H, Jin Y, Liu W, Li F, Fang J, Yin Y, Qian Q, Zhu L, Chu C (2009) DWARF and LOW-TILLERING, a new member of the GRAS family, plays positive roles in brassinosteroid signaling in rice. Plant J 58:803–816. https://doi.org/10.1111/j.1365-313X.2009.03825.x
Tyler L, Thomas SG, Hu J, Dill A, Alonso JM, Ecker JR, Sun TP (2004) DELLA proteins and gibberellin-regulated seed germination and floral development in Arabidopsis. Plant Physiol 135:1008–1019. https://doi.org/10.1104/pp.104.039578
Wang D, Zhang Y, Zhang Z, Zhu J, Yu J (2010a) KaKs_Calculator 2.0: a toolkit incorporating gamma-series methods and sliding window strategies. Genom Proteom Bioinform 8:77–80. https://doi.org/10.1016/S1672-0229(10)60008-3
Wang L, Mai YX, Zhang YC, Luo Q, Yang HQ (2010b) MicroRNA171c-targeted SCL6-II, SCL6-III, and SCL6-IV genes regulate shoot branching in Arabidopsis. Mol Plant 3:794–806. https://doi.org/10.1093/mp/ssq042
Wang Y, Shi S, Zhou Y, Zhou Y, Yang J, Tang X (2016) Genome-wide identification and characterization of GRAS transcription factors in sacred lotus (Nelumbo nucifera). Peer J 4:e2388. https://doi.org/10.7717/peerj.2388
Wang YX, Liu ZW, Wu ZJ, Li H, Wang WL, Cui X, Zhuang J (2018) Genome-wide identification and expression analysis of GRAS family transcription factors in tea plant (Camellia sinensis). Sci Rep 8:3949. https://doi.org/10.1038/s41598-018-22275-z
Welch D, Hassan H, Blilou I, Immink R, Heidstra R, Scheres B (2007) Arabidopsis JACKDAW and MAGPIE zinc finger proteins delimit asymmetric cell division and stabilize tissue boundaries by restricting SHORT-ROOT action. Genes Dev 21:2196–2204. https://doi.org/10.1101/gad.440307
Wysocka-Diller JW, Helariutta Y, Fukaki H, Malamy JE, Benfey PN (2000) Molecular analysis of SCARECROW function reveals a radial patterning mechanism common to root and shoot. Development 127:595–603. https://doi.org/10.1007/PL00008234
Xue L, Cui H, Buer B, Vijayakumar V, Delaux PM, Junkermann S, Bucher M (2015) Network of GRAS transcription factors involved in the control of arbuscule development in Lotus japonicus. Plant Physiol 167:854–871. https://doi.org/10.1104/pp.114.255430
Yang M, Yang Q, Fu T, Zhou Y (2011) Overexpression of the Brassica napus BnLAS gene in Arabidopsis affects plant development and increases drought tolerance. Plant Cell Rep 30:373–388. https://doi.org/10.1007/s00299-010-0940-7
Yoon EK, Dhar S, Lee MH, Song JH, Lee SA, Kim G, Jang S, Choi JW, Choe JE, Kim JH, Lee MM, Lim J (2016) Conservation and diversification of the SHR-SCR-SCL23 regulatory network in the development of the functional endodermis in Arabidopsis shoots. Mol Plant 9:1197–1209. https://doi.org/10.1016/j.molp.2016.06.007
Zhang ZL, Ogawa M, Fleet CM, Zentella R, Zentella R, Hu J, Heo JO, Lim J, Kamiya Y, Yamaguchi S, Sun TP (2011) Scarecrow-like 3 promotes gibberellin signaling by antagonizing master growth repressor DELLA in Arabidopsis. Proc Natl Acad Sci USA 108:2160–2165. https://doi.org/10.1073/pnas.1012232108
Zhang D, Iyer LM, Aravind L (2012) Bacterial GRAS domain proteins throw new light on gibberellic acid response mechanisms. Bioinformatics 28:2407–2411. https://doi.org/10.1093/bioinformatics/bts464
Zhang L, Li Q, Chen JF, Chen WS (2016) Computational identification and systematic classification of novel GRAS genes in Isatis indigotica. Chin J Nat Med 14:161–176. https://doi.org/10.1016/S1875-5364(16)30013-9
Zhang B, Liu J, Yang ZE, Chen EY, Zhang CJ, Zhang XY (2018) Genome-wide analysis of GRAS transcription factor gene family in Gossypium hirsutum L. BMC Genom 19:348. https://doi.org/10.1186/s12864-018-4722-x
Acknowledgements
This work was supported by the National Basic Research Program of China (973 Program; 2015CB150201); the National Natural Science Foundation of China (31671727 and 31471528), the National Key Research and Development Program of China (2018YFD1000900), and the ‘111’ Project of China (B12006).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Pengcheng Guo and Jing Wen contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Guo, P., Wen, J., Yang, J. et al. Genome-wide survey and expression analyses of the GRAS gene family in Brassica napus reveals their roles in root development and stress response. Planta 250, 1051–1072 (2019). https://doi.org/10.1007/s00425-019-03199-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-019-03199-y