Abstract
Exercise training which meets the recommendations set by the National Physical Activity Guidelines ensues a multitude of health benefits towards the prevention and treatment of various chronic diseases. However, not all individuals respond well to exercise training. That is, some individuals have no response, while others respond poorly. Genetic background is known to contribute to the inter-individual (human) and -strain (e.g., mice, rats) variation with acute exercise and exercise training, though to date, no specific genetic factors have been identified that explain the differential responses to exercise. In this review, we provide an overview of studies in human and animal models that have shown a significant contribution of genetics in acute exercise and exercise training-induced adaptations with standardized endurance and resistance training regimens, and further describe the genetic approaches which have been used to demonstrate such responses. Finally, our current understanding of the role of genetics and exercise is limited primarily to the nuclear genome, while only a limited focus has been given to a potential role of the mitochondrial genome and its interactions with the nuclear genome to predict the exercise training-induced phenotype(s) responses. We therefore discuss the mitochondrial genome and literature that suggests it may play a significant role, particularly through interactions with the nuclear genome, in the inherent ability to respond to exercise.
Similar content being viewed by others
Introduction
Regular daily exercise has multiple beneficial health outcomes including reductions in risk for cardiovascular disease (Manson et al. 1999), diabetes (LaMonte et al. 2005), several forms of cancer (Campbell and McTiernan 2007), stroke (Alevizos et al. 2005), neuro-cognitive dysfunctions (Verghese et al. 2003), all-cause mortality rates (Iestra et al. 2005), quality of life (Belardinelli et al. 1999), and lifespan (Moore et al. 2012; Paffenbarger et al. 1986). Such exercise-induced health benefits are due in part to solitary and combined effects of endurance and resistance training regimens which meet recommendations set by the National Physical Activity Guidelines Committee (2008). However, responses to exercise training are not consistent among all individuals, as some individuals respond well to training, while others respond poorly, even when accounting for factors including age, sex, and ethnic origin (Bouchard and Rankinen 2001; Bouchard et al. 1999). This known inter-individual variation in response to exercise training has primarily been characterized by aerobic capacity (VO2max) for endurance training (Bouchard et al. 1999; Pérusse et al. 2001), and by strength and muscle mass measurements for resistance training (Wilmore et al. 1998). With these measures, genetic background has been determined as a strong contributor to inter-individual variation in such measures of endurance and resistance training-induced adaptations. However, our current knowledge regarding links between genetics and inter-individual variation in response to exercise training is limited. The purpose of this review is to bring forth limitations in the literature, and discuss approaches that could be employed to circumvent such limitations.
One key limitation is the lack of a defined phenotype(s) for endurance and resistance training-induced responses, due largely to use of inconsistent research methods among investigators (i.e., modes, frequency, intensity, and duration of study protocols). Furthermore, it is becoming increasingly understood that phenotypes which define exercise training-induced adaptations are complex with numerous structural, functional, physiological, and molecular changes that occur in multiple organs (e.g., heart, skeletal muscle) and systems (cardiovascular, muscular) in response to exercise training. Thus, the complexity of physiological and molecular changes occurring with endurance and resistance training stresses the importance of establishing collaborative efforts among investigators which have expertise in the known phenotypic adaptations with endurance and resistance training.
Another limitation is identification of an appropriate animal modeling system that mimics heterogeneity in humans. Given the need to access specific tissues and organs (e.g., brain, heart) and the need to control confounding variables (e.g., diet, environment, background genetics/epigenetics), animal models remain necessary. The National Institutes of Health recognized this form of “bidirectional translation” (Rubio et al. 2010) as the ideal design for “a highly effective translational research program.” Indeed, multiple mammalian animal models have a long history of being used to understand basic principles of exercise physiology (e.g., Davis et al. 2014, Epp et al. 2006; Ferguson et al. 2014; Kao 1956; Poole et al. 2007; Poole and Erickson 2011; Roberts et al. 2014; Schaeffer et al. 1996) and responses to repeated exercise (e.g., Favier et al. 2016; Firshman et al. 2015; Kuster et al. 2014; Massett et al. 2015). These models are adequate to model exercise responses using molecular techniques; however, it is critical that the model simulates and controls the human genetic heterogeneity underlying variable responses to exercise so that conclusions derived are both translatable and generalizable.
The final section of this review will discuss briefly the mitochondrial genome and its interactions with the nuclear genome, and how different elements of the mitochondrial genome may contribute to the genetic regulation of exercise training adaptations. Mitochondria are cellular organelles responsible for sensing and initiating a multitude of signaling processes involved in energy metabolism, and the function of the mitochondria in eliciting such responses is largely dependent on the mitochondrial genome. Thus, since exercise is a metabolic stressor to the mitochondria, it is likely that the mitochondrial genome may contribute at least partially to the genetic regulation of exercise training-induced adaptations. To date, our primary understanding of genetics and exercise training-induced associations are studied from the nuclear genome, and with only a limited insight to the role of the mitochondrial genome. However, there are multiple aspects of the mitochondrial genome including heteroplasmy and copy number which could have significant influences on an individual’s response to exercise training.
Animal models
For a variety of reasons, including ethical, research design, and systematic control concerns, it is difficult to use humans to examine exercise training alterations in many molecular systems as well as most organ systems. Thus, use of mammalian animal models provide a means to control for a wide variety of research design and environmental factors to investigate the genetic contribution to exercise training-induced adaptations, as well as allowing access to all organ systems. With animal models, however, genetic heterogeneity which mimics that of humans, is an important factor and is a concern for future studies that investigate underlying genetic contributors to exercise training adaptations. Several animal models mimic the heterogeneity of humans, though major limitations exist in most models that complicate the ability to identify underlying genetic factors. Herein, we discuss genetic models that have been used to evaluate the link between genetics and response to exercise training, and end with a discussion of relatively new models that could be used to translate findings between animals and humans.
Inbred strains
Inbred strains (pigs, mice, and rats) provide cohorts of animals that are genetically homozygous, and thus provide the ability to interrogate specific genomes. Further, when multiple inbred strains are tested (i.e., “strain-screen”) for any phenotype, investigators then interrogate the amount of between-strain variation in characteristics and importantly, initial indications of the genomic location associated with that phenotype/trait [i.e., quantitative trait locus (QTL)]. Inbred mouse models have been used extensively to investigate inherent physical activity (Lightfoot et al. 2008, 2010), inherent aerobic capacity (Courtney and Massett 2012, 2014; Lightfoot et al. 2001, 2007), and inbred mouse and rat models have been developed for endurance exercise training (Massett et al. 2009, 2015; Massett and Berk 2005; Koch et al. 2013). However, a weakness of inbred models is the concern that since many of the classic inbred mouse and rat strains arose from common progenitor strains, large regions of the genomes from different inbred strains are actually the same [i.e., identity by descent (IBD)] and thus IBD issues prevent the examination of the full range of genomic variability with a particular trait. For example, within 40 classical inbred mouse lines, it is estimated that IBD ≈ 20% (Roberts et al. 2007; Yang et al. 2011) indicating at least 20% of the genome is not interrogated when using classical inbred strains. To our knowledge, no current estimates of IBD among rat inbred strains exist; however, a recent study compared two related rat inbred lines and estimated levels of IBD approaching 87% (Bell et al. 2011). Thus, higher IBD between inbred strains causes less genomic variation to be interrogated.
Selectively bred animal model
The selectively bred animal model has been used extensively to investigate high-physical activity level in mice (Garland et al. 2011; Garland and Kelly 2006; Swallow et al. 1998b) and rats (Roberts et al. 2013, 2014), inherent endurance exercise capacity in rats (Barbato et al. 1998; Britton and Koch 2001; Koch et al. 1998; Koch and Britton 2001), and endurance exercise training in rats (Koch et al. 2013). Selective breeding generally focuses on identified animals that are high responders and/or low responders in an outbred population, and then breeding those animals within trait (i.e., high-with-high, or low-with-low) for multiple generations. This approach has been successful for a mouse model of high activity [> 127 articles since 1998 (Swallow et al. 1998b)], a rat model of high- and low-aerobic capacity [> 138 articles (Koch et al. 1999)], and a rat model of low- and high-physical activity (Roberts et al. 2013, 2014). Most recently, Xu and Garland (2017) demonstrated in four distinct mouse lines selected as high runners, that each line had differing sets of QTLs associated with activity to suggest that each selected line evolved different physiological approaches to wheel-running activity. As powerful as the selective breeding approach is, at its culmination it examines only one or two inbred strains, and thus has limited genomic investigatory capability. Further, it has been suggested that selectively bred models may be prone to “artificial selective sweeps” in which genome sequences in close proximity to phenotypic alleles evolve, leading to expanding and non-informative genomic information (Atanur et al. 2013).
Recombinant inbred strains
Recombinant inbred strains (RI) are developed from an initial cross of two inbred strains (F0) followed by a cross of their offspring (F1) and then 20 generations of brother–sister mating to develop new inbred lines (usually 15–35 lines; Zou et al. 2005) to capture genetic variability of the two founder strains (Bailey et al. 1971). The strength of using RI strains is the capability to perform genetic mapping among several traits and across varying environmental conditions. To date, no studies have utilized RI strains to determine or confirm genetic contributors to either endurance or resistance training-induced adaptations. While RI strains present a useful model towards increasing our understanding of the genetic determinants of exercise training adaptations, a weakness of RI strains is the usual small number of RI lines available and potentially limited phenotypic differences between the originating two founder lines since, on average, any pair of classical mouse strains is IBD across ≈ 50% of their genomes (Roberts et al. 2007). This weakness has been overcome by development of the Collaborative Cross (CC) mouse model (see below) where underlying genetic architecture is controlled, yet is more representative of the genomic variability present in humans.
Consomic and conplastic inbred strains
A consomic mouse model involves a chromosome substitution from one inbred strain to another (http://www.informatics.jax.org/mgihome/nomen/strains.shtml). An obvious advantage of consomic strains is that they can be utilized as an additional confirmatory measure of strain screen analyses that indicated significant QTLs for a specific trait. However, as with other inbred models, the minimal heterogeneous background is a limitation. Conplastic inbred mice were developed with the intention to analyze interactions between mitochondrial and nuclear genomes (http://www.informatics.jax.org/mgihome/nomen/strains.shtml). Conplastic strains are created by backcrossing the nuclear genome from one inbred strain into the cytoplasm of another (http://www.informatics.jax.org/mgihome/nomen/strains.shtml). Currently, the conplastic approach is the only breeding model that provides an opportunity to analyze interactions between the two genomes. Likewise, the heterogeneity that is needed to develop a bidirectional translation approach between animals and humans is limited. Congenic and consomic inbred strains are two additional rodent models (rats and mice) that have been utilized to determine specific gene regions (congenic) and chromosomal (consomic) contribution to specific phenotypes (Matin et al. 1999).
Collaborative cross mouse model
The CC mice were developed to unravel the genetic architecture of complex traits and behaviors for translation into human models. In short, the CC mice are a large panel of multi-parent RI strains derived from eight founder strains representing the three major mouse subspecies (A/J, C57Bl/6J, 129Sv/ImJ, NOD/LtJ, NZO/H1J, CAST/EiJ, PWK/PhJ, and WSB/EiJ) that capture approximately 90% of the known genetic variation in laboratory mice with the captured variation being randomly distributed across the genome (Roberts et al. 2007; Churchill et al. 2004; Collaborative Cross Consortium 2012). Subsequent characterization has shown that the eight founder lines contributed equally (11.4–13.5%) to the available CC RI lines (Aylor et al. 2011). All current CC mouse lines have been genotyped and their genome sequences imputed (http://csbio.unc.edu/CCstatus/index.py), providing reference to the genetic composition of each mouse. Additionally, CC mice, as opposed to the available inbred mouse and rat strains, were developed using controlled randomization of genetic factors that allows expression of hidden phenotypes (Rogala et al. 2014). Further, F1 crosses of CC lines (CC-RIX lines; Zou et al. 2005) have a statistically significant reduction of within-line phenotypic variability relative to their inbred parents (Zou et al. 2005), support the detection of allele-specific expression differences (Crowley et al. 2015), and importantly model the heterozygous structure of the human genome. Thus, the CC-RIX model could present: (1) genome-wide variation so that all components of exercise-associated systems are interrogated (Collaborative Cross Consortium 2012; Threadgill and Churchill 2012); (2) randomization of genetic variation with no minor alleles so that causative relationships can be identified (Collaborative Cross Consortium 2012); (3) sufficient number of lines to power analyses (Zou et al. 2005, 2014); (4) reproducibility, data integration, robust reproduction, and future use in mechanistic studies (http://csbio.unc.edu/CCstatus/index.py); and (5) a unique platform upon which to determine novel gene functions (Rogala et al. 2014; Bottomly et al. 2012; Ferris et al. 2013; Rasmussen et al. 2014).
Diversity outbred mouse model
Complementing the CC mice is the diversity outbred (DO) mouse breeding system. DO mice are derived from the same eight founder strains of the CC mice; however, the generation of DO mice begins with a randomized and strict breeding scheme of the CC mice so that each DO mouse is genetically unique (Schmidt 2015; Churchill et al. 2012). The DO and CC mouse model schemas can be used in concert with one another to identify and confirm genes governing a specific trait such as with exercise training adaptations. The DO mice, like the CC mice, could be used to translate findings between mice and humans. At this point, however, definitive exercise-related phenotypes must be identified before conducting exercise training studies with DO mice.
We therefore propose that CC and/or DO mouse models be considered in future studies investigating the heritability of exercise training-induced adaptations. While relatively new, the CC model has been successfully used to mimic several human diseases (Bottomly et al. 2012; Ferris et al. 2007) as well as some exercise traits like voluntary wheel-running and energy balance (Mathes et al. 2011). Thus, employing a model like the CC mice, would enable new horizons in the investigation of exercise training adaptations.
The CC (Threadgill and Churchill 2012; Threadgill et al. 2002) and DO (Churchill et al. 2012) mouse models mimic genetic variation in humans (Roberts et al. 2007) and have been used to dissect energy regulation pathways in the healthy condition (Mathes et al. 2011). Some studies suggest translation to humans using the CC mouse model in diseases including influenza (Bottomly et al. 2012; Ferris et al. 2013) and Ebola (Rasmussen et al. 2014). Similarly, the DO mouse model also has suggested translational properties between mice and humans in various drug toxicity-related studies (Harrill 2016; Church et al. 2015), and some have shown that genetically diverse mice can be used to identify phenotypic markers that are unidentifiable in standard inbred mouse models. Thus, CC and/or DO models provide an approach for bidirectional translation between humans and mice, and could be utilized in studies to examine exercise-induced adaptations. Other animal models have been used to investigate exercise-related phenotypes; however, such models pose major limitations that can be overcome with the CC and/or DO mouse models.
Exercise training-induced inter-individual variation in human and animal models
A weakness in the literature is that animal studies have not been designed to match exercise training designs/responses from human studies. While separate studies in animals and humans provide significant and suggestive genetic factors (QTLs, genes) with exercise training-induced responses, such findings are not translatable between animals and humans given the inconsistencies in training designs and phenotypic measures. Thus, a phenotype must be defined for exercise training-induced adaptations that translate between animal models and humans.
Endurance exercise training
Overview
A wide variety of general systemic and organ responses to endurance training in humans have been well documented over the past 70–80 years. In particular, changes in aerobic capacity (VO2max) and extensive changes in metabolic enzymes such as succinate dehydrogenase (SDH), phosphofructokinase (PFK), and hexokinase, along with several other biomarkers, are often used as direct indices of successful adaptation to repeated endurance exercise exposures. Furthermore, numerous structural adaptations in various tissues and organs (e.g., heart, skeletal muscle, vasculature), in addition to functional changes at the cellular and molecular level in systems, respond and adapt to endurance exercise training.
In general, the average change to a treadmill training program in VO2max is approximately 24–25% (Bouchard and Rankinen 2001; Kohrt et al. 1991) in young and older populations without significant influence of weight or sex on the training response. These changes in aerobic capacity are generally indicative of alterations in central (cardiac output) and peripheral (muscle metabolism) factors, and there are well-known alterations in muscle/mitochondrial enzyme levels, capillary density, and increases in ventricular end diastolic volume and contraction strength to increase ventricular stroke volume. At least 50% of these adaptation responses to endurance training are heritable (Bouchard et al. 1999). This genetic influence is clearly shown when considering the broad range of responses to a controlled endurance training program (summarized in Table 1). Thus, it is critical that any attempt to understand the molecular mechanisms responsible for adaptation to endurance training consider underlying genetic architecture so that the widest range of potential causative mechanisms can be captured.
Human studies
In humans, the HERITAGE (HEalth, RIsk factors, exercise Training And Genetics) study has been the primary source of our current understanding of the inter-individual variation in response to endurance exercise training. The hallmark investigation of the HERITAGE study was a twin study of otherwise healthy untrained male identical brothers (Prudhomme et al. 1983). This study found that significant variations in aerobic capacity improvement between pairs were six to nine times greater than within identical twin pairs. The findings were then replicated in four different centers in the U.S. (Indiana University, University of Minnesota, and Texas A&M University) and Canada (Laval University in Quebec). Each center found a similar spectrum of responses in aerobic capacity improvement, where there remained approximately 15% of individuals who responded very little (non-responders), and then another approximate 15% of individuals who responded well to training (high responders) (Bouchard and Rankinen 2001). Another key finding from the HERITAGE study is that family members generally had similar responses to training, where an individual’s ability to respond to endurance training was approximately 50% heritable. The other 50% of an individual’s ability to respond to training is primarily attributed to unique environmental factors, though age, sex, and ethnic origin are not determinants of the training response (Bouchard and Rankinen 2001). The most recent HERITAGE study (Bouchard et al. 1985) indicated that 21 single nucleotide polymorphisms (SNPs) accounted for 49% of the variation in VO2max with training, where ≤ 9 alleles associated with low responders and ≥ 19 alleles associated with high responders. The strongest associations were found in genes including ACSL1, PRDM1, GRIN3A, KCNH8, and ZIC4 (Bouchard et al. 1985).
Contrary to the conclusions from the HERITAGE study that inherent inter-individual variability exists in response to endurance training, is recent discussion that the observed ‘non-response’ is a result of flawed training designs, where some investigators argue that the “non-response” to training is abolished with increased dose of exercise (duration and intensity of training; Montero and Lundby 2017; Lundby et al. 2016; Bonafiglia et al. 2016). For example, a recent study (Montero and Lundby 2017) conducted two-successive 6-week endurance training programs, where subjects were assigned to one of six groups based on the total minutes of exercise per week: 60, 120, 180, 240, and 300 min/week of exercise. The study concluded that, following the first 6 weeks of training, there were 69, 40, and 29% “non-responders” in each respective training group. However, following a second 6-week training session, no “non-responders” remained suggesting that increasing the dose of exercise for the previously classified “non-responders” may be the remedy to prompt a response to increase aerobic capacity with training. A study by Bonafiglia et al. (2016) supports the claim of abolishing non-responders by increasing training. In this study, it was found through a randomized crossover study design of two training protocol types consisting of endurance training (30 min/day at 65% VO2peak) and sprint interval training (eight, 20-s intervals at 170% VO2peak separated by 10-s active rest), the individual response to training was dependent on the training protocol utilized. While there was inter-individual variation within each training protocol, all individuals responded to at least one training protocol to suggest that the “non-response” may be decreased by changing the training stimulus. In totality, these studies suggest that the “non-response” to endurance exercise is abolished through increasing the training stimulus. These findings are interesting, but considerably larger sample sizes are needed to confirm this argument given that there were only 21 and 78 subjects in each study, respectively (Montero and Lundby 2017; Bonafiglia et al. 2016), as well as a lack of diversity of subject background (both sex and ethnicity). Comparatively, the HERITAGE study was conducted across four major institutions and with over 800 individuals representing a wide range of diversity in age, sex, and ethnic background. Thus, the issue of whether “non-responders” exist, when measured by aerobic capacity, to endurance exercise training is not resolved, and deserves further attention.
Rodent studies
Endurance training results similar to humans have been observed using mouse models. For example, Courtney and Massett (2012) and Lightfoot et al. (2001) performed strain screens of acute response to endurance exercise in 34 (Courtney and Massett 2012) and 10 (Lightfoot et al. 2001) inbred mouse strains which demonstrate inter-strain variation in exercise capacity. Other studies have reported (Massett et al. 2009, 2015; Massett and Berk 2005) adaptation to endurance training among several inbred strains as well as several F1, F2, and backcross populations. These results have been expanded recently to include 25 inbred strains, and found a change in endurance capacity after training ranging from − 5.1 ± 3.9% to 20.0 ± 7.3%. Interestingly, when considering the strain screen analysis of inherent endurance exercise capacity by Massett and Berk (2005), the variation in the mouse model is similar to the inter-individual variation in humans (Bouchard et al. 1999). Additionally, when presented as familial data as opposed to individual data, the human familial variation data from Bouchard et al. (1999) and Bouchard and Rankinen (2001) were similar to that found between inbred strains of mice (Massett et al. 2009, 2015; Courtney and Massett 2012; Massett and Berk 2005), with a similar degree of heritability (interclass correlations approximately 65%; coefficient of genetic contribution approximately 47%; Massett and Berk 2005).
The key to successful translatability of results from animal models to humans is to equate the training programs. A weakness of extant rodent literature regarding endurance training is the wide range of training programs used, especially in the duration and intensity of times used. One critical factor listed by the American Physiological Society (Kregel et al. 2006) when designing exercise protocols for animals is animal safety. An ongoing concern is that some protocols (Kemi et al. 2002) may be too long in duration and/or too high in intensity setting leading to less adherence and increased risk of injury and overtraining in the animal subjects. The majority of protocols reported (Massett et al. 2009, 2015; Courtney and Massett 2012; Lightfoot et al. 2001; Massett and Berk 2005; Desai et al. 1997; Petrosino et al. 2016) have intensities and durations more translatable to humans. For example, the training protocols for the mouse endurance model by Massett and colleagues (Massett et al. 2009, 2015; Massett and Berk 2005) were designed to align with the standard, currently recommended human endurance training protocols using a duration of 30 min or more (with 60 min preferred), an intensity between 40–75% VO2max, and a frequency of 3–6 days/week (Pescatello and American College of Sports Medicine 2014).
Exercise-induced adaptations with endurance training vary by the modes of training utilized (e.g., treadmill running, swimming, cycling), the length of training time (weeks; months), and the frequency, duration, and intensity of training programs. Exercise training in rodents has generally consisted of methods to induce continuous aerobic exercise. Treadmills, swimming, and running wheels have been used in training studies in rats and mice (Kregel et al. 2006; Kilikevicius et al. 2013). Acute exercise responses in mice on a treadmill are similar to what is seen with humans (Acosta et al. 2017). Given mice are quadrupeds and humans are bipeds, raises the concern whether treadmill training in mice induces physiological adaptations similar to those found in humans. Therefore, some studies have instead used swimming protocols to mimic similar movements. Swimming, however, does not appear to be an adequate model for continuous aerobic exercise in rodents because some work suggests that swimming protocols for rodents causes a series of hypoxic exposures due to floating and bobbing motions by most rodents when they swim (Kregel et al. 2006). With treadmill aerobic capacity testing, Desai et al. (1997) have shown in four inbred and two outbred mouse strains that a linear increase in heart rate, VO2, VCO2, and RER occurs with an increase in exercise intensity indicative of an escalation in metabolic cost of workload, which is similar to humans. These effects have been observed in several other studies that have measured responses to treadmill running in mice (Fluckey et al. 1996a, b). Thus, while locomotive behavior for running in mice and humans are different, the physiological responses that occur are similar. Therefore, treadmill training is the most applicable mode of training for bidirectional translation of findings between mice and humans.
Resistance training
Overview
A wide literature base exists regarding muscular adaptations from repeated exposure to resistance training in humans. Measures of adaptation in humans involve non-invasive measures including increased strength and muscle size alterations, and direct measures including muscle fiber size changes and contractile mechanism alterations. Additionally, while investigators have long noted inter-individual variation in response to resistance training (Fluckey et al. 1996b), few studies evaluated genetic influence on adaptation to resistance training.
Human studies
Thompson et al. (2004) designed the “Functional Single Nucleotide Polymorphisms Associated with Human Muscle Size and Strength” (FAMuSS) study where they examined responses of ≈ 1300 young adults (men and women) to a 12-week, standardized isometric and dynamic (both concentric and eccentric contractions) resistance training program on the non-dominant arm. As part of the FAMuSS study, in 585 subjects (342 women, 243 men), Hubal et al. (2005) found that participants increased mean (± SEM) muscle size by 18.9 ± 0.4%, increased isometric strength by 19.5 ± 0.8%, and increased 1 repetition max (1RM) strength by 54.1 ± 1.4%. Unlike endurance training, sex appeared to affect the amount of change in strength and size (Hubal et al. 2005). Women had greater percentage increases in isometric strength and 1RM strength compared to males, while males increased muscle size significantly more than females. Further, similar to endurance training (Thomis et al. 1998), Hubal et al. (2005) found extensive inter-individual variation in responses to resistance training, with populations of responders and non-responders in each sex for cross-sectional area, 1RM strength gain, and isometric strength. A limitation of this extensive study was its consideration of a fairly homogenous population of relatively young subjects (ages 18–40 years), with primarily European-American backgrounds. However, while FAMuSS may be limited, it is the only existing population-based resistance training study in either humans or animals.
Rodent studies
In general, there are fewer models of animal resistance training than endurance training. While agriculture has consistently selectively bred livestock for muscle size (especially work horses and beef cattle) over the past several hundred years, mutations targeted in livestock (e.g., myostatin and IGF-2) have not been found to support any of the suggested QTL from the FAMuSS study (Thompson et al. 2004). This lack of support suggests that muscle size and strength as a result of selective breeding and the size and strength adaptations to resistance exercise exposure potentially arise from different molecular pathways.
While no animal resistance training genetic studies exist, multiple acute resistance training models have been developed for rodents (Kregel et al. 2006; Lowe and Alway 2002; Seo et al. 2014). The primary species in these studies are rats, and outcome variables include measurement of muscle mass, fiber cross-sectional area, contractile function, and fractional protein synthesis rates (Bodine and Baar 2012). The ‘jump-squat’ model of resistance training has been used in several studies (Burgess et al. 1993; Klitgaard 1988; Nicastro et al. 2012; Tamaki et al. 1992; Wirth et al. 2003), and is currently the method of choice for maximal translation to human modes (Kregel et al. 2006; Lowe and Alway 2002). Briefly, the jump-squat protocol in rodents involves operant conditioning to avoid a brief foot shock (< 1 mA, 60 Hz) by extending hind limbs fully to press an illuminated bar repeatedly. After conditioning (3–4 training sessions), rats usually do not need shock to engage in resistance training (Fluckey et al. 1995). Following operant conditioning, animals jump-squat in training sessions with prorated weight vests. Thus, training programs control the amount of resistance and number of reps similar to human resistance training programs. In general, with 8–36 weeks of exposure in rats, there is an average 10–30% increase in muscle mass, a 10–30% increase in cross-sectional area, and a 10–60% increase in force production (Lowe and Alway 2002). These responses are similar to those found in humans by Hubal et al. (2005) in the FAMuSS study. Furthermore, rates of protein synthesis are significantly increased in both gastrocnemius and soleus muscle with either short-term (four training session in young, middle-aged, and old rats; Fluckey et al. 1996a) or long-term (8 weeks of training; Farrell et al. 1999) exposures.
These studies suggest that weight-lifting types of exercise are analogous to human resistance training programs and have found similar muscle-physiological adaptation compared to human programs. A weakness of weight-lifting types of exercise is that the exercise is bilateral, removing the ability to have an internal control (Watt et al. 1982), as present in FAMuSS (Hubal et al. 2005). This weakness could be overcome using CC and/or DO mouse models that control for the underlying genetic architecture so that independent control animals can be used that do not undergo resistance training. Lastly, while rodent models can mimic general human muscle adaptation to repeated resistance training programs, a weakness of most of these studies is that they exclusively used male animals. Thus, future research utilizing both sexes is needed.
Potential role of the mitochondrial genome and its interactions with the nuclear genome
Overview
To enable a complete understanding for how genetic background contributes to exercise training adaptations, it is important to also consider the role of the mitochondrial genome. Few studies have investigated links between the mitochondrial genome and endurance training adaptations (Dionne et al. 1991; Rivera et al. 1998; Lihong 2000; Chen et al. 2000; Bray et al. 2009). However, conclusive findings from such studies are limited because they focus only on certain portions of the mitochondrial genome. Recent advances in technologies, including Next-generation Sequencing, have enabled complete and ultra-deep sequencing of mitochondrial DNA in animal and human tissues that contain mitochondria. Furthermore, ultra-deep sequencing capabilities have enabled discovery of other important elements of the mitochondrial genome, including heteroplasmy and insertions/deletion (indels) that could also have significant roles in acute exercise and adaptations to training. To date, the role of the mitochondrial genome and its interactive effects with the nuclear genome on adaptive phenotypes of exercise training are not fully understood. The mitochondrial genome also presents an additional level of complexity due to cell-to-cell variation in mitochondrial DNA copy number that is dependent upon the energy requirement of a cell (Taylor and Turnbull 2005). Since relatively new techniques have allowed for accurate measurement of mitochondrial DNA copy number and heteroplasmy, such factors have not been fully explored. Taken together, studies are needed to assess their association with inherent exercise training adaptations in endurance and resistance training.
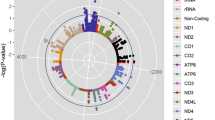
Briefly, mtDNA is circular and spans approximately 16,500 DNA-building base pairs that make up 37 genes: 13 protein-coding genes provide instructions for making enzymes involved in oxidative phosphorylation, 2 transfer RNAs (tRNA), and 22 ribosomal RNAs (rRNA) (Chinnery and Hudson 2013; Schon et al. 2012). One unique characteristic of the mitochondrial genome compared to the nuclear genome is that few to hundreds of thousands of mtDNA copies may exist per mitochondrion. With multiple copies of mtDNA within mitochondria, sequence variants can result due to various factors such as maternal mutation(s) passed to offspring, age, diseases, and other environmental exposures. Such variants in mtDNA lead to a heterogeneous population of mtDNA called heteroplasmy. To date, no study has considered the contribution of mtDNA copy number and/or heteroplasmy as it relates to exercise training-induced adaptations.
The mitochondrial genome is of particular interest with regard to exercise training adaptations because of the crucial role of mitochondrial DNA in regulating overall mitochondrial function. A major function of the mitochondrial genome is to regulate mitochondrial function [the production of energy in the useful form of Adenosine Triphosphate (ATP), and also respond to production of reactive oxygen species (Voet et al. 2013)]. There are also approximately 1,300 nuclear genes that interact with mtDNA and encode mitochondrial proteins (Andersson et al. 2003). Thus, mitochondrial function is dependent on mitochondrial and nuclear DNA interactions. With exercise training, the mitochondrial responses are critical to enable increased aerobic capacity as well as increased muscle strength and size with endurance (Brearley and Zhou 2001) and resistance training (Huffman et al. 2014), respectively. The inability of the mitochondria to respond and adapt in a positive manner to exercise training would inhibit an individual from responding to exercise, and in some cases, potentially decrease the ability to respond to exercise if mitochondrial damage ensued. One major source of mitochondrial DNA damage is through production of reactive oxygen species. With endurance training, aerobic capacity (VO2max)—the primary indicator of how well individuals respond to endurance training—is partly dependent upon how well the mitochondria respond to exercise-induced reactive oxygen species production, and their adaptability to increase efficiency of ATP production (e.g., induce mitochondrial biogenesis). Functions and adaptations of the mitochondria are heavily dependent upon the mitochondrial genome. Because mitochondrial DNA regulates overall mitochondrial function, and mitochondrial function significantly contributes to an individual’s aerobic capacity, investigators have hypothesized that the mitochondrial DNA sequence is associated with the inherent ability to respond to exercise training (Dionne et al. 1991; Scott et al. 2005). In fact, in the 2006–2007 update of the “Human Gene Map for Performance and Health-Related Phenotypes” (Bray et al. 2009), 18 mitochondrial genes were shown across various studies to influence fitness and performance phenotypes.
Mitochondrial DNA sequence variants
Several mitochondrial DNA sequence variants have been associated with exercise intolerance (Hirano et al. 2001; Goethem et al. 2001; Zeviani et al. 1989; Wallace 1992). However, most studies focused on individuals with a mitochondrial DNA mutation-induced diseases that severely inhibit exercise. Such studies may provide insight into specific locations of the mitochondrial genome that influence an individual’s response to exercise training. Using the restriction fragment length polymorphism (RFLP) technique, Dionne et al. (1991) compared mtDNA sequence polymorphisms in the untrained and trained states. Of the mtDNA variants considered, the authors found the subunit 5 of the NADH dehydrogenase gene and one in the tRNA for threonine, had a significantly higher VO2max in the untrained state when compared to non-carriers, while carriers of one mitochondrial DNA morph in subunit 2 of NADH dehydrogenase had a lower initial VO2max. A lower response to endurance training via cycle ergometry was found for three carriers of a variant in subunit five of the NADH dehydrogenase. Thus, while the RFLP technique is limited in terms of fully uncovering the contribution of mtDNA sequence and polymorphisms to the inter-individual responses to exercise training, this study did indicate that the mitochondria genome may have a role in determining VO2max responses which warrants further investigation. Challenging this previous finding, Scott et al. (2005) considered mtDNA haplogroups in a general Ethiopian population and an elite athlete runner Ethiopian population. The study found that the groups did not differ in terms of their mtDNA haplogroup, and therefore hypothesized that running ability was not determined by mitochondrial DNA polymorphisms.
While mitochondrial DNA haplogroups did not associate with exercise training adaptations in this study, there are other important elements of the mitochondrial genome that should be considered, including mtDNA copy number. At the cellular level, mitochondrial biogenesis is a key contributor to increased aerobic capacity with exercise training, and mtDNA copy number has been shown to increase concomitantly with exercise-induced mitochondrial biogenesis occurring in the skeletal muscle with exercise training (Puntschart et al. 1995; Trounce 2000). Thus, there could be significant influential effects of mtDNA replication in each mitochondrion that initiate mitochondrial biogenesis.
There are, however, several studies that support the notion that the mitochondrial genome significantly contributes to inherent exercise capacity (Chow et al. 2007; Robinson et al. 2017). Chow et al. (2007) demonstrated that aerobic exercise training enhances muscle mitochondrial transcription factors, mtDNA abundance, mitochondria-related gene transcript levels, and mitochondrial function. Additionally, the authors suggested that the enhancement in mitochondrial function occurs in association with increased spontaneous physical activity (Chow et al. 2007). Robinson et al. (2017) examined different types of exercise training, and demonstrated an effect of intensity on mitochondrial respiration. When comparing continuous aerobic training to “high intensity interval training” (HIIT), it was found that HIIT induced the greatest changes in gene expression and mitochondrial respiration. Resistance training was also examined, and unlike the endurance exercise training protocols, it did not induce any changes to factors related to mitochondrial function or amount. Another study by Robinson et al. (2017) found after 12 weeks of resistance training, young and old individuals had no changes in mitochondrial function measurements, which also indicated that the mitochondrial genome is unlikely involved with the resulting training adaptations. The authors further investigated factors related to intrinsic mitochondrial function including coupling efficiency and reactive oxygen species production, as well as mitochondrial protein abundance, and the only change with training was mitochondrial protein abundance. That is, the amount and not function, of the mitochondria is a primary contributor to the exercise endurance training response. Thus, from the limited data available on the relationship between genetics and resistance training, it appears that the adaptations to resistance training are independent of the mitochondrial genome.
Protein 53 (p53)
The tumor-suppressor protein, protein 53 (p53) has a role in mitochondrial function and may determine an individual’s response to exercise training. p53 is encoded by the Trp53 gene in mice and TP53 gene in humans, and contributes to inherent aerobic capacity (Bouchard et al. 2015). p53 can regulate mitochondrial function through its role as a nuclear transcription factor, and also through its direct presence in mitochondria via translocation. Bouchard et al. (2015) reviewed such work describing pathways in which p53 participates in mitochondrial biogenesis and thus directly affecting exercise capacity. An important finding about p53, particularly when considering the potential role that the mitochondrial genome may play in determining exercise capacity, is that absence of p53 in mice (p53 knockout mice) results in significant depletion of mtDNA and consequently poor exercise capacity, both at baseline and with training (Matoba et al. 2006; Park et al. 2016; Saleem et al. 2009). In humans, p53 also influences recovery time with exercise. A study by Wang et al. (2013) showed that in individuals with the Li–Fraumeni syndrome, which have a mutation in the TP53 gene, had significantly faster recovery than non-carrier and healthy controls from a 2-min bout of exercise when assessed by phosphocreatine levels in skeletal muscle after exercise. While such studies suggest that genetic alteration to the Trp53 (mice) and TP53 (humans) gene has a necessary role in regulating responses to exercise training through its association with mitochondrial biogenesis and respiration, the precise mechanism through which this occurs is not fully understood (Bouchard et al. 2015).
Another important role of p53 in response to endurance exercise-induced training is to regulate mitochondrial polymerase gamma (POLG1). POLG1 is an enzyme considered to be the only mtDNA repair enzyme, and has been demonstrated to initiate mtDNA mutation repair and mitochondrial biogenesis in response to endurance exercise training (Safdar et al. 2016). By inducing skeletal muscle-specific deletion of p53 in mice, Safdar et al. (2016) showed that endurance training in these mice failed to prevent mtDNA mutations, induced mitochondrial biogenesis, preserved mitochondrial morphology, reversed sarcopenia, or mitigated premature mortality, when compared to controls. From the series of experiments by Safdar et al. (2016), p53 was shown to have a direct role on mtDNA mutation repair and induction of mitochondrial biogenesis. Taken together, the interaction between p53 and exercise capacity highlights a nuclear gene that not only has a direct influence on exercise capacity, but also provides a direct mechanistic pathway through which both the nuclear and mitochondrial genomes associate with exercise capacity.
Bioinformatic and statistical methods are still needed to analyze and determine the interactive and solitary effects of mitochondrial and nuclear genome associations with specific phenotypes, though there are available methods in mice that enable insight into these effects. As noted earlier, conplastic strains have been used to determine the contribution of the mitochondrial genome and its interactions with the nuclear genome, in a variety of phenotypes (http://www.informatics.jax.org/mgihome/nomen/strains.shtml). Another animal, known as the mitochondrial–nuclear eXchange Mice, is a model that allows the combination of the nuclear genome from one mouse strain, via oocyte enucleation, with mtDNA from a different mouse strain, a method similar to the well-known ‘3-parent babies’ (Kesterson et al. 2016). The mitochondrial–nuclear eXchange mice would be particularly interesting to use in mice previously characterized as having high, low, no response, and lowered responses to exercise training (e.g., Massett and Berk 2005; Kilikevicius et al. 2013). For example, determining the effect of mtDNA of a high responder strain on the nuclear genomic background of a low responder strain to exercise, and vice versa. Together with the recent techniques such as next-generation ultra-deep sequencing, such methods have enabled multiple avenues for future research regarding interactions between the mitochondrial and genomes that may associate with exercise training-induced phenotypes.
Conclusions and future directions
The preponderance of evidence that demonstrates an existing association between genetics and physical activity is drawn primarily from considering the nuclear genome while few efforts have considered the mitochondrial genome. From the human and mouse studies by Bouchard et al. (1999) and Massett et al. (2009, 2015), and Massett and Berk (2005), the function and proliferative response of mitochondria (i.e., surface area; density) in energy-demanding tissues (e.g., heart, skeletal muscle) are imperative to enable significant exercise training responses (e.g., increased VO2max). However, attention is yet to be directed on how the mitochondrial genome influences such adaptations. Furthermore, given that genetic background strongly contributes to exercise training-induced responses, and that other lines of work hint towards a significant contribution of the mitochondrial genome (e.g., mediation by p53; Safdar et al. 2016), it is of interest to know whether mitochondrial–nuclear interactions exist to predict mitochondrial adaptations/responses to exercise training.
In conclusion, we suggest that future work investigating the relationship between exercise and genetics consider employing animal models such as the CC or DO mice that are designed to control for the genetic diversity inherent in human models (Roberts et al. 2007; Threadgill and Churchill 2012; Threadgill et al. 2002, 2011). Given that such controlled models are already being successfully used to understand the genetic foundation of several human diseases (Bottomly et al. 2012; Ferris et al. 2013) as well as some exercise traits like voluntary wheel-running and energy balance (Mathes et al. 2011), the use of these models will enable new horizons in the investigation of exercise training adaptations. Furthermore, the CC animal modeling scheme could also be used to delineate the contribution of the mitochondrial genome and its interactive effects with the nuclear genome to exercise responses, given that the breeding strategy is designed in a way to determine solitary and combined associations with each genome.
References
Acosta W, Meek TH, Schutz H, Dlugosz EM, Garland T (2017) Preference for Western diet coadapts in High Runner mice and affects voluntary exercise and spontaneous physical activity in a genotype-dependent manner. Behav Process 135:56–65
Alevizos A, Lentzas J, Kokkoris S, Mariolis A, Korantzopoulos P (2005) Physical activity and stroke risk. Int J Clin Pract 59(8):922–930
Andersson G, Karlberg O, Canbäck B, Kurland CG (2003) On the origin of mitochondria: a genomics perspective. Philos Trans R Soc Lond B 358(1429):165–179
Atanur SS, Diaz AG, Maratou K, Sarkis A, Rotival M, Game L et al (2013) Genome sequencing reveals loci under artificial selection that underlie disease phenotypes in the laboratory rat. Cell 154(3):691–703
Aylor DL, Valdar W, Foulds-Mathes W, Buus RJ, Verdugo RA, Baric RS et al (2011) Genetic analysis of complex traits in the emerging Collaborative Cross. Genome Res 21(8):1213–1222
Bailey D (1971) Recombinant-inbred strains an aid to finding identity, linkage, and function of histocompatibility and other genes. Transplantation 11(3):325–327
Barbato JC, Koch LG, Darvish A, Cicila GT, Metting PJ, Britton SL (1998) Spectrum of aerobic endurance running performance in eleven inbred strains of rats. J Appl Physiol 85(2):530–536
Belardinelli R, Georgiou D, Cianci G, Purcaro A (1999) Randomized, controlled trial of long-term moderate exercise training in chronic heart failure. Circulation 99(9):1173–1182
Bell R, Herring SM, Gokul N, Monita M, Grove ML, Boerwinkle E et al (2011) High resolution identity by descent mapping uncovers the genetic basis for blood pressure differences between SHR lines. Circulation 4:223–231
Bodine SC, Baar K (2012) Analysis of skeletal muscle hypertrophy in models of increased loading. Myogenesis 2012:213–229
Bonafiglia JT, Rotundo MP, Whittall JP, Scribbans TD, Graham RB, Gurd BJ (2016) Inter-individual variability in the adaptive responses to endurance and sprint interval training: a randomized crossover study. PLoS ONE 11(12):e0167790
Bottomly D, Ferris MT, Aicher LD, Rosenzweig E, Whitmore A, Aylor DL et al (2012) Expression quantitative trait Loci for extreme host response to influenza a in pre-collaborative cross mice. G3: Genes Genomes Genet 2(2):213–221
Bouchard C, Rankinen T (2001) Individual differences in response to regular physical activity. Med Sci Sports Exerc 33(6 Suppl):S446–S451 (discussion S52–S53)
Bouchard C, An P, Rice T, Skinner JS, Wilmore JH, Gagnon J et al (1999) Familial aggregation of VO2max response to exercise training: results from the HERITAGE Family Study. J Appl Physiol 87(3):1003–1008
Bouchard C, Sarzynski MA, Rice TK, Kraus WE, Church TS, Sung YJ et al (2011) Genomic predictors of the maximal O2 uptake response to standardized exercise training programs. J Appl Phys 110(5):1160–1170
Bouchard C, Antunes-Correa LM, Ashley EA, Franklin N, Hwang PM, Mattsson CM et al (2015) Personalized preventive medicine: genetics and the response to regular exercise in preventive interventions. Prog Cardiovasc Dis 57(4):337–346
Bray MS, Hagberg JM, Perusse L, Rankinen T, Roth SM, Wolfarth B et al (2009) The human gene map for performance and health-related fitness phenotypes: the 2006–2007 update. Med Sci Sports Exerc 41(1):34–72
Brearley MB, Zhou S (2001) Mitochondrial DNA and maximum oxygen consumption. Sportscience 5(2)
Britton SL, Koch LG (2001) Animal genetic models for complex traits of physical capacity. Exerc Sport Sci Rev 29(1):7–14
Burgess M, Davis J, Wilson S, Borg T, Burgess W, Buggy J (1993) Effects of intracranial self-stimulation on selected physiological variables in rats. Am J Physiol Regul Integr Comp Physiol 264(1):R149–R155
Campbell KL, McTiernan A (2007) Exercise and biomarkers for cancer prevention studies. J Nutr 137(1):161S–169S
Chen Q, Ma L, Chen J (2000) Analysis on genetic polymorphism of mtDNA in endurance athletes and sedentary subjects. Chin J Appl Physiol 16(4):327–330
Chinnery PF, Hudson G (2013) Mitochondrial genetics. Br Med Bull 106(1):135–159
Chow LS, Greenlund LJ, Asmann YW, Short KR, McCrady SK, Levine JA et al (2007) Impact of endurance training on murine spontaneous activity, muscle mitochondrial DNA abundance, gene transcripts, and function. J Appl Physiol 102(3):1078–1089
Church RJ, Gatti DM, Urban TJ, Long N, Yang X, Shi Q et al (2015) Sensitivity to hepatotoxicity due to epigallocatechin gallate is affected by genetic background in diversity outbred mice. Food Chem Toxicol 76:19–26
Churchill GA, Airey DC, Allayee H, Angel JM, Attie AD, Beatty J et al (2004) The Collaborative Cross, a community resource for the genetic analysis of complex traits. Nat Genet 36(11):1133–1137
Churchill GA, Gatti DM, Munger SC, Svenson KL (2012) The diversity outbred mouse population. Mamm Genome 23(9–10):713–718
Collaborative Cross Consortium (2012) The genome architecture of the Collaborative Cross mouse genetic reference population. Genetics 190(2):389–401
Courtney SM, Massett MP (2012) Identification of exercise capacity QTL using association mapping in inbred mice. Physiol Genomics 44(19):948–955
Courtney SM, Massett MP (2014) Effect of chromosome substitution on intrinsic exercise capacity in mice. F1000Research 3
Crowley JJ, Zhabotynsky V, Sun W, Huang S, Pakatci IK, Kim Y et al (2015) Analyses of allele-specific gene expression in highly divergent mouse crosses identifies pervasive allelic imbalance. Nat Genet 47(4):353–360
Davis MS, Bonen A, Snook LA, Jain SS, Bartels K, Geor R et al (2014) Conditioning increases the gain of contraction-induced sarcolemmal substrate transport in ultra-endurance racing sled dogs. PLoS ONE 9(7):e103087
Desai K, Sato R, Schauble E, Barsh G, Kobilka B, Bernstein D (1997) Cardiovascular indexes in the mouse at rest and with exercise: new tools to study models of cardiac disease. Am J Physiol Heart Circ Physiol 272(2):H1053-H61
Dionne FT, Turcotte L, Thibault M-c, Boulay MR, Skinner JS, Bouchard C (1991) Mitochondrial DNA sequence polymorphism, VO2max, and response to endurance training. Med Sci Sports Exerc 23(2):177–185
Epp T, McDonough P, Padilla D, Gentile J, Edwards K, Erickson H et al (2006) Exercise-induced pulmonary haemorrhage during submaximal exercise. Equine Vet J 38(S36):502–507
Farrell PA, Fedele MJ, Hernandez J, Fluckey JD, Miller JL, Lang CH et al (1999) Hypertrophy of skeletal muscle in diabetic rats in response to chronic resistance exercise. J Appl Physiol 87(3):1075–1082
Favier FB, Britto FA, Ponçon B, Begue G, Chabi B, Reboul C et al (2016) Endurance training prevents negative effects of the hypoxia mimetic dimethyloxalylglycine on cardiac and skeletal muscle function. J Appl Physiol 120(4):455–463
Ferguson DP, Dangott LJ, Schmitt EE, Vellers HL, Lightfoot JT (2014) Differential skeletal muscle proteome of high- and low-active mice. J Appl Physiol 116(8):1057–1067
Ferris LT, Williams JS, Shen C-L (2007) The effect of acute exercise on serum brain-derived neurotrophic factor levels and cognitive function. Med Sci Sports Exerc 39(4):728–734
Ferris MT, Aylor DL, Bottomly D, Whitmore AC, Aicher LD, Bell TA et al (2013) Modeling host genetic regulation of influenza pathogenesis in the collaborative cross. PLoS Pathog 9(2):e1003196
Firshman AM, Borgia LA, Valberg SJ (2015) Effects of training at a walk on conventional and underwater treadmills on fiber properties and metabolic responses of superficial digital flexor and gluteal muscles to high-speed exercise in horses. Am J Vet Res 76(12):1058–1065
Fluckey JD, Kraemer WJ, Farrell PA (1995) Pancreatic islet insulin secretion is increased after resistance exercise in rats. J Appl Physiol 79(4):1100–1105
Fluckey JD, Vary TC, Jefferson LS, Evans WJ, Farrell PA (1996a) Insulin stimulation of protein synthesis in rat skeletal muscle following resistance exercise is maintained with advancing age. J Gerontol Ser A: Biol Sci Med Sci 51(5):B323-M30
Fluckey JD, Vary TC, Jefferson LS, Farrell PA (1996b) Augmented insulin action on rates of protein synthesis after resistance exercise in rats. Am J Physiol Endocrinol Metab 270(2):E313–E319
Garland T, Kelly SA (2006) Phenotypic plasticity and experimental evolution. J Exp Biol 209(12):2344–2361
Garland T, Schutz H, Chappell MA, Keeney BK, Meek TH, Copes LE et al (2011) The biological control of voluntary exercise, spontaneous physical activity and daily energy expenditure in relation to obesity: human and rodent perspectives. J Exp Biol 214(2):206–229
Harrill AH (2016) Mouse population-based toxicology for personalized medicine and improved safety prediction. Drug Discov Toxicol 314–329
Hirano M, Marti R, Ferreiro-Barros C, Vilà MR, Tadesse S, Nishigaki Y et al (eds) (2001) Defects of intergenomic communication: autosomal disorders that cause multiple deletions and depletion of mitochondrial DNA. Seminars in cell & developmental biology, Elsevier
Hubal MJ, Gordish-Dressman H, Thompson PD, Price TB, Hoffman EP, Angelopoulos TJ et al (2005) Variability in muscle size and strength gain after unilateral resistance training. Med Sci Sports Exerc 37(6):964–972
Huffman KM, Koves TR, Hubal MJ, Abouassi H, Beri N, Bateman LA et al (2014) Metabolite signatures of exercise training in human skeletal muscle relate to mitochondrial remodelling and cardiometabolic fitness. Diabetologia 57(11):2282–2295
Iestra J, Kromhout D, Van der Schouw Y, Grobbee D, Boshuizen H, Van Staveren W (2005) Effect size estimates of lifestyle and dietary changes on all-cause mortality in coronary artery disease patients. Circulation 112(6):924–934
Kao FF (1956) Regulation of respiration during muscular activity. Am J Physiol 185(1):145–151
Kemi OJ, Loennechen JP, Wisløff U, Ellingsen Ø (2002) Intensity-controlled treadmill running in mice: cardiac and skeletal muscle hypertrophy. J Appl Physiol 93(4):1301–1309
Kesterson RA, Johnson LW, Lambert LJ, Vivian JL, Welch DR, Ballinger SW (2016) Generation of mitochondrial-nuclear eXchange mice via pronuclear transfer. Bio-protocol 6(20)
Kilikevicius A, Venckunas T, Zelniene R, Carroll A, Lionikaite S, Ratkevicius A et al (2013) Divergent physiological characteristics and responses to endurance training among inbred mouse strains. Scand J Med Sci Sports 23(5):657–668
Klitgaard H (1988) A model for quantitative strength training of hindlimb muscles of the rat. J Appl Physiol 64(4):1740–1745
Koch LG, Britton SL (2001) Artificial selection for intrinsic aerobic endurance running capacity in rats. Physiol Genom 5(1):45–52
Koch LG, Meredith TA, Fraker TD, Metting PJ, Britton SL (1998) Heritability of treadmill running endurance in rats. Am J Physiol Regul Integr Comp Physiol 275(5):R1455–R1460
Koch LG, Britton SL, Barbato JC, Rodenbaugh DW, DiCARLO SE (1999) Phenotypic differences in cardiovascular regulation in inbred rat models of aerobic capacity. Physiol Genomics 1(2):63–69
Koch LG, Green CL, Lee AD, Hornyak JE, Cicila GT, Britton SL (2005) Test of the principle of initial value in rat genetic models of exercise capacity. Am J Physiol Regul Integr Comp Physiol 288(2):R466–R472
Koch LG, Pollott GE, Britton SL (2013) Selectively bred rat model system for low and high response to exercise training. Physiol Genomics 45(14):606–614
Kohrt WM, Malley MT, Coggan AR, Spina RJ, Ogawa T, Ehsani A et al (1991) Effects of gender, age, and fitness level on response of VO2max to training in 60–71 year olds. J Appl Physiol 71(5):2004–2011
Kregel KC, Allen DL, Booth FW, Fleshner MR, Henriksen EJ, Musch T et al (2006) Resource book for the design of animal exercise protocols. Am Physiol Soc 152
Kuster DW, Merkus D, Blonden LA, Kremer A, van IJcken WF, Verhoeven AJ et al (2014) Gene reprogramming in exercise-induced cardiac hypertrophy in swine: a transcriptional genomics approach. J Mol Cell Cardiol 77:168–174
LaMonte MJ, Blair SN, Church TS (2005) Physical activity and diabetes prevention. J Appl Physiol 99(3):1205–1213
Lightfoot JT, Turner MJ, Debate KA, Kleeberger SR (2001) Interstrain variation in murine aerobic capacity. Med Sci Sports Exerc 33(12):2053–2057
Lightfoot JT, Turner MJ, Knab AK, Jedlicka AE, Oshimura T, Marzec J et al (2007) Quantitative trait loci associated with maximal exercise endurance in mice. J Appl Physiol 103(1):105–110
Lightfoot JT, Turner MJ, Pomp D, Kleeberger SR, Leamy LJ (2008) Quantitative trait loci for physical activity traits in mice. Physiol Genomics 32(3):401–408
Lightfoot JT, Leamy L, Pomp D, Turner MJ, Fodor AA, Knab A et al (2010) Strain screen and haplotype association mapping of wheel running in inbred mouse strains. J Appl Physiol 109(3):623–634
Lihong M (2000) The mitochondrial DNA D-LOOP polymorphism and VO ~ 2max in Chinese Junior Athletes. Chin J Sports Med 19(4):349–350
Lowe DA, Alway SE (2002) Animal models for inducing muscle hypertrophy: are they relevant for clinical applications in humans? J Orthop Sports Phys Ther 32(2):36–43
Lundby C, Montero D, Joyner M (2016) Biology of VO2max: looking under the physiology lamp. Acta Physiol 220:218–228
Manson JE, Hu FB, Rich-Edwards JW, Colditz GA, Stampfer MJ, Willett WC et al (1999) A prospective study of walking as compared with vigorous exercise in the prevention of coronary heart disease in women. N Engl J Med 341(9):650–658
Massett MP, Berk BC (2005) Strain-dependent differences in responses to exercise training in inbred and hybrid mice. Am J Physiol Regul Integr Comp Physiol 288(4):R1006-R13
Massett MP, Fan R, Berk BC (2009) Quantitative trait loci for exercise training responses in FVB/NJ and C57BL/6J mice. Physiol Genomics 40(1):15–22
Massett MP, Avila JJ, Kim SK (2015) Exercise capacity and response to training quantitative trait loci in a NZW X 129S1 intercross and combined cross analysis of inbred mouse strains. PLoS ONE 10(12):e0145741
Mathes WF, Aylor DL, Miller DR, Churchill GA, Chesler EJ, de Villena FP-M et al (2011) Architecture of energy balance traits in emerging lines of the Collaborative Cross. Am J Physiol Endocrinol Metab 300(6):E1124–E1134
Matin A, Collin GB, Asada Y, Varnum D, Nadeau JH (1999) Susceptibility to testicular germ-cell tumours in a 129. MOLF-Chr 19 chromosome substitution strain. Nat Genet 23(2)
Matoba S, Kang J-G, Patino WD, Wragg A, Boehm M, Gavrilova O et al (2006) p53 regulates mitochondrial respiration. Science 312(5780):1650–1653
Montero D, Lundby C (2017) Refuting the myth of non-response to exercise training: ‘non-responders’ do respond to higher dose of training. J Physiol 595(11):3377–3387
Moore SC, Patel AV, Matthews CE, de Gonzalez AB, Park Y, Katki HA et al (2012) Leisure time physical activity of moderate to vigorous intensity and mortality: a large pooled cohort analysis. PLoS Med 9(11):e1001335
Nicastro H, Zanchi NE, Da Luz CR, Chaves DFS, Lancha AH (2012) An experimental model for resistance exercise in rodents. Biomed Res Int 2012:7
Paffenbarger RS Jr, Hyde R, Wing AL, Hsieh C-c (1986) Physical activity, all-cause mortality, and longevity of college alumni. N Engl J Med 314(10):605–613
Park JH, Zhuang J, Li J, Hwang PM (2016) p53 as guardian of the mitochondrial genome. FEBS Lett 590(7):924–934
Pérusse L, Gagnon J, Province MA, Rao D, Wilmore JH, Leon AS et al (2001) Familial aggregation of submaximal aerobic performance in the HERITAGE Family study. Med Sci Sports Exerc 33(4):597–604
Pescatello LS, American College of Sports Medicine (2014) ACSM’s guidelines for exercise testing and prescription. Wolters Kluwer/Lippincott Williams & Wilkins Health, Philadelphia
Petrosino JM, Heiss VJ, Maurya SK, Kalyanasundaram A, Periasamy M, LaFountain RA et al (2016) Graded maximal exercise testing to assess mouse cardio-metabolic phenotypes. PLoS ONE 11(2):e0148010
Physical Activity Guidelines Advisory Committee (2008) Physical activity guidelines for Americans. US Department of Health Human Services, Washington, DC, pp 15–34
Poole DC, Erickson HH (2011) Highly athletic terrestrial mammals: horses and dogs. Compr Physiol 1:1–37
Poole D, Epp T, Erickson H (2007) Exercise-induced pulmonary haemorrhage (EIPH): mechanistic bases and therapeutic interventions. Equine Vet J 39(4):292–293
Prudhomme D, Bouchard C, Landry F, Fontaine E, Damours Y, Leblanc C (eds) (1983) Sensitivity of maximal aerobic power to training is genotype dependent. In: Medicine and science in sports and exercise. Williams & Wilkins, Baltimore
Puntschart A, Claassen H, Jostarndt K, Hoppeler H, Billeter R (1995) mRNAs of enzymes involved in energy metabolism and mtDNA are increased in endurance-trained athletes. Am J Physiol Cell Physiol 269(3):C619–C625
Rasmussen AL, Okumura A, Ferris MT, Green R, Feldmann F, Kelly SM et al (2014) Host genetic diversity enables Ebola hemorrhagic fever pathogenesis and resistance. Science 346(6212):987–991
Rivera MA, Wolfarth B, Dionne FT, Chagnon M, Simoneau J-A, Boulay MR et al (1998) Three mitochondrial DNA restriction polymorphisms in elite endurance athletes and sedentary controls. Med Sci Sports Exerc 30:687–690
Roberts A, De Villena FP-M, Wang W, McMillan L, Threadgill DW (2007) The polymorphism architecture of mouse genetic resources elucidated using genome-wide resequencing data: implications for QTL discovery and systems genetics. Mamm Genome 18(6–7):473–481
Roberts MD, Brown JD, Oberle LP, Heese AJ, Toedebusch RG, Wells KD et al (2013) Phenotypic and molecular differences between rats selectively bred to voluntarily run high vs. low nightly distances. Am J Physiol Regul Integr Comp Physiol 304(11):R1024–R1035
Roberts MD, Toedebusch RG, Wells KD, Company JM, Brown JD, Cruthirds CL et al (2014) Nucleus accumbens neuronal maturation differences in young rats bred for low versus high voluntary running behaviour. J Physiol 592(10):2119–2135
Robinson MM, Dasari S, Konopka AR, Johnson ML, Manjunatha S, Esponda RR et al (2017) Enhanced protein translation underlies improved metabolic and physical adaptations to different exercise training modes in young and old humans. Cell Metab 25(3):581–592
Rogala AR, Morgan AP, Christensen AM, Gooch TJ, Bell TA, Miller DR et al (2014) The Collaborative Cross as a resource for modeling human disease: CC011/Unc, a new mouse model for spontaneous colitis. Mamm Genome 25(3–4):95–108
Rubio DM, Schoenbaum EE, Lee LS, Schteingart DE, Marantz PR, Anderson KE et al (2010) Defining translational research: implications for training. Acad Med: J Assoc Am Med Coll 85(3):470
Safdar A, Khrapko K, Flynn JM, Saleem A, Lisio M, Johnston AP et al (2016) Exercise-induced mitochondrial p53 repairs mtDNA mutations in mutator mice. Skelet Muscle 6(1):7
Saleem A, Adhihetty PJ, Hood DA (2009) Role of p53 in mitochondrial biogenesis and apoptosis in skeletal muscle. Physiol Genomics 37(1):58–66
Schaeffer P, Conley K, Lindstedt S (1996) Structural correlates of speed and endurance in skeletal muscle: the rattlesnake tailshaker muscle. J Exp Biol 199(2):351–358
Schmidt CW (2015) Diversity outbred: a new generation of mouse model. Environ Health Perspect 123(3):A64–A67
Schon EA, DiMauro S, Hirano M (2012) Human mitochondrial DNA: roles of inherited and somatic mutations. Nat Rev Genet 13(12):878–890
Scott RA, Wilson RH, Goodwin WH, Moran CN, Georgiades E, Wolde B et al (2005) Mitochondrial DNA lineages of elite Ethiopian athletes. Comp Biochem Physiol B 140(3):497–503
Seo DY, Lee SR, Kim N, Ko KS, Rhee BD, Han J (2014) Humanized animal exercise model for clinical implication. Pflügers Arch Eur J Physiol 466(9):1673–1687
Swallow JG, Garland T, Carter PA, Zhan W-Z, Sieck GC (1998a) Effects of voluntary activity and genetic selection on aerobic capacity in house mice (Mus domesticus). J Appl Physiol 84(1):69–76
Swallow JG, Carter PA, Garland T (1998b) Artificial selection for increased wheel-running behavior in house mice. Behav Genet 28(3):227–237
Tamaki T, Uchiyama S, Nakano S (1992) A weight-lifting exercise model for inducing hypertrophy in the hindlimb muscles of rats. Med Sci Sports Exerc 24(8):881–886
Taylor RW, Turnbull DM (2005) Mitochondrial DNA mutations in human disease. Nat Rev Genet 6(5):389–402
Thomis M, Beunen GP, Maes HH, Blimkie CJ, Van Leemputte M, Claessens AL et al (1998) Strength training: importance of genetic factors. Med Sci Sports Exerc 30(5):724–731
Thompson PD, Moyna N, Seip R, Price T, Clarkson P, Angelopoulos T et al (2004) Functional polymorphisms associated with human muscle size and strength. Med Sci Sports Exerc 36(7):1132–1139
Threadgill DW, Churchill GA (2012) Ten years of the collaborative cross. G3: Genes Genomes Genet 2(2):153–156
Threadgill DW, Hunter KW, Williams RW (2002) Genetic dissection of complex and quantitative traits: from fantasy to reality via a community effort. Mamm Genome 13(4):175–178
Threadgill DW, Miller DR, Churchill GA, de Villena FP-M (2011) The collaborative cross: a recombinant inbred mouse population for the systems genetic era. ILAR J 52(1):24–31
Trounce I (2000) Genetic control of oxidative phosphorylation and experimental models of defects. Hum Reprod 15(suppl_2):18–27
Van Goethem G, Dermaut B, Löfgren A, Martin J-J, Van Broeckhoven C (2001) Mutation of POLG is associated with progressive external ophthalmoplegia characterized by mtDNA deletions. Nat Genet 28(3):211–212
Verghese J, Lipton RB, Katz MJ, Hall CB, Derby CA, Kuslansky G et al (2003) Leisure activities and the risk of dementia in the elderly. N Engl J Med 348(25):2508–2516
Voet D, Voet JGP, Charlotte W, Judith GV, Charlotte WP (2013) Fundamentals of biochemistry: life at the molecular level. Wiley, Hoboken
Wallace DC (1992) Diseases of the mitochondrial DNA. Ann Rev Biochem 61(1):1175–1212
Wang P-y, Ma W, Park J-Y, Celi FS, Arena R, Choi JW et al (2013) Increased oxidative metabolism in the Li–Fraumeni syndrome. N Engl J Med 368(11):1027–1032
Watt PW, Kelly F, Goldspink DF, Goldspink G (1982) Exercise-induced morphological and biochemical changes in skeletal muscles of the rat. J Appl Physiol 53(5):1144–1151
Wilmore JH, Stanforth PR, Hudspeth LA, Gagnon J, Daw EW, Leon AS et al (1998) Alterations in resting metabolic rate as a consequence of 20 wk of endurance training: the HERITAGE Family Study. Am J Clin Nutr 68(1):66–71
Wirth O, Gregory EW, Cutlip RG, Miller GR (2003) Control and quantitation of voluntary weight-lifting performance of rats. J Appl Physiol 95(1):402–412
Xu S, Garland T (2017) A mixed model approach to genome-wide association studies for selection signatures, with application to mice bred for voluntary exercise behavior. Genetics 207:785–799
Yang H, Wang JR, Didion JP, Buus RJ, Bell TA, Welsh CE et al (2011) Subspecific origin and haplotype diversity in the laboratory mouse. Nat Genet 43(7):648–655
Zeviani M, Servidei S, Gellera C, Bertini E, DiMauro S, DiDonato S (1989) An autosomal dominant disorder with multiple deletions of mitochondrial DNA starting at the D-loop region. Nature 339(6222):309–311
Zou F, Gelfond JA, Airey DC, Lu L, Manly KF, Williams RW et al (2005) Quantitative trait locus analysis using recombinant inbred intercrosses. Genetics 170(3):1299–1311
Zou F, Sun W, Crowley JJ, Zhabotynsky V, Sullivan PF, de Villena FP-M (2014) A novel statistical approach for jointly analyzing RNA-Seq data from F 1 reciprocal crosses and inbred lines. Genetics 197(1):389–399
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Vellers, H.L., Kleeberger, S.R. & Lightfoot, J.T. Inter-individual variation in adaptations to endurance and resistance exercise training: genetic approaches towards understanding a complex phenotype. Mamm Genome 29, 48–62 (2018). https://doi.org/10.1007/s00335-017-9732-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00335-017-9732-5