Abstract
The increasing amount of genomic data currently available is expanding the horizons of population genetics inference. A wide range of methods have been published allowing to detect and date major changes in population size during the history of species. At the same time, there has been an increasing recognition that population structure can generate genetic data similar to those generated under models of population size change. Recently, Mazet et al. (Heredity 116(4):362–371, 2016) introduced the idea that, for any model of population structure, it is always possible to find a panmictic model with a particular function of population size-change having an identical distribution of \(T_{2}\) (the time of the first coalescence for a sample of size two). This implies that there is an identifiability problem between a panmictic and a structured model when we base our analysis only on \(T_2\). In this paper, based on an analytical study of the rate matrix of the ancestral lineage process, we obtain new theoretical results about the joint distribution of the coalescence times \((T_3,T_2)\) for a sample of three haploid genes in a n-island model with constant size. Even if, for any \(k \ge 2\), it is always possible to find a size-change scenario for a panmictic population such that the marginal distribution of \(T_k\) is exactly the same as in a n-island model with constant population size, we show that the joint distribution of the coalescence times \((T_3,T_2)\) for a sample of three genes contains enough information to distinguish between a panmictic population and a n-island model of constant size.






Similar content being viewed by others
References
Beaumont MA (1999) Detecting population expansion and decline using microsatellites. Genetics 153(4):2013–2029
Beaumont MA (2004) Recent developments in genetic data analysis: what can they tell us about human demographic history? Heredity 92(5):365–379
Bhaskar A, Song YS (2014) Descartes’ rule of signs and the identifiability of population demographic models from genomic variation data. Ann Stat 42(6):2469–2493
Chikhi L, Sousa VC, Luisi P, Goossens B, Beaumont MA (2010) The confounding effects of population structure, genetic diversity and the sampling scheme on the detection and quantification of population size changes. Genetics 186(3):983–995
Chikhi L, Rodríguez W, Grusea S, Santos P, Boitard S, Mazet O (2018) The IICR (inverse instantaneous coalescence rate) as a summary of genomic diversity: insights into demographic inference and model choice. Heredity 120:13–24
Goossens B, Chikhi L, Ancrenaz M, Lackman-Ancrenaz I, Andau P, Bruford MW et al (2006) Genetic signature of anthropogenic population collapse in orang-utans. PLoS Biol 4(2):285
Griffiths RC, Tavaré S (1994) Sampling theory for neutral alleles in a varying environment. Philos Trans R Soc B Biol Sci 344(1310):403–410
Griffiths RC, Tavaré S (1998) The age of a mutation in a general coalescent tree. Stoch Models 14(1–2):273–295
Heller R, Chikhi L, Siegismund HR (2013) The confounding effect of population structure on Bayesian skyline plot inferences of demographic history. PLoS ONE 8(5):e62992
Herbots HM (1994) Stochastic models in population genetics: genealogy and genetic differentiation in structured populations. PhD thesis, University of London. https://qmro.qmul.ac.uk/xmlui/handle/123456789/1482?show=full
Hudson RR (1983) Properties of a neutral allele model with intragenic recombination. Theor Popul Biol 23(2):183–201
Hudson RR (2002) Generating samples under a Wright–Fisher neutral model of genetic variation. Bioinformatics 18(2):337–338
Hudson RR et al (1990) Gene genealogies and the coalescent process. Oxf Surv Evol Biol 7(1):44
Kim J, Mossel E, Rácz MZ, Ross N (2015) Can one hear the shape of a population history? Theor Popul Biol 100:26–38 ISSN 0040-5809
Kingman JFC (1982) The coalescent. Stoch Process Appl 13(3):235–248
Lang S (2002) Algebra, rev. 3 edn. Springer, Berlin
Lapierre M, Lambert A, Achaz G (2017) Accuracy of demographic inferences from the site frequency spectrum: the case of the Yoruba population. Genetics 206(1):439–449
Mazet O, Rodríguez W, Chikhi L (2015) Demographic inference using genetic data from a single individual: separating population size variation from population structure. Theor Popul Biol 104:46–58
Mazet O, Rodríguez W, Grusea S, Boitard S, Chikhi L (2016) On the importance of being structured: instantaneous coalescence rates and human evolution-lessons for ancestral population size inference? Heredity 116(4):362–371
Myers S, Fefferman C, Patterson N (2008) Can one learn history from the allelic spectrum? Theor Popul Biol 73(3):342–348
Nei M, Takahata N (1993) Effective population size, genetic diversity, and coalescence time in subdivided populations. J Mol Evol 37(3):240–244
Nickalls RWD (1993) A new approach to solving the cubic: Cardan’s solution revealed. Math Gazette 77:354–359. http://www.nickalls.org/dick/papers/maths/cubic1993.pdf
Nielsen R, Wakeley J (2001) Distinguishing migration from isolation: a Markov Chain Monte Carlo approach. Genetics 158(2):885–896
Norris JR (1998) Markov Chains. Cambridge series in statistical and probabilistic mathematics. Cambridge University Press, Cambridge
Notohara M (1990) The coalescent and the genealogical process in geographically structured population. J Math Biol 29(1):59–75
Peter BM, Wegmann D, Excoffier L (2010) Distinguishing between population bottleneck and population subdivision by a Bayesian model choice procedure. Mol Ecol 19(21):4648–4660
Quéméré E, Amelot X, Pierson J, Crouau-Roy B, Chikhi L (2012) Genetic data suggest a natural prehuman origin of open habitats in northern Madagascar and question the deforestation narrative in this region. Proc Natl Acad Sci USA 109(32):13028–13033
Rodríguez W (2016) Estimation de l’histoire démographique des populations à partir de génomes entièrement séquencés. PhD thesis, University Paul Sabatier, Toulouse, June 2016
Rogers AR, Harpending H (1992) Population growth makes waves in the distribution of pairwise genetic differences. Mol Biol Evol 9(3):552–569
Schiffels S, Durbin R (2014) Inferring human population size and separation history from multiple genome sequences. Nat Genet 46(8):919–925
Slatkin M (1991) Inbreeding coefficients and coalescence times. Genet Res 58(02):167–175
Storz JF, Beaumont MA (2002) Testing for genetic evidence of population expansion and contraction: an empirical analysis of microsatellite DNA variation using a hierarchical Bayesian model. Evolution 56(1):154–166
Tajima F (1983) Evolutionary relationship of DNA sequences in finite populations. Genetics 105:437–460
Terhorst J, Song YS (2015) Fundamental limits on the accuracy of demographic inference based on the sample frequency spectrum. Proc Natl Acad Sci USA 112(25):7677–7682
Vallander SS (1973) Calculation of the Wasserstein distance between probability distributions on the line. Theory Probab Appl 18:784–786
Wakeley J (1999) Nonequilibrium migration in human history. Genetics 153(4):1863–1871
Weissman DB, Hallatschek O (2017) Minimal-assumption inference from population-genomic data. eLife 6:e24836
Wilkinson-Herbots HM (1998) Genealogy and subpopulation differentiation under various models of population structure. J Math Biol 37(6):535–585
Wright S (1931) Evolution in mendelian populations. Genetics 16(2):97
Acknowledgements
The authors wish to thank Josué M. Corujo Rodríguez for very interesting discussions in preparing this article. The authors also thank the anonymous reviewers for their reading and for valuable suggestions. This research was funded through the 2015–2016 BiodivERsA COFUND call for research proposals, with the national funders ANR (ANR-16-EBI3-0014), FCT (Biodiversa/0003/2015) and PT-DLR (01LC1617A), under the INFRAGECO (Inference, Fragmentation, Genomics, and Conservation) Project (https://infrageco-biodiversa.org/). The research was also supported by the LABEX entitled TULIP (ANR-10-LABX-41), as well as the Pôle de Recherche et d’Enseignement Suprieur (PRES) and the Région Midi-Pyrénées, France. We finally thank the LIA BEEG-B (Laboratoire International Associé—Bioinformatics, Ecology, Evolution, Genomics and Behaviour) (CNRS) and the PESSOA program for facilitating travel and collaboration between EDB, IMT and INSA in Toulouse and the IGC, in Portugal.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Appendices
Appendix A Proofs for Sections 2 and 3
Proof of Lemma 1
We first observe that
Then the calculus gives
The intermediate value theorem applied to \(p(\mu )\) thus provides a proof of the result. \(\square \)
Proof of Proposition 2
In order to simplify computations, we make the following change of parameters:
The new parameters a et N verify \(a > 0\) and \(N > 0\) and the matrix Q becomes
The matrix Q has the double eigenvalue 0 and it is easy to check that the corresponding eigenspace is generated by the following non colinear vectors:
We then consider the change of basis matrix
whose inverse is
We have thus obtained a partial diagonalization (by blocks) of the matrix Q:
and we put
The characteristic polynomial of R is the polynomial \(p(\mu )\), which has the following expressions using the new parameters:
which has three strictly negative real roots. These roots are all distinct by Lemma 1.
If \(\mu \) is an eigenvalue of R, we then determine a corresponding eigenvector \(W(\mu )\) by solving the equation \(R W(\mu ) = \mu W(\mu )\).
The computations show that we can choose
We then consider the \(3 \times 3\) passage matrix \(P_2\), whose column vectors are the \(W(\mu _i), \, i=1,2,3\), where the \(\mu _i\) are the three eigenvalues of R:
Some easy computations on the rows show that the determinant of \(P_2\) is a Van der Monde determinant, and hence: \(\mathrm {det}(P_2) = (\mu _1 - \mu _2)(\mu _1-\mu _3)(\mu _2 - \mu _3) \ne 0\).
The computations of the inverse \(P_2^{-1}\) gives
where
with
We further obtain
We then introduce the matrices \(A(\mu )\) and B defined by
where \(\bar{W}_j(\mu )\) (resp. \(\bar{Z}(\mu )\)) is a vector of length 5 obtained by adding two null coordinates to \(W(\mu )\) (resp. \(Z(\mu )\)), and
Emphasizing the rank one property of matrix \(A(\mu )\), let us define the vectors
where \(\mu \) is an eigenvalue of R, and u, v are functions of \(\mu \) defined by
Note that, since
\(\displaystyle -\frac{3M}{n-1}\) and \(\displaystyle -\frac{3(M+2)}{2}\) cannot be eigenvalues and thus \(u \ne 0\) et \(v \ne 0\).
With \(\delta (M,n,\mu ) = 2(n-1)^2 p'(\mu )\), we have
which gives the expression of \(A(\mu )\) in Proposition 2.
The stated result easily follows. \(\square \)
Proof of Proposition 4
Using Eq. (15) and Proposition 2, we deduce that
Note that the matrix \(E_1 E_2^T\) introduced in the proof of Proposition 2 can also be factorized in a different manner. If we define the \(3 \times 3\) matrix C, the \(5 \times 3\) matrix \(D_1\) and the \(3 \times 5\) matrix \(D_2\) by
we may check that, for any value of \(\mu \), we have \(E_1 E_2^T = D_1 C D_2\).
The vectors \(V_1\) and \(V_2\), defined by
verify \(D_2 V_j = 0\) for \(j=1,2\), and hence, using Eq. (25), \(A(\mu )V_j = 0\), for \(j=1,2\).
Therefore, for \(i=1,2,3\),
and the result follows. \(\square \)
Proof of Lemma 2
Note that \(-3\alpha \) and \(-3\beta \), with \(\alpha , \beta \) defined in Eq. (10), are the roots of the polynomial \(q_1(X)= q(X/3)\), where
and on the open subset \(D = \left\{ (n,M):\; n> 2, \; M>0 \right\} \) of \(\mathbb {R}^2\) the polynomials p(X) and \(q_1(X)\) have a common root if and only their resultant \(R(n,M) = \text{ Res }(p(X),q_1(X),X)\) with respect to X is null [see e.g. Lang (2002, Chapter IV–8)].
Because \(R(n,M) = - \frac{M^3}{9(n-1)^2} < 0\) on D, and because the roots \(-3 \alpha \) and \(\mu _3\) are continuous functions of (n, M) on D, the inequality \(-3 \alpha < \mu _3\) for one value of \((n,M) \in D\) implies the inequality everywhere on D. The same is true for the inequality \(\mu _1 < -3 \beta \).
This achieves the proof of the lemma. \(\square \)
Proof of Proposition 5
We will only give the proof of (i), which corresponds to the case of three genes sampled from the same deme. The proofs ot the analogous results (ii) and (iii), corresponding to the two other sample schemes, are similar.
When \(t \rightarrow 0\), using the well-known relation \(P_t = I_5 + tQ + o(t)\), where \(I_5\) denotes the identity matrix, we have the following Taylor expansions:
In particular, we have
Using Eqs. (7)–(11), and a Taylor expansion of order 2 of the exponential in the neighborhood of 0, we easily obtain
When substituting the above expressions into Eq. (11), straightforward calculations give
Further, using Eq. (18), let us denote
We will write a Taylor expansion of order 2 of h(u, t) in the neighborhood of (0, 0). We have \(h(0,0)=0\) and direct computations give
Using the fact that \(P_0 = I_5,\) and \(Q(1,4)=3, Q(1,5)=0\), we obtain
We further compute the second partial derivatives of h in (0, 0). With \(Q^2\) being the square matrix of the rate matrix Q, and using the fact that \(Q^2(1,4) = -9\left( \frac{M}{2}+1\right) \) and \(Q^2(1,5) = \frac{3M}{2},\) we easily derive
The Taylor expansion of order 2 of h(u, t) near (0, 0) finally gives
which finishes the proof. \(\square \)
Proof of Proposition 6
Because \(0< \beta < \alpha \), using Eqs. (7) and (8), we get
and
Therefore, using Eq. (11), we obtain
where \(K_{1,i}(n,M,t)\), given by (20), is strictly positive because \(0< a < 1\) and \(c < 0\).
Using Proposition 2 we get
Therefore, using Eq. (18) and relation \(1 - P_t(i,4) - P_t(i,5) = \sum _{j=1}^3 P_t(i,j)\), we obtain
where
Because \(\frac{\mu _1}{3} < - \beta \) from Lemma 2, we have \( \displaystyle e^{\frac{\mu _1}{3} u} = o(e^{-\beta u}), \) which achieves the proof of (19). \(\square \)
Proof of Proposition 7
We will first prove a useful lemma. \(\square \)
Lemma 3
Let us define \(\phi (\mu )\) by
where the matrix \(A(\mu )\) is given in Proposition 2.
Then,
-
(i)
\(0< \phi (\mu _i)< - \frac{\mu _i}{3} < \alpha , \; i=1,2,3.\)
-
(ii)
Defining h(n) by
$$\begin{aligned} h(n) = \frac{(n-1)\left( 25 - 9n + 5 \sqrt{9n^2 - 18n + 25}\right) }{12 n^2}, \end{aligned}$$we have the following results:
-
(a)
If \(M > h(n)\), then \(\phi (\mu _1)< \beta< \phi (\mu _2)< \phi (\mu _3) < \alpha .\)
-
(b)
If \(M = h(n)\), then \(\phi (\mu _1)< \beta = \phi (\mu _2)< \phi (\mu _3) < \alpha .\)
-
(c)
If \(M < h(n)\), then \(\phi (\mu _1)< \phi (\mu _2)< \beta< \phi (\mu _3) < \alpha .\)
-
(a)
Proof
From Proposition 2 and using \(p(\mu _i)=0\), we get
From Lemma 1, it follows that \(\mu n + 3 >0\). Because \( -\alpha< \displaystyle \frac{\mu _i}{3} < -\beta \) from Lemma 2, we get \(q\left( \frac{\mu _i}{3}\right) < 0\), that proves \(\phi (\mu _i) < - \frac{\mu _i}{3}\).
Now, using again that \(p(\mu _i)=0\), we obtain the following expression for \(\phi (\mu _i)\)
Introducing the new variable \(\nu = \displaystyle \frac{3 M}{2(n-1)\mu + 3nM + 6(n-1)}\), and thus \(\mu = \displaystyle - \frac{3}{2} \frac{(nM + 2n-2))\nu - M}{(n-1) \nu }\), we get that \(\phi (\mu _i)\), for \(i=1,2,3\) are the three real roots of the polynomial
The coefficient signs of r(X) show that r(X) has no negative roots, implying that \(\phi (\mu _i) >0, \; i=1,2,3\), which achieves the proof of (i).
The set of (n, M) such that r(X) and \(q(-X)\) have a common root is obtained by computing their resultant with respect to X, denoted by R(n, M):
In the domain \(D = \{(n,M),\; n>2, \; M>0 \}\) the curve \(6n^2 M^2 + (n-1)(9n-25) M - 6(n-1)^2 = 0\) is identical to the graph of the function \(M = h(n), \; n > 2\).
The set \(D {\setminus } \{(n,h(n)): \; n > 2 \}\) has two connex open components in which the relative position of \(\beta , \alpha \) and \(\phi (\mu _i), \; i=1,2,3\) are the same.
In the component \(D_1 := \{(n,M):\; n>2,\; M > h(n)\}\), one may choose \(n=3, \; M=\frac{4}{3}\) for which
that proves \((ii-a)\).
In the component \(D_2 := \{(n,M):\; n>2,\; M < h(n)\}\), one may choose \(n=3, \; M= \frac{1}{2} \) for which
that proves \((ii-c)\).
The curve arc \(\{(n,h(n)): n > 2 \}\) may be parametrized by
and we obtain
and \((ii-b)\) is proved. \(\square \)
Lemma 3 is now used to prove Proposition 7. First note that we have
for every \(i=1,2,3\).
When \(t \rightarrow + \infty \), \(P_t(i,j) = A(\mu _1)(i,j) e^{\mu _1 t} + o\left( e^{\mu _1 t}\right) ,\; j=1,2\) and thus
Using the definitions of \(F_{2,s}(u)\) and \(F_{2,d}(u)\) in Eqs. (7) and (8), we get
where \(c_1\) and \(c_2\) are given by (23).
On another hand, we have \(P_t(i,j) = A(\mu _1)(i,j) e^{\mu _1 t} + o\left( e^{\mu _1 t}\right) ,\; j=1,2,3\), we obtain
Using the fact that \(1 - P_t(i,4) - P_t(i,5) = \sum _{j=1}^3 P_t(i,j)\), this implies
and thus (21) holds.
It remains to show that \(K_3(n,M,u) >0\).
Using \(c_1 + c_2 = 1\), we may write
From Lemma 2, \(-\alpha< \frac{\mu _1}{3} < -\beta \) and thus \(e^{-\alpha u}< e^{\frac{\mu _1}{3} u} < e^{-\beta u}\). On the other hand, from Lemma 3 we have \(\phi (\mu _1)< \beta < \alpha \). Therefore \(K_3(n,M,u) >0\). \(\square \)
Appendix B The case of \(n=2\) islands
If we now consider the ancestral lineage process for a sample of three genes in the case of a symmetrical n-island model with \(n=2\) two islands, we only have the following four possible configurations:
-
1.
the three lineages are in the same island,
-
2.
two lineages are in the same island and the third one is in the other island,
-
3.
there are only two ancestral lineages left and they are in the same island,
-
4.
there are only two ancestral lineages left and they are in different islands.
The corresponding transition rate matrix is:
The characteristic polynomial of Q is \(\chi _Q(\mu ) = -\mu ^2 p(\mu )\), with
The matrix Q has the double eigenvalue 0 and the corresponding eigenspace of dimension 2 can be generated by the vectors \(\displaystyle \left[ \frac{M}{2},\frac{M}{2}-1, M, -1\right] ^T\) and \(\displaystyle [1,1,1,1]^T\). The two other eigenvalues are \(\mu _1 = - M - 2 + \sqrt{M^2+M+1}\) and \(\mu _2 = - M - 2 - \sqrt{M^2+M+1}\).
An eigenvector for \(\mu _1\) (resp. \(\mu _2\)) is \([3M, 2 \mu _1+3M+6, 0, 0]^T\) (resp. \([3M, 2 \mu _2+3M+6, 0, 0]^T\)), and we may consider the following change of basis matrix P given by
From
and the equality
a proof of the following proposition is obtained.
Proposition 8
The transition kernel \(P_t = e^{t Q}\) is given by
where
with \(\delta (M,\mu ) = 4(\mu + M+2)\) and \(b= \displaystyle \frac{M+2}{2(M+1)}\).
The IICRs \(\lambda _i(\cdot )/3\), for the initial sample configurations \(i=1\) and \(i=2\), correspond to the size-change functions
and the following proposition is verified.
Proposition 9
When \(t \rightarrow \infty \), \(\lambda _i(\cdot ),\; i=1,2\), have the following limit
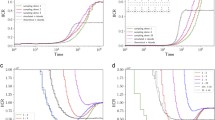
In Fig. 7 we plot the two functions \(\lambda _i(\cdot ),\; i=1,2\) for \(n=2\) demes and \(M=1\) (left), respectively \(M=0.1\) (right). The dashed line indicates the common asymptotic value \(- \frac{3}{\mu _1}\). We can also show, using Proposition 8, that \(\displaystyle \lim _{M \rightarrow \infty } \lambda _i(t) = 2\) for every \(t > 0\) and \(i=1,2\). This corresponds to the dotted line in the left panel of Fig. 7.
Proof
Using Proposition 8 we get
Then the relations \(3 A(\mu _1)(i,1) = \mu _1 A(\mu _1)(i,3)\) and \(A(\mu _1)(i,2) = \mu _1 A(\mu _1)(i,4)\) allow to prove the result. \(\square \)
The conditional cumulative distribution functions \(\mathbb {P}\left( T_2^{(3),\lambda _i} \le \cdot | T_3^{(3),\lambda _i} = t\right) \) and \(\mathbb {P}_i(T_2^{(3),2,M} \le \cdot | T_3^{(3),2,M} = t) \) are given, for every \(t > 0\), by formulas analogous to (11) and (18):
In order to compare, for \(u, t > 0\), the conditional cumulative distribution functions given in Eqs. (27) and (28), let us introduce the functions
Proposition 10
The functions \(g_i(u,t),\; i=1,2\) have the following asymptotic behaviour:
-
1.
For (u, t) in the neighborhood of (0, 0), we have
$$\begin{aligned} g_1(u,t)= & {} \displaystyle \frac{M}{2} u t + o\left( u^2+t^2\right) , \\ g_2(u,t)= & {} \displaystyle -\frac{u}{3} + \frac{6 M + 1}{18} u^2 + \frac{7 M}{6} u t + o\left( u^2+t^2\right) . \end{aligned}$$ -
2.
For fixed \(t > 0\), when \(u \rightarrow +\infty \),
$$\begin{aligned} g_i(u,t) = - K_{1,i} (M,t) e^{-\beta u} + o(e^{-\beta u}),\; i=1,2, \end{aligned}$$where \(K_{1,i}(M,t) > 0\) is given by
$$\begin{aligned} K_{1,i}(M,t) = \displaystyle \frac{3 P_t(i,1)}{3P_t(i,1) + P_t(i,2)} \frac{1-a}{\beta } - \displaystyle \frac{P_t(i,2)}{3P_t(i,1) + P_t(i,2)} \frac{c}{\beta }, \end{aligned}$$where constants \(\beta , a, c\) are defined in Eqs. (9) and (10) with \(n=2\), i.e.
$$\begin{aligned}&\beta = M + \frac{1}{2}\left( 1 - \left( 4M^2+1\right) ^{1/2}\right) ,\quad a = \frac{1}{2} \left( 1 + \left( 4M^2+1\right) ^{-1/2}\right) ,\\&\quad c = -M \left( 4M^2+1\right) ^{-1/2}. \end{aligned}$$ -
3.
For fixed \(u \ge 0\), we have
$$\begin{aligned} \displaystyle \lim _{t \rightarrow +\infty } g_i(u,t) = -K_3(M,u),\quad i=1,2, \end{aligned}$$where \(K_3(M,u) > 0\) is given by
$$\begin{aligned} K_3(M,u) = c_1 e^{-\beta u} + c_2 e^{-\alpha u} - e^{\frac{\mu _1}{3} u}, \end{aligned}$$with
$$\begin{aligned} \alpha= & {} M + \frac{1}{2}\left( 1 + \left( 4M^2+1\right) ^{1/2}\right) ,\quad \beta = M + \frac{1}{2}\left( 1 - \left( 4M^2+1\right) ^{1/2}\right) ,\; \\ \phi \left( \mu _1\right)= & {} \displaystyle \frac{3M}{2\left( \mu _1 + 3M + 3\right) },\quad c_1 = \frac{\phi \left( \mu _1\right) - \alpha }{\beta -\alpha },\quad c_2 = \frac{\beta - \phi \left( \mu _1\right) }{\beta -\alpha }. \end{aligned}$$
The proof is similar to the one given in the case \(n > 2\).
Most of the calculations of “Appendices A and B” section were made and/or verified using the computer algebra system Maple: programs and tracks of their execution are available from the authors.
Rights and permissions
About this article
Cite this article
Grusea, S., Rodríguez, W., Pinchon, D. et al. Coalescence times for three genes provide sufficient information to distinguish population structure from population size changes. J. Math. Biol. 78, 189–224 (2019). https://doi.org/10.1007/s00285-018-1272-4
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00285-018-1272-4
Keywords
- Inverse instantaneous coalescence rate (IICR)
- Population structure
- Population size change
- Demographic history
- Rate matrix
- Structured coalescent