Abstract
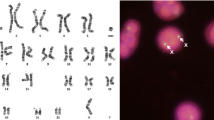
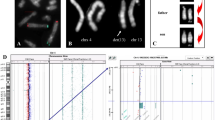
The study of de novo balanced chromosome rearrangements associated with phenotypic alterations has provided an important tool for the understanding of pathogenic mechanisms, for they can reveal gene disruptions and their effects in patients. Because balanced rearrangements show no copy number alterations, standard array techniques are inefficient in studying these cases. For this reason, a variation of the array technique, named array painting, is required for determining the chromosome breakpoints. Using this technique, only the rearranged chromosomes, or chromosome segments including the rearranged region, are separated by microdissection, differentially labeled, and hybridized in the array CGH. The chromosome region corresponding to transition point from one hybridization fluorochrome signal to the other reveals the breakpoint. The resolution of the technique depends mainly on the array type and the density of probes as well as on the repetitiveness of the DNA sequence at the breakpoint.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Menten B, Buysse K, Vermeulen S et al (2007) Report of a female patient with mental retardation and tall stature due to a chromosomal rearrangement disrupting the OPHN1 gene on Xq12. Eur J Med Genet 50:446–454
Moysés-Oliveira M, Guilherme RS, Meloni VA et al (2015) X-linked intellectual disability related genes disrupted by balanced X-autosome translocations. Am J Med Genet B 168:669–677
Schneider A, Puechberty J, Ng BL et al (2015) Identification of disrupted AUTS2 and EPHA6 genes by array painting in a patient carrying a de novo balanced translocation t(3;7) with intellectual disability and neurodevelopment disorder. Am J Med Genet A 167:3031–3037
Di Gregorio E, Bianchi FT, Schiavi A et al (2013) A de novo X;8 translocation creates a PTK2-THOC2 gene fusion with THOC2 expression knockdown in a patient with psychomotor retardation and congenital cerebellar hypoplasia. J Med Genet 50:543–551
Córdova-Fletes C, Domínguez MG, Delint-Ramirez I et al (2015) A de novo t(10;19)(q22.3;q13.33) leads to ZMIZ1/PRR12 reciprocal fusion transcripts in a girl with intellectual disability and neuropsychiatric alterations. Neurogenetics 16:287–298
Fonseca AC, Bonaldi A, Bertola DR et al (2013) The clinical impact of chromosomal rearrangements with breakpoints upstream of the SOX9 gene: two novel de novo balanced translocations associated with acampomelic campomelic dysplasia. BMC Med Genet 14:50
Crippa M, Bestetti I, Perotti M et al (2014) New case of trichorinophalangeal syndrome-like phenotype with a de novo t(2;8)(p16.1;q23.3) translocation which does not disrupt the TRPS1 gene. BMC Med Genet 15:52
Velagaleti GV, Bien-Willner GA, Northup JK et al (2005) Position effects due to chromosome breakpoints that map approximately 900 Kb upstream and approximately 1.3 Mb downstream of SOX9 in two patients with campomelic dysplasia. Am J Hum Genet 76:652–662
Moysés-Oliveira M, Guilherme R dos S, Dantas AG et al (2015) Genetic mechanisms leading to primary amenorrhea in balanced X-autosome translocations. Fertil Steril 103:1289–1296
Gribble SM, Prigmore E, Burford DC et al (2005) The complex nature of constitutional de novo apparently balanced translocations in patients presenting with abnormal phenotypes. J Med Genet 42:8–16
Bugge M, Bruun-Petersen G, Brøndum-Nielsen K et al (2000) Disease associated balanced chromosome rearrangements: a resource for large scale genotype-phenotype delineation in man. J Med Genet 37:858–865
Koenig M, Hoffman EP, Bertelson CJ et al (1987) Complete cloning of the Duchenne muscular dystrophy (DMD) cDNA and preliminary genomic organization of the DMD gene in normal and affected individuals. Cell 50:509–517
Alkuraya FS, Saadi I, Lund JJ et al (2006) SUMO1 haploinsufficiency leads to cleft lip and palate. Science 313:1751
Gu W, Zhang F, Lupski JR (2008) Mechanisms for human genomic rearrangements. Pathogenetics 1:4
Le Scouarnec S, Gribble SM (2012) Characterising chromosome rearrangements: recent technical advances in molecular cytogenetics. Heredity 108:75–85
Talkowski ME, Ernst C, Heilbut A et al (2011) Next-generation sequencing strategies enable routine detection of balanced chromosome rearrangements for clinical diagnostics and genetic research. Am J Hum Genet 88:469–481
Gribble SM, Kalaitzopoulos D, Burford DC et al (2007) Ultra-high resolution array painting facilitates breakpoint sequencing. J Med Genet 44:51–58
Fiegler H, Gribble SM, Burford DC et al (2003) Array painting: a method for the rapid analysis of aberrant chromosomes using DNA microarrays. J Med Genet 40:664–670
Carter NP, Ferguson-Smith MA, Perryman MT et al (1992) Reverse chromosome painting: a method for the rapid analysis of aberrant chromosomes in clinical cytogenetics. J Med Genet 29:299–307
Backx L, Van Esch H, Melotte C et al (2007) Array painting using microdissected chromosomes to map chromosomal breakpoints. Cytogenet Genome Res 116:158–166
Obenauf AC, Schwarzbraun T, Auer M et al (2010) Mapping of balanced chromosome translocation breakpoints to the basepair level from microdissected chromosomes. J Cell Mol Med 14:2078–2084
Weise A, Timmermann B, Grabherr M et al (2010) High-throughput sequencing of microdissected chromosomal regions. Eur J Hum Genet 18:457–462
Sobreira NL, Gnanakkan V, Walsh M et al (2011) Characterization of complex chromosomal rearrangements by targeted capture and next-generation sequencing. Genome Res 21:1720–1727
Shluth-Bolard C, Labalme A, Cordier MP et al (2013) Breakpoint mapping by next generation sequencing reveals causative gene disruption in patients carrying apparently balanced chromosome rearrangements with intellectual deficiency and/or congenital malformations. J Med Genet 50:144–150
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer-Verlag Berlin Heidelberg
About this protocol
Cite this protocol
Melaragno, M.I., Moysés-Oliveira, M. (2017). Breakpoint Mapping of Balanced Chromosomal Rearrangements Using Array CGH of Microdissection-Derived FISH Probes. In: Liehr, T. (eds) Fluorescence In Situ Hybridization (FISH). Springer Protocols Handbooks. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-662-52959-1_56
Download citation
DOI: https://doi.org/10.1007/978-3-662-52959-1_56
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-662-52957-7
Online ISBN: 978-3-662-52959-1
eBook Packages: Springer Protocols