Abstract
DNA integrity is constantly challenged by endogenous and exogenous factors that can alter the DNA sequence, leading to mutagenesis, aberrant transcriptional activity, and cytotoxicity. Left unrepaired, damaged DNA can ultimately lead to the development of cancer. To overcome this threat, a series of complex mechanisms collectively known as the DNA damage response (DDR) are able to detect the various types of DNA damage that can occur and stimulate the appropriate repair process. Each DNA damage repair pathway leads to the recruitment, upregulation, or activation of specific proteins within the nucleus, which, in some cases, can represent attractive targets for molecular imaging. Given the well-established involvement of DDR during tumorigenesis and cancer therapy, the ability to monitor these repair processes non-invasively using nuclear imaging techniques may facilitate the earlier detection of cancer and may also assist in monitoring response to DNA damaging treatment. This review article aims to provide an overview of recent efforts to develop PET and SPECT radiotracers for imaging of DNA damage repair proteins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The DNA double helix within every cell of the human body is constantly exposed to damaging agents, and, consequently, tens of thousands of DNA lesions occur per cell each day [1]. If a lesion is not correctly repaired, it may cause the cell to become senescent, apoptotic, or even malignant. Over the last few decades, a multitude of endogenous and exogenous causes of DNA damage have been identified [2, 3]. Endogenous processes are responsible for the vast majority of DNA damage and can be divided into three main categories: oxidative (i.e. produced by reactive oxygen species), hydrolytic (e.g. deamination of cytosine to uracil), and alkylation reactions (e.g. methylation of the N7-position of guanine residues) [4]. As exogenous sources of DNA damage, ultraviolet light and ionizing radiation have been found to be among the most prevailing factors, causing single-strand breaks (SSBs) and double-strand breaks (DSBs), respectively [5]. In addition, certain chemotherapy drugs used for cancer treatment, industrial chemicals, and carcinogens associated with tobacco products are well recognised to cause DNA damage.
The recognition and repair of DNA damage is achieved by a set of complex yet finely tuned DNA damage response (DDR) signalling pathways that inhibit cell cycle progression and repair DNA lesions by a variety of mechanisms (Fig. 1). The excellent level of control over these processes ultimately minimises genomic instability and impedes tumorigenesis [6]. Defects in this defensive mechanism have been found to occur with significantly higher prevalence in many human cancers compared to normal tissues [7–11]. As a consequence, extensive DNA damage and DDR signalling is present and critically important in virtually all stages of tumour development, from dysplasia to advanced metastatic disease [12, 13].
A simplified overview of the DNA damage response and the main targets involved. BER, Base Excision Repair; HR, Homologous Recombination; NHEJ, Non-Homologous End Joining; LIG3, DNA Ligase 3; XRCC1, X-ray repair cross-complementing protein 1; PARP-1, poly (ADP-ribose) polymerase 1; BRCA1/2, Breast Cancer 1/2; ATM, Ataxia Telangiectasia Mutated; DNA-PK, DNA-dependent protein kinase catalytic subunit
Therefore, the ability to monitor DDR in vivo in a non-invasive manner via molecular imaging is an attractive prospect as the information obtained from these techniques could facilitate the earlier detection of several cancer types. Furthermore, as most cancer therapies are designed to cause DNA damage, these techniques may also provide a rapid and broadly applicable means of evaluating response to therapy. In this review article, we discuss potentially valuable biomarkers, which arise during the major cellular responses to SSBs and DSBs, and we also evaluate recent efforts to monitor these biomarkers non-invasively in vivo using PET and SPECT radiotracers.
Single-strand break repair mechanisms
The repair of SSBs in DNA is mainly facilitated by base excision repair (BER) [14]. Deficiencies and mutations of proteins in this pathway are linked to genomic instability, aging, and cancer [15]. BER has two sub-pathways referred to as short-patch repair (Fig. 2) and long-patch repair; the former being responsible for up to 90% of all BER [16]. In brief, short-patch BER is based on five major steps: (i) recognition of the damaged base by a DNA glycosylase and the consequent removal of the base, creating an apurinic or apyrimidinic (AP) site intermediate, (ii) incision of the abasic site by an AP endonuclease (APE) or AP lyase, (iii) removal of the remaining sugar fragment by a lyase or phosphodiesterase, (iv) filling of the remaining gap by a DNA polymerase (commonly, DNA polymerase β, POLB) with the correct nucleotide, and finally (v) sealing of the remaining nick by a DNA ligase (LIG1 or LIG3/XRCC1 complex) [17].
Among the many sensors involved in the BER repair of SSBs, poly(ADP-ribose) polymerase 1 (PARP-1) is particularly influential in this process [18–21]. Upon binding to nicked DNA, PARP-1 cleaves nicotinamide adenine dinucleotide (NADb), whereupon it catalyses the polymerisation of ADP-ribose units into long, branched chains of poly(ADP-ribose) (PAR) [22]. While the chief target for poly(ADP-ribosylation) is PARP-1 itself via automodification, other DNA damage repair proteins and histones are also PARylated. PAR is thereafter responsible for the recruitment of additional DDR proteins which cooperate for the completion of SSB repair [23–25]. Small molecule inhibitors of PARP-1 have the ability to disrupt the BER repair pathway, leading to collapsed replication forks and ultimately DSBs upon replication [26–30]. Several key studies have since shown that normal cells can compensate in these circumstances by relying on other repair mechanisms, such as homologous recombination (HR), which act to restore the original DNA sequence [31]. In accordance with these observations, studies focused on the genetic removal of PARP-1 have found no significant effect upon the frequency of tumour development [31]. Notably, however, cancer cells with defects in HR (most commonly arising from mutations of BRCA1 and BRCA2 proteins) have exhibited vastly increased sensitivity to PARP-1 inhibitors [32, 33]. In such cases, damaged DNA either persists unrepaired or is subjected to a more error-prone DNA repair mechanism (e.g. non-homologous end joining [NHEJ], or single-strand annealing [SSA]) [34]. Both scenarios will ultimately trigger cell death via apoptosis. These discoveries have stimulated intensive research efforts focused on evaluating the therapeutic potential of PARP-1 inhibitors, principally for breast and ovarian cancers in BRCA-mutation carriers [35].
Expression levels of the PARP1 enzyme are significantly elevated in a variety of cancer types [36–42] compared with normal tissues, due to genomic stress, rapid proliferation, and abnormal metabolism. Furthermore, this enzyme has been found to have value as a prognostic indicator, particularly as the upregulation of PARP1 has been linked with reduced overall survival [38]. This is most notably the case in brain malignancies which frequently contain elevated levels of PARP1 while healthy brain tissue has extremely low basal expression of this enzyme [40, 41]. These observations strongly indicate that PARP1 is a valuable biomarker of DNA damage which could be detected by PET or SPECT imaging. Furthermore, as PARP1 is a well-established therapeutic target, radiotracers based on PARP-1 inhibitors could find application as companion diagnostics during therapy, since they could provide useful information during the drug development process regarding important aspects of in vivo behaviour, such as biodistribution, pharmacokinetics, and target engagement.
Double-strand break repair mechanisms
Double-strand breaks are the most harmful form of DNA damage as just a single occurrence can potentially result in chromosomal translocation or cell death [43]. The repair of DSBs is executed by two main pathways: HR and NHEJ (Fig. 3) [44].
HR is initiated when ataxia telangiectasia mutated (ATM) kinase protein binds to a DSB, whereupon it is activated and triggers the DNA damage response [45]. In the HR pathway, there are several protagonists, including the MRN complex (Mre11, Rad50, Nbs1), RPA, Rad51, and BRCA1/2. The MRN complex is responsible for the resection of 5’-3’ ends upon DSB recognition which are then coated with RPA [46]. Recombination is performed by Rad51, which replaces RPA in a BRCA1/2-dependant manner to assemble presynaptic Rad51 filaments [47]. A displacement loop (D-Loop) containing the novel heteroduplex DNA is then formed via DNA strand exchange between the target DNA and the Rad51 filament. Lastly, the broken 3’ end primes DNA synthesis using the duplex DNA as a template [48]. Several HR proteins are mutated in cancer, including BRCA1/2 in breast and ovarian cancer. These mutations can lead to inactivation of sub-pathways of HR, thereby driving other genetic effects responsible for the development of cancer.
NHEJ involves binding of the Ku70-Ku80 protein heterodimer to the DNA lesion, followed by the attachment of DNA-dependent protein kinase catalytic subunit (DNA-PKcs). The resulting DNA-PK holoenzyme binds and phosphorylates the protein Artemis, which cleaves the single-stand overhangs of DNA. Lastly, a complex of proteins, including DNA ligase 4 (LIG4), XRCC4 and XLF then complete the process by joining the DNA ends [49]. While NHEJ is less accurate than HR, it can be performed in the absence of undamaged sister chromatid DNA [50]. As with HR, NHEJ is important for genomic integrity since alterations of the Ku complex or LIG4 can cause genome rearrangements [51].
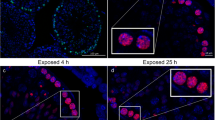
Shortly after a DSB event, the X isoform of the histone H2A is phosphorylated at the serine-139 position by members of the phosphoinositide 3-kinase (PI3K)-related protein kinase (PIKK) family such as ATM, ATR, and DNA-PKcs [52, 53]. The resulting protein, known as γH2AX, forms foci (Fig. 4 [54]) containing hundreds of copies (measuring up to 40 Mbp) [55] around each individual break site. Here, γH2AX is involved in the recruitment of most of the other DNA repair proteins discussed above, whereupon it promotes re-joining of DNA remnants [56, 57]. In addition, γH2AX regulates cell cycle checkpoints to ensure completion of DNA repair and chromatin structure around the affected site. After the damage is repaired, γH2AX is removed, restoring the affected parts of chromatin and preserving both genetic and epigenetic information [58]. The mechanism of removal is still not fully understood, but it has been proposed that it is mediated by dephosphorylation of γH2AX by phosphatases and through histone exchange in the chromatin [59, 60].
Immunostaining of fine-needle aspiration tumour specimens from a patient with non-Hodgkin’s lymphoma deposits reveals the appearance of γH2AX (green) and 53BP1 foci within the nucleus (DAPI, blue) 20 min following irradiation. Reproduced with permission from [54]
Phosphorylation of H2AX at serine-139 is abundant, rapid, and correlates well with each DSB, and consequently γH2AX has become the most commonly probed marker of DNA DSBs. γH2AX has added clinical value as it has been found to be expressed during the early development of most cancers. This includes bladder, breast, cervix, colon, lung, ovaries, pancreas, and skin cancers [57, 61, 62]. Furthermore, in clinical samples, high numbers of γH2AX foci have been correlated with poor outcomes [63]. Taken together, these properties render γH2AX an attractive target for quantitative, high sensitivity molecular imaging techniques.
Methods for DNA damage detection: state-of the-art
One of the most well-established methods of probing DNA damage in vitro is pulsed-field gel electrophoresis (PFGE) [64]. While conventional gel electrophoresis techniques can resolve DNA fragments up to roughly 50 kb, the introduction of an alternating voltage gradient in PFGE is advantageous as it permits resolution of larger DNA fragments up to 10 Mb [65]. Another common electrophoresis-based method is the comet assay which, following separation of DNA fragments, leads to comet-like shapes which can be observed by fluorescence microscopy [66, 67]. The relative intensities of the head (undamaged DNA) and tail (damaged DNA) regions of each comet can be used to quantify SSBs and DSBs in individual eukaryotic cells; although this assay does not reveal the size of individual DNA fragments. Both PGFE and the comet assay have a common drawback as they rely on the extraction of damaged DNA from lysed cells prior to analysis.
The development of confocal immunofluorescence microscopy has since permitted visualisation of DDR proteins within the nuclear compartment of single cells and, in doing so, has helped to elucidate several key DNA repair mechanisms. For example, γH2AX and 53BP1 foci are routinely used to enumerate the extent of DSB repair following genotoxic stimuli [68, 69].
More recently, live cell imaging has provided valuable information on the kinetics of DNA repair in vitro [70]. Most live cell imaging experiments rely on transfection of the cell-of-interest with a gene coding for a chimeric fusion protein of the DDR protein-of-interest coupled to a fluorescent protein such as GFP, YFP, or mCherry. Examples include the use of a 53BP1-mCherry construct to study the kinetics of 53BP1 recruitment to DSB repair foci and their dissolution [71], and the use of Mdc1-, ATR-, and Chk1-GFP fusion proteins to probe the effects of ultraviolet laser radiation [72]. In 2009, Hilario et al. characterised the dynamics of Rad51 filaments and their assembly and disassembly to DNA by single molecule fluorescence microscopy [73]. This was one of the first demonstrations of a real-time study of Rad51 nucleoprotein filament formation, providing details on the rate of filament growth as well as the rate of DNA extension upon Rad51 association.
All of these studies have provided tremendous insight toward the spatiotemporal dynamics of DNA damage repair. The various genetic, biochemical, and molecular biological approaches that have been used to date have characterised in detail the different repair pathways involved. However, much more effort is still required to gain important insight such as how functional pathways are formed by coordination between different repair players, as well as the mechanisms involved in the interaction between these pathways and other cellular processes.
In an effort to extend some of the live cell DDR imaging techniques to an in vivo preclinical setting, Li et al. transfected H322 lung cancer cells with N- and C-terminal fragments of firefly luciferase genes fused with H2AX and MDC1, respectively [74]. Upon irradiation of the cells and formation of DSBs, MDC1 is recruited to phosphorylated H2AX in foci around the DSB, thereby bringing into close proximity both halves of the luciferase protein and allowing the formation of a visible light signal upon addition of luciferin. The same authors also showed that this approach is viable for use in mice bearing subcutaneous xenografts of transfected H322 cells [74]. An initial DDR response was observed in the first day after irradiation (6 Gy) of xenografts. A consecutive apoptotic response, reaching a maximum at 10 days post-irradiation, was also observed.
Later, a luciferase-based reporter was developed to non-invasively test ATM activity in cells and was found to undergo increased activation upon ATM inhibition in a dose-dependent manner, thus enabling the validation of ATM inhibitors in addition to quantifying ATM kinase activity [75]. This method could potentially allow the successful characterisation of ATM inhibitors used in therapeutic regimes, and was also evaluated for in vivo use. Both of these elegant methods allow repeated probing of aspects of the DDR, yet have the main disadvantage that they cannot be translated into clinical use, since transfection of the target cells is a necessity.
A much more advantageous prospect for translation to the clinic is the use of PET or SPECT imaging agents, particularly as they do not rely on modification of the target cell. These functional imaging techniques have excellent sensitivity in comparison to other clinical imaging modalities and are routinely used for in vivo tracking of biomolecular processes. The principal advantages that in vivo imaging of DDR can offer over conventional tissue biopsies are: (1) the ability to analyse larger tissue volumes compared with a small, potentially unrepresentative sample, (2) an improved insight into tumour heterogeneity, (3) the lack of need for an invasive operation to access the area of interest, which completely removes the risk of serious complications related to infection, haemorrhaging etc., and (4) the option to perform repeated imaging of the same area which would allow longitudinal assessment.
Imaging PARP-1 with PET and SPECT
Because of the well-established role of PARP-1 as a mediator in the repair of DNA SSBs, it represents an attractive biomarker for PET and SPECT imaging. Consequently, over the last decade, there have been several attempts to develop radiolabelled imaging agents in order to permit visualisation of this DDR protein:
[11C]-PARP-1 radiotracers
The first example of a PET radiotracer based on a PARP-1 inhibitor was reported by Tu et al. in 2005 [76]. In this case, a phenanthridinone derivative known as PJ34 was selected as it can block NAD+ from its binding site on the activated form of the PARP-1 enzyme. PJ34 was radiolabelled with carbon-11 via a base-catalysed reaction with [11C]methyl iodide and the resulting radiotracer, [11C]PJ34 (Table 1), was used in a rat model of type 1 diabetes to assess its ability to detect early stages of necrosis in pancreatic islets. Promisingly, [11C]PJ34 accumulated significantly more in necrotic pancreases compared with the pancreases of healthy rats at both 5 and 30 min after injection of the radiotracer.
[18F]-PARP-1 radiotracers
A handful of PARP inhibitors radiolabelled with fluorine-18 have also been evaluated and, in some cases, have shown good potential. Initial studies were focused on the preparation of radiofluorinated pirenzepine derivatives and related metabolites due to their ability to inhibit PARP-1 activity [84]. An early example, [18F]-4 (Table 1), reported by Riss et al. in 2009 exhibited a moderate binding affinity (K i) of 200 nM, good stability in human serum, and a logD7.4 value of 1.4, which would likely aid in penetration of cellular membranes although subsequent in vivo evaluation of this radiotracer has not been reported [84].
Weissleder and colleagues subsequently reported an 18F-radiolabeled derivative of the much-studied PARP inhibitor Olaparib, [18F]-BO (Fig. 5, Table 1), which was synthesised via a [4+2] inverse-electron-demand Diels-Alder cycloaddition reaction between a tetrazine-modified Olaparib derivative and an 18F prosthetic group based on trans-cyclooctene [81–83]. The half maximal inhibitory concentration of the resulting compound (IC50 = 17.9±1.1 nM) was only moderately reduced relative to unmodified Olaparib (IC50 = 5 nM) [87], which indicates that minor chemical modifications to the piperazine moiety of Olaparib are only minimally disruptive to PARP-1 binding. In addition to showing PARP-1-mediated cellular uptake in in vitro assays, [18F]-BO was also shown in PET experiments to accumulate specifically in PARP-1 overexpressing MDA-MB-468 breast cancer xenografts in mice [82]. Reiner et al. later demonstrated that [18F]-BO could accurately measure the extent of PARP-1 expression in a variety of xenograft tumour models in mice and showed that uptake of [18F]-BO in ovarian (A2780) xenograft tumours was markedly reduced after administration of Olaparib (Fig. 6) [83]. These compelling findings indicate the potential of this radiotracer to be used as a companion diagnostic for measuring therapeutic drug inhibition in an in vivo setting.
Reiner et al. demonstrated that measuring response to Olaparib treatment is possible using [18F]-BO [83]. Left: In mice bearing A2780 tumour xenografts, tumour-to-muscle contrast ratios markedly reduce following administration of Olaparib. Right: Representative PET/CT images pre- and post-Olaparib administration. Reproduced with permission from [83]
In an effort to create a dual-modality PARP imaging agent, Carlucci et al. developed an Olaparib derivative ([18F]PARPi-FL, Fig. 5, Table 1) containing both BODIPY-FL dye and a fluorine-18 atom which was introduced via an 18F/19F trans-fluorination exchange reaction [77]. While relatively facile, this method of radiolabeling led to low specific activities (2.9±0.7 or 9 mCi/μmol using manual and automated synthesis methods, respectively) compared to the other PARP inhibitors discussed herein. Furthermore, [18F]PARPi-FL was subject to rapid metabolic defluorination, which resulted in high bone uptake (∼10-15%ID/g). This imaging agent was capable, however, of distinguishing U87 glioblastoma xenografts (0.78±0.1%ID/g at 90 min post injection) in small animal PET imaging and ex vivo biodistribution experiments, which was effectively blocked (0.15±0.06%ID/g) with an excess of Olaparib.
While [18F]-BO and [18F]PARPi-FL each contain bulky chemical substituents, a more recent study has resulted in an [18F]-radiolabeled compound which is structurally more consistent with the parent molecule, Olaparib [78]. This radiotracer, [18F]PARPi (Fig. 5, Table 1), has an attractive IC50 value of 2.83 nM and, in contrast to [18F]PARPi-FL, the aromatic carbon-[18F]fluorine bond exhibited high stability, remaining largely intact in human serum samples over 4 h. In ex vivo biodistribution experiments performed at 2 h post injection, this radiotracer achieved uptake values of 1.82±0.21%ID/g in subcutaneous U87 xenograft tumours, which was proven to be mediated by PARP-1. In an orthotopic glioblastoma model, [18F]PARPi was also shown to selectively accumulate in PARP-1-expressing U251-MG tumours (Fig. 7), indicating the ability of this radiotracer to pass through the blood–brain barrier. In a similar manner to the other radiotracers derived from Olaparib discussed herein, the majority of [18F]PARPi excretion occurs via the hepatobiliary clearance pathway.
Top: In orthotopic glioblastoma-bearing mice, PET/MRI images showed pronounced uptake of [18F]PARPi at 2 h post-injection. Bottom: Pre-injection with a 500-fold excess of Olaparib effectively reduced tumour uptake of [18F]PARPi, providing evidence of specificity of the imaging agent for PARP-1. Reproduced with permission from [78]
In 2014, Zhou et al. showed that radiofluorinated derivatives of the benzimidazole NU1085 and its structural near-relative AG014361 also had high inhibitory potency against the PARP-1 enzyme [80]. One of these structures, known as [18F]FluorThanatrace ([18F]FTT; Fig. 8, Table 1), was prepared in considerably higher specific activities compared to the other PARP-1 radiotracers discussed herein (5,500-18,000 mCi/μmol) and was also found to have a reasonable IC50 value of 6.3±1.3 nM. Small animal PET/CT experiments comparing MDA-MB-231 (low PARP-1 expressing) and MDA-MB-468 (high PARP-1 expressing) xenograft tumours revealed PARP-1 mediated tumour uptake which could be blocked following pre-injection of either Olaparib or [19F]FTT. Edmonds et al. also recently used this compound, albeit at a significantly lower specific activity (<2,200 mCi/μmol), and showed that uptake of this radiotracer in a variety of xenograft tumour models in mice could be correlated with intrinsic PARP-1 expression levels. This study also provided compelling evidence that uptake of [18F]FTT is mediated solely by PARP-1, which was concluded after in vitro uptake experiments in PARP-2 knock-out cells revealed specific binding at levels comparable to wild-type cells [79].
Radioiodinated PARP-1 radiotracers
There have been several recent examples of PARP-1 inhibitors labeled with radioisotopes of iodine for both PET and SPECT imaging applications [85, 86]. Two of these reports describe the development of multiple radioiodinated Olaparib derivatives; however, each focuses on the evaluation of a single structurally identical compound, [123/124/131I]-I2-PARPi (Table 1). The Reiner group demonstrated the feasibility of using this compound to detect glioblastoma by targeting orthotopic U251 MG xenografts in mice [86]. SPECT/CT and PET/CT studies involving [131I]-I2-PARPi and [124I]-I2-PARPi, respectively, enabled visualisation of PARP-expressing tumour tissue that could be readily delineated from normal brain tissue. In particular, [124I]-I2-PARPi yielded attractive tumour-to-brain and tumour-to-muscle ratios of 40.0±6.3 and 13.7±4.1, respectively, at 2 h post-injection. Zmuda et al. also demonstrated in a subcutaneous U87MG-Luc2 glioblastoma xenograft model (WHO grade IV) the ability of [123I]-I2-PARPi to be retained within tumour tissue and correlated this uptake to the expression of PARP and the proliferative marker Ki67 [85]. In this case, a peak tumour-to-muscle ratio of 5.61±1.99 was achieved at 2 h post injection.
Mach and colleagues have reported two radioiodinated benzimidazole derivatives, [125I]KX1 [88] and [125I]KX-02-019 [89] (Fig. 8), which bear close structural resemblance to FluorThanatrace. The ability of [125I]KX1 to measure PARP-1 expression in vivo was tested in mice bearing subcutaneous HCC1937 (high PARP-1) and MDA-MB-231 (low PARP-1) human breast cancer xenografts [88]. At 2 h post injection, significantly higher uptake of [125I]KX1 was observed in HCC1937 tumours (reaching approximately 5%ID/g); however, no blocking effect was observed following administration of olaparib suggesting a lack of specificity. It was postulated that differences in the pharmacokinetic profiles of these two agents could prevent a blocking effect, although no additional attempts at blocking with unlabelled KX1 were reported. Autoradiography analysis of HCC1937 tumour tissue at 2 h post injection did however reveal a reduction of signal following Olaparib treatment compared to non-treated mice. It could, therefore, be envisaged that this agent, and analogous agents containing iodine-123/124/131, could serve as companion diagnostic agents during therapy and may assist in patient stratification.
Imaging γH2AX with PET and SPECT
The ability to monitor γH2AX expression in vivo may help to detect certain cancers earlier in their development compared with existing diagnostic methods, facilitating timelier intervention and improved survival. Furthermore, it would also allow indirect monitoring of the DSBs caused by radiotherapy and some chemotherapeutic agents, thus permitting rapid determination of therapeutic efficacy. Consequently, a concerted effort is now underway to develop a non-invasive means of quantifying γH2AX expression levels in vivo using both PET and SPECT imaging techniques.
As an imaging biomarker, γH2AX has several advantages compared with other DDR proteins. For example, H2AX can be phosphorylated throughout the cell cycle, whereas 53BP1, MRE11, and NBS1 are dissociated from DNA damage foci during mitosis [56, 90, 91]. Furthermore, 53BP1 is known to form DNA repair foci via translocation during, which its expression levels do not change dramatically. While PET and SPECT imaging are very sensitive imaging techniques, they cannot distinguish between the various intracellular locations of proteins such as 53BP1. In stark contrast, γH2AX is a new species which is induced by phosphorylation following DSB formation. This on/off switch-like behaviour renders the γH2AX a much more attractive imaging target, i.e. under physiological conditions, cells express little to no yH2AX, whereas upon DSB formation, γH2AX is formed very rapidly.
γH2AX radiotracer development
While anti-γH2AX antibodies are now used routinely in ex vivo assays to quantify the number of γH2AX foci within permeabilised cell populations, the translation of such antibody-based imaging agents into an in vivo setting requires the addition of the cell penetrating peptide (CPP) [92, 93] named “TAT”, which is derived from the transactivator of transcription protein of the HIV-1 virus [94–98]. This arginine-rich peptide has been shown to promote the cellular internalisation of antibodies and a variety of other species, including peptides [99], nanoparticles [100], and liposomes [101]. The precise mechanism(s) of internalisation have been the focus of several studies, many of which have provided strong evidence that electrostatic interaction of the positively charged CPP with negatively charged heparin sulfate proteoglycans on the periphery of the cell membrane plays an important role in promoting internalisation via endocytosis [102, 103]. However, it is worth noting that none of these studies have been able to inhibit completely this process, which suggests that other mechanisms, including energy-independent direct translocation, could also be a contributing factor to this phenomenon. Of the endocytotic pathways that could be responsible, virtually all known possibilities have been implicated, including macropinocytosis [104], clathrin- [105], and caveolin-mediated endocytosis [106]. It is of course possible, if not likely, that more than one mechanism of TAT-internalisation exists and therefore these studies are not necessarily contradictory. The TAT peptide is also known to contain a nuclear localisation sequence (NLS), which, through binding to importins [107], is further trafficked into the nuclear compartment of the cell whereupon the anti-γH2AX antibody can bind to its target.
It is important to note that the CPP does not impart specificity for any particular cancer biomarker and, therefore, it is crucial that the whole antibody construct is able to be externalised so that it can have further opportunity to reach its target. An externalisation mechanism for the TAT protein has been ascertained in a series of elegant experiments which implicate binding to phosphatidylinositol-(4,5)-bisphosphate on the inner leaflet of the cell membrane [108]. The net result of these concurrent cellular import/export mechanisms is that cells in a more active state of DNA damage repair will retain the anti-γH2AX-TAT construct for longer time periods compared with healthy cells with low basal expression levels of γH2AX.
The addition of TAT to the antibody is typically achieved using conventional EDC/sulfo-NHS coupling reagents which promote the formation of an amide bond between the terminal primary amine of TAT and any accessible carboxylic acid residue on the antibody. Alternative bioconjugation methods involving modification of carbohydrate groups on the Fc chain have also been employed successfully [109].
Examples of γH2AX imaging with PET and SPECT
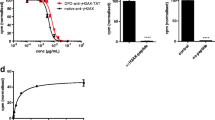
Our first report of a TAT-modified anti-γH2AX antibody emerged in 2011 [110]. Here, the construct was labelled with either a fluorophore or the SPECT radioisotope 111In. When labelled with the fluorophore Alexa Fluor® 488 (λex/em: 495/519 nm) this construct was shown in in vitro experiments to gradually internalise over the course of 23 h into MDA-MB-468 human breast cancer cells which had been exposed to DNA damaging radiation (4 Gy). Promisingly, this compound formed discrete foci within the nuclear compartment which strongly co-localised with staining for γH2AX. The radioactive 111In-anti-γH2AX-TAT compound was also found to be retained significantly longer within irradiated cells compared with a series of experimental controls. The in vivo evaluation of 111In-anti-γH2AX-TAT showed an ability to track DNA damage using a MDA-MB-468 xenograft tumour model in mice (Fig. 9). Here, DNA damage within the tumours was induced by either irradiation or via administration of bleomycin. In both cases, higher uptake of 111In-anti-γH2AX-TAT was observed in the tumours of mice that had received therapy. Taken together, these experiments provided the first compelling evidence of the feasibility of using CPP-modified antibody constructs to image intracellular DDR targets.
Cornelissen et al. showed that uptake of 111In-anti-γH2AX-TAT in MDA-MB-468 breast cancer tumours increased following irradiation in a dose-dependent manner. Reproduced with permission from [110]
The ability of 111In-anti-γH2AX-TAT to image non-invasively DDR during oncogenesis has since been evaluated in a genetically engineered mouse model of HER2/neu-overexpression driven breast cancer [111]. This model results in the development of multiple palpable carcinomas in the mammary fat pads when mice reach 130 days old. In a longitudinal study, SPECT images were acquired on a weekly basis at 24 h post-injection of 111In-anti-γH2AX-TAT. In SPECT images acquired from mice between 76–110 days old, uptake of the radiotracer in the mammary fat pads was markedly higher compared with a non-specific isotype-matched antibody which was modified in an identical manner. Immunohistochemical analysis of resected mammary fats pads showed that the number of γH2AX foci per cell reached a peak within this age range and was significantly greater compared with mice <76 or >106 days old. Encouragingly, it was found that the median time to the detection of positive tissue with SPECT imaging (96 days) was much earlier compared with the detection of lesions >150 μm by DCE-MRI (120 days) or by palpation (131 days) (Fig. 10).
While advances in SPECT technology (specifically relating to improvements to collimators, quantitation, and reconstruction software) are leading to a revival in this modalities appeal, PET/CT imaging has gained acceptance as the standard of care in the management of cancer. This is due in part to the high resolution and sensitivity that clinical PET has so far offered compared with SPECT, and the ability during processing of PET images to accurately correct for signal attenuation. These important advantages have resulted in higher quality images from which more meaningful data can be extracted. Consequently, a PET radiotracer based on the anti-γH2AX-TAT antibody construct has been developed containing the radiometal zirconium-89.
The performance of 89Zr-anti-γH2AX-TAT has largely shown consistency with its indium-111 radiolabelled analogue [112]. In cells exposed to DNA damaging radiation (4 Gy), 89Zr-anti-γH2AX-TAT exhibits considerably (eightfold) longer retention compared with control experiments involving non-irradiated cells or a non-specific IgG. Furthermore, in subcutaneous MDA-MB-468 xenograft tumours in mice, higher uptake of 89Zr-anti-γH2AX-TAT (0.5 MBq, 5 μg) was found following exposure to radiation (10 Gy) compared with experimental controls (12.1 ± 1.6%ID/g, P < 0.001).
Radiotracers based on anti-γH2AX-TAT constructs have shown promise in a variety of preclinical models based on early detection and therapy evaluation. However, prior to clinical translation, some important issues require consideration. Firstly, it will be important to integrate a humanised version of the anti-γH2AX antibody (the antibody used in these studies is raised in rabbit) in order to prevent the invocation of an immune response. Secondly, as non-specific tumour uptake (resulting from the enhanced permeability and retention [EPR] effect [113]) is responsible for a substantial contribution to overall tumour uptake, it will be desirable to amplify γH2AX-mediated contrast. We hypothesise that this could be achieved by improving delivery of the construct to tumours by, for example, attachment of tumour-targeting peptides [114]. This may also be achieved by reducing non-specific uptake resulting from EPR by either using smaller antibody fragments (minibodies, diabodies, etc.) [115] or by adopting a pretargeted imaging approach [116].
Lastly, it is worth noting that γH2AX is a secondary marker of DNA DSBs and its expression or foci number is not a direct 1:1 measure of DSBs. This leads to difficulty in quantifying precise numbers of DSBs that are being visualised in vivo, especially when considering the relatively slow pharmacokinetic profile of antibody-based imaging agents. Therefore, it is important that the biology of yH2AX is taken into account when interpreting images and image quantitation. At present, no in vivo imaging modality exists that is sufficiently sensitive to image the DSBs directly.
Conclusions
Our understanding of the cellular processes that are invoked in response to DNA damage has improved considerably over recent years. Advancements in this area have revealed attractive biomarkers, which could be used to improve upon existing methods for non-invasive early cancer detection and therapy evaluation. With these aims in mind, a range of PET/SPECT imaging agents are currently under development and, in some cases, are poised for evaluation in clinical settings. So far, these imaging agents have mostly consisted of small molecule inhibitors of the PARP-1 enzyme or antibody-based constructs targeting yH2AX.
Most of the reports described herein have demonstrated the ability to measure PARP-1 or yH2AX expression levels in preclinical in vivo experiments and some studies have been able to detect changes in expression following chemo- or radiotherapy. While these findings are very promising, it should be noted that uptake of these agents within tumours is generally low, particularly in comparison to what can often be achieved through targeting of cancer biomarkers situated on the cell-surface. This can be attributed to a variety of factors, including the transient nature of DDR proteins, and the inefficient internalisation/nuclear translocation of these agents. While the first of these challenges is an inherent and unavoidable obstacle in this endeavour, the latter is an area which can conceivably be improved upon through advances in the understanding of the various internalisation mechanisms and by improved chemical design.
There is also a need to identify which of the multitude of proteins involved in the DDR process represent the most valuable targets. This complex task will involve measuring intranuclear concentrations of target proteins at various stages during oncogenesis (as well as during cancer therapy), and determining relative abundances in cancerous versus healthy tissues. The duration of the targets’ existence is also an important parameter, particularly if it will be used to assess DDR activation following radiotherapy where expression levels of many biomarkers, including PARP-1 and γH2AX, disappear within days.
Of equal importance is the need to develop improved PET and SPECT imaging agents which can bind to these targets with high affinity and specificity, while also possessing the necessary chemical properties that would promote efficient cellular internalisation. Certainly, this will be aided by recent advances in radiofluorination chemistry which allow far greater freedom in the design of novel PET radiotracers compared to what has previously been possible with conventional radiosynthetic approaches.
More broadly, non-invasive imaging of intracellular targets is an important research endeavour with implications which extend beyond imaging the DNA damage response. This ability also opens the door to a multitude of other intracellular biomarkers with relevancy to additional aspects of cancer biology.
References
Jackson SP, Bartek J. The DNA-damage response in human biology and disease. Nature. 2009;461(7267):1071–8. http://www.nature.com/nature/journal/v461/n7267/suppinfo/nature08467_S1.html.
Hoeijmakers JHJ. DNA damage, aging, and cancer. N Engl J Med. 2009;361(15):1475–85. doi:10.1056/NEJMra0804615.
Sirbu BM, Cortez D. DNA damage response: three levels of dna repair regulation. Cold Spring Harb Perspect Biol. 2013;5(8). doi:10.1101/cshperspect.a012724.
Brenerman BM, Illuzzi JL, Wilson DM. Base excision repair capacity in informing healthspan. Carcinogenesis. 2014;35(12):2643–52. doi:10.1093/carcin/bgu225.
Lord CJ, Ashworth A. The DNA damage response and cancer therapy. Nature. 2012;481(7381):287–94.
Li X, Xu H, Xu C, Lin M, Song X, Yi F, et al. The Yin-Yang of DNA damage response: roles in tumorigenesis and cellular senescence. Int J Mol Sci. 2013;14(2):2431–48. doi:10.3390/ijms14022431.
Halazonetis TD, Gorgoulis VG, Bartek J. An oncogene-induced DNA damage model for cancer development. Science. 2008;319(5868):1352–5. doi:10.1126/science.1140735.
Koorstra JBM, Hong SM, Shi C, Meeker AK, Ryu JK, Offerhaus GJA, et al. Widespread activation of the DNA damage response in human pancreatic intraepithelial neoplasia. Modern Pathol. 2009;22(11):1439–45. doi:10.1038/modpathol.2009.114.
Kshirsagar M, Jiang W, Shih IM. DNA damage response is prominent in ovarian high-grade serous carcinomas, especially those with Rsf-1 (HBXAP) overexpression. J Oncol. 2012. doi:10.1155/2012/621685.
Matsuda Y, Wakai T, Kubota M, Osawa M, Takamura M, Yamagiwa S et al. DNA damage sensor γ-H2AX is increased in preneoplastic lesions of hepatocellular carcinoma. Scientific World J. 2013;2013. doi:10.1155/2013/597095.
Marteijn JA, Lans H, Vermeulen W, Hoeijmakers JHJ. Understanding nucleotide excision repair and its roles in cancer and ageing. Nat Rev Mol Cell Biol. 2014;15(7):465–81. doi:10.1038/nrm3822.
Gorgoulis VG, Vassiliou LVF, Karakaidos P, Zacharatos P, Kotsinas A, Liloglou T, et al. Activation of the DNA damage checkpoint and genomic instability in human precancerous lesions. Nature. 2005;434(7035):907–13. doi:10.1038/nature03485.
Sedelnikova OA, Bonner WM. γH2AX in cancer cells: a potential biomarker for cancer diagnostics, prediction and recurrence. Cell Cycle. 2006;5(24):2909–13.
Seeberg E, Eide L, Bjørås M. The base excision repair pathway. Trends Biochem Sci. 1995;20(10):391–7. doi:10.1016/S0968-0004(00)89086-6.
Dianov GL, Hübscher U. Mammalian base excision repair: the forgotten archangel. Nucleic Acids Res. 2013;41(6):3483–90. doi:10.1093/nar/gkt076.
Dexheimer ST. DNA repair pathways and mechanisms. In: Mathews AL, Cabarcas MS, Hurt ME, editors. DNA Repair of Cancer Stem Cells. Dordrecht: Springer Netherlands; 2013. p. 19–32.
Kim Y-J, Wilson DM. Overview of base excision repair biochemistry. Curr Mol Pharmacol. 2012;5(1):3–13.
D’Amours D, Desnoyers S, D’Silva I, Poirier GG. Poly(ADP-ribosyl)ation reactions in the regulation of nuclear functions. Biochem J. 1999;342(2):249–68. doi:10.1042/0264-6021:3420249.
Schreiber V, Dantzer F, Amé JC, De Murcia G. Poly(ADP-ribose): novel functions for an old molecule. Nat Rev Mol Cell Biol. 2006;7(7):517–28. doi:10.1038/nrm1963.
Amé JC, Spenlehauer C, De Murcia G. The PARP superfamily. BioEssays. 2004;26(8):882–93. doi:10.1002/bies.20085.
de Murcia G, de Murcia JM. Poly(ADP-ribose) polymerase: a molecular nick-sensor. Trends Biochem Sci. 1994;19(4):172–3. doi:10.1016/0968-0004(94)90280-1.
Lindahl T, Satoh MS, Poirier GG, Klungland A. Post-translational modification of poly(ADP-ribose) polymerase induced by DNA strand breaks. Trends Biochem Sci. 1995;20(10):405–11. doi:10.1016/S0968-0004(00)89089-1.
de Murcia G, Schreiber V, Molinete M, Saulier B, Poch O, Masson M, et al. Structure and function of poly(ADP-ribose) polymerase. Mol Cell Biochem. 1994;138(1):15–24. doi:10.1007/BF00928438.
El‐Khamisy SF, Masutani M, Suzuki H, Caldecott KW. A requirement for PARP‐1 for the assembly or stability of XRCC1 nuclear foci at sites of oxidative DNA damage. Nucleic Acids Res. 2003;31(19):5526–33. doi:10.1093/nar/gkg761.
Malanga M, Althaus FR. The role of poly(ADP-ribose) in the DNA damage signaling network. Biochem Cell Biol. 2005;83(3):354–64. doi:10.1139/o05-038.
Virág L, Szabó C. The therapeutic potential of poly(ADP-Ribose) polymerase inhibitors. Pharmacol Rev. 2002;54(3):375–429.
Curtin NJ, Szabo C. Therapeutic applications of PARP inhibitors: Anticancer therapy and beyond. Mol Aspects Med. 2013;34(6):1217–56. doi:10.1016/j.mam.2013.01.006.
Ferraris DV. Evolution of poly(ADP-ribose) polymerase-1 (PARP-1) inhibitors. from concept to clinic. J Med Chem. 2010;53(12):4561–84. doi:10.1021/jm100012m.
Rouleau M, Patel A, Hendzel MJ, Kaufmann SH, Poirier GG. PARP inhibition: PARP1 and beyond. Nat Rev Cancer. 2010;10(4):293–301. doi:10.1038/nrc2812.
Javle M, Curtin NJ. The potential for poly (ADP-ribose) polymerase inhibitors in cancer therapy. Ther Adv Med Oncol. 2011;3(6):257–67. doi:10.1177/1758834011417039.
Shall S, de Murcia G. Poly(ADP-ribose) polymerase-1: what have we learned from the deficient mouse model? Mutat Res-DNA Repair. 2000;460(1):1–15. doi:10.1016/S0921-8777(00)00016-1.
Bryant HE, Schultz N, Thomas HD, Parker KM, Flower D, Lopez E, et al. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature. 2005;434(7035):913–7. http://www.nature.com/nature/journal/v434/n7035/suppinfo/nature03443_S1.html.
Farmer H, McCabe N, Lord CJ, Tutt ANJ, Johnson DA, Richardson TB, et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature. 2005;434(7035):917–21. http://www.nature.com/nature/journal/v434/n7035/suppinfo/nature03445_S1.html.
Patel AG, Sarkaria JN, Kaufmann SH. Nonhomologous end joining drives poly(ADP-ribose) polymerase (PARP) inhibitor lethality in homologous recombination-deficient cells. Proc Natl Acad Sci U S A. 2011;108(8):3406–11. doi:10.1073/pnas.1013715108.
Sonnenblick A, De Azambuja E, Azim HA, Piccart M. An update on PARP inhibitors - Moving to the adjuvant setting. Nat Rev Clin Oncol. 2015;12(1):27–41. doi:10.1038/nrclinonc.2014.163.
Ossovskaya V, Koo IC, Kaldjian EP, Alvares C, Sherman BM. Upregulation of poly (ADP-Ribose) polymerase-1 (PARP1) in triple-negative breast cancer and other primary human tumor types. Genes Cancer. 2010;1(8):812–21. doi:10.1177/1947601910383418.
Bièche I, De Murcia G, Lidereau R. Poly(ADP-ribose) polymerase gene expression status and genomic instability in human breast cancer. Clin Cancer Res. 1996;2(7):1163–7.
Rojo F, García-Parra J, Zazo S, Tusquets I, Ferrer-Lozano J, Menendez S, et al. Nuclear PARP-1 protein overexpression is associated with poor overall survival in early breast cancer. Ann Oncol. 2012;23(5):1156–64. doi:10.1093/annonc/mdr361.
Alanazi M, Pathan AAK, Abduljaleel Z. Association between PARP-1 V762A polymorphism and breast cancer susceptibility in Saudi population. PLoS One. 2013;3:92360.
Galia A, Calogero AE, Condorelli R, Fraggetta F, La Corte A, Ridolfo F et al. PARP-1 protein expression in glioblastoma multiforme. Eur J Histochem. 2012;56(1).
Barton VN, Donson AM, Kleinschmidt-DeMasters BK, Gore L, Liu AK, Foreman NK. PARP1 expression in pediatric central nervous system tumors. Pediatr Blood Cancer. 2009;53(7):1227–30. doi:10.1002/pbc.22141.
Staibano S, Pepe S, Muzio LL, Somma P, Mascolo M, Argenziano G, et al. Poly(adenosine diphosphate-ribose) polymerase 1 expression in malignant melanomas from photoexposed areas of the head and neck region. Hum Pathol. 2005;36(7):724–31. doi:10.1016/j.humpath.2005.04.017.
Khanna KK, Jackson SP. DNA double-strand breaks: signaling, repair and the cancer connection. Nat Genet. 2001;27(3):247–54.
McKinnon PJ, Caldecott KW. DNA strand break repair and human genetic disease. Annu Rev Genomics Hum Genet. 2007;8:37–55.
Cerbinskaite A, Mukhopadhyay A, Plummer ER, Curtin NJ, Edmondson RJ. Defective homologous recombination in human cancers. Cancer Treat Rev. 2012;38(2):89–100.
Krajewska M, Fehrmann RSN, De Vries EGE, van Vugt MATM. Regulators of homologous recombination repair as novel targets for cancer treatment. Front Genet. 2015;6:1–15.
Helleday T. Homologous recombination in cancer development, treatment and development of drug resistance. Carcinogenesis. 2010;31(6):955–60.
Renkawitz J, Lademann CA, Jentsch S. Mechanisms and principles of homology search during recombination. Nat Rev Mol Cell Biol. 2014;15(6):369–83.
Bunting SF, Nussenzweig A. End-joining, translocations and cancer. Nat Rev Cancer. 2013;13(7):443–54. doi:10.1038/nrc3537.
O’Connor MJ. Targeting the DNA damage response in cancer. Mol Cell. 2015;60(4):547–60.
Yan CT, Boboila C, Souza EK, Franco S, Hickernell TR, Murphy M, et al. IgH class switching and translocations use a robust non-classical end-joining pathway. Nature. 2007;449(7161):478–82.
Kuo LJ, Yang L-X. γ-H2AX - A novel biomarker for DNA double-strand breaks. In Vivo. 2008;22(3):305–9.
Sharma A, Singh K, Almasan A. Histone H2AX phosphorylation: a marker for DNA damage. In: Bjergbæk L, editor. DNA Repair Protocols. Totowa: Humana Press; 2012. p. 613–26.
Shah K, Boghozian RA, Kartsonaki C, Shah KA, Vallis KA. γH2AX expression in cytological specimens as a biomarker of response to radiotherapy in solid malignancies. Diagn Cytopathol. 2016;44(2):141–6. doi:10.1002/dc.23396.
Banáth JP, Olive PL. Expression of phosphorylated histone H2AX as a surrogate of cell killing by drugs that create DNA double-strand breaks. Cancer Res. 2003;63(15):4347–50.
Nakamura AJ, Rao VA, Pommier Y, Bonner WM. The complexity of phosphorylated H2AX foci formation and DNA repair assembly at DNA double-strand breaks. Cell Cycle. 2010;9(2):389–97. doi:10.4161/cc.9.2.10475.
Bonner WM, Redon CE, Dickey JS, Nakamura AJ, Sedelnikova OA, Solier S, et al. γH2AX and cancer. Nat Rev Cancer. 2008;8(12):957–67. doi:10.1038/nrc2523.
Mah LJ, El-Osta A, Karagiannis TC. [gamma]H2AX: a sensitive molecular marker of DNA damage and repair. Leukemia. 2010;24(4):679–86.
Kusch T, Florens L, Macdonald WH, Swanson SK, Glaser RL, Yates 3rd JR, et al. Acetylation by Tip60 is required for selective histone variant exchange at DNA lesions. Science. 2004;306(5704):2084–7.
Downs JA, Allard S, Jobin-Robitaille O, Javaheri A, Auger A, Bouchard N, et al. Binding of chromatin-modifying activities to phosphorylated histone H2A at DNA damage sites. Mol Cell. 2004;16(6):979–90. doi:10.1016/j.molcel.2004.12.003.
Bartkova J, Horejsi Z, Koed K, Kramer A, Tort F, Zieger K, et al. DNA damage response as a candidate anti-cancer barrier in early human tumorigenesis. Nature. 2005;434(7035):864–70. http://www.nature.com/nature/journal/v434/n7035/suppinfo/nature03482_S1.html.
Bartek J, Bartkova J, Lukas J. DNA damage signalling guards against activated oncogenes and tumour progression. Oncogene. 2007;26(56):7773–9.
Dickey JS, Redon CE, Nakamura AJ, Baird BJ, Sedelnikova OA, Bonner WM. H2AX: functional roles and potential applications. Chromosoma. 2009;118(6):683–92. doi:10.1007/s00412-009-0234-4.
Cedervall B, Wong R, Albright N, Dynlacht J, Lambin P, Dewey WC. Methods for the quantification of DNA double-strand breaks determined from the distribution of DNA fragment sizes measured by pulsed-field gel electrophoresis. Radiat Res. 1995;143(1):8–16. doi:10.2307/3578920.
Herschleb J, Ananiev G, Schwartz DC. Pulsed-field gel electrophoresis. Nat Protoc. 2007;2(3):677–84. http://www.nature.com/nprot/journal/v2/n3/suppinfo/nprot.2007.94_S1.html.
Collins AR. The comet assay for DNA damage and repair: Principles, applications, and limitations. Appl Biochem Biotech - Part B Mol Biotech. 2004;26(3):249–61. doi:10.1385/MB:26:3:249.
Olive PL, Banáth JP. The comet assay: a method to measure DNA damage in individual cells. Nat Protoc. 2006;1(1):23–9. doi:10.1038/nprot.2006.5.
Löbrich M, Shibata A, Beucher A, Fisher A, Ensminger M, Goodarzi AA, et al. γH2AX foci analysis for monitoring DNA double-strand break repair: strengths, limitations and optimization. Cell Cycle. 2010;9(4):662–9.
Panier S, Boulton SJ. Double-strand break repair: 53BP1 comes into focus. Nat Rev Mol Cell Biol. 2014;15(1):7–18. doi:10.1038/nrm3719.
Rizzuto R, Brini M, Pizzo P, Murgia M, Pozzan T. Chimeric green fluorescent protein as a tool for visualizing subcellular organelles in living cells. Curr Biol. 1995;5(6):635–42.
Loewer A, Karanam K, Mock C, Lahav G. The p53 response in single cells is linearly correlated to the number of DNA breaks without a distinct threshold. BMC Biol. 2013;11:114. doi:10.1186/1741-7007-11-114.
Bekker-Jensen S, Lukas C, Kitagawa R, Melander F, Kastan MB, Bartek J, et al. Spatial organization of the mammalian genome surveillance machinery in response to DNA strand breaks. J Cell Biol. 2006;173(2):195–206. doi:10.1083/jcb.200510130.
Hilario J, Amitani I, Baskin RJ, Kowalczykowski SC. Direct imaging of human Rad51 nucleoprotein dynamics on individual DNA molecules. Proc Natl Acad Sci U S A. 2009;106(2):361–8. doi:10.1073/pnas.0811965106.
Li W, Li F, Huang Q, Shen J, Wolf F, He Y, et al. Quantitative, noninvasive imaging of radiation-induced DNA double-strand breaks in vivo. Cancer Res. 2011;71(12):4130–7. doi:10.1158/0008-5472.CAN-10-2540.
Williams TM, Nyati S, Ross BD, Rehemtulla A. Molecular imaging of the ATM kinase activity. Int J Radiat Oncol. 2013;86(5):969–77. doi:10.1016/j.ijrobp.2013.04.028.
Tu Z, Chu W, Zhang J, Dence CS, Welch MJ, Mach RH. Synthesis and in vivo evaluation of [11C]PJ34, a potential radiotracer for imaging the role of PARP-1 in necrosis. Nucl Med Biol. 2005;32(5):437–43. doi:10.1016/j.nucmedbio.2005.03.001.
Carlucci G, Carney B, Brand C, Kossatz S, Irwin CP, Carlin SD, et al. Dual-modality optical/PET imaging of PARP1 in glioblastoma. Mol Imaging Biol. 2015;17(6):848–55. doi:10.1007/s11307-015-0858-0.
Carney B, Carlucci G, Salinas B, Di Gialleonardo V, Kossatz S, Vansteene A, et al. Non-invasive PET imaging of PARP1 expression in glioblastoma models. Mol Imaging Biol. 2015. doi:10.1007/s11307-015-0904-y.
Edmonds CE, Makvandi M, Lieberman BP, Xu K, Zeng C, Li S, et al. [18F]FluorThanatrace uptake as a marker of PARP1 expression and activity in breast cancer. Am J Nucl Med Mol Imaging. 2016;6(1):94–101.
Zhou D, Chu W, Xu J, Jones LA, Peng X, Li S, et al. Synthesis, [18F] radiolabeling, and evaluation of poly (ADP-ribose) polymerase-1 (PARP-1) inhibitors for in vivo imaging of PARP-1 using positron emission tomography. Bioorg Med Chem. 2014;22(5):1700–7. doi:10.1016/j.bmc.2014.01.019.
Keliher EJ, Reiner T, Turetsky A, Hilderbrand SA, Weissleder R. High-yielding, two-step 18F labeling strategy for 18F-PARP1 inhibitors. ChemMedChem. 2011;6(3):424–7. doi:10.1002/cmdc.201000426.
Reiner T, Keliher EJ, Earley S, Marinelli B, Weissleder R. Synthesis and in vivo imaging of a 18F-labeled PARP1 inhibitor using a chemically orthogonal scavenger-assisted high-performance method. Angew Chem Int Ed. 2011;50(8):1922–5. doi:10.1002/anie.201006579.
Reiner T, Lacy J, Keliher EJ, Yang KS, Ullal A, Kohler RH, et al. Imaging therapeutic PARP inhibition in vivo through bioorthogonally developed companion imaging agents. Neoplasia. 2012;14:169–77.
Riss PJ, Soskic V, Schrattenholz A, Roescha F. Synthesis and radiosynthesis of N5-[18F]fluoroethyl- Pirenzepine and its metabolite N5-[18F]fluoroethyl-LS 75. J Labelled Comp Radiopharm. 2009;52(14):576–9. doi:10.1002/jlcr.1687.
Zmuda F, Malviya G, Blair A, Boyd M, Chalmers AJ, Sutherland A, et al. Synthesis and evaluation of a radioiodinated tracer with specificity for poly(ADP-ribose) polymerase-1 (PARP-1) in vivo. J Med Chem. 2015;58(21):8683–93. doi:10.1021/acs.jmedchem.5b01324.
Salinas B, Irwin CP, Kossatz S, Bolaender A, Chiosis G, Pillarsetty N et al. Radioiodinated PARP1 tracers for glioblastoma imaging. Eur J Nucl Med Mol Imaging Res. 2015;5(1). doi:10.1186/s13550-015-0123-1.
Menear KA, Adcock C, Boulter R, Cockcroft X-L, Copsey L, Cranston A, et al. 4-[3-(4-Cyclopropanecarbonylpiperazine-1-carbonyl)-4-fluorobenzyl]-2H-phthalazin-1-one: a novel bioavailable inhibitor of poly(ADP-ribose) polymerase-1. J Med Chem. 2008;51(20):6581–91. doi:10.1021/jm8001263.
Makvandi M, Xu K, Lieberman BP, Anderson R-C, Effron SS, Winters HD, et al. A radiotracer strategy to quantify PARP-1 expression in vivo provides a biomarker that can enable patient selection for PARP inhibitor therapy. Cancer Res. 2016;76(15):4516.
Anderson R-C, Makvandi M, Xu K, Lieberman BP, Zeng C, Pryma DA, et al. Iodinated benzimidazole PARP radiotracer for evaluating PARP1/2 expression in vitro and in vivo. Nucl Med Biol. 2016. doi:10.1016/j.nucmedbio.2016.08.007.
Giunta S, Belotserkovskaya R, Jackson SP. DNA damage signaling in response to double-strand breaks during mitosis. J Cell Biol. 2010;190(2):197–207. doi:10.1083/jcb.200911156.
Nelson G, Buhmann M, Von Zglinicki T. DNA damage foci in mitosis are devoid of 53BP1. Cell Cycle. 2009;8(20):3379–83.
Kersemans V, Kersemans K, Cornelissen B. Cell penetrating peptides for in vivo molecular imaging applications. Curr Pharm Des. 2008;14(24):2415–27. doi:10.2174/138161208785777432.
Kersemans V, Cornelissen B. Targeting the tumour: cell penetrating peptides for molecular imaging and radiotherapy. Pharmaceuticals. 2010;3(3):600–20. doi:10.3390/ph3030600.
Zhao M, Weissleder R. Intracellular cargo delivery using tat peptide and derivatives. Med Res Rev. 2004;24(1):1–12. doi:10.1002/med.10056.
Vives E. Cellular utake of the Tat peptide: an endocytosis mechanism following ionic interactions. J Mol Recogn. 2003;16(5):265–71. doi:10.1002/jmr.636.
Mishra A, Lai GH, Schmidt NW, Sun VZ, Rodriguez AR, Tong R, et al. Translocation of HIV TAT peptide and analogues induced by multiplexed membrane and cytoskeletal interactions. Proc Natl Acad Sci U S A. 2011;108(41):16883–8. doi:10.1073/pnas.1108795108.
Ziegler A, Nervi P, Dürrenberger M, Seelig J. The cationic cell-penetrating peptide CPPTAT derived from the HIV-1 protein TAT is rapidly transported into living fibroblasts: optical, biophysical, and metabolic evidence. Biochemistry. 2005;44(1):138–48. doi:10.1021/bi0491604.
Brooks H, Lebleu B, Vivès E. Tat peptide-mediated cellular delivery: back to basics. Adv Drug Deliv Rev. 2005;57(4):559–77. doi:10.1016/j.addr.2004.12.001.
Jones SW, Christison R, Bundell K, Voyce CJ, Brockbank SMV, Newham P, et al. Characterisation of cell-penetrating peptide-mediated peptide delivery. Br J Pharmacol. 2005;145(8):1093–102. doi:10.1038/sj.bjp.0706279.
Pan L, He Q, Liu J, Chen Y, Ma M, Zhang L, et al. Nuclear-targeted drug delivery of TAT peptide-conjugated monodisperse mesoporous silica nanoparticles. J Am Chem Soc. 2012;134(13):5722–5. doi:10.1021/ja211035w.
Torchilin VP, Rammohan R, Weissig V, Levchenko TS. TAT peptide on the surface of liposomes affords their efficient intracellular delivery even at low temperature and in the presence of metabolic inhibitors. Proc Natl Acad Sci U S A. 2001;98(15):8786–91. doi:10.1073/pnas.151247498.
Pang H-B, Braun GB, Ruoslahti E. Neuropilin-1 and heparan sulfate proteoglycans cooperate in cellular uptake of nanoparticles functionalized by cationic cell-penetrating peptides. Sci Adv. 2015;1(10). doi:10.1126/sciadv.1500821.
Tyagi M, Rusnati M, Presta M, Giacca M. Internalization of HIV-1 tat requires cell surface heparan sulfate proteoglycans. (0021–9258 (Print)).
Kaplan IM, Wadia JS, Dowdy SF. Cationic TAT peptide transduction domain enters cells by macropinocytosis. J Control Release. 2005;102(1):247–53. doi:10.1016/j.jconrel.2004.10.018.
Kawaguchi Y, Takeuchi T, Kuwata K, Chiba J, Hatanaka Y, Nakase I, et al. Syndecan-4 is a receptor for clathrin-mediated endocytosis of arginine-rich cell-penetrating peptides. Bioconjugate Chem. 2016;27(4):1119–30. doi:10.1021/acs.bioconjchem.6b00082.
Ferrari A, Pellegrini V, Arcangeli C, Fittipaldi A, Giacca M, Beltram F. Caveolae-mediated internalization of extracellular HIV-1 tat fusion proteins visualized in real time. Mol Ther. 2003;8(2):284–94.
Truant R, Cullen BR. The arginine-rich domains present in human immunodeficiency virus Type 1 Tat and Rev function as direct importin β-dependent nuclear localization signals. Mol Cell Biol. 1999;19(2):1210–7. doi:10.1128/mcb.19.2.1210.
Rayne F, Debaisieux S, Yezid H, Lin YL, Mettling C, Konate K, et al. Phosphatidylinositol-(4,5)-bisphosphate enables efficient secretion of HIV-1 Tat by infected T-cells. EMBO J. 2010;29(8):1348–62. doi:10.1038/emboj.2010.32.
Cornelissen B, McLarty K, Kersemans V, Scollard DA, Reilly RM. Properties of [111In]-labeled HIV-1 tat peptide radioimmunoconjugates in tumor-bearing mice following intravenous or intratumoral injection. Nucl Med Biol. 2008;35(1):101–10. doi:10.1016/j.nucmedbio.2007.09.007.
Cornelissen B, Kersemans V, Darbar S, Thompson J, Shah K, Sleeth K, et al. Imaging DNA damage in vivo using γH2AX-targeted immunoconjugates. Cancer Res. 2011;71(13):4539–49. doi:10.1158/0008-5472.CAN-10-4587.
Cornelissen B, Able S, Kartsonaki C, Kersemans V, Allen PD, Cavallo F, et al. Imaging DNA damage allows detection of preneoplasia in the BALB-neuT model of breast cancer. J Nucl Med. 2014;55(12):2026–31. doi:10.2967/jnumed.114.142083.
Knight JC, Topping C, Mosley M, Kersemans V, Falzone N, Fernández-Varea JM, et al. PET imaging of DNA damage using 89Zr-labelled anti-γH2AX-TAT immunoconjugates. Eur J Nucl Med Mol Imaging. 2015;42(11):1707–17. doi:10.1007/s00259-015-3092-8.
Maeda H. The enhanced permeability and retention (EPR) effect in tumor vasculature: the key role of tumor-selective macromolecular drug targeting. Adv Enzym Regul. 2001;41(1):189–207. doi:10.1016/S0065-2571(00)00013-3.
Cornelissen B, Waller A, Able S, Vallis KA. Molecular radiotherapy using cleavable radioimmunoconjugates that target eGFR and γH2AX. Mol Cancer Ther. 2013;12(11):2472–82. doi:10.1158/1535-7163.
Olafsen T, Wu AM. Antibody vectors for imaging. Semin Nucl Med. 2010;40(3):167–81. doi:10.1053/j.semnuclmed.2009.12.005.
Knight JC, Cornelissen B. Bioorthogonal chemistry: implications for pretargeted nuclear (PET/SPECT) imaging and therapy. Am J Nucl Med Mol Imaging. 2014;4(2):96–113.
Acknowledgements
This research was supported by the CRUK/MRC Oxford Institute for Radiation Oncology.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This study was funded by Cancer Research UK
Conflict of interest
James C. Knight declares that he has no conflict of interest. Sofia Koustoulidou declares that she has no conflict of interest. Bart Cornelissen declares that he has no conflict of interest.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed. This article does not contain any studies with human participants performed by any of the authors.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Knight, J.C., Koustoulidou, S. & Cornelissen, B. Imaging the DNA damage response with PET and SPECT. Eur J Nucl Med Mol Imaging 44, 1065–1078 (2017). https://doi.org/10.1007/s00259-016-3604-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-016-3604-1