

Volume 22, issue 1, December 2021
351 articles in this issue
-

-
Author Correction: The SEQC2 epigenomics quality control (EpiQC) study
Authors (first, second and last of 57)
- Jonathan Foox
- Jessica Nordlund
- Christopher E. Mason
- Content type: Author Correction
- Open Access
- Published: 23 December 2021
- Article: 350
-
An accessible, efficient and global approach for the large-scale sequencing of bacterial genomes
Authors (first, second and last of 19)
- Blanca M. Perez-Sepulveda
- Darren Heavens
- The 10KSG consortium
- Content type: Method
- Open Access
- Published: 21 December 2021
- Article: 349
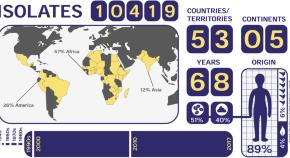
-
Quantitative-enhancer-FACS-seq (QeFS) reveals epistatic interactions among motifs within transcriptional enhancers in developing Drosophila tissue
Authors (first, second and last of 6)
- Colin T. Waters
- Stephen S. Gisselbrecht
- Martha L. Bulyk
- Content type: Method
- Open Access
- Published: 20 December 2021
- Article: 348
This is part of 1 collection:
-
Hidden biases in germline structural variant detection
Authors (first, second and last of 11)
- Michael M. Khayat
- Sayed Mohammad Ebrahim Sahraeian
- Fritz J. Sedlazeck
- Content type: Research
- Open Access
- Published: 20 December 2021
- Article: 347
This is part of 1 collection: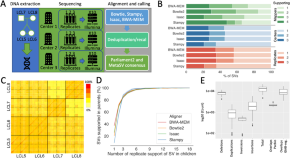
-
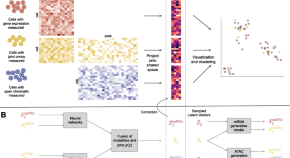
MultiMAP: dimensionality reduction and integration of multimodal data
Authors (first, second and last of 13)
- Mika Sarkin Jain
- Krzysztof Polanski
- Sarah A. Teichmann
- Content type: Method
- Open Access
- Published: 20 December 2021
- Article: 346
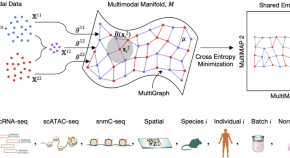
-
The economics of organellar gene loss and endosymbiotic gene transfer
Authors
- Steven Kelly
- Content type: Research
- Open Access
- Published: 20 December 2021
- Article: 345
-
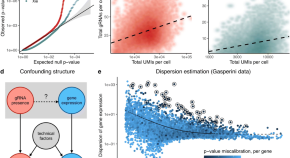
SCEPTRE improves calibration and sensitivity in single-cell CRISPR screen analysis
Authors (first, second and last of 5)
- Timothy Barry
- Xuran Wang
- Eugene Katsevich
- Content type: Method
- Open Access
- Published: 20 December 2021
- Article: 344
-
Chronos: a cell population dynamics model of CRISPR experiments that improves inference of gene fitness effects
Authors (first, second and last of 8)
- Joshua M. Dempster
- Isabella Boyle
- James M. McFarland
- Content type: Method
- Open Access
- Published: 20 December 2021
- Article: 343

-
A benchmark of structural variation detection by long reads through a realistic simulated model
Authors (first, second and last of 4)
- Nicolas Dierckxsens
- Tong Li
- Zhi Xie
- Content type: Method
- Open Access
- Published: 15 December 2021
- Article: 342
This is part of 1 collection: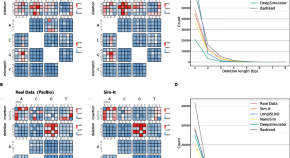
-
splatPop: simulating population scale single-cell RNA sequencing data
Authors (first, second and last of 4)
- Christina B. Azodi
- Luke Zappia
- Davis J. McCarthy
- Content type: Method
- Open Access
- Published: 15 December 2021
- Article: 341

-
Reprogramming of RNA silencing triggered by cucumber mosaic virus infection in Arabidopsis
Authors
- Maria Luz Annacondia
- German Martinez
- Content type: Research
- Open Access
- Published: 15 December 2021
- Article: 340
This is part of 1 collection: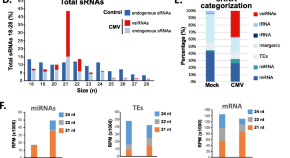
-
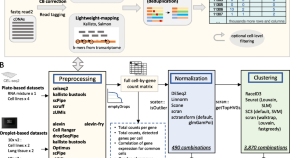
Benchmarking UMI-based single-cell RNA-seq preprocessing workflows
Authors (first, second and last of 7)
- Yue You
- Luyi Tian
- Matthew E. Ritchie
- Content type: Research
- Open Access
- Published: 14 December 2021
- Article: 339
-
Flimma: a federated and privacy-aware tool for differential gene expression analysis
Authors (first, second and last of 15)
- Olga Zolotareva
- Reza Nasirigerdeh
- Jan Baumbach
- Content type: Method
- Open Access
- Published: 14 December 2021
- Article: 338

-
CIDER: an interpretable meta-clustering framework for single-cell RNA-seq data integration and evaluation
Authors
- Zhiyuan Hu
- Ahmed A. Ahmed
- Christopher Yau
- Content type: Method
- Open Access
- Published: 13 December 2021
- Article: 337

-
Microbial co-occurrence complicates associations of gut microbiome with US immigration, dietary intake and obesity
Authors (first, second and last of 17)
- Zheng Wang
- Mykhaylo Usyk
- Qibin Qi
- Content type: Research
- Open Access
- Published: 10 December 2021
- Article: 336

-
A cis-regulatory-directed pipeline for the identification of genes involved in cardiac development and disease
Authors (first, second and last of 15)
- Hieu T. Nim
- Louis Dang
- Mirana Ramialison
- Content type: Research
- Open Access
- Published: 15 December 2021
- Article: 335
This is part of 1 collection: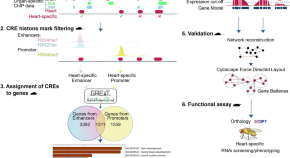
-
Programmed genomic instability regulates neural transdifferentiation of human brain microvascular pericytes
Authors (first, second and last of 5)
- Saba Rezaei-Lotfi
- Filip Vujovic
- Ramin M. Farahani
- Content type: Research
- Open Access
- Published: 09 December 2021
- Article: 334

-
geneBasis: an iterative approach for unsupervised selection of targeted gene panels from scRNA-seq
Authors (first, second and last of 12)
- Alsu Missarova
- Jaison Jain
- John C. Marioni
- Content type: Method
- Open Access
- Published: 06 December 2021
- Article: 333

-
The SEQC2 epigenomics quality control (EpiQC) study
Authors (first, second and last of 57)
- Jonathan Foox
- Jessica Nordlund
- Christopher E. Mason
- Content type: Research
- Open Access
- Published: 06 December 2021
- Article: 332
This is part of 1 collection: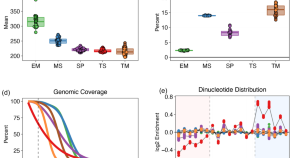
-
Single-cell characterization of CRISPR-modified transcript isoforms with nanopore sequencing
Authors (first, second and last of 5)
- Heon Seok Kim
- Susan M. Grimes
- Hanlee P. Ji
- Content type: Method
- Open Access
- Published: 06 December 2021
- Article: 331

-
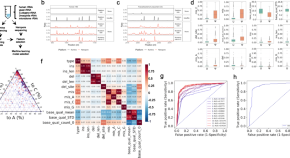
Interferon inducible pseudouridine modification in human mRNA by quantitative nanopore profiling
Authors (first, second and last of 9)
- Sihao Huang
- Wen Zhang
- Tao Pan
- Content type: Short Report
- Open Access
- Published: 06 December 2021
- Article: 330
-
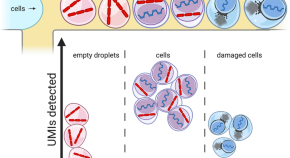
DropletQC: improved identification of empty droplets and damaged cells in single-cell RNA-seq data
Authors
- Walter Muskovic
- Joseph E. Powell
- Content type: Short Report
- Open Access
- Published: 02 December 2021
- Article: 329
-
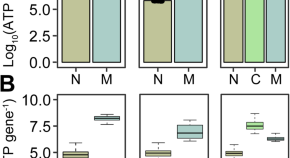
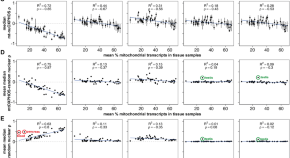
Malignancy and NF-κB signalling strengthen coordination between expression of mitochondrial and nuclear-encoded oxidative phosphorylation genes
Authors
- Marcos Francisco Perez
- Peter Sarkies
- Content type: Research
- Open Access
- Published: 02 December 2021
- Article: 328

-
Functional enrichment of alternative splicing events with NEASE reveals insights into tissue identity and diseases
Authors (first, second and last of 11)
- Zakaria Louadi
- Maria L. Elkjaer
- Olga Tsoy
- Content type: Method
- Open Access
- Published: 02 December 2021
- Article: 327

-
Wheat in vivo RNA structure landscape reveals a prevalent role of RNA structure in modulating translational subgenome expression asymmetry
Authors (first, second and last of 14)
- Xiaofei Yang
- Haopeng Yu
- Huakun Zhang
- Content type: Research
- Open Access
- Published: 30 November 2021
- Article: 326
This is part of 1 collection: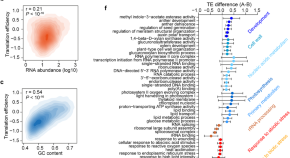
-
Heterochromatin: did H3K9 methylation evolve to tame transposons?
Authors
- Manisha Kabi
- Guillaume J. Filion
- Content type: Editorial
- Open Access
- Published: 03 December 2021
- Article: 325
This is part of 1 collection: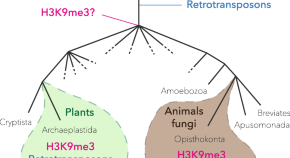
-
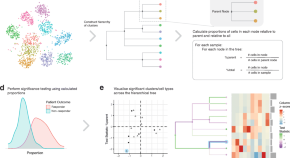
treekoR: identifying cellular-to-phenotype associations by elucidating hierarchical relationships in high-dimensional cytometry data
Authors (first, second and last of 5)
- Adam Chan
- Wei Jiang
- Ellis Patrick
- Content type: Method
- Open Access
- Published: 29 November 2021
- Article: 324
-
recount3: summaries and queries for large-scale RNA-seq expression and splicing
Authors (first, second and last of 15)
- Christopher Wilks
- Shijie C. Zheng
- Ben Langmead
- Content type: Database
- Open Access
- Published: 29 November 2021
- Article: 323

-
Landscape of transcription termination in Arabidopsis revealed by single-molecule nascent RNA sequencing
Authors (first, second and last of 13)
- Weipeng Mo
- Bo Liu
- Jixian Zhai
- Content type: Research
- Open Access
- Published: 25 November 2021
- Article: 322
This is part of 1 collection: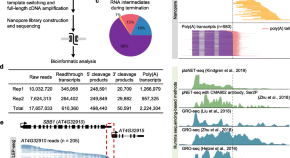
-
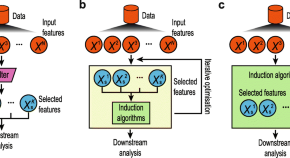
Feature selection revisited in the single-cell era
Authors
- Pengyi Yang
- Hao Huang
- Chunlei Liu
- Content type: Review
- Open Access
- Published: 01 December 2021
- Article: 321
-
Roadblock: improved annotations do not necessarily translate into new functional insights
Authors (first, second and last of 4)
- Nicola A. L. Hall
- Becky C. Carlyle
- Elizabeth M. Tunbridge
- Content type: Editorial
- Open Access
- Published: 22 November 2021
- Article: 320
-
Transcriptional landscape of highly lignified poplar stems at single-cell resolution
Authors (first, second and last of 12)
- Yang Chen
- Shaofei Tong
- Tao Ma
- Content type: Research
- Open Access
- Published: 22 November 2021
- Article: 319
This is part of 1 collection:
-
Promotech: a general tool for bacterial promoter recognition
Authors
- Ruben Chevez-Guardado
- Lourdes Peña-Castillo
- Content type: Method
- Open Access
- Published: 17 November 2021
- Article: 318
This is part of 1 collection: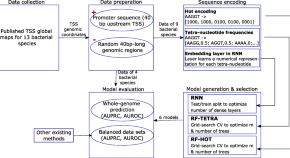
-
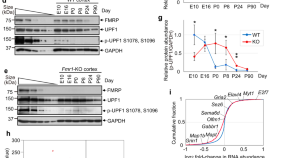
NMD abnormalities during brain development in the Fmr1-knockout mouse model of fragile X syndrome
Authors (first, second and last of 6)
- Tatsuaki Kurosaki
- Hitomi Sakano
- Lynne E. Maquat
- Content type: Short Report
- Open Access
- Published: 16 November 2021
- Article: 317
-
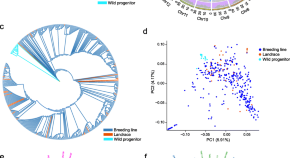
Resequencing of 388 cassava accessions identifies valuable loci and selection for variation in heterozygosity
Authors (first, second and last of 34)
- Wei Hu
- Changmian Ji
- Kaimian Li
- Content type: Research
- Open Access
- Published: 16 November 2021
- Article: 316
-
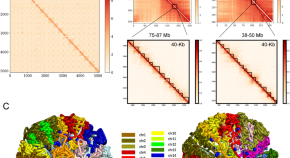
Chromatin spatial organization of wild type and mutant peanuts reveals high-resolution genomic architecture and interaction alterations
Authors (first, second and last of 11)
- Xingguo Zhang
- Manish K. Pandey
- Dongmei Yin
- Content type: Research
- Open Access
- Published: 16 November 2021
- Article: 315
-
Correction: Surfaceome CRISPR screen identifies OLFML3 as a rhinovirus-inducible IFN antagonist
Authors (first, second and last of 10)
- Hong Mei
- Zhao Zha
- Jia Liu
- Content type: Author Correction
- Open Access
- Published: 15 November 2021
- Article: 314
-
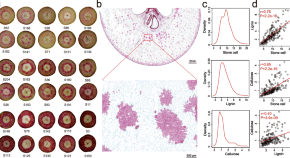
A systems genetics approach reveals PbrNSC as a regulator of lignin and cellulose biosynthesis in stone cells of pear fruit
Authors (first, second and last of 19)
- Runze Wang
- Yongsong Xue
- Jun Wu
- Content type: Research
- Open Access
- Published: 14 November 2021
- Article: 313
-
Accurate long-read de novo assembly evaluation with Inspector
Authors (first, second and last of 5)
- Yu Chen
- Yixin Zhang
- Zechen Chong
- Content type: Method
- Open Access
- Published: 14 November 2021
- Article: 312
This is part of 1 collection:
-
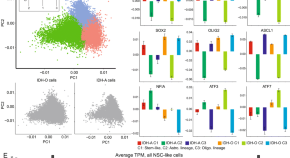
ATRX regulates glial identity and the tumor microenvironment in IDH-mutant glioma
Authors (first, second and last of 9)
- Husam Babikir
- Lin Wang
- Aaron A. Diaz
- Content type: Research
- Open Access
- Published: 11 November 2021
- Article: 311
-
Comprehensive characterization of single-cell full-length isoforms in human and mouse with long-read sequencing
Authors (first, second and last of 31)
- Luyi Tian
- Jafar S. Jabbari
- Matthew E. Ritchie
- Content type: Method
- Open Access
- Published: 11 November 2021
- Article: 310
This is part of 1 collection: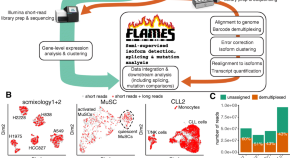
-
TAD-like single-cell domain structures exist on both active and inactive X chromosomes and persist under epigenetic perturbations
Authors (first, second and last of 4)
- Yubao Cheng
- Miao Liu
- Siyuan Wang
- Content type: Research
- Open Access
- Published: 08 November 2021
- Article: 309

-
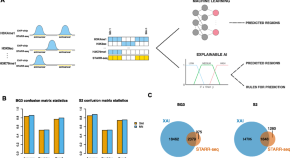
An explainable artificial intelligence approach for decoding the enhancer histone modifications code and identification of novel enhancers in Drosophila
Authors (first, second and last of 4)
- Jareth C. Wolfe
- Liudmila A. Mikheeva
- Nicolae Radu Zabet
- Content type: Research
- Open Access
- Published: 08 November 2021
- Article: 308
-
IRFinder-S: a comprehensive suite to discover and explore intron retention
Authors (first, second and last of 6)
- Claudio Lorenzi
- Sylvain Barriere
- William Ritchie
- Content type: Software
- Open Access
- Published: 08 November 2021
- Article: 307

-
The Sequencing Quality Control 2 study: establishing community standards for sequencing in precision medicine
Authors (first, second and last of 5)
- Tim R. Mercer
- Joshua Xu
- on behalf of the MAQC/SEQC2 Consortium
- Content type: Editorial
- Open Access
- Published: 08 November 2021
- Article: 306
This is part of 1 collection: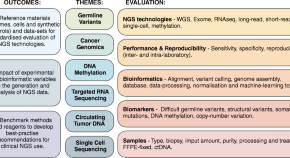
-
Lamin C is required to establish genome organization after mitosis
Authors (first, second and last of 6)
- Xianrong Wong
- Victoria E. Hoskins
- Karen L. Reddy
- Content type: Research
- Open Access
- Published: 15 November 2021
- Article: 305

-
High-quality reference genome sequences of two coconut cultivars provide insights into evolution of monocot chromosomes and differentiation of fiber content and plant height
Authors (first, second and last of 24)
- Shouchuang Wang
- Yong Xiao
- Jie Luo
- Content type: Research
- Open Access
- Published: 04 November 2021
- Article: 304

-
Sex without crossing over in the yeast Saccharomycodes ludwigii
Authors (first, second and last of 9)
- Ioannis A. Papaioannou
- Fabien Dutreux
- Michael Knop
- Content type: Research
- Open Access
- Published: 03 November 2021
- Article: 303

-
Enhanced chromatin accessibility contributes to X chromosome dosage compensation in mammals
Authors (first, second and last of 14)
- Irene Talon
- Adrian Janiszewski
- Vincent Pasque
- Content type: Research
- Open Access
- Published: 01 November 2021
- Article: 302


