Abstract
This study focusses on the molecular and morphological variation among Scandinavian species within the Syntrichia ruralis complex (S. calcicola, S. norvegica, S. ruraliformis, S. ruralis) of the moss family Pottiaceae, plus the similar-looking S. princeps. Molecular variation was explored based on the internal transcribed spacer 1 (ITS1), and the plastid atpB-rbcL spacer and rpl16 G2 intron. The relationships of the S. ruralis complex taxa were evaluated by including twelve additional, morphologically defined Syntrichia species in the ingroup, including S. subpapillosissima that was here shown to occur in Scandinavia. The molecular evidence favours a wide circumscription of the S. ruralis complex, including the species around S. caninervis and some other ones but excluding S. princeps, and that these species are closely related. ITS1 paralogues were revealed in almost one-third of the samples, and for those cloned between 2 and 8 variants were found, including specimens with paralogues belonging to two (2 cases) or three (1) different species. Together with several cases of discrepancy between ITS1 and plastid relationships, this could suggest an exchange of genetic material between species and may explain the extensive and partly overlapping morphological variation among some of them. Syntrichia subpapillosissima and S. ruralis var. epilosa may represent special phenotypes within S. ruraliformis or S. ruralis, but studies of more material of these are required to decide their correct status.
Similar content being viewed by others
Introduction
The genus Syntrichia Brid. in the family Pottiaceae includes around 90 species (Frey and Stech 2009), distributed almost all over the terrestrial world. The history of the genus is almost as long as that of moss nomenclature (for a review, see Gallego 2005). However, it was only with its typification (Zander 1989) and later use in the encyclopaedic overview of the Pottiaceae by Zander (1993) that the name Syntrichia became widely established in the 1990s for a group of species that had earlier mostly been included in Tortula Hedw. Zander’s treatment of Syntrichia as distinct from Tortula was later supported also by molecular data (e.g. Spagnuolo et al. 1999; Werner et al. 2002).
Within Syntrichia, the treatment of several taxa has been debated, and perhaps especially several of the species around S. ruralis (Hedw.) F.Weber & D.Mohr have attracted much attention over the last 40 years (Gallego 2005; Gallego et al. 2002a; Kramer 1980; Mishler 1985; Vanderpoorten 2001). However, there is still no consensus regarding which taxa to accept, or at which level to recognize several of these. For Scandinavia, the ‘S. ruralis complex’ is from here onwards understood to include four taxa that are easily confused with each other based on their similar morphologies, S. calcicola J.J.Amann, S. norvegica F.Weber, S. ruraliformis (Besch.) Mans., and S. ruralis. In Europe, where the morphological diversity encompassed by these names is greatest, different authors recognize between one and four species (Gallego 2006; Gallego et al. 2002a; Hallingbäck et al. 2008; Hill et al. 2006; Ignatov et al. 2006; Kramer 1980; Nyholm 1991; Ochyra et al. 2003; Smith 2004; Vanderpoorten 2001). Sometimes, a fifth taxon is recognized at the species level within the complex, under one of the names S. densa (Velen.) J.-P.Frahm or S. glabra J.-P.Frahm & M.T.Gallego (Frahm and Gallego 2001; Frahm and Sabovljević 2006). Finally, S. princeps (De Not.) Mitt. has a similar habit and habitat and is frequently confused with species of the S. ruralis complex in Scandinavia. Considering the variable numbers of species recognized by different authors, it is evident that morphological, anatomical, and biosystematic information is not sufficient to settle several of the species-level problems.
The focus of this study is on resolving remaining issues within the S. ruralis complex and S. princeps in Scandinavia. A full understanding of positions and relationships among lineages in this complex within the genus requires studies of related species, and representatives of twelve additional Syntrichia species are, therefore, included. We use molecular information from numerous specimens of the S. ruralis complex and S. princeps to seek for potential congruence with morphology. This study (1) identifies which Scandinavian lineages are distinct and correspond with taxa within the S. ruralis complex and S. princeps, (2) tests whether distinct molecular lineages differ in a selection of morphological features, and (3) discusses the variation within species and the problems encountered in distinguishing them from each other.
Materials and methods
Study species and material
We studied mainly specimens collected by the first author and kept at S, with the addition of herbarium specimens from GOET, MUB, and S. The Scandinavian members of the S. ruralis complex, S. calcicola, S. norvegica, S. ruraliformis, and S. ruralis, typically occur in relatively base-rich to calcareous habitats that are dry during long periods. They occur on rocks, boulders, thin soil overlayering rocks, on sand or sandy soil, and occasionally as epiphytes on trees. They occur in forests as well as in environments without trees, including alvar, heaths, and sand dunes. Representatives of this species complex grow throughout Scandinavia, although less commonly so in large portions of the boreal and boreo-nemoral zones, and some of its species occur mainly in relatively small regions. In Scandinavia, members of the S. ruralis complex are especially abundant on the two Baltic Sea islands Öland and Gotland, where their phenotypic variation is also great. Scandinavian species of the S. ruralis complex are widely distributed in Europe, Africa, Asia, North and South America, and Australia (Hill et al. 2006; Ignatov et al. 2006; Kramer 1980, 1988; Mishler 2007; Streimann and Klazenga 2002), although individual species have more restricted distributions. Thus, S. calcicola occurs only in Europe, N Africa (incl. Macaronesia) and SW Asia (Gallego et al. 2002a; Ignatov et al. 2006; Kramer 1980). Members of the S. ruralis complex are dioicous (separate male and female plants). The vegetative leaves are broad, tongue shaped or upwards gradually narrowed, not constricted at mid-leaf, and have recurved leaf margins. The leaves are usually recurved when moist. They have a single costa that lacks hydroids and usually ends in a short or long, spinose to denticulate hairpoint. Like in other Syntrichia species, the leaf lamina consists of isodiametric or shortly elongate and more or less strongly papillose cells in its middle and upper portions, whereas the cells in the basal portion are differentiated into rectangular or longly rectangular, thin-walled, inflated, and more or less hyaline cells, with narrow non-hyaline cells both towards the costa and leaf margin. The similar-looking S. princeps can be synoicous or dioicous, has wide hydroids in the costa, and the leaves are usually constricted at the middle. Its global distribution is similar to that of the S. ruralis complex (Gallego 2005), but in Scandinavia it occurs only on the Baltic Sea islands Bornholm (Kramer 1980), Öland (this paper) and Gotland (Hedenäs 1993). This species typically grows on stable, leached calcareous sand or gravel. On Gotland, the species is most frequent along the coasts, but it has also scattered inland occurrences.
We sampled 174 specimens of Syntrichia in the ingroup, including 139 of members of the S. ruralis complex, as understood in Scandinavia. This covers the variation in Scandinavia, but includes also selected samples from other portions of Europe and a few samples from other continents. We sampled 28 specimens morphologically identifiable as S. calcicola, 32 as S. norvegica, 37 as S. ruraliformis, and 42 as S. ruralis [including two specimens of var. epilosa (Venturi) J.J.Amann]. We sampled twelve specimens of S. princeps. To understand the position of the Scandinavian S. ruralis complex in relation to other members of the genus, we included 1–3 specimens of S. caninervis Mitt. (3), S. handelii (Schiffn.) S.Agnew & Vondr. (2), S. laevipila Brid. (2), S. latifolia (Hartm.) Huebener (2), S. montana Nees (2), S. papillosa (Spruce) Spruce (2), S. papillosissima (Copp.) Loeske (2), S. pseudohandelii (J. Froehl.) S.Agnew & Vondr. (1), S. rigescens (Broth. & Geh.) Ochyra (2), S. sinensis (Müll.Hal.) Ochyra (1), S. subpapillosissima (W.A.Kramer) M.T.Gallego & J. Guerra (2), and S. virescens (De Not.) Ochyra (2). We selected five more distantly related Pottiaceae members as outgroup, based on overview studies by Inoue et al. (2012) and Werner et al. (2004), one specimen per species: Cinclidotus fontinaloides (Hedw.) P.Beauv., Leptophascum leptophyllum (Müll.Hal.) J.Guerra & M.J.Cano, Microbryum davallianum (Sm.) R.H.Zander, Tortella tortuosa (Hedw.) Limpr., and Tortula acaulon (With.) R.H.Zander. Specimen data are provided in “Appendix”.
Molecular methods
Total DNA was extracted using the DNeasy® Plant Mini Kit for DNA isolation from plant tissue (QIAGEN). Double-stranded DNA templates were prepared by polymerase chain reaction (PCR). PCR was performed using IllustraTM Hot Start Mix RTG (GE Healthcare) in a 25-µl reaction volume according to the manufacturer’s instructions.
Initially, variation in the nuclear internal transcribed spacer 1 (ITS1), and the plastid atpB-rbcL spacer (atpB-rbcL), rpl16 G2 intron (rpl16), rps4 gene + trnS-rps4 spacer (rps4), trnGUCC G2 intron (trnG), and trnLUAA-trnFGAA spacer (trnL-trnF) were explored for 2 specimens with the morphology of S. calcicola, 6 of S. norvegica, 5 of S. ruraliformis, and 7 of S. ruralis. The three most variable ones, ITS1, atpB-rbcL, and rpl16 were selected for the investigation. For the three used molecular markers, the PCR programs given below were initiated by a denaturation step of 5 min at 95°C and were followed by a final extension period of 10 min at 72°C. For ITS1, the PCR program employed was 40 cycles of 30 s at 95°C, 30 s at 52°C, and 1 min at 72°C, with various combinations of the primers ‘ITS4-bryo’ (Stech 1999), ‘ITSbryoR’ (Hedenäs 2014), ‘5.8SC’ (Bartish et al. 2005), and occasionally ‘18SF’ and ‘26SR’ (Rydin et al. 2004). For atpB-rbcL, 40 cycles of 30 s at 94°C, 30 s at 52°C, and 40 s at 72°C were employed, with the primers ‘ATPB-1’ and ‘RBCL-1′ (Chiang et al. 1998). For rpl16, 5 cycles of 30 s 94°C, 30 s 57–53°C, 30 s 72°, followed by 30 cycles of 30 s at 94°C, 30 s at 52°C, and 1 min 30 s at 72°C were employed, with the primers ‘F71’ (Jordan et al. 1996) and ‘rpl16-antR2’ (Hedenäs 2008) were used.
Twenty millilitres of each amplified fragment was cleaned using a mixture of 20 units of Exonuclease I from E.coli and 4 units of FastAP TM Thermosensitive Alkaline Phosphatase (Fermentas LIFE SCIENCE), mixed and incubated at 37°C for 30 min and inactivated at 80°C for 15 min. Cycle sequencing was performed using the ABI BigDye Terminator Kit (Applied Biosystems) according to the instructions on the kit (BDT ver. 3.1). The sequencing products were cleaned using the DyeEx® 96 Kit (QIAGEN). The same primers as for the initial PCR were used. Sequencing products were resolved on an ABI3130xl automated sequencer. Double-stranded sequencing was performed.
In 55 of the total 179 samples (31%), the amplified ITS1 yielded only messy sequence curves. Thus, for four of the outgroup taxa, one of S. caninervis, and 28 specimens of the S. ruralis complex, representing the different morphological units, DNA was cloned in order to check whether ITS paralogues caused the observed patterns and, if so, the potential origin of different paralogues.
The amplified fragment was ligated into a pTZ57R/T TA cloning vector using the Thermo Scientific InsTAclone PCR Cloning Kit (Thermo Fisher Scientific, Waltham, MA, USA), according to the manufacturer’s instructions. Escherichia coli K12 JM101 cells (New England Biolabs, Ipswich, MA, USA) were made chemically competent and were then transformed with the recombinant pTZ57R/T vector as described in the InsTAclone PCR Cloning Kit protocol. Colony PCR was performed on 6–8 successfully transformed colonies per sample using the primers ‘ITS-bryoR’ and ‘ITS-5.8SC’, and then sequenced as described above.
Sequence editing and analysis
We edited and assembled nucleotide sequence fragments for each DNA region and aligned the assembled sequences manually, using PhyDE® 0.9971 (http://www.phyde.de/index.html; accessed 6 Jan 2017). We identified regions of partially incomplete data in the beginning and end of the sequences and excluded these from subsequent analyses. Insertions and deletions were coded using Simple Indel Coding (Simmons and Ochoterena 2000) in the program SeqStat (Müller 2005). The indels provided additional evidence, and we present the analyses with these included. The sequence alignments used in the analyses are found in Online Resources 1 and 2. GenBank accession numbers are listed “Appendix”.
Preliminary NeighborNet (NN) split network analyses with SplitsTree 4.12.6 (Huson and Bryant 2006) showed that networks based on ITS1 and plastid data, respectively, place several specimens among members of different species in the two data sets. Thus, the two data sets provide incongruent results and, in addition, reticulation is frequent. Therefore and because the numerous specimens with several ITS1 paralogues cannot be combined with corresponding plastid sequences, we ran the final analyses separately for ITS1 and plastid data, and generated NN split networks using SplitsTree 4.12.6. Jackknife analyses (1000 replications) were performed with the program TNT (Goloboff et al. 2003) to test whether there exists supported lineages in a tree context.
Each paralogue variant was included in the ITS1 analyses. When we retrieved two or more representatives of a paralogue variant, we excluded all except one. In the analyses of relationships, only specimens with complete plastid data (both markers) were included in the overall analysis based on plastid data. The positions in the networks of specimens, for which only one plastid marker could be generated, were checked in supplementary analyses based on a single plastid marker for consistency regarding the relation to specimens where both markers were available. The results of these supplementary results were included here only for S. papillosa, for which no plastid information would otherwise have been available (Fig. 1b).
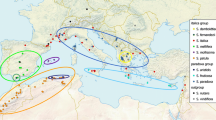
NeighborNet split network for the sampled Syntrichia species, based on ITS1 (a) and the plastid markers atpB-rbcL and rpl16 (b) specimens with both markers available are included in the network, the approximate position of S. papillosa is estimated from atpB-rbcL only. Thick black lines across lineages indicate Jackknife support of 95–100. Species of the S. ruralis complex and S. princeps are in bold coloured fonts, with the portions of the network corresponding with the respective morphologically defined species encircled in the same colour. Different lineages (occasionally groups) within a species are encircled and indicated by numbers (1, 2, etc.). When encircled by a broken line, only different ITS1 paralogues belong to this lineage. Lineage 3 in 2b with the two samples of var. epilosa (P325, P435), includes also ten specimens with hairpoints. Other Syntrichia species and outgroup ones Cinclidotus fontinaloides, Leptophascum leptophyllum, Microbryum davallianum, Tortella tortuosa and Tortula acaulon are in normal fonts. When more than one sample of such species were included, the number studied is indicated in parenthesis after the species name and when samples had to be cloned, this is indicated by a ‘p’ after the number of paralogues in this sample; for S. caninervis, one out of two samples was cloned. A detailed resolution of the S. princeps lineage is found in Fig. 4
Morphology
After clarifying the molecular relationships among the studied Syntrichia ruralis complex specimens, we studied the morphology of selected specimens belonging to the distinguished molecular entities to confirm that the molecular patterns correspond with morphologically circumscribed species. We used an Olympus-BX43 light microscope (LM) with an Olympus SC50 digital camera (DC) mounted on this microscope. An approach including both standard comparisons of qualitative and quantitative characters and the quantification of sizes of vegetative leaf features and leaf lamina cells was used (cf., Hedenäs 2017a). We applied the typical anatomical and morphological methods used for the Pottiaceae (Zander 1993). Further, among molecularly identified entities of the S. ruralis complex, the main entities S. calcicola, two plastid variants of S. norvegica, three plastid variants of S. ruralis, and two ITS variants of S. ruraliformis we sampled 10–11 specimens each. We also sampled six specimens which molecular signal was incongruent between ITS1 and the plastid markers at the species level, to test whether such specimens are morphologically intermediate between those lineages suggested by the different molecular markers. The specimens which leaves and cells were measured are indicated with an asterisk (*) in “Appendix”. For each of these specimens, we sampled four vegetative leaves from two shoots (two leaves from each shoot, to avoid sampling all leaves from an untypical shoot for the specimen). For each leaf, we measured length and maximal width, width of the costa, and length of the hairpoint. Further, we recorded the length, width, and length to width ratio of 20 cells in the middle of the upper lamina and 20 in the differentiated portion of the basal lamina. We produced temporary images of the leaves with the LM and DC equipment mentioned above, and the Olympus cellSens Standard 1.13 software (Olympus Corporation) for automatic and continuous image stacking. We then measured the features from these leaf and cell images, using the Olympus cellSens Standard 1.13 software. While sporophytic features are not very variable (Gallego 2005), except that the basal membrane of the peristome is shorter in S. norvegica and to some degree S. calcicola than in S. ruralis and S. ruraliformis, and sporophytes are often absent from specimens, we scored only gametophytic characters.
We base our comparisons among the entities within the S. ruralis complex on two approaches. First, we compared the measurements between the molecularly identified entities, both for raw measurements and for measurements standardized to a leaf length, excluding hairpoint, of 2 mm, and a leaf width of 1 mm. The latter adjustment was made to compensate for potential effects of leaf size on cell size, of the kind found by Hedenäs (1996). Shapiro Wilks W-test (normality) and Levenes test (homogeneity of variance) were both statistically significant, and inspection of the distributions of residuals in preliminary Anovas (normality) showed that the data do not meet the criteria of normality and homogeneity of variance. Thus, we used the nonparametric Kruskal–Wallis test for multiple comparisons to compare the cell measurements among the entities. Second, we subjected the measurements of the individual leaves (leaf length, leaf width, costa width near base, mean leaf lamina cell length, mean leaf lamina cell with, at mid-leaf and leaf base; in total seven parameters) to a Principal Component Analysis (PCA) to see whether the combined morphological leaf information corresponds with the molecularly identified entities. All statistical calculations were made in STATISTICA 12 (StatSoft 2013).
Results
The total number of aligned ITS1 sites in the 174 studied Syntrichia specimens, and five outgroup specimens (257 terminals including paralogues), after deletion of regions at the beginnings and ends that were incomplete for some specimens, was 1005. Of these, 35.6% were variable (24.5% in Syntrichia; 14.2% in the S. ruralis complex), and 28.1% (17.7%; 7.3%) parsimony-informative; 676 (450; 233) indels were present, with 97.2% (95.8%; 79.4%) parsimony-informative. For atpB-rbcL the length was 729, 19.6% (4.5%; 1.4%) were variable, and 18.6% (3.2%; 1.1%) parsimony-informative; 87 (21; 4) indels with 97.7% (100%; 100%) parsimony-informative. For rpl16 the length was 711, 25.0% (12.1%; 9.3) were variable, and 9.4% (5.5%; 2.1%) parsimony-informative; 96 (38; 21) indels with 100% (100%; 100%) parsimony-informative. The sequence lengths for the species were (numbers of samples and ITS1 paralogues, see “Appendix”): S. calcicola: 329–401 (ITS1), 535 (atpB-rbcL), 619–621 (rpl16); S. caninervis: 403–404, 534, 621; S. handelii: 396, 534, 621; S. laevipila: 373, 538, 622; S. latifolia: 412, 535, 623; S. montana: 395–575, 535, 620; S. norvegica: 396–430, 535–536, 620–621; S. papillosa: 377, 540, –; S. papillosissima: 413, 535, 621; S. princeps: 326–329, 536–538, 617; S. pseudohandelii: 414, 536, 620; S. rigescens: 397, 535, 621; S. ruraliformis: 377–420, 535, 621; S. ruralis: 386–416, 534–538, 621; S. sinensis: 359, 536, 618–623; S. subpapillosissima: 390, 535, 621; S. virescens: 399, 534, 624; and the outgroup species Cinclidotus fontinaloides: 353–421, 519, 635; Leptophascum leptophyllum: 280, 603, 629; Microbryum davallianum: 291–294, 534, 608; Tortella tortuosa: 303, 543, 621; Tortula acaulon: 196, 537, 623.
The NeighborNet split networks for both ITS1 and the plastid markers place the Scandinavian members of the S. ruralis complex and S. princeps firmly within Syntrichia (Fig. 1a). Syntrichia handelii, S. latifolia, S. montana, S. rigescens, S. papillosissima, S. pseudohandelii, S. subpapillosissima, and S. virescens are closely related to the Scandinavian S. ruralis complex species, whereas S. laevipila, S. papillosa, S. princeps, and S. sinensis are more distantly related to this complex. Although S. caninervis appears somewhat separate from the other members of the S. ruralis complex according to the ITS1 network, its position with the complex has a high Jackknife support (100), and the structure of the plastid network supports this (Fig. 1b). According to the plastid markers, the Scandinavian S. ruralis complex specimens form four distinct lineages or groups, corresponding with the four morphologically defined species S. calcicola, S. norvegica (except one outlying specimen), S. ruraliformis, and S. ruralis (Fig. 1b). ITS1 data also place such specimens in four groups of closely related lineages, but except for S. calcicola these groups appear more closely related than according to the plastid network (Fig. 1a). Neither of the Scandinavian S. ruralis complex species has a high Jackknife support, but a close relationship between one S. calcicola lineage and S. montana is well-supported (99). Two specimens of S. ruralis var. epilosa are nested within S. ruralis according to both ITS1 and the plastid markers. In both networks, but especially in the ITS1 one, some specimens have a morphology that mainly agrees with that based on the other data set, and some ITS1 paralogue(s) of a specimen belong to other species than the morphologically defined one. Such incongruent patterns are described in further detail in the following.
Specimens with messy ITS1 curves, suggesting the occurrence of paralogues, occurred both in the outgroup and ingroup, and in the S. ruralis complex they were found throughout the sampled European area. The samples that were cloned were thus selected from throughout the area. All cloned samples for which more than one clone was successfully amplified (all species except Tortula acaulon) yielded at least two different ITS1 paralogues or sets of paralogues. In most cases the different variants belonged to the same intraspecific ITS1 lineage, but in twelve of the S. ruralis complex specimens the clones belonged to different ITS1 lineages, and in three cases the different paralogues could be referred to two (2 cases) or three (1) different species (Fig. 2, Table 1). In the three cases where clones belonged to more than one species, at least one of these was the same as that suggested by the plastid markers. In nine cases, including P364 where the plastid markers suggests S. calcicola and ITS1 S. princeps, ITS1 and plastid sequences suggested different species (Figs. 1b, 3, Table 1).
The Syntrichia ruralis complex portion of the ITS1 NeighborNet split network in Fig. 1a, showing where different paralogues of the 28 cloned specimens belong. ‘P’ numbers are sample numbers (“Appendix”); the number of paralogues that belong to a specific lineage is provided after the hyphen following the number. Dashed lines connect paralogues that belong to different lineages within a specimen
The Syntrichia ruralis complex portion of the ITS1 NeighborNet split network in Fig. 1a, showing where specimens of ambiguous molecular identity belong according to their ITS1 and plastid markers, respectively. The plastid markers place the specimens with violet rings and black numbers in S. ruraliformis and those with yellow rings and grey numbers in S. ruralis. ‘P’ numbers are sample numbers (“Appendix”); the underlined numbers include paralogues belonging to lineages of different species according to ITS1
Syntrichia norvegica samples belonging to plastid marker lineages 1 and 2 (CHL1 + 2) were found only in the Scandinavian mountain range and the Alps, whereas those belonging to lineage 3 (CHL3) were collected only in the Scandinavian mountain range and on the Baltic Sea islands Gotland and Öland. Within S. ruraliformis, specimens belonging to ITS1 lineages 1 and 2 (ITS1 + 2) are almost exclusively coastal and grow predominantly in sandy habitats, whereas ITS1 lineage 3 (ITS3) tends to grow in inland habitats and mainly on limestone or thin soil over mainly calcareous rocks. Besides the minor quantitative morphological differences described below, ITS3 plants of S. ruraliformis tend to have a more strongly papillose dorsal costa than ITS1 + 2 plants, including more frequently branched papillae. In ITS3 plants the lamina also tends to extend less longly up along the basal hairpoint and this extending portion is also less strongly hyaline than in ITS1 + 2 plants. The S. ruralis plastid marker lineage 3 (CHL3) occurs in the Baltic Sea islands Öland and Gotland, in the Stockholm archipelago, and in an exposed limestone area in Västmanland (S Swedish mainland), whereas the other two S. ruralis plastid marker groups are more widespread. We found no habitat differentiation among these three lineages.
In S. princeps, ITS1 and the plastid markers yield congruent results (Fig. 4), except for one German specimen (P364; Fig. 4a) that has ITS1 of S. princeps and morphology as well as plastid markers of S. calcicola (Fig. 1b).
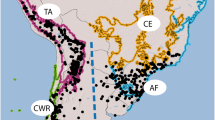
The Syntrichia princeps lineages of the ITS1 (a) and plastid (b) NeighborNet split networks of Fig. 1. The geographic origin of each sample is indicated by two capital letters, as explained in the box, and ‘P’ numbers are sample numbers (“Appendix”). The underlined sample GE-P364 belongs to S. calcicola according to morphology and the plastid markers (Fig. 1b)
The correspondence between identified molecular lineages or groups within species and measured quantitative morphological features was weak (Fig. 5). Even when there was statistically significant differentiation in leaf cell size between molecular entities within a species, the differences were slight and the overlaps great. At the species level, the differences were larger, even if the overlap also here was often relatively large. Thus, S. norvegica and S. calcicola had shorter leaves and, especially S. calcicola, shorter hairpoints than most of the entities within the other two species, whereas S. ruraliformis had larger and wider leaves with a longer hairpoint than most of the other three (Fig. 5). As regards cell measurements, S. norvegica and S. calcicola have larger and slightly more elongate mid-leaf cells than the other two species, whereas these cells are smaller in S. ruraliformis than in the other three. The size differences are clearest when cell sizes are adjusted for the size of the measured leaf. Cell size differentiation was found also for the basal leaf lamina cells, but the pattern is less distinct than for the upper cells (Fig. 5).
Boxplots with median values, quartiles, and whiskers from maximum to minimum values, leaf and cell sizes in different Syntrichia lineages. Lineage abbreviations: norv-C3: norvegica chloroplast lineage 3; norv-C1 + 2: norvegica chloroplast lineages 1 and 2; rurf-I3: ruraliformis ITS lineage 3; rurfI-1 + 2: ruraliformis ITS lineages 1 and 2; calc: calcicola; rura-C1 + 4: ruralis chloroplast lineages 1 and 4; rura-C2: ruralis chloroplast lineage 2; rura-C3: ruralis chloroplast lineage 3. Lineage numbers correspond with those in Fig. 1. From each molecularly identified specimen, four leaves were arbitrarily selected from two different stems; in each leaf 20 cells were measured in mid-leaf, and 20 in the differentiated basal groups. The measurements are based on ten specimens per lineage, except for rurf-I1 + 2 (n = 11). In the leaf 20 cells were measured. Thus, the number of measured leaves (leaf length, leaf width, hairpoint length, costa width) is 40 and the number of measured cells is 800 per lineage, except for ruraliformis ITS lineages 1 and 2 (44, 880). Lineages, which values differ significantly from each other, have different letters to the right of the bars. Details of the statistic comparisons is found in Online Resource 3
The significant overlap in the measured features is evident also from the PCA (Fig. 6a). The samples of the four species are grouped with S. norvegica in the lower left, followed by S. calcicola, S, ruralis, and finally S. ruraliformis in the upper right. This reflects the differentiation between small leaves and large cells in the left or lower left of the diagram and large leaves and small cells in the opposite end of the gradient (Fig. 6b). Neither of the lineages or groups within the species can be clearly distinguished in the PCA, although the lineages/groups do not overlap totally within the respective species’ distribution along axes 1 and 2. Specimens with incongruent ITS and plastid sequences at the species level, marked in grey in Fig. 6, are indicated with detailed information in Fig. 7. Although the general morphology of the specimens agrees best with the plastid marker identity, their measurements in some cases place them outside these species, or in some cases even outside all four species.
a The positions of four leaves from each of ten molecularly identified specimens per lineage (except for ruraliformis ITS lineages 1 and 2 = 11 specimens), along the first two axes in a PCA. The PCA is based on each leaf’s length (LL) and width (LW), costa width near base (CW) and leaf lamina cell length and width in mid-leaf (ML, MW) and differentiated basal portion (BL, BW). Cell measurements are the mean values of 20 cells per position in each leaf. Factors 1 and 2 explain 40.44% and 33.21% of the variation. b Explanatory factors in the plane of factors 1 and 2
Because the two different marker sets or different ITS1 paralogues suggested that several specimens have genetic material from more than one species, we briefly describe morphological peculiarities of such specimens below. Except for three specimens mentioned below with S. ruralis morphology and S. ruraliformis plastid markers, their morphology is consistent or almost consistent with the species suggested by the plastid information.
One German specimen with S. calcicola morphology had ITS1 from S. princeps. This specimen, P364, is morphologically typical S. calcicola. Two specimens with S. ruralis morphology have ITS1 paralogues belonging to both S. ruralis and S. norvegica. One of these, P403 from northern Norway, lacks dorsal stereids in the upper third of some young leaves, like in S. norvegica, but the size of the leaf lamina cells agrees with S. ruralis. In the other one, P407 from the middle of the Scandinavian mountain range, plants with normal S. ruralis appearance, having leaves spirally curved when dry and with long hyaline hairpoints, give rise to young and somewhat flageliform branches that often have characteristics of S. norvegica. Thus, the leaf lamina cells are wider and the dorsal costal stereids disappear in the uppermost costa.
For three specimens with S. ruralis morphology, ITS1 rather than plastid data suggest this species. In specimen P357 from California, the margins are not recurved all the way up to the apical region. Because a MUB duplicate (MUB45428) of this specimen belongs to S. princeps (synoicous), several species were likely growing together and included in the original collection that was divided into duplicates. Specimen P413 from the middle of the Scandinavian mountain range looks like a typical S. ruralis. P470 from the Baltic Sea island Öland has mostly the morphology of S. ruralis, but in several plants some leaves have an excurrent leaf apex and the leaf lamina cells are 10.0–13.0 (16.0) µm, which overlaps more with S. ruraliformis than with S. ruralis.
P395 from northern Norway and P434 from the Baltic Sea island Öland have the morphology of S. ruraliformis but ITS1 implies S. ruralis. The lamina extends up along the hairpoint, and the lamina cells are slightly wider than normal in this species (P395: 12.5–17.0 µm; P434: 12.5–14.0 µm). P391 from northern Norway has ITS1 paralogues from both S. norvegica, S. ruraliformis, and S. ruralis. It is morphologically similar to P395, except that in some young leaves the dorsal costal stereids disappear in the upper part of the leaf, like in S. norvegica. In P400 from Hungary, with ITS1 from S. norvegica, the leaf lamina extends up along the hairpoint as in S. ruraliformis, but the extending lamina is not hyaline. The mid-leaf cells are smaller than in S. norvegica, 11.0–15.0 µm wide, and the dorsal costa stereids do not disappear near the leaf apex.
Two specimens from the Baltic Sea Island Öland had the morphology of S. subpapillosissima (P454) or were morphologically most similar to this species (P463). Specimen P454 grew on a horizontal rock, whereas P463 grew on deep soil, and was intermixed with S. calcicola and S. ruralis. According to our molecular results, specimens with S. subpapillosissima morphology are nested within S. ruralis (ITS1) or S. ruraliformis (plastid markers) (Fig. 1).
Discussion
Scandinavian members of the Syntrichia ruralis complex
The Scandinavian S. ruralis complex is part of a larger group of species, of which only some occur in our study area. This group is supported both by the NN split network structures and by a high ITS1 Jackknife support, and includes at least S. calcicola, S. caninervis, S. handelii, S. latifolia, S. montana, S. norvegica, S. papillosissima, S. pseudohandelii, S. rigescens, S. ruraliformis, S. ruralis, S. subpapillosissima, and S. virescens. The group is currently circumscribed by molecular evidence only, since we have not been able to find morphological or anatomical synapomorphies to support it. Where species sampling overlaps, the circumscription of this group agrees with the ITS2 relationships of Afonina et al. (2014). On the other hand, it is at variance with ideas on relationships based on morphology and anatomy presented by Kramer (1980) and Mishler (1985), who included S. princeps in this group, and Gallego et al. (2002a) and Gallego et al. (2002b), who separated the S. ruralis and S. caninervis complexes of species. Neither of the two latter complexes are monophyletic, and their species occur mixed among each other in the NeighborNet split networks (Fig. 1). The character states that supposedly define either group, such as the bistratose leaf lamina, ovate to lanceolate leaves, usually not constricted at the middle, costa with hydroids, and strongly spinose hairpoint in the S. caninervis complex (Gallego et al. 2002b), are, therefore, not synapomorphies, but in the case of the S. caninervis complex may represent independently evolved analogous states. However, because our study addresses the Scandinavian species referred to the S. ruralis complex, we will continue to use the term S. ruralis complex in this narrow sense here. We are aware that additional Scandinavian as well as extra-Scandinavian species belong in a more widely circumscribed group of Syntrichia species, and some of our observations may, therefore, be valid for additional species and other geographical regions.
The status of the sometimes accepted, non-Scandinavian S. densa (Velen.) J.-P.Frahm, of which S. glabra J.-P.Frahm & M.T.Gallego was claimed to represent juvenile plants (Frahm and Sabovljević 2006), as distinct from S. calcicola seems doubtful (Gallego et al. 2002a; Vanderpoorten 2001). Frahm and Sabovljević (2006) studied its molecular relationships with S. calcicola and S. ruralis. Because their study was based on ITS, we had originally planned to compare S. densa with Scandinavian material, but surprisingly the GenBank sequences of Frahm and Sabovljević (2006) actually belong to trnL-trnF. In view of this puzzling discrepancy, we decided not to pursue this comparison.
Chloroplast information place our focal members of the S. ruralis complex in four entities that almost invariably correspond with the morphologically defined species S. calcicola, S. norvegica, S. ruraliformis, and S. ruralis (see below, and Gallego et al. 2002a). One N Italian specimen of S. norvegica (P386) appears at a position outside these groups, due to its deviating rpl16 sequence, but ITS1 as well as morphology place it firmly in this species, although its leaf lamina sometimes extends slightly up along the hairpoint. ITS1 distinguishes the same four species, but except for S. calcicola the species appear relatively more closely related. Syntrichia subpapillosissima from Spain (P436) and two southern Scandinavian specimens that morphologically clearly belong to this species (P454), or is most similar to this species (P463), are nested within S. ruraliformis according to plastid information and within S. ruralis according to ITS1 information. This suggests that S. subpapillosissima may represent a specific phenotype of either of these two species, or possibly it is at least partly of hybridogen origin (see below), rather than being a species of its own. The taxon clearly requires further studies before its status can be finally resolved.
Moreover, almost a third of the specimens have sequence curve patterns suggesting two or more ITS1 paralogues. Among the cloned specimens, the retrieved ITS1 sequences sometimes belong to more than one species. In addition, several specimens appear in a different species by ITS1 than by plastid or morphological information. Paralogues were observed earlier in the family Pottiaceae, in the genus Tortula (Košnar et al. 2012), and our study shows that they are widespread in this family. Outside the Scandinavian S. ruralis complex, they were found in S. caninervis, Cinclidotus fontinaloides, Leptophascum leptophyllum, and Microbryum davallianum, but on the other hand appear to be rare or absent in members of the Trichostomoideae (Alonso et al. 2016; Hedenäs 2015a; Köckinger and Hedenäs 2017; Werner et al. 2005).
Both ITS1 and the plastid markers suggest that S. princeps includes two lineages (Fig. 4), possibly of different origins. One includes most European specimens together with those from Cyprus and Turkey, and the other includes the German and Californian specimens. Based on the present sampling it cannot be decided whether the eastern Mediterranean-southern Scandinavian group differs from the entire more western European population. This requires additional sampling in the western Mediterranean to Britain. However, disjunctions between Europe and Western North America are well-known (Schofield and Crum 1972), and small genetic differences between populations of these areas (Hedenäs 2008; Shaw et al. 2003; Werner et al. 2003) suggest that relatively recent long-distance dispersal explains this pattern. Morphology of the studied plants provides no additional evidence on the differentiation between the lineages. The single North American plant included (P356) has slightly narrower leaf lamina cells, 9–14 µm, than European plants, mostly 12 -> 16 µm (Gallego 2006; Nyholm 1991; Smith 2004). Most of the European specimens are synoicous, but the North American and one of the Scandinavian specimens (P440) have inflorescences that are either synoicous or female; in the North American specimen, some individual plants even vary between years.
Morphology of the Scandinavian Syntrichia ruralis complex species
Earlier studies of the S. ruralis group that included species of the Scandinavian S. ruralis complex compared morphology alone, or contrasted plants cultivated under different conditions (Gallego 2005; Gallego et al. 2002a, b; Kramer 1980; Mishler 1985; Vanderpoorten 2001). Comparative cultivation experiments provide extremely valuable information on the phenotypic, habitat-related variation in several quantitative characters, including leaf shape and leaf lamina cell size (Mishler 1985). However, such studies may be misleading if the genetic identity of the study plants is unknown or the phenotypic variation of species is very wide. Likewise, studies exclusively of morphological features may be misleading because the identity of extreme plants of strongly variable species can be unclear and may confuse the delimitation of taxa. Here we compare a set of morphological characters in molecularly unambiguously identified specimens.
We found some differences in cell size between molecular entities within species, but the differences were slight and could not distinguish such entities in practical identification work. However, together with the mentioned differences between S. norvergica lineages in geographical distributions, or habitat preferences and relative strength of abaxial costa ornamentation in S. ruraliformis, these could be signs of incipient speciation. The species-level differences were greater, with S. calcicola and S. norvegica having relatively short leaves, large leaf lamina cells, and short hairpoints (less distinct in S. calcicola), which agrees with the conditions in the Mediterranean area (Gallego 2005; Gallego et al. 2002a), and S. ruraliformis having large leaves, a wide costa, a long hairpoint, and small leaf lamina cells. The cell size differences were clearer for the upper leaf lamina cells and with cell size adjusted for leaf size. Our results show that cell size at least partly correlates with leaf size, in agreement with earlier observations in mosses (e.g. Hedenäs 1996; Hedenäs et al. 2014; Loeske 1907). The species distinguishing quantitative characters agree roughly with the circumscriptions of earlier authors, but the smaller leaf cells in S. ruraliformis than in S. ruralis, despite its relatively larger leaves, are significant. This characteristic was noted by some earlier authors (Kramer 1980; Vanderpoorten 2001), but was not considered significant enough for recognition at the species level by Gallego (2005) and Gallego et al. (2002a), and was overlooked by Mishler (1985). The qualitative characters that distinguish the species totally agree with those of earlier authors. Specimens molecularly grouped under S. norvegica, according to both ITS1 and plastid markers, have dorsal costal stereids disappearing at the apical part of the leaves, have mostly orange hairpoints, and have not hydroids in the costa. On the other hand, specimens molecularly grouped under S. ruraliformis, according to both marker sets, have usually an acuminate hyaline leaf apex tapering into the hairpoint.
Despite their striking appearance, S. ruralis specimens lacking hairpoints (var. epilosa) are nested among S. ruralis specimens with hairpoints. However, plants without hairpoints often grow together with plants having hairpoints, without intermediate plants. Our results, therefore, preliminarily suggest that lack of hairpoint represents a minor genetic variant within S. ruralis. Because similar plants occur in S. caninervis and S. montana (Gallego et al. 2002b) and S. norvegica (Gallego et al. 2018; Kramer 1980) in the S. ruralis group, hairpoint loss could be due to a simple mechanism. On the other hand, S. latifolia within the same species group is characterized by a constant lack of hairpoint. While it is evident that each case requires individual investigation, molecular studies including larger numbers of epilose specimens are required to explore both their evolution and how to recognize them taxonomically.
The PCA based on the measured quantitative characters shows that S. calcicola overlaps with S. norvegica, both these small species with large lamina cells overlap significantly with S. ruralis but hardly with the much larger S. ruraliformis, which in addition has small lamina cells, and S. ruralis and S. ruraliformis overlap to a relatively high degree. The overlap between intraspecific molecular entities within species is even greater, and it is obvious that neither of these are distinguishable by the quantitative features in combination. The considerable overlap in the PCA also illustrates the danger in trusting such information to explore which taxa to recognize. Without independent evidence, either from other morphological characters or from molecular data, the interpretation of the PCA results is extremely difficult. Fortunately, in the present case there exists both molecular and other morphological features that on their own or together effectively distinguish the four species.
Understanding intraspecific variation and incongruences
ITS1 paralogues from a single specimen mostly differ in few bases or belong to different lineages within a species, as found for paralogues in other species by Shaw et al. (2002). Especially differences within lineages could potentially be an effect of mutations originating faster than can be countered by concerted evolution (Košnar et al. 2012). However, in cases where paralogues belong to different lineages, or when different ITS1 paralogues belong to two or even three different species, hybridization or other exchange of genetic material seems more likely. If the most widely different ITS1 paralogue identities found in individual specimens are instead a result of incomplete lineage sorting, then concerted evolution must be very inefficient. The numerous cases where ITS1 species identities deviate from those suggested by morphology and plastid information do not contradict an explanation involving hybridization or other exchange of genetic material. Morphological hybrids are known in other Pottiaceae genera (Natcheva and Cronberg 2004), and molecular patterns similar to those in Syntrichia occur in other mosses (Draper and Hedenäs 2009; Draper et al. 2007, 2015; Hedenäs 2015b, 2017b; Natcheva and Cronberg 2007b; Shaw and Goffinet 2000). Allopolyploids (including hybrids between members of different conspecific lineages), could also explain some of the morphological variation within Syntrichia species (Morrison 2014; Vanderpoorten et al. 2004; Wyatt et al. 1988). The varying chromosome numbers found among the studied members of the S. ruralis group (n = 12, 12 + m, 13, 13 + m, 24 + m, 26) as well as among Syntrichia species outside this group (n = 6, 12, 24, 24 + m26, 28, 36 + m) clearly suggest different ploidy levels in different species (Fritsch 1991). Causes for species-level incongruence include, beside hybridization, for example, insufficient data, rapid diversification, horizontal gene transfer, incomplete lineage sorting, convergence caused by natural selection, and variation in evolutionary rates (Harris 2008; Morrison 2014; Wendel and Doyle 1998). Also for plastids, horizontal transfer has been shown to occur in vascular plants (Acosta and Premoli 2010; Stegemann et al. 2012). Although this could potentially explain plastid reticulation or incongruences with nuclear data in bryophytes (see also Hedenäs 2012, 2015a, b, 2017b), we refrain from further speculation on this until firm evidence for bryophytes is at hand. However, evidence additional to results of phylogenetic analyses is mostly required to decide which explanation is most likely (Morrison 2014; Wendel and Doyle 1998).
In combination with our observations on specimens having morphological traits from more than one species when they appear in positions incongruent between the ITS1 and plastid networks, certain features of the reproductive biology of Syntrichia species suggest that hybridization may indeed explain the observed incongruences and different ITS1 paralogue identities. If hybridization caused the found patterns, and moss chloroplasts are maternally inherited (Duckett et al. 1983; McDaniel et al. 2007; Natcheva and Cronberg 2007a), the male parent contributed the ITS1 sequence. Male expressing plants are much rarer than female expressing ones in the Scandinavian S. ruralis complex, especially in relatively dry habitats. Scandinavian non-sporophytic specimens with female expressing shoots are between 7.5 and ≥ 10 times as common as male ones, depending on the species and geographic area (total n = 214), except for S. ruralis in the S Swedish mainland (3.2 times as common; n = 53) (Hedenäs, full data to be published elsewhere). A strong female bias was found also in S. caninervis (Baughman et al. 2017; Bowker et al. 2000), especially strong for expressed sex, suggesting a phylogenetic component in explaining this imbalance within the larger group around S. ruralis (cf., Bisang et al. 2014). Sporophytes are occasional or relatively rare in our species. We, therefore, suggest that a possible proximate mechanism that could explain frequent hybridization within the S. ruralis complex is that when males are very rare there is a great risk that the genes of female (and consequently male) plants will not be propagated through sexual reproduction. It is then advantageous for the genes if the reproductive barriers are incomplete and allow not only conspecific sexual reproduction; as a precaution females can occasionally also be fertilised by males of closely related species that may be available. Interestingly, such potential hybridization is not restricted to within the S. ruralis complex, because one German specimen with S. calcicola morphology and plastid identity (P364) had ITS1 of S. princeps type. If the hypothesis on imperfect reproductive barriers is correct, this suggests that Syntrichia species outside the group around S. ruralis are not completely reproductively isolated from the latter. We believe that further studies including more material of those species presently represented by few specimens and from other geographical regions are required to clarify if our suggested hypothesis is likely to be correct. Experiments aiming at producing interspecific hybrids would also be valuable.
When we know that Syntrichia species are plastic (Mishler 1985) and likely through some mechanism exchange DNA between intraspecific lineages as well as species, sometimes even distantly related ones, the strong overlap in the measured features is easier to understand. The positions of measured specimens with molecular markers from more than one species are frequently outside one or both parental species in the PCA (Fig. 7), suggesting that transfer of genetic material between species may lead to the development of features beyond the parental phenotypes. If hybridization or other exchange of genetic material is as common in the S. ruralis complex as suggested here, coding genetic material that has moved between species through introgression could thus contribute to the wide phenotypic range of the species. Beside phenotypic plasticity, this could explain the rather wide overlap seen between the species in quantitative features in the PCA. Obviously, the entire S. ruralis group requires further detailed studies to evaluate whether it can function as a model for studies of the exchange of genetic material in bryophytes.
Brief characterizations of the Scandinavian species of the Syntrichia ruralis complex
Based on our observations of qualitative as well as quantitative features in the molecularly distinguished specimens, the four Scandinavian species of the S. ruralis complex can be well characterized, and mostly the differentiating features agree with those found for the Mediterranean area (Gallego et al. 2002a). If S. subpapillosissima is accepted as a species, this would be a fifth Scandinavian species. Several of the differentiating features are quantitative and overlap significantly between species (see Fig. 5); these require some experience to apply in practical identification work. As a rule of thumb, S. ruraliformis is the largest species, S. calcicola and S. norvegica the smallest, with the size of S. ruralis in between.
Syntrichia ruralis
The leaves are recurved when moist, with margins recurved up to near the apex, have a rounded or acute apex and the lamina does not extend up along the basal hairpoint. The average size of the leaf lamina cells is smaller than in S. calcicola and S. norvegica, and larger than in S. ruraliformis, but due to a wide overlap, comparisons of measurements are most reliable from mixed collections or if adjusted for the leaf size. Rarely, occasional leaves in S. ruralis have a costa that disappears near the apex (P403).
Syntrichia ruraliformis
The leaves taper to the apex, with the upper lamina portions that extend more or less broadly up along the basal hairpoint mainly hyaline. In rare S. ruralis specimens, occasional leaves may have a lamina that extends up narrowly along the costa and even be partly hyaline, but this is never as consistent as in S. ruraliformis. The average lamina cells size, especially when adjusted for leaf size, is smaller than in S. ruralis, but because of the great overlap between these two species, Gallego et al. (2002a) treated S. ruraliformis as a variety of S. ruralis. Two samples with S. subpapillosissima morphology were found from Öland in southern Sweden (P454, P463); one of these, P463, was somewhat untypical for this taxon. Syntrichia subpapillosissima has high and pedicellate leaf lamina papillae, a strongly papillose dorsal surface of the costa, and a strongly hyaline apex tapering to the apical portion of the leaf (Fig. 8). Like the Öland specimens, Mediterranean S. subpapillosissima have longer hairpoints and a more strongly papillose dorsal surface of the costa than S. ruraliformis. All these features indicate a possible adaptation of S. subpapillosissima to arid conditions.
Illustration of Syntrichia subpapillosissima (W.A.Kramer) M.T.Gallego & J.Guerra based on specimen P454 (Sweden. Öland, L.Hedenäs; S, reg. no. B236078). a Transition between leaf lamina and hairpoint. b Leaf lamina cells from mid-leaf in surface view. c Leaf lamina cells from mid-leaf in transverse section. d Transverse section of costa in mid-leaf
Syntrichia norvegica
Moist leaves are, on the average, more strongly recurved than in S. ruralis. It deviates from the other three species in that the stereids of the upper dorsal costa disappear just below the hairpoint; it looks as if lamina cells cover the costa. Otherwise, this species is rather similar to S calcicola in size, in their leaf margin typically recurved in its lower 50–65%, and in having relatively large leaf lamina cells and short hairpoints, but the hairpoints are typically orange-red throughout or in large portions (orange-red hairpoints are rare in S. calcicola).
Syntrichia calcicola
This species differs from S. norvegica in its less strongly recurved moist leaves, a mostly hyaline hairpoint, and the dorsal costa has stereids throughout its length. The hairpoint in S. calcicola is sometimes longer than in S. norvegica. It is less strongly spinose and when dry more strongly crisped than in other S. ruralis complex species, and the leaves tend to have a shorter hyaline basal area of the leaf than the other three species.
Taxonomic treatment
Key to Scandinavian Syntrichia species
-
1a.
Plants with vegetative diaspores … 2
-
1b.
Plants without vegetative diaspores … 4
-
2a.
Vegetative diaspores in the form of brood leaves, at the stem apex or in the axils of the upper leaves, often forming a rosette at the shoot apex … S. laevipila
-
2b.
Vegetative diaspores globular, ovate or rounded, smooth, arising on the ventral side of the leaf (laminar or costal gemmae) … 3
-
3a.
Leaves without hairpoints; lamina with papillae on the dorsal and ventral surface; gemmae laminar … S. latifolia
-
3b.
Leaves with hairpoints, typically smooth; lamina with papillae only on dorsal surface; gemmae costal … S. papillosa
-
4a.
Costa with hydroids … 5
-
4b.
Costa without hydroids … 7
-
5a.
Hairpoints smooth, rarely weakly spinulose; leaves bordered or not, with border usually consisting of 2 rows of thick-walled less papillose cells, sometimes smooth; dioicous or autoicous … S. laevipila
-
5b.
Hairpoints spinose; leaves not bordered; dioicous or synoicous … 6
-
6a.
Mid-leaf lamina cells 5.0–10.0 (12.5) µm wide; dioicous … S. montana
-
6b.
Mid-leaf lamina cells 12.5–17.5 (20.0) µm wide; synoicous or more rarely dioicous … S. princeps
-
7a.
Costa transverse section with 1–2(3) dorsal stereid rows; leaves constricted at the middle; margins plane or lightly recurved up to the mid-leaf … S. virescens
-
7b.
Costa transverse section with (2)3–6 dorsal stereid rows; leaves not constricted at the middle; margins revolute up to the middle, upper third, or up to near the leaf apex … 8
-
8a.
Dorsal costa stereids disappearing near the leaf apex; hairpoints typically orange-red throughout or in large portions … S. norvegica
-
8b.
Dorsal costa stereids not disappearing near the leaf apex; hairpoints typically hyaline, sometimes brown at base or very rarely reddish brown in larger portions … 9
-
9a.
Leaf margins revolute up to the upper third, rarely up to the middle; mid-leaf lamina cells (9.5)13.5–16.0(21.0) µm wide; hairpoints spinulose … S. calcicola
-
9b.
Leaf margins revolute up to near the apex, sometimes up to upper third; mid-leaf lamina cells (6.5)10.0–14.5(21.0) µm wide; hairpoints strongly spinose … 10
-
10a.
Papillae on mid-leaf lamina cells pedicellate, 5.0–7.5 μm high … S. subpapillosissima
-
10b.
Papillae on mid-leaf lamina cells not pedicellate, c. 2.5 μm high … 11
-
11a.
Leaf apex usually rounded, sometimes acute, not tapering to the apex, mainly chlorophyllose … S. ruralis
-
11b.
Leaf apex acuminate, tapering to the apex, mainly hyaline … S. ruraliformis
References
Acosta MC, Premoli AC (2010) Evidence of chloroplast capture in South American Nothofagus (subgenus Nothofagus, Nothofagaceae). Molec Phylogen Evol 54:235–242. https://doi.org/10.1016/j.ympev.2009.08.008
Afonina OM, Ignatova EA, Fedosov VE, Kuznetsova OI (2014) Toward a new understanding of Syntrichia submontana (Pottiaceae, Bryophyta). Arctoa 23:11–24. https://doi.org/10.15298/arctoa.23.03
Alonso M, Jiménez JA, Nylinder S, Hedenäs L, Cano MJ (2016) Disentangling generic limits in Chionoloma, Oxystegus, Pachyneuropsis and Pseudosymblepharis (Bryophyta: Pottiaceae): an inquiry into their phylogenetic relationships. Taxon 65:3–18. https://doi.org/10.12705/651.1
Bartish IV, Swenson U, Munzinger J, Anderberg AA (2005) Phylogenetic relationships among New Caledonian Sapotaceae (Ericales): molecular evidence for generic polyphyly and repeated dispersal. Amer J Bot 92:667–773
Baughman JT, Payton AC, Paasch AE, Fisher KM, McDaniel SF (2017) Multiple factors influence population sex ratios in the Mojave Desert moss Syntrichia caninervis. Amer J Bot 104:733–742. https://doi.org/10.3732/ajb.1700045
Bisang I, Ehrlén J, Persson C, Hedenäs L (2014) Family affiliation, sex ratio and sporophyte frequency in unisexual mosses. Bot J Linn Soc 174:163–172. https://doi.org/10.1111/boj.12135
Bowker MA, Stark LR, McLetchie DN, Mishler BD (2000) Sex expression, skewed sex ratio, and microhabitat distribution in the dioecious desert moss Syntrichia caninervis (Pottiaceae). Amer J Bot 87:517–526
Chiang TI, Schaal BA, Peng C-I (1998) Universal primers for amplification and sequencing a noncoding spacer between the atpB and rbcL genes of chloroplast DNA. Bot Bull Acad Sin 39:245–250
Draper I, Hedenäs L (2009) Circumscription of European taxa within the Sciuro-hypnum reflexum complex (Brachytheciaceae, Bryophyta), based on molecular and morphological data. Taxon 58:572–584
Draper I, Hedenäs L, Grimm GW (2007) Molecular and morphological incongruence in European species of Isothecium (Bryophyta). Molec Phylogen Evol 42:700–716. https://doi.org/10.1016/j.ympev.2006.09.021
Draper I et al (2015) How many species of Isothecium (Lembophyllaceae, Bryophyta) are there in Macaronesia? A survey using integrative taxonomy. Bot J Linn Soc 177:418–438. https://doi.org/10.1111/boj.12250
Duckett JG, Carothers ZB, Miller CCJ (1983) Gametogenesis. In: Schuster RM (ed) New manual of bryology, vol. 1. The Hattori Botanical Laboratory, Nichinan, pp 232–275
Frahm J-P, Gallego MT (2001) Syntrichia glabra, a new moss from Germany. J Bryol 23:119–122. https://doi.org/10.1179/jbr.2001.23.2.119
Frahm J-P, Sabovljević M (2006) Preliminary results of the taxonomic value of Tortula densa (Velen.) J.-P. Frahm inferred from the Internal Transcribed Spacer (ITS) of the nrDNA. Cryptogam Bryol 27:405–412
Frey W, Stech M (2009) Division of Bryophyta Schimp. (Musci, Mosses). In: Frey W (ed) Syllabus of plant families. Adolf Engler’s Syllabus der Pflanzenfamilien, 13th edition. Part 3. Bryophytes and seedless vascular plants. Gebrüder Borntraeger, Berlin, pp 116–257
Fritsch R (1991) Index to bryophyte chromosome counts. Bryophyt Biblioth 40:1–352
Gallego MT (2005) A taxonomic study of the genus Syntrichia Brid. (Pottiaceae, Musci) in the Mediterranean region and Macaronesia. J Hattori Bot Lab 98:47–122
Gallego MT (2006) 20. Syntrichia brid. In: Guerra J, Cano MJ, Ros RM (eds) Flora Briofítica Ibérica, volumen III. Pottiales: Pottiaceae. Encalyptales: Encalyptaceae. Universidad de Murcia/Sociedad Española de Briología, Murcia, pp 120–145
Gallego MT, Cano MJ, Ros RM, Guerra J (2002a) An overview of Syntrichia ruralis complex (Pottiaceae: Musci) in the Mediterranean region and neighbouring areas. Bot J Linn Soc 138:209–224
Gallego MT, Cano MJ, Ros RM, Guerra J (2002b) New taxonomic data on a circum-Tethyan group of Syntrichia (Pottiaceae, Bryophyta): the S. caninervis complex. Syst Bot 27:643–653
Gallego MT, Hugonnot V, Cano MJ (2018) Taxonomic resurrection of an awnless variety of Syntrichia ruralis and comparison with other European muticous taxa in this genus. J Bryol 40:244–250. https://doi.org/10.1080/03736687.2018.1468971
Goloboff P, Farris J, Nixon K (2003) Tree analysis using new technology. Available at: http://www.lillo.org.ar/phylogeny/tnt/. Accessed 3 May 2017
Hallingbäck T, Lönnell N, Weibull H, von Knorring P, Korotynska M, Reisborg C, Birgersson M (2008) Nationalnyckeln till Sveriges flora och fauna. Bladmossor: Kompaktmossor-kapmossor. Bryophyta: Anoectangium-Orthodontium. ArtDatabanken, SLU, Uppsala
Harris ESJ (2008) Paraphyly and multiple causes of phylogenetic incongruence in the moss genus Plagiomnium (Mniaceae). Taxon 57:417–433
Hedenäs L (1993) Tortula princeps, stäppskruvmossa, i Sverige. Svensk Bot Tidskr 87:71–79
Hedenäs L (1996) On the interdependence of some leaf characters within the Drepanocladus aduncus-polycarpus complex. J Bryol 19:311–324
Hedenäs L (2008) Molecular variation and speciation in Antitrichia curtipendula s. l. (Leucodontaceae, Bryophyta). Bot J Linn Soc 156:341–354. https://doi.org/10.1111/j.1095-8339.2007.00775.x
Hedenäs L (2012) Global phylogeography in Sanionia uncinata (Amblystegiaceae, Bryophyta). Bot J Linn Soc 168:19–42. https://doi.org/10.1111/j.1095-8339.2011.01189.x
Hedenäs L (2014) Intraspecific genetic variation in selected mosses of Scandinavian interglacial refugia suggests contrasting distribution history patterns. Bot J Linn Soc 176:295–310. https://doi.org/10.1111/boj.12210
Hedenäs L (2015a) Tortella rigens (Bryophyta, Pottiaceae): relationships, regional variation, and conservation aspects. Pl Syst Evol 301:1361–1375. https://doi.org/10.1007/s00606-014-1159-9
Hedenäs L (2015b) Molecular and morphological incongruence among the genera around Sarmentypnum (Bryophyta: Calliergonaceae). Nova Hedwigia 100:279–292. https://doi.org/10.1127/nova_hedwigia/2014/0226
Hedenäs L (2017a) Scandinavian Oncophorus (Bryopsida, Oncophoraceae): species, cryptic species, and intraspecific variation. Eur J Taxon 315:1–34. https://doi.org/10.5852/ejt.2017.315
Hedenäs L (2017b) Three molecular markers suggest different relationships among three Drepanocladus species (Bryophyta: Amblystegiaceae). Pl Syst Evol 303:521–529. https://doi.org/10.1007/s00606-017-1389-8
Hedenäs L et al (2014) Three species for the price of one within the moss Homalothecium sericeum s.l. Taxon 63:249–257. https://doi.org/10.12705/632.16
Hill MO et al (2006) An annotated checklist of the mosses of Europe and Macaronesia. J Bryol 28:198–267
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Molec Biol Evol 23:254–267. https://doi.org/10.1093/molbev/msj030
Ignatov MS, Afonina OM, Ignatova EA (2006) Check-list of mosses of East Europe and North Asia. Arctoa 15:1–130
Inoue Y, Tsubota H, Sato H, Yamaguchi T (2012) Phylogenetic note on Pachyneuropsis miyagii T. Yamag. (Pottiaceae, Bryophyta). Hikobia 16:221–228
Jordan WC, Courtney MW, Neigel JE (1996) Low levels of infraspecific genetic variation at a rapidly evolving chloroplast DNA locus in North American Duckweeds (Lemnaceae). Amer J Bot 83:430–439
Köckinger H, Hedenäs L (2017) A farewell to Tortella bambergeri (Pottiaceae) as understood over the last decades. J Bryol 39:213–225. https://doi.org/10.1080/03736687.2017.1307313
Košnar J, Herbstová M, Kolář F, Koutecký P, Kučera J (2012) A case of intragenomic ITS variation in bryophytes: assessment of gene flow and role of plyploidy in the origin of European taxa of the Tortula muralis (Musci: Pottiaceae) complex. Taxon 61:709–720
Kramer W (1980) Tortula Hedw. sect. Rurales De Not. (Pottiaceae, Musci) in der östlichen Holarktis. Bryophyt Biblioth 21:1–165, Taf. 161–129
Kramer W (1988) Beiträge zur Systematik und Bryogeographie einiger Sippen von Tortula Hedw. sect. Rurales De Not. (Pottiaceae, Musci) unter besonderer Berücksichtigung der Südhemisphäre. J Hattori Bot Lab 65:81–144
Loeske L (1907) Ueber Parallelformen und Veränderlichkeit der Zellenlänge bei Laubmoosen. Allg Bot Z Syst 7:119–122
McDaniel SF, Willis HJ, Shaw AJ (2007) A linkage map reveals a complex basis for segregation distortion in an interpopulation cross in the moss Ceratodon purpureus. Genetics 176:2489–2500. https://doi.org/10.1534/genetics.107.075424
Mishler BD (1985) Biosystematic studies of the Tortula ruralis complex. I: variation of taxonomic characters in culture. J Hattori Bot Lab 58:225–253
Mishler BD (2007) 33. Syntrichia Bridel, J. Bot. (Schrader) 1801(1): 299. 1801. In: Committee FoNAE (ed) Flora of North America North of Mexico, vol. 27. Bryophyta, part 1. Oxford University Press, New York, pp 618–627
Morrison DA (2014) Phylogenetic networks: a review of methods to display evolutionary history. Annual Res Rev Biol 4:1518–1543
Müller K (2005) SeqState. Appl Bioinf 4:65–69. https://doi.org/10.2165/00822942-200504010-00008
Natcheva R, Cronberg N (2004) What do we know about hybridization among bryophytes in nature? Canad J Bot 82:1687–1704. https://doi.org/10.1139/b04-139
Natcheva R, Cronberg N (2007a) Maternal transmission of cytoplasmic DNA in interspecific hybrids of peat mosses, Sphagnum (Bryophyta). J Evol Biol 20:1613–1616. https://doi.org/10.1111/j.1420-9101.2007.01341.x
Natcheva R, Cronberg N (2007b) Recombination and introgression of nuclear and chloroplast genomes between the peat mosses, Sphagnum capillifolium and Sphagnum quinquefarium. Molec Ecol 16:811–818. https://doi.org/10.1111/j.1365-294X.2006.03163.x
Nyholm E (1991) Illustrated flora of Nordic mosses. Fasc. 2 Pottiaceae-Splachnaceae-Schistostegaceae. Nordic Bryological Society, Copenhagen and Lund
Ochyra R, Zarnowiec J, Bednarek-Ochyra H (2003) Census catalogue of Polish mosses. Biodivers Poland 3:1–372
Rydin C, Pedersen KR, Friis EM (2004) On the evolutionary history of Ephedra: cretaceous fossils and extant molecules. Proc Natl Acad Sci USA 101:16571–16576. https://doi.org/10.1073/pnas.0407588101
Schofield WB, Crum HA (1972) Disjunctions in bryophytes. Ann Missouri Bot Gard 59:174–202. https://doi.org/10.2307/2394752
Shaw AJ, Goffinet B (2000) Molecular evidence of reticulate evolution in the peatmosses (Sphagnum), including S. ehyalinum sp. nov. Bryologist 103:357–374
Shaw AJ, McDaniel SF, Werner O, Ros RM (2002) New frontiers in bryology and lichenology: phylogeography and phylodemography. Bryologist 105:373–383. https://doi.org/10.1639/0007-2745(2002)105%5b0373:PAP%5d2.0.CO;2
Shaw AJ, Werner O, Ros RM (2003) Intercontinental Mediterranean disjunct mosses: morphological and molecular patterns. Amer J Bot 90:540–550
Simmons MP, Ochoterena H (2000) Gaps as characters in sequence-based phylogenetic analyses. Syst Biol 49:369–381. https://doi.org/10.1093/sysbio/49.2.369
Smith AJE (2004) The moss flora of Britain and Ireland, 2nd edn. Cambridge University Press, Cambridge
Spagnuolo V, Caputo P, Cozzolino S, Castaldo R, De Luca P (1999) Patterns of relationships in Trichostomoideae (Pottiaceae, Musci). Pl Syst Evol 216:69–79
StatSoft I (2013) STATISTICA (data analysis software system), version 12. Available at: https://www.statsoft.com
Stech M (1999) Molekulare Systematik haplolepider Laubmoose (Dicranaceae, Bryopsida). PhD Thesis, Freie Universität Berlin, Berlin
Stegemann S, Keuthe M, Greiner S, Bock R (2012) Horizontal transfer of chloroplast genomes between plant species. Proc Natl Acad Sci USA 109:2434–2438. https://doi.org/10.1073/pnas.1114076109
Streimann H, Klazenga N (2002) Catalogue of Australian mosses. Fl Austral Suppl Ser 17:1–259
Vanderpoorten A (2001) The Syntrichia ruralis complex in Belgium. Cryptogam Bryol 22:71–84
Vanderpoorten A, Shaw AJ, Cox CJ (2004) Evolution of multiple paralogous adenosine kinase genes in the moss genus Hygroamblystegium: phylogenetic implications. Molec Phylogen Evol 31:505–516. https://doi.org/10.1016/j.ympev.2003.09.020
Wendel JF, Doyle JJ (1998) Phylogenetic incongruence: window into genome history and molecular evolution. In: Soltis DE, Soltis PS, Doyle JJ (eds) Molecular systematics of plants II. DNA sequencing. Chapman and Hall, New York, pp 265–296
Werner O, Ros RM, Cano MJ, Guerra J (2002) Tortula and some related genera (Pottiaceae, Musci): phylogenetic relationships based on chloroplast rps4 sequences. Pl Syst Evol 235:197–207. https://doi.org/10.1007/s00606-002-0230-0
Werner O, Ros RM, Guerra J, Shaw AJ (2003) Molecular data confirm the presence of Anacolia menziesii (Bartramiaceae, Musci) in southern Europe and its separation from Anacolia webbii. Syst Bot 28:483–489
Werner O, Ros RM, Cano MJ, Guerra J (2004) Molecular phylogeny of Pottiaceae (Musci) based on chloroplast rps4 sequence data. Pl Syst Evol 243:147–164
Werner O, Ros RM, Grundmann M (2005) Molecular phylogeny of Trichostomoideae (Pottiaceae, Bryophyta) based on nrITS sequence data. Taxon 54:361–368
Wyatt R, Odrzykoski IJ, Stoneburner A, Bass HW, Galau GA (1988) Allopolyploidy in bryophytes: multiple origins of Plagiomnium medium. Proc Natl Acad Sci USA 85:5601–5604
Zander RH (1989) Seven new genera in Pottiaceae (Musci) and a lectotype for Syntrichia. Phytologia 65:424–436
Zander RH (1993) Genera of the Pottiaceae: mosses of harsh environments. Bull Buffalo Soc Nat Sci 32:1–378
Acknowledgements
Open access funding provided by Swedish Museum of Natural History. We thank the curators of GOET and MUB for loans of material, and Rasa Bukontaite and Bodil Cronholm for their efficient laboratory work. We appreciate comments from an anonymous reviewer. This research was funded by the Swedish Taxonomy Initiative (Svenska artprojektet, dha 2016-20 4.3) and the Spanish Flora Briofítica Ibérica project CGL2015/64068-P (MINECO/FEDER).
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: Terry A.J. Hedderson.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Lars Hedenäs and Jochen Heinrichs initiated this study. After the unexpected decease of Jochen in 2018 [PSE 304(8):937–941] it was continued and finalised by Lars and María Teresa Gallego in memory of Jochen, who was one of the journal’s associate editors.
Jochen Heinrichs: Deceased
Information on Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Appendix
Appendix
Specimen data and GenBank accession numbers for the studied Syntrichia and outgroup sequences. Data format: Sample No. [* = specimens which leaves and cells were measured]: Locality; Coll. Year, Collector [LH = L. Hedenäs] [collector’s No.]; Herbarium acronym: registration No.; GenBank accession numbers for ITS1 [paralogue A/paralogue B/etc. (paralogue labels/number of clones)], atpB-rbcL, rpl16 [NA = Not available].
Syntrichia calcicolaJ.J.Amann. P306*: Sweden. Gotland, Langs hage; 2002, T.Hallingbäck 38745; S: B184804; MH919981, MK007615, MK007794. P354: Turkey. Prov. Icel, NNW of Mut; 1978, B.Bremer & E.Nyholm 279/78; S: B61657; MH920034, MK007662, MK007839. P362*: Sweden. Gotland, Othem, N of Othemars; 2015, LH; S: B234292; MH920050, MK007670, MK007847. P363*: Sweden. Öland, Böda, Björnsnabben; 1960, A.C.Crundwell & E.Nyholm; S: B219042; MH920051, MK007671, MK007848. P364: Germany. Nordrhein-Westf., Niederrh., Bucht; 1992, J.Heinrichs & P.Kolshorn 347; S: B136160; MH920052, MK007672, MK007849. P370*: Sweden. Södermanland, Utö, N of Kroka; 2014, LH; S: B211229; MH920058, MK007678, MK007855. P425*: Sweden. Södermanland, Utö, N portion of Utö; 2015, LH; S: B222162; MH920138, MK007733, MK007907. P433*: Sweden. Öland, Resmo, NE of Resmo; 2015, LH & I.Bisang; S: B222108; MH920162, MK007741, MK007915. P441*: Sweden. Gotland, Bunge, Bungenäs; 2016, LH; S: B235960; MH920170, MK007749, MK007921. P442*: Sweden. Öland, Alböke, SE of Lindkärr; 2016, LH; S: B236091; MH920171/MH920172 (A,H/2), MK007750, MK007922. P443*: Sweden. Gotland, Östergarn, Kaupungsklint; 2016, LH; S: B235926; MH920173/MH920174/MH920175/MH920176/MH920177/MH920178/MH920179 (A,B,D,E,F,G,H/7), MK007751, MK007923. P444*: Sweden. Öland, Persnäs, Lilla Horn; 2016, LH; S: B236145; MH920180, MK007752, MK007924. P445: Sweden. Öland, Ventlinge, E of Ventlinge; 2016, LH; S: B236114; MH920181, MK007753, MK007925. P446: Sweden. Gotland, Etelhem, E of Bjärby; 2016, LH; S: B235925; MH920182, MK007754, MK007926. P447: Sweden. Öland, Ventlinge, W of Sebberneby; 2016, LH; S: B236125; MH920183/MH920184/MH920185 (B,C,H/3), MK007755, MK007927. P448: Sweden. Gotland, Östergarn, S of Falhammars; 2016, LH; S: B235935; MH920186/MH920187/MH920188/MH920189/MH920190 (B,C,E,F,G/5), MK007756, MK007928. P465: Sweden. Gotland, Etelhem, NNW of Villbärsmyr; 2016, LH; S: B235909; MH920211, MK007773, MK007945. P466: Sweden. Gotland, Bunge, S end of Bungenäs; 2016, LH; S: B235962; NA, MK007774, MK007946. P467: Sweden. Gotland, Gammelgarn, Klintklinten; 2016, LH; S: B235950; MH920212, MK007775, MK007947. P469: Sweden. Öland, Persnäs, N of Sandvik; 2016, LH; S: B236163; MH920217, MK007777, MK007949. P472: Sweden. Öland, Räpplinge, W of Greby; 2016, LH; S: B236187; MH920225, MK007780, MK007952. P473: Sweden. Öland, Räpplinge, W of Greby; 2016, LH; S: B236177; NA, MK007781, MK007953. P480: Sweden. Gotland, Fröjel, Ansarve; 1993, J.Heinrichs; GOET: Herb. Heinrichs 1224; NA, MK007788, MK007959. P481: Germany. Nordrhein-Westf., Niederrh., Vierssen; 1995, Abts 6370c; GOET: Herb. Heinrichs; MH920233, MK007789, MK007960. P482: Belgium. West-Flandern, Nordseeküste, De Panne; 1995, Abts 6254a; GOET: Herb. Heinrichs; MH920234, MK007790, MK007961. P483: Germany. Rheinland-Pfalz, Bad Neuenahr-Ahrweiler; 1999, J.Heinrichs; GOET: Herb. Heinrichs 4251; MH920235, MK007791, MK007962. P484: Germany. Nordrhein-Westf., Niederrh. Tiefland; 1995, J.Heinrichs; GOET: Herb. Heinrichs 2689; MH920236, MK007792, MK007963. Syntrichia caninervisMitt. P341: Turkey. Prov. Van, N of the town Van; 1974, E.Nyholm 69/74; S: B61898; MH920022, MK007649, MK007826. P344: United States. Colorado, Montezuma Co., East Rock Creek trail; 2008, LH; S: B152429; MH920024/MH920025/MH920026 (A,D,G/3), MK007652, MK007829. P345: United States. California, S Sierra Nevada, Sequoia NF; 2000, J.Shevock et al. 20262; S: B205969; NA, MK007653, MK007830. Syntrichia handelii(Schiffn.) S.Agnew & Vondr. P342: Turkey. Prov. Konya, Karaman; 1978, B.Bremer & E.Nyholm 322a/78; S: B62268; NA, MK007650, MK007827. P343: Turkey. Prov. Içel, NNW of Mut; 1978, B.Bremer & E.Nyholm 292/78; S: B62418; MH920023, MK007651, MK007828. Syntrichia laevipilaBrid. P328: Norway. Hordaland, Kvinnherad, Eid; 1991, N.Hakelier; S: B233936; MH920012, MK007637, MK007816. P329: Norway. Hordaland, Förde, Valestrand; 1991, N.Hakelier; S: B233937; MH920013, MK007638, MK007817. Syntrichia latifolia(Hartm.) Huebener. P326: Sweden. Uppland, Börje, Broby bro; 1986, N.Hakelier; S: B233935; MH920010, MK007635, MK007814. P327: United Kingdom. England, Shropshire, Preston Montford; 2008, LH; S: B144774; MH920011, MK007636, MK007815. Syntrichia montanaNees. P338: Sweden. Öland, Räpplinge, Borgholms slottsruin; 2005, T.Hallingbäck 43447; S: B184817; MH920020, MK007646, MK007823. P339: Sweden. Gotland, Sundre, N of Hoburgsgubben; 2014, LH et al.; S: B205523; NA, MK007647, MK007824. P462: Sweden. Öland, Ventlinge, E of Ventlinge; 2016, LH; S: B236120; MH920208, MK007770, MK007942. Syntrichia norvegicaF.Weber. P307*: Sweden. Öland, St Alvaret på väg mot Alby; 1999, T.Hallingbäck 45914; S: B184805; MH919982, MK007616, MK007795. P308*: Norway. Finnmark, Hammerfest, Sørøya (Sállan); 2010, LH; S: B176568; MH919983, MK007617, MK007796. P309*: Sweden. Pite Lappmark, Arjeplog, N of Mávasjávrre; 2015, LH et al.; S: B223718; MH919984/MH919985/MH919986 (A,B,C/6), MK007618, MK007797. P310: Sweden. Härjedalen, Ljusnedal, River Mittån;2007, LH; S: B122831; MH919987/MH919988/MH919989/MH919990 (A,B,C,D/6), MK007619, MK007798. P311: Sweden. Öland, Kastlösa, St. Dalby; 2014, LH; S: B209861; MH919991, MK007620, MK007799. P312*: Sweden. Gotland, Stenkyrka, N of Ekebys; 2013, LH; S: B196939; MH919992, MK007621, MK007800. P313*: Sweden. Gotland, Linde, Lindeberget; 2014, LH et al.; S: B204352; MH919993, MK007622, MK007801. P323*: Sweden. Åsele Lappmark, Dorotea, Mt Kalvberget; 2004, LH; S: B100569; NA, MK007632, MK007811. P365*: Sweden. Öland, Torslunda, NE of Lenstad; 2015, LH; S: B209915; MH920053, MK007673, MK007850. P366: Sweden. Gotland, Stenkyrka, E of Gräne; 2001, LH; S: B62346; MH920054, MK007674, MK007851. P367*: Sweden. Gotland, Endre, Ölbäck; 1989, LH G89-235; S: B65935; MH920055, MK007675, MK007852. P368: Sweden. Gotland, Othem, Filehajdar; 1995, LH; S: B65926; MH920056, MK007676, MK007853. P369: Sweden. Gotland, Hejdeby, Torsalvret; 1989, LH; S: B65931; MH920057, MK007677, MK007854. P372: Sweden. Härjedalen, Ljusnedal, River Mittån; 2007, LH; S: B122847; MH920060, MK007680, MK007857. P373: Sweden. Jämtland, Frostviken, Mt Brakkfjället; 2009, LH; S: B164563; MH920061, MK007681, MK007858. P374*: Sweden. Jämtland, Frostviken, NW of Mt Lappkojberget; 2009, LH; S: B164638; MH920062, MK007682, MK007859. P375*: Sweden. Jämtland, Undersåker, Välliste; 1993, N.Hakelier; S: B218742; MH920063, MK007683, MK007860. P376*: Sweden. Åsele Lappmark, Vilhelmina, Stuore Gämo; 2004, LH; S: B99799; MH920064, MK007684, MK007861. P377*: Sweden. Pite Lappmark, Arjeplog, N of Mávasjávrre; 2015, LH et al.; S: B226184; MH920065, MK007685, MK007862. P378*: Sweden. Pite Lappmark, Arjeplog, N of Mávasjávrre; 2015, LH et al.; S: B227641; MH920066, MK007686, MK007863. P379: Sweden. Torne Lappmark, Jukkasjärvi, Kärkevagge; 1990, LH; S: B218807; MH920067, MK007687, MK007864. P380*: Sweden. Torne Lappmark, Vittangi, Masugnsbyn; 2008, T.Hallingbäck 46184; S: B184821; MH920068, MK007688, MK007865. P381*: Norway. Finnmark, Söröysund, Seiland (Siev’jo), Hönsebybotn; 2001, LH; S: B59853; MH920069, MK007689, MK007866. P382*: Norway. Troms, Lyngen, Vardu, River Storelva; 2003, LH; S: B82988; NA, MK007690, MK007867. P383*: Norway. Oppland, Lom, Bövertunholet; 1988, N.Hakelier; S: B218830; NA, MK007691, MK007868. P384*: Switzerland. Kt. Luzern, Flühli, Schrattenflue; 1999, LH; S: B11904; MH920070/MH920071 (A,D/6), MK007692, MK007869. P385*: Switzerland. Graubünden, Poschiavo, Ospizio Bernina; 2011, LH; S: B184378; MH920072, MK007693, MK007870. P386*: Italy. Lombardia, Sondrio, Pass da Val Viola; 2011, LH; S: B184383; MH920073/MH920074/MH920075/MH920076 (A,B,C,F/6), MK007694, MK007871. P449: Sweden. Gotland, Fleringe, W of Hau; 2016, LH; S: B235980; MH920191, MK007757, MK007929. P450: Sweden. Öland, Ventlinge, NNW of Lunda; 2016, LH; S: B236105; MH920192/MH920193/MH920194/MH920195 (A,B,C,D/6), MK007758, MK007930. P451: Sweden. Gotland, Bro, Torsalvret; 1989, LH G89-221; S: B65932; MH920196, MK007759, MK007931. P478: Austria. Tirol, Silvretta, Fimbertal; 1993, J.Heinrichs; GOET: Herb. Heinrichs 1513; NA, MK007786, NA. Syntrichia papillosa(Spruce) Spruce. P336: Sweden. Bohuslän, Tossene; 1992, LH; S: B11336; MH920018, MK007644, NA. P337: Poland. W Carpathians, Beskid Wysp. Mts; 2004, A.Stebel (M Macror Merid Pol Exs 1393); S: B99105; MH920019, MK007645, NA. Syntrichia papillosissima(Copp.) Loeske. P334: Turkey. Prov. Içel, NNW of the town Mut; 1978, B.Bremer & E.Nyholm 300/78; S: B62273; MH920016, MK007642, MK007821. P335: Turkey. Eskisehir, Sarickaya; 1986, E.Yücel 37/0; S: B233941; MH920017, MK007643, MK007822. Syntrichia princeps(De Not.) Mitt. P346: Sweden. Gotland, Boge, Tjälderholm; 1992, LH; S: B222803; MH920027, MK007654, MK007831. P347: Sweden. Gotland, Visby, Muramaris; 1992, LH; S: B222802; MH920028, MK007655, MK007832. P348: Sweden. Gotland, Rute, Furilden; 1992, LH; S: B222800; MH920029, MK007656, MK007833. P349: Sweden. Gotland, Boge, NNE of Gyle; 2011, LH; S: B183357; MH920030, MK007657, MK007834. P350: Sweden. Gotland, Etelhem, Lake Sigvaldeträsk; 2014, LH et al.; S: B205004; MH920031, MK007658, MK007835. P351: United Kingdom. Skottland, Sutherland, Assynt; 1973, N.Hakelier; S: B233942; NA, MK007659, MK007836. P352: Cyprus. Trodos Mts, Trodos; 1995, LH; S: B87602; MH920032, MK007660, MK007837. P353: Turkey. Prov. Icel, Cakmak SE Camliyayla; 1978, B.Bremer & E.Nyholm 217/78; S: B61598; MH920033, MK007661, MK007838. P355: Turkey. Prov. Bolu, Abant Gölü, SW Bolu; 1974, TB.Engelmark & E.Nyholm 940/74; S: B61023; MH920035, MK007663, MK007840. P356: United States. California, Peninsular R., Cleveland NF; 2013, J.Shevock et al. 41925; S: B205980; MH920036, MK007664, MK007841. P440: Sweden. Öland, Räpplinge, W of Greby; 2016, LH; S: B236184; MH920169, MK007748, MK007920. P479: Germany. Rheinland-Pf., Koblenz; 1995, S.Caspari & J.Heinrichs; GOET: Herb. Heinrichs 2827; MH920232, MK007787, MK007958. Syntrichia pseudohandelii(J.Froehl.) S.Agnew & Vondr. P340: Turkey. Prov. Van, N of Van; 1974, E.Nyholm 61/74; S: B61915; MH920021, MK007648, MK007825. Syntrichia rigescens(Broth. & Geh.) Ochyra. P438: Morocco. Jbel Touchka, Imaouzzer-Ida-Outanen; 2001, M.J.Cano & J.Munoz; MUB: Bryo 11378; MH920167, MK007746, MK007919. P439: Jordania. Karak, Tafila, Dana; 1987, A.El-Oqlah 771; MUB: Bryo 22255; MH920168, MK007747, NA. Syntrichia ruraliformis(Besch.) Mans. P318*: Sweden. Öland, Kastlösa, betw. St. Dalby and Bårby; 2014, LH; S: B209856; MH920003, MK007627, MK007806. P319*: Sweden. Gotland, Othem, N of Othemars; 2015, LH; S: B220597; MH920004, MK007628, MK007807. P320*: Sweden. Gotland, Eksta, N of Kronvald; 1992, LH; S: B51998; MH920005, MK007629, MK007808. P321*: Sweden. Södermanland, Utö, Sandviken; 2015, LH; S: B211883; MH920006, MK007630, MK007809. P322*: Sweden. Gotland, Fårö, S of Ullahau; 2003, LH; S: B84459; MH920007, MK007631, MK007810. P357: United States. California, C Sierra Nevada, Sierra NF; 2000, J.Shevock & K.Kellman 19757; S: B213957; MH920037, MK007665, MK007842. P387: Sweden. Skåne. Degeberga, Forsakar-Bökestorp; 1944, S.Waldheim; S: B193919; NA, MK007695, NA. P388*: Sweden. Öland, Kastlösa, St. Dalby; 2014, LH; S: B209863; MH920077, MK007696, MK007872. P389*: Sweden. Öland, Karums alvar, SW of Karum; 2006, T.Hallingbäck 43909; S: B184829; MH920078, MK007697, MK007873. P390*: Sweden. Gotland, Gothem, Gothemsån; 1987, LH; S: B219355; MH920079, MK007698, MK007874. P391*: Norway. Nordland, Flakstad, Krystad; 2015, LH; S: B222133; MH920080/MH920081/MH920082/MH920083/MH920084 (A,B,D,E,F/6), MK007699, MK007875. P392*: Denmark. E Jutland, Djursland, Glatved strand; 2009, M.H.G.Gustafsson 1125; S: B172547; MH920085, MK007700, MK007876. P393: Sweden. Uppland, Djurö, Runmarö, Bredvarpen; 2012, LH; S: B194453; MH920086/MH920087/MH920088/MH920089/MH920090/MH920091/MH920092/MH920093 (A,B,C,D,E,G,H,I/10), MK007701, MK007877. P394*: Denmark. Jutland, Ålbaek, Napstjert; 2012, LH; S: B193445; MH920094, MK007702, MK007878. P395*: Norway. Nordland, Vestvågøy, Haukland; 2015, LH; S: B221729; MH920095, MK007703, MK007879. P396*: Norway. Nordland, Vestvågøy, Unstad; 2015, LH; S: B222128; MH920096, MK007704, MK007880. P397*: Netherlands. Zuid-Holland, De Zilk; 1992, LH; S: B4619; MH920097, MK007705, MK007881. P398: Ireland, Mayo, Louisburgh, Old Head; 1992, N.Hakelier; S: B234468; MH920098, MK007706, MK007882. P399*: Estonia. Kingissepa, Kõinastu; 1989, LH E-40; S: B234469; MH920099, MK007707, MK007883. P400*: Hungary. W of Kecskémet, Fülöpháza; 2005, LH; S: B104372; MH920100, MK007708, MK007884. P401: Germany. Nordrhein-Westf., Niederrh., Mönchengladbach-Damm; 1994, Abts 5072; S: B136175; NA, MK007709, MK007885. P420*: Sweden. Uppland, Djurö, Runmarö, Fladen; 2011, LH; S: B187053; MH920131, MK007728, MK007902. P430: Sweden. Uppland, Djurö, Runmarö, SW of Byholmen; 2015, LH; S: B225281; MH920150/MH920151/MH920152/MH920153/MH920154 (B,C,E,F,G/6), MK007738, MK007912. P434: Sweden. Öland, Resmo, NE of Resmo; 2015, LH & I.Bisang; S: B222114; MH920163/MH920164 (A,B/6), MK007742, MK007916. P452*: Sweden. Öland, Högby, Sandby; 2016, LH; S: B236133; MH920197, MK007760, MK007932. P453: Sweden. Gotland, Östergarn, S of Falhammars; 2016, LH; S: B235932; MH920198, MK007761, MK007933. P455*: Sweden. Öland, Alböke, W of Lindkärr; 2016, LH; S: B236090; MH920200, MK007763, MK007935. P456: Sweden. Gotland, Bunge, Bungenäs; 2016, LH; S: B235958; NA, MK007764, MK007936. P457: Sweden. Gotland, Bunge, Bungenäs; 2016, LH; S: B235961; MH920201, MK007765, MK007937. P458*: Sweden. Öland, Persnäs, Lilla Horn; 2016, LH; S: B236147; MH920202, MK007766, MK007938. P459*: Sweden. Gotland, Gammelgarn, Klintklinten; 2016, LH; S: B235953; MH920203, MK007767, MK007939. P461*: Sweden. Öland, Räpplinge, W of Greby; 2016, LH; S: B236165; MH920207, MK007769, MK007941. P464*: Sweden. Öland, Persnäs, Lilla Horn; 2016, LH; S: B236144; MH920210, MK007772, MK007944. P470*: Sweden. Öland, Persnäs, N of Sandvik; 2016, LH; S: B236152; MH920218/MH920219/MH920220/MH920221/MH920222/MH920223 (A,B,C,D,E,F/6), MK007778, MK007950. P475: Sweden. Gotland, N Ardre, Torsburgen; 1993, J.Heinrichs; GOET: Herb. Heinrichs 1310; MH920229, MK007783, MK007955. P476*: Sweden. Gotland, Hemmor; 1993, J.Heinrichs; GOET: Herb. Heinrichs 1239; MH920230, MK007784, MK007956. P477: Germany. Nordrhein-Westf., Niederrh., Xanten; 1992, Abts & Heinrichs 563/92; GOET: Herb. Heinrichs; MH920231, MK007785, MK007957. Syntrichia ruralis(Hedw.) F.Weber & D.Mohr. P314*: Sweden. Gotland, Hejnum; 2015, LH; S: B220794; MH919994/MH919995/MH919996/MH919997/MH919998/MH919999 (B,C,D,E,F,H/6), MK007623, MK007802. P315*: Sweden. Västmanland, Linde, Östra Öskevik; 2015, LH; S: B226631; MH920000, MK007624, MK007803. P316*: Sweden. Jämtland, Kall, Mt Stavattsberget; 2005, LH; S: B104724; MH920001, MK007625, MK007804. P317*: Sweden. Pite Lappmark, Arjeplog, N of Mávasjávrre; 2015, LH et al.; S: B223755; MH920002, MK007626, MK007805. P324*: Sweden. Öland, Torslunda, E of Eriksöre; 2015, LH & I.Bisang; S: B222100; MH920008, MK007633, MK007812. P371*: Sweden. Södermanland, Grödinge, Norrga; 1998, G.Odelvik & R.Karlsson; S: B36489; MH920059, MK007679, MK007856. P402*: Norway. Finnmark, Söröysund, Seiland (Siev’jo), Hönsebybotn; 2001, LH; S: B59852; MH920101, MK007710, MK007886. P403*: Norway. Nordland, Vestvågøy, Haukland; 2015, LH; S: B221740; MH920102/MH920103/MH920104/MH920105 (A,D,E,F/6), MK007711, MK007887. P404: Sweden. Pite Lappmark, Arjeplog, N of Mávasjávrre; 2015, LH et al.; S: B223774; NA, MK007712, MK007888. P405*: Sweden. Pite Lappmark, Arjeplog, N of Mávasjávrre; 2015, LH et al.; S: B224601; MH920106, MK007713, MK007889. P406*: Sweden. Åsele Lappmark, Dorotea, Storbäck; 2004, LH; S: B93241; NA, MK007714, MK007890. P407*: Sweden. Åsele Lappmark, Dorotea, Mt Kalvberget; 2004, LH; S: B95783; MH920107/MH920108/MH920109/MH920110/MH920111/MH920112 (A,B,C,D,E,F/6), MK007715, MK007891. P408*: Sweden. Norrbotten, Pajala, Jupukka; 1990, LH & M.Aronsson NT90-57; S: B42363; MH920113, MK007716, MK007892. P409: Sweden. Ångermanland, Högsjö, Rö; 2013, LH et al.; S: B200791; NA, MK007717, MK007893. P410*: Sweden. Medelpad, Borgsjö, Mt Öberget; 2006, LH; S: B115548; MH920114/MH920115/MH920116 (A,B,C/6), MK007718, MK007894. P411*: Sweden. Medelpad, Torp, Ljungaverk, Västerhångsta; 2006, LH; S: B116586; MH920117, MK007719, MK007895. P412: Sweden. Medelpad, Liden, Dacke, Sillre såg; 2013, LH; S: B198584; NA, MK007720, NA. P413: Sweden. Jämtland, Frostviken, Hovdbäcken; 1988, LH J88-60; S: B42359; MH920118, MK007721, NA. P414*: Sweden. Jämtland, Frostviken, N of Raurenjaure; 1988, LH J88-596; S: B42360; MH920119, MK007722, MK007896. P415*: Sweden. Jämtland, Ragunda, Mt Prästberget; 2014, LH; S: B204297; MH920120/MH920121 (A,C/6), MK007723, MK007897. P416: Sweden. Härjedalen, Sveg, Mt Bådhushammaren; 1989, LH HD89-167; S: B221580; MH920122/MH920123/MH920124/MH920125 (A,B,D,G/7), MK007724, MK007898. P417: Sweden. Härjedalen, Tännäs, Tännäs village; 2005, LH; S: B104211; NA, MK007725, MK007899. P418*: Sweden. Härjedalen, Linsell, Glöte, Mt Dyckesberget; 2007, LH et al.; S: B123120; MH920126, MK007726, MK007900. P419*: Sweden. Härjedalen, Tännäs, Mt Joltere (Lill-Mittåkläppen); 2014, LH; S: B207533; MH920127/MH920128/MH920129/MH920130 (A,B,C,D/6), MK007727, MK007901. P421*: Sweden. Södermanland, Botkyrka, Ekholmen NR; 2011, LH; S: B188311; MH920132, MK007729, MK007903. P422*: Sweden. Östergötland, Krokek, ESE of Oxåker; 2015, LH; S: B225249; MH920133, MK007730, MK007904. P423*: Sweden. Dalarna, Husby, E of Kloster; 2011, LH et al.; S: B186569; MH920134/MH920135/MH920136/MH920137 (A,B,D,E/6), MK007731, MK007905. P424: Sweden. Uppland, Väddö, Nothamn; 2015, LH; S: B227327; NA, MK007732, MK007906. P426*: Sweden. Småland, Virserum, SW of Rydet; 2014, T.Nilsson 2134; S: B221128; MH920139, MK007734, MK007908. P427*: Sweden. Västmanland, Nora, Bergsmanshyttan, Kalkudden; 2015, LH; S: B226646; MH920140/MH920141/MH920142/MH920143/MH920144 (A,B,C,E,F/6), MK007735, MK007909. P428*: Sweden, Värmland, Filipstad, Saxån; 2010, LH & G.Odelvik; S: B178198; MH920145/MH920146/MH920147/MH920148 (A,B,C,F/6), MK007736, MK007910. P429*: Sweden. Uppland, Djurö, Runmarö, SW of Byholmen; 2015, LH; S: B225281; MH920149, MK007737, MK007911. P431*: Sweden. Gotland, Hejnum, Hejnum hällar; 2015, LH; S: B220812; MH920155, MK007739, MK007913. P432*: Sweden. Gotland, Hejnum, Hejnum hällar; 2015, LH; S: B220812; MH920156/MH920157/MH920158/MH920159/MH920160/MH920161 (A,B,C,D,E,F/6), MK007740, MK007914. P460*: Sweden. Gotland, Etelhem, SE of Branden; 2016, LH; S: B235917; MH920204/MH920205/MH920206 (B,C,E/6), MK007768, MK007940. P468*: Sweden. Gotland, Gammelgarn, Klintklinten; 2016, LH; S: B235951; MH920213/MH920214/MH920215/MH920216 (A,C,D,E/6), MK007776, MK007948. P471*: Sweden. Öland, Räpplinge, W of Greby; 2016, LH; S: B236187; MH920224, MK007779, MK007951. P474: Sweden. Gotland, Etelhem, SE of Branden; 2016, LH; S: B235913; MH920226/MH920227/MH920228 (A,C,D/6), MK007782, MK007954. Syntrichia ruralisvar.epilosa(Venturi) J.J.Amann. P325*: Sweden. Södermanland, Ornö, Sundby NR; 2015, LH; S: B226607; MH920009, MK007634, MK007813. P435*: Sweden. Uppland, Djurö, Runmarö, SW of Noreträsk; 2016, LH & I.Bisang; S: B234849; MH920165, MK007743, MK007917. Syntrichia sinensis(Müll.Hal.) Ochyra. P333: Russia. Altai Mts., Teletskoye Lake, Yailyu; 1989, M.Ignatov 87; S: B233940; MH920015, MK007641, MK007820. Syntrichia subpapillosissima(W.A.Kramer) M.T.Gallego & J.Guerra. P436: Spain. Murcia, Moratalla, Sierra de Villafuerte; 2009, J.Guerra et al.; MUB: Bryo 34105; MH920166, MK007744, MK007918. P437: Spain. Guadalajara, Checa, Rio de la Hoz Seca; 2004, M.T.Gallego; MUB: Bryo 18001; NA, MK007745, NA. P454: Sweden. Öland, Alböke, W of Alböke; 2016, LH; S: B236078; MH920199, MK007762, MK007934. P463: Sweden. Öland, Alböke, W of Alböke; 2016, LH; S: B236077; MH920209, MK007771, MK007943. Syntrichia virescens(De Not.) Ochyra. P330: Sweden. Gotland, Roma kloster and kungsgård; 2002, T.Hallingbäck 38749; S: B184841; MH920014, MK007639, MK007818. P331: Sweden. Närke, Örebro, Nikolai; 1993, N.Hakelier; S: B233938; NA, MK007640, MK007819. OUTGROUP:Cinclidotus fontinaloides(Hedw.) P.Beauv. P358: United Kingdom. Scotland. Strathclyde, Dumbarton; 1996, LH; S: B233956; MH920038/MH920039/MH920040/MH920041/MH920042 (A,B,C,E,F/6), MK007666, MK007843. Leptophascum leptophyllum(Müll.Hal.) J.Guerra & M.J.Cano. P359: Réunion. E coast, St. Benoit; 1998, Schäfer-Verwimp & Verwimp 20171; S: B97392; MH920043/MH920044/MH920045 (B,F,G/6), MK007667, MK007844. Microbryum davallianum(Sm.) R.H.Zander. P360: Sweden. Gotland, Eksta, Bopparve, Ajvide; 1992, LH; S: B11338; MH920046/MH920047/MH920048 (B,C,D/6), MK007668, MK007845. Tortella tortuosa(Hedw.) Limpr. P150: United Kingdom. N Ireland, Fermanagh co., Crossmurrin; 1993, T.Hallingbäck 42536; S: B240756; KM020632, KM020522, MK007793. Tortula acaulon(With.) R.H.Zander. P361: Sweden. Södermanland, Västerhaninge, N of Ekeby; 2015, LH; S: B216407; MH920049 (D/1), MK007669, MK007846.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Hedenäs, L., Heinrichs, J. & Teresa Gallego, M. The Scandinavian Syntrichia ruralis complex (Musci, Pottiaceae): a chaos of diversification. Plant Syst Evol 305, 639–661 (2019). https://doi.org/10.1007/s00606-019-01596-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-019-01596-0