Abstract
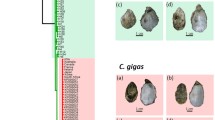
The Kumamoto oyster (Crassostrea sikamea) shows a spatially restricted distribution, favoring estuarine tideland environment. On the other hand, the Pacific oyster (C. gigas) has a broader range of habitat. The present study compared the mitochondrial population structure between the two closely related species. For accurate species identification of oysters sampled from Japanese and East Asian continental coasts, we performed sequencing analysis of the mitochondrial DNA (mtDNA) and PCR-RFLP assay of the first internal transcribed spacer of nuclear rRNA genes. Then, we estimated the extent of population differentiation within each of C. sikamea and C. gigas based on the mtDNA data. Few haplotypes were shared among the sites of sampling in C. sikamea, which contrasted with an extensive haplotype sharing among C. gigas samples. We discuss the mechanisms of elevated population differentiation observed in C. sikamea in light of the ecology and the ancient ocean geography around the present-day habitats.






Similar content being viewed by others
References
Akashi H (1995) Inferring weak selection from patterns of polymorphism and divergence at “silent” sites in Drosophila DNA. Genetics 139:1067–1076
Amemiya I (1928) Ecological studies of Japanese oysters, with special reference to the salinity of their habitats. J Coll Agric Univ Tokyo 9:333–382
An HK, Jee YJ, Park DW, Ryu HY, Min KS (2000) Genetic variation in populations of the Pacific oyster (Crassostrea gigas), based on the mitochondrial COI gene sequence. Korean J Genet 22:249–255
Andolfatto P (2005) Adaptive evolution of non-coding DNA in Drosophila. Nature 437:1149–1152
Aris-Brosou S, Excoffier L (1996) The impact of population expansion and mutation rate heterogeneity on DNA sequence polymorphism. Mol Biol Evol 13:494–506
Avise JC (2000) Phylogeography: the history and formation of species. Harvard University Press, Cambridge
Avise JC, Arnold J, Ball RM, Bermingham E, Lamb T, Neigell JE, Reeb CA, Saunders NC (1987) Intraspecific phylogeography: the mitochondrial DNA bridge between population genetics and systematics. Annu Rev Ecol Syst 18:489–522
Bandelt HJ, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16:37–48
Banks MA, Hedgecock D (1993) Discrimination between closely related Pacific oyster species (Crassostrea) via mitochondrial DNA sequences coding for large subunit rRNA. Mol Mar Biol Biotechnol 2:129–136
Banks MA, McGoldrick DJ, Borgeson W, Hedgecock D (1994) Gametic incompatibility and genetic divergence of Pacific and Kumamoto oysters, Crassostrea gigas and C. sikamea. Mar Biol 121:127–135
Bazin E, Glémin S, Galtier N (2006) Population size does not influence mitochondrial genetic diversity in animals. Science 312:570–572
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B 57:289–300
Bermingham E, Moritz C (1998) Comparative phylogeography: concepts and applications. Mol Ecol 7:367–369
Bernatchez L, Wilson CC (1998) Comparative phylogeography of Nearctic and Palearctic fishes. Mol Ecol 7:431–452
Bertness MD, Gaines SD (1993) Larval dispersal and local adaptation in acorn barnacles. Evolution 47:316–320
Buroker NE, Hershberger WK, Chew KK (1979) Population genetics of the family Ostreidae. I. Intraspecific studies of Crassostrea gigas and Saccostrea commercialis. Mar Biol 54:157–169
Camara MD, Davis JP, Sekino M, Hedgecock D, Li G, Langdon CJ, Evans S (2008) The Kumamoto oyster Crassostrea sikamea is neither rare nor threatened by hybridization in the northern Ariake Sea, Japan. J Shellfish Res 27:313–322
Carlini DB, Stephan W (2003) In vivo introduction of unpreferred synonymous codons into the Drosophila Adh gene results in reduced levels of ADH protein. Genetics 163:239–243
Chen Z, Song B, Wang Z, Cai Y (2000) Late Quaternary evolution of the sub-aqueous Yangtze Delta, China: sedimentation, stratigraphy, palynology, and deformation. Mar Geol 162:423–441
Chew KK (1990) Global bivalve shellfish introductions. World Aquac 21:9–22
Choi KS (2008) Oyster capture-based aquaculture in the Republic of Korea. In: FAO Fisheries Technical Paper, vol 508, pp 271–286
Cordes JF, Xiao J, Reece KS (2008) Discrimination of nine Crassostrea oyster species based upon restriction fragment-length polymorphism analysis of nuclear and mitochondrial DNA markers. J Shellfish Res 27:1155–1161
Dowling DK, Friberg U, Lindell J (2008) Evolutionary implications of non-neutral mitochondrial genetic variation. Trends Ecol Evol 23:546–554
Egea R, Casillas S, Barbadilla A (2008) Standard and generalized McDonald-Kreitman test: a website to detect selection by comparing different classes of DNA sites. Nucleic Acids Res 36: W157–W162 (web server issue)
Emery KO, Niino H, Sullivan B (1971) Post-Pleistocene levels of East China Sea. In: Trekian KK (ed) Late Cenozoic glacial ages. Yale University Press, New Heaven, pp 381–390
Ernande B, Clobert J, Mccombie H, Boudry P (2003) Genetic polymorphism and trade-off in the early life-history strategy of the Pacific oyster, Crassostrea gigas (Thunberg, 1795): a quantitative genetic study. J Evol Biol 16:399–414
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Fan D, Li C (2008) Timing of the Yangtze initiation draining the Tibetan Plateau throughout to the East China Sea: a review. Front Earth Sci China 2:302–313
FAO (2005) Cultured Aquatic Species Information Programme (Crassostrea gigas). In: Helm MM (ed) FAO Fisheries and Aquaculture Department (online), Rome http://www.fao.org/fishery/culturedspecies/Crassostrea_gigas/en
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R (1994) DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrate. Mol Mar Biol Biotechnol 3:294–299
Fu Y-X (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147:915–925
Fu Y-X, Li W-H (1993) Statistical tests of neutrality of mutations. Genetics 133:693–709
Grant WS, Bowen BW (1998) Shallow population histories in deep evolutionary lineages of marine fishes: insights from sardines and anchovies and lessons for conservation. J Hered 89:415–426
Guo X, Ford SE, Zhang F (1999) Molluscan aquaculture in China. J Shellfish Res 18:19–31
Heads M (2005) Towards a panbiogeography of the seas. Biol J Linn Soc 84:675–723
Hedgecock D (1994) Does variance in reproductive success limit effective population size of marine organisms? In: Beaumont A (ed) Genetics and evolution of aquaculture organisms. Chapman & Hall, London, pp 122–134
Hedgecock D, Pudovkin AI (2011) Sweepstakes reproductive success in highly fecund marine fish and shellfish: a review and commentary. Bull Mar Sci 87:971–1002
Hedgecock D, Li G, Banks MA, Kain Z (1999) Occurrence of the Kumamoto oyster Crassostrea sikamea in the Ariake Sea, Japan. Mar Biol 133:65–68
Hedrick PW (2000) Genetics of populations. Jones and Bartlett Publishers, Sudbury
Hewitt GM (2000) The genetic legacy of the Quaternary ice ages. Nature 405:907–913
Hewitt GM (2004) Genetic consequences of climatic oscillations in the Quaternary. Philos Trans R Soc Lond B 359:183–195
Hong J-S, Sekino M, Sato S (2012) Molecular species diagnosis confirmed the occurrence of Kumamoto oyster Crassostrea sikamea in Korean waters. Fish Sci 78:259–267
Hori K, Saito Y, Zhao Q, Wang P (2002) Architecture and evolution of the tide-dominated Changjiang (Yangtze) River delta, China. Sediment Geol 146:249–264
Hughes AL (2007) Looking for Darwin in all the wrong places: the misguided quest for positive selection at the nucleotide sequence level. Heredity 99:364–373
Hughes AL (2008) The origin of adaptive phenotypes. Proc Natl Acad Sci USA 105:13193–13194
Ijiri A, Wang L, Oba T, Kawahata H, Huang C-Y, Huang C-Y (2005) Paleoenvironmental changes in the northern area of the East China Sea during the past 42,000 years. Palaeogeogr Palaeoclimateol Palaeoecol 219:239–261
Jukes TH, Cantor CR (1969) Evolution of protein molecules. Academic Press, New York, pp 21–132
Lam K, Morton B (2004) The oysters of Hong Kong (Bivalvia: Ostreidae and Gryphaeidae). Raffles B Zool 52:11–28
Lambeck K, Esat TM, Potter E-K (2002) Links between climate and sea levels for the past three million years. Nature 419:199–206
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Liu K-B, Sun S, Jiang X (1992) Environmental change in the Yangtze River delta since 12,000 years BP. Quaternary Res 38:32–45
Liu J-X, Gao T-X, Yokogawa K, Zhang Y-P (2006a) Differential population structuring and demographic history of two closely related fish species, Japanese Sea bass (Lateolabrax japonicus) and spotted sea bass (Lateolabrax maculatus) in Northwestern Pacific. Mol Phyl Evol 39:799–811
Liu J-X, Gao T-X, Yokogawa K, Zhang Y-P (2006b) Late Pleistocene divergence and subsequent population expansion of two closely related fish species, Japanese anchovy (Engraulis japonicus) and Australian anchovy (Engraulis austrails). Mol Phyl Evol 40:712–723
McDonald JH, Kreitman M (1991) Adaptive protein evolution at the Adh locus in Drosophila. Nature 351:652–654
Meiklejohn CD, Montooth KL, Rand DM (2007) Positive and negative selection on the mitochondrial genome. Trends Genet 23:259–263
Milbury CA, Gaffney PM (2005) Complete mitochondrial DNA sequence of the eastern oyster Crassostrea virginica. Mar Biotechnol 7:697–712
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, New York
Nielsen R (2001) Statistical tests of selective neutrality in the age of genomics. Heredity 86:641–647
Numachi K (1971) Biological research on the oyster. In: Imai T (ed) Aquaculture in shallow seas: progress in shallow sea culture. Koseisha Koseikaku, Tokyo, pp 52–105 (in Japanese)
Ó Foighil D, Gaffney PM, Wilbur AE, Hilbish TJ (1998) Mitochondrial cytochrome oxidase I gene sequences support an Asian origin for the Portuguese oyster Crassostrea angulata. Mar Biol 131:497–503
Ohta T (1972a) Evolutionary rate of cistrons and DNA divergence. J Mol Evol 1:150–157
Ohta T (1972b) Population size and rate of evolution. J Mol Evol 1:305–314
Ohta T (1973) Slightly deleterious mutant substitutions in evolution. Nature 246:96–98
Oizumi S, Ito S, Koganezawa A, Sakai S, Sato R, Kanno H (1971) Techniques of oyster culture. In: Imai T (ed) Aquaculture in shallow seas: progress in shallow sea culture. Koseisha Koseikaku, Tokyo, pp 149–185 (in Japanese)
Oliveira DCSG, Raychoudhury R, Lavrov DV, Werren JH (2008) Rapidly evolving mitochondrial genome and directional selection in mitochondrial genes in the parasitic wasp Nasonia (Hymenoptera: Pteromalidae). Mol Biol Evol 25:2167–2180
Petit RJ, El Mousadik A, Pons O (1998) Identifying populations for conservation on the basis of genetic markers. Conserv Biol 12:844–855
Polzin T, Daneschmand SV (2003) On Steiner trees and minimum spanning trees in hypergraphs. Oper Res Lett 31:12–20
Posada D, Crandall KA (2001) Intraspecific gene genealogies: trees grafting into networks. Trends Ecol Evol 16:37–45
Provan J, Bennett KD (2008) Phylogeographic insights into cryptic glacial refugia. Trends Ecol Evol 23:564–571
Rand DM (2001) The units of selection on mitochondrial DNA. Annu Rev Ecol Syst 32:415–448
Rand DM, Kann LM (1996) Excess amino acid polymorphism in mitochondrial DNA: contrasts among genes from Drosophila, mice, and humans. Mol Biol Evol 13:735–748
Rand DM, Haney RA, Fry AJ (2004) Cytonuclear coevolution: the genomics of cooperation. Trends Ecol Evol 19:645–653
Reece KS, Cordes JF, Stubbs JB, Hudson KL, Francis EA (2008) Molecular phylogenies help resolve taxonomic confusion with Asian Crassostrea oyster species. Mar Biol 153:709–721
Richardson NJ, Densmore AL, Seward D, Wipf M, Yong L (2010) Did incision of the three Gorges begin in the Eocene? Geology 38:551–554
Rozen S, Skaletsky HJ (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana Press, Totowa, pp 365–386
Saitoh K, Chen WJ (2008) Reducing cloning artifacts for recovery of allelic sequences by T7 endonuclease I cleavage and single re-extension of PCR products—a benchmark. Gene 423:92–95
Sambrook J, Fritsch EF, Manitaris T (1989) Molecular cloning, 2nd edn. Cold Spring Harbor Laboratory Press, Plainview
Sargsyan O, Wakeley J (2008) A coalescent process with simultaneous multiple mergers for approximating the gene genealogies of many marine organisms. Theor Pop Biol 74:104–114
Sato S (2000) Bivalves: focusing on the species distributed in Isahaya Bay. In: Sato M (ed) Life in Ariake Sea: biodiversity in tidal flats and estuaries. Kaiyusha, Tokyo, pp 150–183 (in Japanese)
Sato M, Takita T (2000) Biota and environment of Ariake Sea. In: Sato M (ed) Life in Ariake Sea: biodiversity in tidal flats and estuaries. Kaiyusha, Tokyo, pp 10–36 (in Japanese)
Sekino M (2009) In search of the Kumamoto oyster Crassostrea sikamea (Amemiya, 1928) based on molecular markers: is the natural resource at stake? Fish Sci 75:319–331
Sekino M, Kobayashi T, Hara M (2006) Segregation and linkage analysis of 75 novel microsatellite DNA markers in pair crosses of Japanese abalone (Haliotis discus hannai) using the 5’-tailed primer method. Mar Biotechnol 8:453–466
Selkoe KA, Toonen RJ (2011) Marine connectivity: a new look at pelagic larval duration and genetic metrics of dispersal. Mar Ecol Prog Ser 436:291–305
Shimoyama S (2000) Geographic history of the Ariake Sea and development of Ariake-endemic species. In: Sato M (ed) Life in Ariake Sea: biodiversity in tidal flats and estuaries. Kaiyusha, Tokyo, pp 37–48 (in Japanese)
Simonsen KL, Churchill GA, Aquadro CF (1995) Properties of statistical tests of neutrality for DNA polymorphism data. Genetics 141:413–429
Stewart JR, Lister AM (2001) Cryptic northern refugia and the origins of the modern biota. Trends Ecol Evol 16:608–613
Sugawara Y, Koganezawa A (1994) Advances in aquaculture of Pacific oyster, Japanese scallop, and Ezo abalones: Oyster (Crassostrea gigas). In: Nomura M (ed) Oyster, scallop and abalone: advances in culture of molluscs and related fields of research. Koseisha Koseikaku, Tokyo, pp 1–17 (in Japanese)
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tanaka M (2007) Relict estuarine ecosystem isolated from the continental coastal waters. Aquabiology 29:3–9 (in Japanese with English abstract)
Thompson JD, Higgins DJ, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Uehara K, Saito Y (2003) Late Quaternary evolution of the Yellow/East China Sea tidal regime and its impacts on sediments dispersal and seafloor morphology. Sediment Geol 162:25–38
Uehara K, Saito Y, Hori K (2002) Paleotidal regime in the Changjiang (Yantze) Estuary, the East China Sea, and the Yellow Sea at 6 ka and 10 ka estimated from a numerical model. Mar Geol 183:179–192
Usuki H (2002) Evaluation of characteristics and preservation of Pacific oyster, Crassostrea gigas, in view of the genetic resources. Bull Fish Res Agency 4:40–104 (in Japanese with English abstract)
Wang P (1999) Response of Western Pacific marginal seas to glacial cycles: paleoceanographic and sedimentological features. Mar Geol 156:5–39
Wang Y, Aubrey DG (1987) The characteristics of the China coastline. Cont Shelf Res 7:329–349
Wang J, Wang P (1980) Relationship between sea-level changes and climatic fluctuations in East China Sea since late Pleistocene. Acta Geogr Sin 35:299–312 (in Chinese with English abstract)
Wang H, Zhang G, Liu X, Guo X (2008) Classification of common oysters from north China. J Shellfish Res 27:495–503
Wu X, Xu X, Yu Z, Wei Z, Xia J (2010) Comparison of seven Crassostrea mitogenomes and phylogenetic analysis. Mol Phyl Evol 57:448–454
Xiao S, Li A, Jiang F, Li T, Wan S, Huang P (2004) The history of the Yangtze River entering sea since the Last Glacial Maxima: a review and look forward. J Coast Res 20:599–604
Xu X, Oda M (1999) Surface-water evolution of the eastern East China Sea during the last 36,000 years. Mar Geol 156:285–304
Yu H, Li Q (2012) Complete mitochondrial DNA sequence of Crassostrea nippona: comparative and phylogenomic studies on seven commercial Crassostrea species. Mol Biol Rep 39:999–1009
Zhao B, Guo H, Yan Y, Wang Q, Li B (2008) A simple waterline approach for tidelands using multi-temporal satellite images: a case study in the Yangtze Delta. Estuar Coast Shelf Sci 77:134–142
Acknowledgments
We are grateful to professor D. Hedgecock, University of Southern California, for kindly reading the earlier draft of the manuscript and providing many insightful comments. Many constructive suggestions from two anonymous reviewers and handling editor helped improve the manuscript. We also thank Mr. K. Kitahara of Seikai Yoshoku Giken and Dr. R. Fuseya of National Research Institute of Fisheries Engineering, Fisheries Research Agency of Japan, for their assistance to MS’s expedition to Ariake Sea and Omura Bay for sample collection. QL’s graduate students helped MS to collect oysters in Qingdao.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by T. Reusch.
Appendices
Appendix 1
See Table 6.
Appendix 2
See Table 7.
Appendix 3
See Table 8.
Rights and permissions
About this article
Cite this article
Sekino, M., Sato, S., Hong, JS. et al. Contrasting pattern of mitochondrial population diversity between an estuarine bivalve, the Kumamoto oyster Crassostrea sikamea, and the closely related Pacific oyster C. gigas . Mar Biol 159, 2757–2776 (2012). https://doi.org/10.1007/s00227-012-2037-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00227-012-2037-z