Abstract
The oleaginous yeast Yarrowia lipolytica is widely used for the production of both bulk and fine chemicals, including organic acids, fatty acid-derived biofuels and chemicals, polyunsaturated fatty acids, single-cell proteins, terpenoids, and other valuable products. Consequently, it is becoming increasingly popular for metabolic engineering applications. Multiple gene manipulation tools including URA blast, Cre/LoxP, and transcription activator-like effector nucleases (TALENs) have been developed for metabolic engineering in Y. lipolytica. However, the low efficiency and time-consuming procedures involved in these methods hamper further research. The emergence of the CRISPR/Cas system offers a potential solution for these problems due to its high efficiency, ease of operation, and time savings, which can significantly accelerate the genomic engineering of Y. lipolytica. In this review, we summarize the research progress on the development of CRISPR/Cas systems for Y. lipolytica, including Cas9 proteins and sgRNA expression strategies, as well as gene knock-out/knock-in and repression/activation applications. Finally, the most promising and tantalizing future prospects in this area are highlighted.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Yarrowia lipolytica is a well-known non-conventional yeast, which is generally recognized as safe (Liu et al. 2015). Due to its strong lipogenesis capability and high protein expression levels, Y. lipolytica is widely researched for the production of both bulk and fine chemicals, including organic acids, fatty acid-derived biofuels and chemicals, polyunsaturated fatty acids, single-cell proteins, terpenoids, and other valuable products (Rymowicz et al. 2010; Cui et al. 2011; Yin et al. 2012; Xue et al. 2013; Kamzolova et al. 2014; Blazeck et al. 2015; Sun et al. 2016; Liu et al. 2017a, b; Gao et al. 2017). Meanwhile, a large range of substrates can be effectively utilized by Y. lipolytica, including not only glucose and glycerol but also xylose, cellobiose, and other industrial wastes, which has made it into a hot topic of recent biorefinery research (Ledesma-Amaro and Nicaud 2016; Zeng et al. 2018). Metabolic engineering is a rapidly developing field that purposely uses genetic recombination technologies to modify cellular metabolic pathways, change cell characteristics, and combines with other technologies such as biochemical engineering to construct new metabolic pathways for the synthesis of specific products (Stephanopoulos 2012; Nielsen and Keasling 2016; Chen et al. 2017). For instance, the overexpression of the endogenous acetyl-CoA carboxylase (ACC1) and diacylglycerol acyltransferase (DGA1) genes in Y. lipolytica increased the lipid content to 41.4%, a 4.7-fold improvement over the parental strain (Tai and Stephanopoulos 2013). Subsequently, the aldehyde dehydrogenase gene was introduced to improve the strain’s resistance to oxidative stress, after which the lipid content reached up to 90% (Xu et al. 2017). Metabolic engineering was also employed to produce other products in Y. lipolytica in recent years with good results, which attracted increasing attention to the metabolic engineering research of this yeast (Yin et al. 2012; Beopoulos et al. 2014; Rutter et al. 2015; Blazeck et al. 2015; Kildegaard et al. 2017; Liu et al. 2017a, b). With the increasing number of metabolic engineering studies in Y. lipolytica, various genetic engineering tools have been developed to meet the demands (Hussain et al. 2016). These tools include URA blast, TRP1 blast, Cre/LoxP systems for recycling selection markers, and transcription activator-like effector nucleases (TALENs) for gene knock-out and knock-in (Cheon et al. 2003; Fickers et al. 2003; Rigouin et al. 2017; Gao et al. 2017). While these tools have been established in Y. lipolytica, they suffer from various limitations and remain not fully conducive to efficient, high-throughput genetic engineering.
The CRISPR/Cas system, which emerged at an opportune time, to some extent solves the traditional problems. The CRISPR/Cas system consists of mainly two components, a Cas9 protein and the corresponding sgRNA (Shi et al. 2017). As shown in Fig. 1, CRISPR/Cas systems based on different types of Cas proteins can be classified into three groups—knock-out/in-oriented CRISPR/Cas9, CRISPR interference (CRISPRi), and CRISPR activation (CRISPRa) (Sharma et al. 2017). When the sgRNA recognizes the targeted sequence, the Cas9 protein catalyzes a double-strand break (DSB) in the targeted DNA, which induces either random deletion and insertion or the introduction of heterologous genes through partially complementary donor DNA (O’Connell et al. 2014; Ran et al. 2013). CRISPRi is used for gene repression via a catalytically deactivated Cas9 (dCas9), which has no cleavage activity, but can nevertheless bind the DNA and repress the expression of the gene targeted by the gRNA (Larson et al. 2013). In order to enhance the repression activity, transcriptional repressors, such as Krüppel associated box (KRAB) domain, is usually expressed as a fusion with the Cas9 protein (Zhang et al. 2018). Similarly, CRISPRa was developed for targeted gene activation by fusing dCas9 to transcriptional activators that bind promoters of targeted genes and improve gene expression levels (Simeonov et al. 2017). These technologies offer important solutions, including multi-gene targeting and marker-free integration, which promote the development of metabolic engineering in Y. lipolytica.
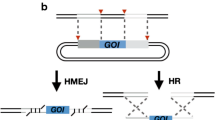
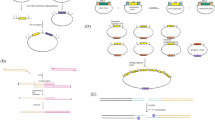
The CRISPR/Cas genome editing platform for Yarrowia lipolytica. a CRISPR/Cas9 method for gene knock-out/knock-in. When the sgRNA recognizes the targeted sequence, which is located before a protospacer adjacent motif (PAM) site, the Cas9 protein will catalyze the formation of a double-strand break (DSB) in the targeted DNA. In order to repair the genome, two kinds of repair mechanisms can be used. The non-homologous end-joining (NHEJ) repair mechanism, which is dominant in Y. lipolytica, can quickly repair the genome at the expense of the deletion or insertion of a few nucleotides, which can lead to the frameshift mutations in the targeted gene. Additionally, in the presence of a homologous sequence, cells can use the donor DNA to introduce nested heterologous genes or disrupt a targeted gene by homologous recombination (HR), homology-mediated end-joining (HMEJ), and microhomology-mediated end-joining (MMEJ) repair mechanisms. b CRISPRi and CRISPRa methods for gene interference and activation, respectively. A catalytically deactivated Cas9 (dCas9), which has no cleavage activity, can be fused with different effector domains to control gene expression. When the targeted region is recognized, the dCas9 fusion protein with the transcriptional repressor domain binds the DNA to repress gene expression. Similarly, the fusion protein of dCas9 and a transcriptional activator domain binds to targeted regions to improve the gene expression level
In this review, we summarize the expression strategies and recent applications of the CRISPR/Cas system in Y. lipolytica, followed by a brief discussion of future prospects of this system. We hope to provide a practical reference for genome editing in Y. lipolytica.
Development of a CRISPR/Cas9 system for Y. lipolytica
Cas9/dCas9 expression strategies
As the first CRISPR research in Y. lipolytica by Schwartz et al. (2016a), both expression of the active Cas9 protein in the CRISPR/Cas9 system and dCas9 in CRISPRi and CRISPRa systems after that has been engineered using a strong constitutive promoter as well as a SV40 nuclear localization signal (Gao et al. 2016; Schwartz et al. 2017; Schwartz et al. 2018; Schwartz and Wheeldon 2018; Holkenbrink et al. 2018; Zhang et al. 2018). There are two general strategies for the expression of Cas9/dCas9—one based on plasmids and the other on chromosomal integration. In the plasmid-based setup, Cas9/dCas9 can be cloned into an autonomously replicating plasmid (ARP) for recycling of marker genes, or a non-ARP for transient expression, both of which showed a high editing efficiency in Y. lipolytica (Schwartz et al. 2016a). In addition, Holkenbrink et al. (2018) established an EasyCloneYALI genetic toolbox in which Cas9 is integrated into the genome for easier transformation protocols. With this system, highly efficient genome editing only requires an sgRNA expression cassette, a strategy that is also of interest for CRISPRi and CRISPRa systems.
sgRNA expression strategies
Promoter engineering to improve the genome editing efficiency
Efficient genome editing mainly depends on the level of sgRNA transcription. Therefore, many studies have focused on promoter engineering in recent years. Gao et al. (2016) and Wong et al. (2017) used a polymerase II promoter (Pol II) to transcribe sgRNAs in Y. lipolytica. In this system, sgRNAs were flanked by hammerhead (HH) and hepatitis delta virus (HDV) ribozymes. Due to the strength of Pol II and self-processed RNA cleavage, sgRNAs can be successfully released. Similar research was also previously carried out by Schwartz et al. (2016a). However, the efficiency was quite low, and in order to further improve the editing efficiency, Schwartz et al. (2016a) adopted a synthetic RNA polymerase III promoter (Pol III) strategy in which differently designed Pol III promoters were individually and combinatorically used to optimize the knockout efficiency. The SCR1-tRNAGly hybrid Pol III achieved an efficiency of nearly 100% after 4 days of outgrowth, and was consequently frequently utilized to express sgRNAs in later studies. In very recent research, Morse et al. (2018) established a T7 polymerase-based CRISPR/Cas system in different yeasts, including Y. lipolytica. Here, the sgRNAs were expressed from a T7 promoter which was transcribed by a heterologous mutant T7 polymerase, and the genome editing efficiency reached nearly 60%. The establishment of promoter engineering strategies has laid the foundation for further development of the CRISPR/Cas system in Y. lipolytica.
Multiplex sgRNA expression strategy
Multiplex sgRNA expression strategies have been widely applied to the CRISPR/Cas system in Y. lipolytica. Gao et al. (2016) investigated the efficiency of simultaneous single, double, and triple gene disruption in Y. lipolytica. The final result showed that the frequency of single-disruption events approached 100%, while for double disruption, it was nearly 36%, and reached 19% for triple disruption. Holkenbrink et al. (2018) established a multi-sgRNA expression system, in which multiplex sgRNAs that target different genes can be constructed rapidly through the BioBricks assembly. Additionally, a multiplex sgRNA expression strategy has been used for CRISPRi and CRISPRa (Zhang et al. 2018; Schwartz et al. 2017); however, in these cases, the multiplex sgRNAs were usually designed to target a single desired gene rather than multiplex different genes. The reasoning for this is that when the targeting is biased towards a single gene, the efficiency can be lower, and multi-gene targeting has a better efficacy for gene interference and activation. These examples demonstrate that multiplex sgRNA expression plays an important role in the CRISPR/Cas system.
CRISPRS/Cas system for knock-out/knock-in and repression or activation of genes
Gene knock-out or knock-in
In the field of metabolic engineering, a highly efficient homologous recombination (HR) system of microbes is essential for gene knock-out/knock-in, which in turn is a prerequisite for investigating the functions of the targeted genes. However, the system is in direct competition with the stronger non-homologous end-joining (NHEJ) mechanism in most organisms including Y. lipolytica (Ueno et al. 2007; Decottignies 2007). Consequently, a high HR efficiency usually requires homologous arms with a length of more than 1 kb. In order to enhance the HR efficiency, chemical and biological approaches for inactivation of the NHEJ repair pathway in Y. lipolytica were adopted. Verbeke et al. (2013) identified the ku70 and ku80 genes in Y. lipolytica, which play a role in the NHEJ repair mechanism after the formation of DSB. While the disruption of ku80 did not affect the HR efficiency, it was significantly improved by ku70 disruption. In the corresponding knockout strain, the efficiency of HR mediated by only 50-bp homologous arms can be increased to 43%. Interestingly, Kretzschmar et al. (2013) proved that ku80 disruption can also increase the HR efficiency to 80% with long homologous arms 1 kb in length. Moreover, they observed the highest HR efficiency up to 85% with ku70 disruption. In addition, based on the ku70 disruption, Jang et al. (2018) added hydroxyurea into the medium to synchronize the cell cycle to the S-phase, which has been proved to induce the HR in Y. lipolytica (Tsakraklides et al. 2015). The experiment demonstrated that 50-bp homologous arms can yield an HR efficiency of 46% and 100-bp homologous arms can reach up to 100%. Although these strategies have been developed to improve HR efficiency, it is quite difficult to knock out multiple genes simultaneously even in an NHEJ-knockout strain. Moreover, the use of multiple selection markers is not conducive to further metabolic engineering and industrial utilization of the resulting strains (Wagner and Alper 2016). Fortunately, the emergence of the CRISPR/Cas system offers the possibility to solve these problems.
CRISPR/Cas9 system for Y. lipolytica was first established by Schwartz et al. (2016a). In their research, more attention was paid to finding the best Pol III promoter to improve the genome editing efficiency. The final result indicated that a combination of SCR1 and tRNA Pol III promoters was the best choice. Based on these results, a standardized markerless gene integration tool for pathway engineering was subsequently established by Schwartz et al. (2016b). By knocking out 17 genes using an autonomously replicating CRISPR/Cas9 plasmid as well as the repair fragment, five loci which offer efficient gene integration were identified. These repair fragments as well as the CRISPR/Cas9 plasmids together comprise a standardized tool that allows efficient genome editing in any of the five loci. In order to verify the practical applicability of this tool, the multigene pathway for lycopene biosynthesis was subsequently integrated into the Y. lipolytica genome. Importantly, repair fragments were designed to easily insert any targeted genes, and these plasmids can be removed in a day, which has the potential to significantly accelerate the construction of any metabolic pathway in Y. lipolytica.
Comparable to the above approach, Holkenbrink et al. (2018) established a CRISPR/Cas9-based toolbox for engineering Y. lipolytica. In the system, Cas9 and sgRNA were separately expressed from two different plasmids, and the Cas9 protein was subsequently integrated into the genome. A total of 11 loci which did not affect cell growth were selected to design the sgRNAs and repair fragment. Both marker-mediated integration and CRISPR/Cas9-based marker-free genome editing had a high efficiency. Additionally, the use of this toolbox for multiplex gene knockouts was tested. For single gene disruption, the efficiency was above 80%, and it varied from 6 to 66% for double gene disruption. However, for triple gene disruption, no successful transformants were found. To simplify the plasmid construction process, 90-bp double-stranded oligonucleotides were used as the template to repair the DSB by HR, and the editing efficiency reached 100%, which further demonstrated the validity of this toolbox.
Interestingly, Gao et al. (2018) recently established a dual-sgRNA-mediated gene knockout and integration strategy for Y. lipolytica. By designing paired sgRNAs for single genes, both non-coding and coding regions of the targeted gene could be cleaved precisely. The result was further confirmed by knocking out six genes. Moreover, based on a new homology-mediated end-joining (HMEJ) strategy, which was recently established in animal embryo and tissue cells (Xuan et al. 2017; Yao et al. 2018), researchers also applied this HMEJ strategy to Y. lipolytica. Strikingly, the efficiency was twice as high as that of HR.
Taken together, fast developments of the CRISPR/Cas9 system provide a great deal of convenience for metabolic engineering in Y. lipolytica. Both marker-free integration and multi-gene editing are powerful tools to overcome traditional shortcomings in HR systems, facilitating further metabolic engineering of this yeast.
Sequence-specific repression or activation of genes
In the CRISPR/Cas9 system, the original purpose of the Cas9 protein was to bind the DNA and cleave the targeted gene sequence. However, it was found that a dCas9 variant which has no cleavage activity can also specifically bind the targeted DNA (Ma et al. 2016). Importantly, dCas9 can be fused to transcriptional repressors and activators to further repress or activate gene expression. Subsequently, CRISPRi and CRISPRa manipulations have been quickly applied to many different organisms, including Y. lipolytica.
Schwartz et al. (2017) were the first to establish the CRISPRi system in Y. lipolytica. The purpose was to repress NHEJ to enhance HR efficiency. In the verification experiments, eight of nine target genes were efficiently repressed. In order to further improve the HR efficiency, a multiplex sgRNA expression strategy as well as a dCas9 fusion protein with the Mxi1 repressor was adopted to repress the ku70 and ku80 genes. The subsequent rate of HR was nearly 90%. Additionally, a microhomology-mediated end-joining (MMEJ) mechanism, which is independent of the NHEJ mechanism, was found in Y. lipolytica. Homology regions of only 8 bp can be used to repair the genome with the MMEJ after DSB formation. Subsequently, Zhang et al. (2018) used the four different repression proteins dCpf1, dCas9, dCas9-KRAB, and dCpf1-KRAB, to rapidly develop the CRISPRi system in Y. lipolytica. Notably, the researchers found that there was no explicit relation between target sites and repression efficiency. Therefore, multiplex sgRNAs were simultaneously expressed to improve the system’s efficiency, and rates of gene repression as high as 85% and 92% were achieved using dCpf1 and dCas9, respectively. Furthermore, the possibility of double and triple gene interference using the CRISPRi system was explored in this research. The final results showed that the combined repression events successfully occurred in Y. lipolytica, which demonstrated that CRISPRi was indeed a powerful tool for metabolic engineering of Y. lipolytica.
In addition to the gene interference system, the dCas9 protein has also been fused to transcription activators to activate the target genes, which is known as CRISPRa. Based on the previously established CRISPRi technology, Schwartz and Wheeldon (2018) rapidly developed CRISPRa manipulation in Y. lipolytica. Considering that transcription activators have an enormous influence on the activation efficiency, researchers firstly compared four different activators and found that the synthetic tripartite activator VPR yielded the highest activation. After that, the dCas9-VPR fusion protein was used to activate two β-glucosidase genes—BGL1 and BGL2—which enabled Y. lipolytica to grow on cellobiose robustly. By designing multiplex sgRNAs targeting the promoters of the two β-glucosidase genes, researchers eventually found that sgRNAs near to the core promoter region could greatly increase the activation degree. The expression level of BGL1 increased 112-fold, while that of BGL2 increased 43-fold. Moreover, the activation of both genes simultaneously also yielded a high efficiency.
In summary, dCas9-mediated gene repression and activation is playing increasingly more important roles. Consequently, more silent regions in the genome of Y. lipolytica can be activated to explore their encoded functions. Furthermore, the correlation between gene expression and cellular phenotypes can be understood in significantly more detail, which will further deepen metabolic engineering research in Y. lipolytica.
Applications of the CRISPR/Cas system in Y. lipolytica
As shown in Table 1, the CRISPR/Cas system has been quickly applied in metabolic engineering of Y. lipolytica following its introduction.
Rodriguez et al. (2016) used CRISPR/Cas9 to knock-out the xylulose kinase and xylitol dehydrogenase genes in Y. lipolytica in the xylose metabolic pathway. The knockout strains demonstrated that both genes are essential for xylitol/xylose metabolism. Based on these result as well as further study in Escherichia coli, researchers engineered xylose utilization in Y. lipolytica, which enabled it to grow on xylose robustly. Bhutada et al. (2017) used the CRISPR/Cas9 system to knock out the glycogen synthase gene in Y. lipolytica because they found it was too challenging to knock out this gene using HR in a triacylglycerol (TAG) synthesis-deficient strain. The final result showed that glycogen synthesis played a competing role in the TAG accumulation process and the deletion of this gene improved the lipid content by 60%. Markham et al. (2018) engineered Y. lipolytica to produce triacetic acid lactone. In this research, the TRP1 gene was disrupted by CRISPR/Cas9 to introduce an available selection marker. Similar to this research, Wagner et al. (2018) also knocked out TRP1 with CRISPR/Cas9 in order to establish a piggyBac transposon system in Y. lipolytica.
Compared to the gene knock-out/knock-in-oriented CRISPR/Cas9 method, both CRISPRi and CRISPRa methods are still in their infancy. Although Schwartz et al. (2017) and Zhang et al. (2018) individually established the CRISPRi system, and Schwartz and Wheeldon (2018) subsequently established the CRISPRa system, there are few applications of these two methods in Y. lipolytica. However, we believe that the need to engineer Y. lipolytica for the tailored production of specific chemicals will greatly expand the use of CRISPRi and CRISPRa systems in metabolic engineering research of this organism in the near future.
Conclusions and perspectives
Compared to the widely used yeast model organism Saccharomyces cerevisiae, the non-conventional yeast Y. lipolytica has a stronger lipogenesis ability. Therefore, Y. lipolytica has been increasingly explored for the production of lipid-related products via metabolic engineering. The adaptation of genetic tools from S. cerevisiae to Y. lipolytica would enable more rapid and convenient strain engineering and facilitate reaching the full potential of Y. lipolytica. The emergence and application of the CRISPR/Cas system undoubtedly accelerates the rate of metabolic engineering for Y. lipolytica strain improvement. Although the CRISPR/Cas system has been firmly established in this yeast, there still is much room for further improvement. For instance, problems related to multi-gene editing efficiency, more precise site-directed mutagenesis in the genome as well as high-throughput screening technology after genome editing need to be addressed. However, we believe that these problems can be solved and that increasing numbers of applications of CRIPSR/Cas in Y. lipolytica will quickly come available in the near future.
References
Beopoulos A, Verbeke J, Bordes F, Guicherd M, Bressy M, Marty A, Nicaud JM (2014) Metabolic engineering for ricinoleic acid production in the oleaginous yeast Yarrowia lipolytica. Appl Microbiol Biotechnol 98:251–262
Bhutada G, Kavšcek M, Ledesma-Amaro R, Thomas S, Rechberger GN, Nicaud JM, Natter K (2017) Sugar versus fat: elimination of glycogen storage improves lipid accumulation in Yarrowia lipolytica. FEMS Yeast Res 17:fox020
Blazeck J, Hill A, Jamoussi M, Pan A, Miller J, Alper HS (2015) Metabolic engineering of Yarrowia lipolytica for itaconic acid production. Metab Eng 32:66–73
Chen X, Cong G, Liang G, Hu G, Luo Q, Liu J, Nielsen J, Chen J, Liu L (2017) DCEO biotechnology: tools to design, construct, evaluate, and optimize the metabolic pathway for biosynthesis of chemicals. Chem Rev 118:4–72
Cheon SA, Han EJ, Kang HA, Ogrydziak DM, Kim JY (2003) Isolation and characterization of the TRP1 gene from the yeast Yarrowia lipolytica and multiple gene disruption using a TRP blaster. Yeast 20:677–685
Cui W, Wang Q, Zhang F, Zhang SC, Chi ZM, Madzak C (2011) Direct conversion of inulin into single cell protein by the engineered Yarrowia lipolytica carrying inulinase gene. Process Biochem 46:1442–1448
Decottignies A (2007) Microhomology-mediated end joining in fission yeast is repressed by pku70 and relies on genes involved in homologous recombination. Genetics 176:1403–1415
Fickers P, Gaillardin C, Nicaud JM, Dall MTL, Thonart P (2003) New disruption cassettes for rapid gene disruption and marker rescue in the yeast Yarrowia lipolytica. J Microbiol Methods 55:727–737
Gao S, Tong Y, Wen Z, Zhu L, Ge M, Chen D, Jiang Y, Yang S (2016) Multiplex gene editing of the Yarrowia lipolytica genome using the CRISPR-Cas9 system. J Ind Microbiol Biotechnol 43:1085–1093
Gao S, Tong Y, Zhu L, Ge M, Zhang Y, Chen D, Jiang Y, Yang S (2017) Iterative integration of multiple-copy pathway genes in Yarrowia lipolytica for heterologous β-carotene production. Metab Eng 41:192–201
Gao D, Smith S, Spagnuolo M, Rodriguez G, Blenner M (2018) Dual CRISPR-Cas9 cleavage mediated gene excision and targeted integration in Yarrowia lipolytica. Biotechnol J 13:1700590
Holkenbrink C, Dam MI, Kildegaard KR, Beder J, Dahlin J, Belda DD, Borodina I (2018) EasyCloneYALI: CRISPR/Cas9-based synthetic toolbox for engineering of the yeast Yarrowia lipolytica. Biotechnol J 13:1700543
Hussain MS, Rodriguez GM, Gao D, Spagnuolo M, Gambill L, Blenner M (2016) Recent advances in bioengineering of the oleaginous yeast Yarrowia lipolytica. AIMS Bioeng 3:493–514
Jang IS, Yu BJ, Jang JY, Jegal J, Lee JY (2018) Improving the efficiency of homologous recombination by chemical and biological approaches in Yarrowia lipolytica. PLoS One 13:e0194954
Kamzolova SV, Vinokurova NG, Dedyukhina EG, Samoilenko VA, Lunina JN, Mironov AA, Allayarov RK, Morgunov IG (2014) The peculiarities of succinic acid production from rapeseed oil by Yarrowia lipolytica yeast. Appl Microbiol Biotechnol 98:4149–4157
Kildegaard KR, Adiegopérez B, Doménech DB, Khangura JK, Holkenbrink C, Borodina I (2017) Engineering of Yarrowia lipolytica for production of astaxanthin. Synth Syst Biotechnol 2:287–294
Kretzschmar A, Otto C, Holz M, Werner S, Hübner L, Barth G (2013) Increased homologous integration frequency in Yarrowia lipolytica strains defective in non-homologous end-joining. Curr Genet 59:63–72
Larson MH, Gilbert LA, Wang X, Lim WA, Weissman JS, Qi LS (2013) CRISPR interference (CRISPRi) for sequence-specific control of gene expression. Nat Protoc 8:2180–2196
Ledesma-Amaro R, Nicaud JM (2016) Metabolic engineering for expanding the substrate range of Yarrowia lipolytica. Trends Biotechnol 34:798–809
Liu HH, Ji XJ, Huang H (2015) Biotechnological applications of Yarrowia lipolytica: past, present and future. Biotechnol Adv 33:1522–1546
Liu HH, Madzak C, Sun ML, Ren LJ, Song P, Huang H Ji XJ (2017a) Engineering Yarrowia lipolytica for arachidonic acid production through rapid assembly of metabolic pathway. Biochem Eng J 119:52–58
Liu HH, Zeng SY, Shi TQ, Ding Y, Ren LJ, Song P, Huang H, Madzak C, Ji XJ (2017b) A Yarrowia lipolytica strain engineered for arachidonic acid production counteracts metabolic burden by redirecting carbon flux towards intracellular fatty acid accumulation at the expense of organic acids secretion. Biochem Eng J 128:201–209
Ma H, Tu LC, Naseri A, Huisman M, Zhang S, Grunwald D, Pederson T (2016) Multiplexed labeling of genomic loci with dCas9 and engineered sgRNAs using CRISPRainbow. Nat Biotechnol 34:528–530
Markham KA, Palmer CM, Chwatko M, Wagner JM, Murray C, Vazquez S, Swaminathan A, Chakravarty I, Lynd NA, Alper HS (2018) Rewiring Yarrowia lipolytica toward triacetic acid lactone for materials generation. P Natl Acad Sci U S A 115:2096–2101
Morse NJ, Wagner JM, Reed KB, Gopal MR, Lauffer LH, Alper HS (2018) T7 polymerase expression of guide RNAs in vivo allows exportable CRISPR-Cas9 editing in multiple yeast hosts. ACS Synth Biol 7:1075–1084
Nielsen J, Keasling JD (2016) Engineering cellular metabolism. Cell 164:1185–1197
O’Connell MR, Oakes BL, Sternberg SH, Eastseletsky A, Kaplan M, Doudna JA (2014) Programmable RNA recognition and cleavage by CRISPR/Cas9. Nature 516:263–266
Ran FA, Hsu PD, Jason W, Vineeta A, Scott DA, Feng Z (2013) Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8:2281–2308
Rigouin C, Guéroult M, Croux C, Dubois G, Borsenberger V, Barbe S, Marty A, Daboussi F, André I, Bordes F (2017) Production of medium chain fatty acids by Yarrowia lipolytica: combining molecular design and Talen to engineer the fatty acid synthase. ACS Synth Biol 6:1870–1879
Rodriguez GM, Hussain MS, Gambill L, Gao D, Yaguchi A, Blenner M (2016) Engineering xylose utilization in Yarrowia lipolytica by understanding its cryptic xylose pathway. Biotechnol Biofuels 9:149–163
Rutter CD, Zhang S, Rao CV (2015) Engineering Yarrowia lipolytica for production of medium-chain fatty acids. Appl Microbiol Biotechnol 99:7359–7368
Rymowicz W, Fatykhova AR, Kamzolova SV, Rywińska A, Morgunov IG (2010) Citric acid production from glycerol-containing waste of biodiesel industry by Yarrowia lipolytica in batch, repeated batch, and cell recycle regimes. Appl Microbiol Biotechnol 87:971–979
Schwartz C, Wheeldon I (2018) CRISPR-Cas9-mediated genome editing and transcriptional control in Yarrowia lipolytica. Methods Mol Biol 1772:327–345
Schwartz CM, Hussain MS, Blenner M, Wheeldon I (2016a) Synthetic RNA polymerase III promoters facilitate high efficiency CRISPR-Cas9 mediated genome editing in Yarrowia lipolytica. ACS Synth Biol 5:356–359
Schwartz C, Shabbirhussain M, Frogue K, Blenner M, Wheeldon I (2016b) Standardized markerless gene integration for pathway engineering in Yarrowia lipolytica. ACS Synth Biol 6:402–409
Schwartz C, Frogue K, Ramesh A, Misa J, Wheeldon I (2017) CRISPRi repression of nonhomologous end-joining for enhanced genome engineering via homologous recombination in Yarrowia lipolytica. Biotechnol Bioeng 114:2896–2906
Schwartz C, Curtis N, Löbs AK, Wheeldon I (2018) Multiplexed CRISPR activation of cryptic sugar metabolism enables Yarrowia lipolytica growth on cellobiose. Biotechnol J 13:1700584
Sharma S, Kaur R, Singh A (2017) Recent advances in CRISPR/Cas mediated genome editing for crop improvement. Plant Biotechnol Rep 11:193–207
Shi TQ, Liu GN, Ji RY, Shi K, Song P, Ren LJ, Huang H, Ji XJ (2017) CRISPR/Cas9-based genome editing of the filamentous fungi: the state of the art. Appl Microbiol Biotechnol 101:7435–7443
Simeonov DR, Gowen BG, Boontanrart M, Roth TL, Gagnon JD, Mumbach MR, Satpathy AT, Lee Y, Bray NL, Chan AY, Lituiev DS, Nguyen ML, Gate RE, Subramaniam M, Li Z, Woo JM, Mitros T, Ray GJ, Curie GL, Naddaf N, Chu JS, Ma H, Boyer E, Van Gool F, Huang H, Liu R, Tobin VR, Schumann K, Daly MJ, Farh KK, Ansel M, Ye CJ, Greenleaf WJ, Anderson MS, Bouestone JA, Chang HY, Corn JE, Marson A (2017) Discovery of stimulation-responsive immune enhancers with CRISPR activation. Nature 549:111–115
Stephanopoulos G (2012) Synthetic biology and metabolic engineering. ACS Synth Biol 1:514–525
Sun ML, Madzak C, Liu HH, Song P, Ren LJ, Huang H, Ji XJ (2016) Engineering Yarrowia lipolytica for efficient γ-linolenic acid production. Biochem Eng J 117:172–180
Tai M, Stephanopoulos G (2013) Engineering the push and pull of lipid biosynthesis in oleaginous yeast Yarrowia lipolytica for biofuel production. Metab Eng 15:1–9
Tsakraklides V, Brevnova E, Stephanopoulos G, Shaw AJ (2015) Improved gene targeting through cell cycle synchronization. PLoS One 10:e0133434
Ueno K, Uno J, Nakayama H, Sasamoto K, Mikami Y, Chibana H (2007) Development of a highly efficient gene targeting system induced by transient repression of YKU80 expression in Candida glabrata. Eukaryot Cell 6:1239–1247
Verbeke J, Beopoulos A, Nicaud JM (2013) Efficient homologous recombination with short length flanking fragments in ku70 deficient Yarrowia lipolytica strains. Biotechnol Lett 35:571–576
Wagner JM, Alper HS (2016) Synthetic biology and molecular genetics in non-conventional yeasts: current tools and future advances. Fungal Genet Biol 89:126–136
Wagner JM, Williams EV, Alper HS (2018) Developing a piggyBac transposon system and compatible selection markers for insertional mutagenesis and genome engineering in Yarrowia lipolytica. Biotechnol J 13:1800022
Wong L, Engel J, Jin E, Holdridge B, Peng X (2017) YaliBricks, a versatile genetic toolkit for streamlined and rapid pathway engineering in Yarrowia lipolytica. Metab Eng Commun 5:68–77
Xu P, Qiao K, Stephanopoulos G (2017) Engineering oxidative stress defense pathways to build a robust lipid production platform in Yarrowia lipolytica. Biotechnol Bioeng 114:1521–1530
Xuan Y, Xing W, Hu X, Zhen L, Liu J, Zhou H, Shen X, Wei Y, Huang Z, Ying W, Wang Y, Nie YH, Zhang CC, Li S, Cheng L, Wang Q, Wu Y, Huang P, Sun Q, Shi L, Yang H (2017) Homology-mediated end joining-based targeted integration using CRISPR/Cas9. Cell Res 27:801–814
Xue Z, Sharpe PL, Hong SP, Yadav NS, Xie D, Short DR, Damude HG, Rupert RA, Seip JE, Wang J, Pollak DW, Bostick MW, Bosak MD, Macool DJ, Hollerbach DH, Zhang H, Arcilla DM, Bledsoe SA, Croker K, McCord EF, Tyreus BD, Jackson EN, Zhu Q (2013) Production of omega-3 eicosapentaenoic acid by metabolic engineering of Yarrowia lipolytica. Nat Biotechnol 31:734–740
Yao X, Liu Z, Wang X, Wang Y, Nie YH, Lai L, Sun R, Shi L, Sun Q, Yang H (2018) Generation of knock-in cynomolgus monkey via CRISPR/Cas9 editing. Cell Res 28:379–382
Yin X, Madzak C, Du G, Zhou J, Chen J (2012) Enhanced alpha-ketoglutaric acid production in Yarrowia lipolytica, WSH-Z06 by regulation of the pyruvate carboxylation pathway. Appl Microbiol Biotechnol 96:1527–1537
Zeng SY, Liu HH, Shi TQ, Song P, Ren LJ, Huang H, Ji XJ (2018) Recent advances in metabolic engineering of Yarrowia lipolytica for lipid overproduction. Eur J Lipid Sci Technol 120:1700352
Zhang JL, Peng YZ, Liu D, Liu H, Cao YX, Li BZ, Li C, Yuan YJ (2018) Gene repression via multiplex gRNA strategy in Y. lipolytica. Microb Cell Factories 17:62–74
Funding
This work was financially supported by the National Natural Science Foundation of China (Grant Nos. 21776131 and 21476111), the Program for Innovative Research Teams in Universities of Jiangsu Province, the Priority Academic Program for the Development of Jiangsu Higher Education Institutions, the Jiangsu Synergetic Innovation Center for Advanced Bio-Manufacture (No. XTD1814), and the Åforsk Foundation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no conflicts of interest.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Shi, TQ., Huang, H., Kerkhoven, E.J. et al. Advancing metabolic engineering of Yarrowia lipolytica using the CRISPR/Cas system. Appl Microbiol Biotechnol 102, 9541–9548 (2018). https://doi.org/10.1007/s00253-018-9366-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9366-x