Abstract
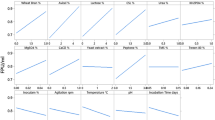
The ever increasing energy demands of modern civilization and rapidly dwindling fossil fuels point towards a renewable substitute like biofuels. However, higher costs associated with biofuel productions is the major bottleneck for its commercialization. The present study demonstrates the use of a statistical approach called response surface methodology (RSM) to investigate the optimum parameters for maximum production of cellulase by Bacillus tequilensis G9. The Plackett–Burman design (PB) of the RSM analysis indicated grass straw (GS) concentration, pH, FeSO4, inoculum, MgSO4, incubation period and NH4Cl as significant variables that influence the cellulase production. Further, to propose the best medium for the maximum production of cellulase by B. tequilensis G9, the most influential parameters, namely concentrations of GS as substrate, FeSO4, pH, inoculum size, etc. were fine-tuned by central composite design (CCD) involving four factors and five levels. The CCD analysis demonstrated 8% substrate concentration, 1.5% of inoculum along with 10 ppm FeSO4 and a pH of 5.5 in media as optimum conditions for highest enzyme production. The field emission scanning electron microscopic analysis of the treated GS showed structural alterations depicting significant deconstruction caused by B. tequilensis G9. The yield of the partially purified cellulase proteins were found to be 21% revealing molecular mass between 30 and 97 kDa. The enhanced cellulase production by B. tequilensis G9 demonstrated in our study brands its applications in many industrial processes like biorefinery, biofuels, etc.




Similar content being viewed by others
References
Shajahan S, Moorthy IG, Sivakumar N, et al. Statistical modeling and optimization of cellulase production by Bacillus licheniformis NCIM 5556 isolated from the hot spring, Maharashtra India. J King Saud Univ Sci. 2017;29:302–10. https://doi.org/10.1016/j.jksus.2016.08.001.
Lynd LR, Laser MS, Bransby D, et al. How biotech can transform biofuels. Nat Biotechnol. 2008;26:169–72. https://doi.org/10.1038/nbt0208-169.
Sukumaran RK, Singhania RR, Pandey A. Microbial cellulases: production, applications and challenges. J Sci Ind Res. 2005;64:832–44.
Bhat MK. Cellulases and related enzymes in biotechnology. Biotechnol Adv. 2000;18:355–83. https://doi.org/10.1016/S0734-9750(00)00041-0.
Binod P, Palkhiwala P, Gaikaiwari R, et al. Industrial enzymes: present status and future perspectives for India: present scenario and perspectives. J Sci Ind Res. 2013;72: 271–86. http://hdl.handle.net/123456789/17451.
Kuhad RC, Gupta R, Singh A. Microbial cellulases and their industrial applications. Enz Res. 2011;2011:1–10. http://dx.doi.org/10.4061/2011/280696.
Immanuel G, Dhanusa R, Prema P, et al. Effect of different growth parameters on endoglucanase enzyme activity by bacteria isolated from coir retting effluents of estuarine environment. Int J Environ Sci Technol. 2006;3:25–34. https://doi.org/10.1007/BF03325904.
Galbe M, Sassner P, Wingren A, et al. Process engineering economics of bioethanol production. Adv Biochem Eng/Biotechnol. 2007;108:303–27. https://doi.org/10.1007/10_2007_063.
Gigras P, Sahai V, Gupta R. Statistical media optimizationand production of ITS α-amylase from Aspergillusoryzae in a bioreactor. Curr Microbiol. 2002;45:203–8. https://doi.org/10.1007/s00284-001-0107-4.
Mukhopadhyay A, Redding AM, Rutherford BJ, et al. Importance of systems biology in engineering microbes for biofuel production. Curr Opin Biotechnol. 2008;19:228–34. https://doi.org/10.1016/j.copbio.2008.05.003.
Moorthy IMG, Baskar R. Statistical modeling and optimization of alkaline protease production from a newly isolated alkalophilic Bacillus species BGS using response surface methodology and genetic algorithm. Prep Biochem Biotechnol. 2013;43:293–314. https://doi.org/10.1080/10826068.2012.719850.
Brijwani K, Obero HS, Vadlani PV. Production of cellulolytic enzyme system in mixed culture solid state fermentation of soya bean hulls supplemented with wheat bran. Process Biochem. 2010;45:120–8. https://doi.org/10.1016/j.procbio.2009.08.015.
Thakkar A, Saraf M. Application of statistically based experimental designs to optimize cellulase production and identification of gene. Nat Prod Bioprospect. 2014;4:341–51. https://doi.org/10.1007/s13659-014-0046-y.
Ferreira S, Duarte AP, Ribeiro MHL, et al. Response surface optimization of enzymatic hydrolysis of Cistusladanifer and Cytisusstriatus for bioethanol production. Biochem Eng J. 2009;45:192–200. https://doi.org/10.1016/j.bej.2009.03.012.
Zambare V. Optimization of amylase production from Bacillus sp. using statistics based experimental design. Emir J Food Agric. 2011;23:37–47. https://doi.org/10.9755/ejfa.v23i1.5311.
Dar MA, Pawar KD, Pandit RS. Prospecting the gut fluid of giant African land snail Achatina fulica for cellulose degrading bacteria. Int Biodeterior Biodegrad. 2018;126:103–11. https://doi.org/10.1016/j.ibiod.2017.10.006.
Dar MA, Pawar KD, Jadhav JP, et al. Isolation of cellulolytic bacteria from the gastro-intestinal tract of Achatina fulica (Gastropoda: Pulmonata) and their evaluation for cellulose biodegradation. Int Biodeterior Biodegrad. 2015;98:73–80. https://doi.org/10.1016/j.ibiod.2014.11.016.
Plackett RL, Burman JP. The design of optimum multifactorial experiments. Biometrika. 1946;33:305–25.
Miller GL. Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem. 1959;31:426–8. https://doi.org/10.1021/ac60147a030.
Dar MA, Shaikh AA, Pawar KD, et al. Exploring the gut of Helicoverpa armigera for cellulose degrading bacteria and evaluation of a potential strain for lignocellulosic biomass deconstruction. Process Biochem. 2018;73:12–153. https://doi.org/10.1016/j.procbio.2018.08.001.
Laemmli UK. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970;227:680–5. https://doi.org/10.1038/227680a0.
Liang YL, Zhang Z, Wu M, et al. Isolation screening and identification of cellulolytic bacteria from natural reserves in the subtropical region of China and optimization of cellulase production by Paenibacillus terrae ME27-1. Biomed Res Int. 2014;512497:13. https://doi.org/10.1155/2014/512497.
Rastogi G, Bhalla A, Adhikari A, et al. Characterization of thermostable cellulases produced by Bacillus and Geobacillusstrains. Biores Technol. 2010;101:8798–806. https://doi.org/10.1016/j.biortech.2010.06.001.
Otajevwo FD, Aluyi HAS. Cultural conditions necessary for optimal cellulase yield by cellulolytic bacterial organisms as they relate to residual sugars released in broth medium. Mod Appl Sci. 2011;5:141–51. https://doi.org/10.5539/mas.v5n3p141.
Shankar T, Isaiarasu L. Statistical optimization for cellulase production by Bacillus pumilus EWBCM1 using response surface methodology. Global J Biotechnol Biochem. 2012;7:1–6. https://doi.org/10.5829/idosi.gjbb.2012.7.1.626.
Lee YJ, Kim HJ, Gao W, et al. Statistical optimization for production of carboxymethyl cellulase of Bacillus amyloliquefaciens DL-3 by a recombinant Escherichia coli JM109/DL-3 from rice bran using response surface method. Biotechnol Bioprocess Eng. 2012;17:227–35.
Irfan M, Safdar A, Syed Q, et al. Isolation and screening of cellulolytic bacteria from soil and optimization of cellulase production and activity. Turk J Biochem. 2012;37:287–93. https://doi.org/10.5505/tjb.2012.09709.
Chen H, Wu M, Chen Z, et al. Enhancing production of a 24-membered ring macrolide compound by a marine bacterium using response surface methodology. J Zhejiang univ sci B. 2013;14:346–54. https://doi.org/10.1631/jzus.B1200153.
Rajeswari A, Jose PA, Amiya R, et al. Characterization of saltern based Streptomyces sp. and statistical media optimization for its improved antibacterial activity. Front Microbiol. 2015;5:753. https://doi.org/10.3389/fmicb.2014.00753.
Wang YX, Lu ZX. Optimization of processing parameters for the mycelial growth and extracellular polysaccharide production by Boletus spp. ACCC 50328. Process Biochem. 2005;40:1043–51. https://doi.org/10.1016/j.procbio.2004.03.012.
Yang W, Meng F, Peng J, et al. Isolation and identification of a cellulolytic bacterium from the Tibetan pig’s intestine and investigation of its cellulase production. Electron J Biotechnol. 2014;17:262–7. https://doi.org/10.1016/j.ejbt.2014.08.002.
Singh S, Moholkar VS, Goyal A. Optimization of carboxymethyl cellulase production from Bacillus amyloliquefaciens SS35. 3 Biotech. 2014;4:411–24. https://doi.org/10.1007/s13205-013-0169-6.
Singh D, Kaur G. Optimization of different process variables for the production of an indolizidine alkaloid, swainsonine from Metarhiziumanisopliae. J Basic Microbiol. 2012;52:590–7. https://doi.org/10.1002/jobm.201100255.
Jose PA, Jebakumar SRD. Successive nonstatistical and statistical approaches for the improved antibiotic activity of rare actinomycetes Nonomuraea sp. JAJ18. Biomed Research. International. 2014;2014:1–11. https://doi.org/10.1155/2014/906097
Wang Q, Ma H, Xu W, et al. Ethanol production from kitchen garbage using response surface methodology. Biochem Eng J. 2008;39:604–10. https://doi.org/10.1016/j.bej.2007.12.018.
Muralidhar RV, Chirumamila RR, Marchant R, et al. A response surface approach for the comparison of lipase production by Candida cylindracea using two different carbon sources. Biochem Eng J. 2001;9:17–23. https://doi.org/10.1016/S1369-703X(01)00117-6.
Patil SA, Surwase SN, Jadhav SB, et al. Optimization of medium using response surface methodology for L-DOPA production by Pseudomonas sp. SSA Biochem Eng J. 2013;74:36–45. https://doi.org/10.1016/j.bej.2013.02.021.
Li M, Wang B, Zhang M, et al. Symbiotic gut microbes modulate human metabolic phenotypes. Proc Natl Acad Sci USA. 2008;105:2117–22. https://doi.org/10.1073/pnas.0712038105.
Geetha K, Gunasekaran P. Optimization of nutrient medium containing agricultural waste for xylanase production by Bacillus pumilus B20. Biotechnol Bioprocess Eng. 2010;15:882–9. https://doi.org/10.1007/s12257-009-3094-0.
Deka D, Bhargavi P, Sharma A, et al. Enhancement of cellulase activity from a new strain of Bacillus subtilis by medium optimization and analysis with various cellulosic substrates. Enzyme Research. 2011;2011:1–8. https://doi.org/10.4061/2011/151656
Kim HJ, Lee YJ, Gao W, et al. Statistical optimization of fermentation conditions and comparison of their influences on production of cellulases by a psychrophilic marine bacterium Psychrobacteraquimaris LBH-10 using orthogonal array method. Biotechnol Bioprocess Eng. 2011;16:542–8. https://doi.org/10.1007/s12257-010-0457-5.
Santhi VS, Bhagat AK, Saranya S, et al. Seaweed (Eucheumacottonii) associated microorganisms, a versatile enzyme source for the lignocellulosic biomass processing. Int Biodeterior Biodegrad. 2014;96:144–51. https://doi.org/10.1016/j.ibiod.2014.08.007.
Deka D, Das SP, Sahoo N, et al. Enhanced cellulase production from Bacillus subtilis by optimizing physical parameters for bioethanol production. ISRN. Biotechnology. 2013;2013:1–11. https://doi.org/10.5402/2013/965310.
Premalatha N, Gopal NO, Jose PA, et al. Optimization of cellulase production by Ehydrobacter sp ACCA2 and its application in biomass saccharification. Front Microbiol. 2015;6:1046. https://doi.org/10.3389/fmicb.2015.01046.
Sadhu S, Maiti TK. Cellulase production by bacteria: a review. B Microbiol Res J. 2013;3:235–58. https://doi.org/10.9734/BMRJ/2013/2367.
Kamble R, Jadhav AR. Xylanase production under solid state and submerged fermentation conditions by bacterial strains. Afr J Microbiol Res. 2010;6:4292–7. https://doi.org/10.5897/AJMR11.823.
Schwarz WH. The cellulosome and cellulose degradation by anaerobic bacteria. Appl Microbiol Biotechnol. 2001;56:634–49. https://doi.org/10.1007/s002530100710.
Maki ML, Idrees A, Leung KT, et al. Newly isolated and characterized bacteria with great application potential for decomposition of lignocellulosic biomass. J Mol Microbiol Biotechnol. 2012;22:156–66. https://doi.org/10.1159/000341107.
Barton-Pudlik J, Czaja K, Grzymek M, et al. Evaluation of wood-polyethylene composites biodegradability caused by filamentous fungi. Int Biodeterior Biodegrad. 2017;118:10–8. https://doi.org/10.1016/j.ibiod.2017.01.014.
Xu K, Feng J, Zhong T, et al. Effects of volatile chemical components of wood species on mould growth susceptibility and termite attack resistance of wood plastic composites. Int Biodeterior Biodegrad. 2015;100:106–15. https://doi.org/10.1016/j.ibiod.2015.02.002.
Schröpfer SB, Bottene MK, Bianchin L, et al. Biodegradation evaluation of bacterial cellulose, vegetable cellulose and poly (3-hydroxybutyrate) in soil. Polmeros. 2015;25:154–60. https://doi.org/10.1590/0104-1428.1712.
Jiménez DJ, de Lima Brossi MJ, Schückel J, et al. Characterization of three plant biomass-degrading microbial consortia by metagenomics- and metasecretomics-based approaches. Appl Microbiol Biotechnol. 2016;100:10463–77. https://doi.org/10.1007/s00253-016-7713-3.
Lu WJ, Wang HT, Yang SJ, et al. Isolation and characterization of mesophilic cellulose-degrading bacteria from flower stalks-vegetable waste co-composting system. J Gen Appl Microbiol. 2005;51:353–60. https://doi.org/10.2323/jgam.51.353.
Kim BK, Lee BH, Lee YJ, et al. Purification and characterization of carboxymethyl cellulase isolated from a marine bacterium, Bacillus subtilis sub sp. subtilis A-53. Enzyme Microb Technol. 2009;44:411–6. https://doi.org/10.1016/j.enzmictec.2009.02.005.
Mawadza C, Hatti-Kaul R, Zvauya R, et al. Purification and characterization of cellulases produced by two Bacillus strains. J Biotechnol. 2000;83:177–87. https://doi.org/10.1016/S0168-1656(00)00305-9.
Saha BC. Production, purification and properties of endoglucanase from newly isolated strain of Mucorcircinelloides. Process Biochem. 2004;39:1871–6. https://doi.org/10.1016/j.procbio.2003.09.013.
Lucas R, Robles A, Garcia MT, et al. Production, purification and properties of an endoglucanase produced by the hyphomyceteChalara (syn.Thielaviopsis) paradoxa CH32. J Agric Food Chem. 2001;49:79–85. https://doi.org/10.1021/jf000916p.
Robson LM, Chambliss GH. Endo-1, 4-β-glucanase gene of Bacillus subtilis DLG. J Bacteriol. 1987;169:2017–25. https://doi.org/10.1128/jb.169.5.2017-2025.1987.
Eppinger M, Bunk B, Johns MA, et al. Genome sequences of the biotechnologically important Bacillus megaterium strains QM B1551 and DSM319. J Bacteriol. 2011;193:4199–213. https://doi.org/10.1128/JB.00449-11.
Kunst F, Ogasawara N, Moszer I, et al. The complete genome sequence of the Gram-positive bacterium Bacillus subtilis. Nature. 1997;390:249–56. https://doi.org/10.1038/36786.
Acknowledgements
MD is indebted to University Grants Commission (UGC), New Delhi, India, for providing senior research fellowship. Partial funding for this work was received from University Grants Commission, New Delhi, India under Start-Up scheme (F.30-121/2015BSR). RSP acknowledges theUPE-II (nanobiotechnology), UoP-BCUD grant (15-SCI-001422), as well as DRDP and DST-PURSE schemes provided to the department. The authors thank Mr. Harishchandra Nikule (Central Instrumentation facility (CIF), Savitribai Phule Pune University, Pune) for technical assistance provided during FESEM analysis. We also acknowledge Dr. Mohd. Shahnawaz for language corrections and anonymous reviewers whose critical comments significantly improved this manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest with respect to the research, authorship, and publication of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dar, M.A., Pawar, K.D., Chintalchere, J.M. et al. Statistical optimization of lignocellulosic waste containing culture medium for enhanced production of cellulase by Bacillus tequilensis G9. Waste Dispos. Sustain. Energy 1, 213–226 (2019). https://doi.org/10.1007/s42768-019-00016-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42768-019-00016-w