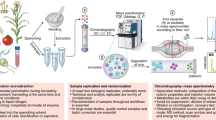
Abstract
Structural mass spectrometry (MS) is a chemical structure-revealing analytical technique that measures mass/charge ratio of ions. Biochemical analyses in the context of metabolomics—which seeks chemical descriptions of cellular, organismal, and even community biology—pose challenges distinct from past applications of MS. The scientific need for both far greater depth and breadth of metabolic analytes is manifested analytically in two general approaches: the “Biomarker” approach and the “Pathway” approach. In this chapter, the basic principles of mass spectrometry relevant to these two approaches to metabolomics are reviewed. The emphasis is on practical aspects of how experimental design interacts with instrument designs and capabilities. Both simulated theoretical and real-life analyses are used to illustrate the key concepts and their ramifications.
Similar content being viewed by others
Key words
1 Introduction to MS
1.1 What is MS? The MS Experiment
Structural mass spectrometry (MS) is a chemical structure-revealing analytical technique that measures mass/charge ratio of ions. If the ion is singly charged, then it measures the putative molecular mass. Often, the ions decompose in the MS, leading to fragmentation that is rich in chemical structure information (see Fig. 1), sometimes revealing even the subatomic composition. All of these properties are extremely useful in metabolomics, as will be seen. The use of mass spectrometers for nonstructural analyses, such as ICP-MS and isotope-ratio MS, is beyond the scope of this chapter; ICP-MS is treated in Chap. 5.
Schematic illustration of the mass spectrometry (MS) “experiment.” Ionized molecules are fragmented, and the fragment ions of a single mass/charge ratio (m/z) are passed by a “quadrupole” MS instrument; this acts as a filter and all other ions are rejected during that period. By rapidly changing the m/z that is to be passed, a spectrum of intensity vs. time is generated. The time abscissa is then calibrated to m/z to produce a mass spectrum. Here, the molecule MOLECULE is fragmented into a pattern that is representative of the original structure. Unlike most forms of spectroscopy, MS entails degradative physical–chemical reactions; experimental control over these reactions produces the desired type of data.
Most structure-revealing analytical measurements are spectroscopies that use ranges of the electromagnetic spectrum; for example, NMR is in the radio frequency (RF) range, and ESR is in the microwave region. In fact, the names of the techniques are often descriptive of their frequency range (IR, near IR, visible, UV, X-ray, etc.).
In contrast, MS is found nowhere in the electromagnetic spectrum. As illustrated in Fig. 1, it is a technique that is based on physical–chemical reactions and ionic separations, and therefore is an “experiment” in the same sense of the NMR experiment. One does not just “take a spectrum” or “analyze by MS,” but rather has to plan the MS experiment to conduct. MS is also distinguished by the fact that it necessarily destroys the sample. This in part accounts for the use of the term “spectrometry” to describe the technique, and is usually distinct from other forms of “spectroscopy.”
1.2 Why MS for Metabolomics? Perspective from Modern MS
It is vital to note that this chapter covers MS usage for “metabolomics.” Although the century-long history of MS can provide valuable lessons and insights, the marriage of MS with metabolomics is a comparatively recent event. Thus, the historical perspective is evoked only when deemed necessary. For an appreciation of historic MS development, the reader is referred to older books (1). For over half a century, MS has been a tool of choice for the study of the “metabolism” of drugs, pesticides, nutrients, pollutants, and other xenobiotics; for those perspectives, the reader is referred to the many excellent books and reviews on these topics.
Biochemical analyses in the context of metabolomics—which seeks chemical descriptions of cellular, organismal, and even community biology—pose challenges distinct from past applications of MS. First, the problem in metabolomics is to distinguish small differences in metabolite patterns in the presence of an overwhelmingly greater background of the same metabolites. Second, this feat must be accomplished on literally many orders of magnitude more analytes than in past applications of MS. Third, the concentration ranges needed to solve metabolomic problems can span 12 orders of magnitude. These analytical problems are far greater than any single analysis can address.
This scientific need for both far greater depth and breadth of metabolic analytes is manifested analytically in two general approaches: the “Biomarker” approach and the “Pathway” approach. In the former, MS is utilized for its capability of high sensitivity and selectivity, with acceptable precision. This helps to statistically distinguish concentration profiles of mixtures of analytes, which is the chemical signature that is sought. In the Pathway approach, MS is used to distinguish among isotopologues (sometimes also called “mass isotopomers”) of metabolites with high sensitivity, which yields the biochemical signatures. In both cases, the information generated by MS is highly complementary to NMR, and in effect is the only practical analytical partner to NMR for the Pathway approach. Generally, these two are the only analytical techniques that can distinguish and quantify the necessarily large number of metabolites together with their isotopomers, across the necessary span of concentrations.
The current state of MS instrument development enables interfacing to most separation techniques (see below), with high sensitivity down to the sub-pM range, structural information from pmole amounts of analyte, and rapid generation of spectral data. Moreover, the technology is experiencing extremely active developments on all fronts, generally leading to greater performance at lower costs or extended performance at the high end of MS instrumentation, and improved ease of use and reliability.
2 Information Encoded by the Mass Spectrum
2.1 The Mass Spectrum
In order to understand the uses of MS in metabolomics, it is crucial to first understand the types of information provided by MS. It is instructive to start with the traditional graphical conveyance of MS information: the mass spectrum.
Shown in Fig. 2 is a mass spectrum of the reduced form of nicotinamide adenine dinucleotide reduced form (NADH) along with the historically more prevalent representation, the mass histogram. The histogram is summary of the actual spectrum via a process called “centroiding.” As will become clear later, this distinction is important. The abscissa in Fig. 2 reflects the second central concept that an MS instrument measures mass-to-charge ratios (m/z), not mass. A third central concept is that only charged ions are measured; neutral molecules are not measured by MS instruments.
Figure 3 illustrates several aspects of a fourth concept: fragmentation. The mass histogram consists of many ions of different m/z from a single molecule, described in Fig. 3 and its legend. Fragmentation of the molecular ion conveys structural information, the pattern of which is used for identification by comparison with fragmentation patterns of authentic chemical standards or for interpretation of structure. The use of fragment information varies enormously by discipline. For example, in drug analysis, fragment patterns are the most important information for identification, since the structures are known and authentic standards are available; thus, interpretation of fragment patterns is rarely performed. In contrast, MS-based proteomics uses fragment patterns almost exclusively for sequence structure interpretation, and comparison with authentic standards is generally not done because it is often extremely impractical.
Fragmentation of mugineic acid illustrating how fragmentation (Fig. 1) conveys structural information, including isotopic composition. The black-line portion of the figure represents the mass histogram of unlabeled mugineic acid, with the corresponding fragment structures assigned above each bar. Such mass histograms are often not interpreted for fragment structures at all, but instead the pattern is used for confirming identity by comparison with authentic standards. The red- and blue-colored lines represent the mass histogram if both nitrogens were labeled with 15N, with the resulting m/z of the fragment ions listed below the abscissa. Note that some of the ions shift 1 m/z (red or blue) while others shift 2 m/z (red + blue lines denoted by purple values) because they contain one or two 15N, respectively. In this fashion, it is possible to determine if a metabolite had zero, one, or two 15N, and the relative amount of label in each position. Redrawn from 10.
In Biomarker and Pathway metabolomics approaches, both uses of fragment patterns are practiced for confirming identities of known metabolites, as well as attempting to identify unknown or unexpected metabolites. In addition, there is a third use of fragment patterns for only the Pathway approach: submolecular localization of isotopic incorporation. Figure 3 illustrates how the locations and number of 15N atoms incorporated can be discerned, which are direct pathway clues. This has been used, for example, to follow the fate of N from nitrate by rice coleoptiles (2).
The overarching importance of fragmentation led to the development of “tandem” MS or, more generally, “MS to the nth” (MSn). Figure 4 and its legend illustrate this fifth concept, whereby a series of related MS spectra are generated, roughly analogous to 1D vs 2D-NMR. The practice of MSn attains sequential control over fragmentation, allowing unambiguous assignment of fragment ions to their source molecule. For example, two phosphatidylcholine (PC) metabolites with acyl chains of C16:O and C18:3 (PC1) or C16:1 and C18:2 (PC2) have the identical molecular ion exact-mass of 756.554321 m/z (M + H), thus unresolvable in a single-spectrum MS. However, the two PC lipids have different tandem MS fragments from, e.g., loss of different acyl chains. Namely, the ions of 501.321916 m/z (loss of C16:0) and 479.337566 m/z (loss of C18:3) are indicative of PC lipid with C16:0 and C18:3 acyl chains while the mass of 503.337566 m/z (loss of C16:1) and 477.321916 m/z (loss of C18:2) can be assigned to PC lipid with C16:1 and C18:2 acyl chains. Thus, tandem MS can distinguish different structures with the same molecular ion (identical molecular formulae).
Illustration of tandem MSn. These spectra were obtained directly from a crude extract of wheat plant under continuous nanoelectrospray ion trap MS, total acquisition time was <30 s, and the precursor ion of 637.1 m/z is an unknown metabolite. MSn attains sequential control of fragmentation, thus establishing the provenance of the m/z ions labeled in the MS1 spectrum. Figure redrawn from 10.
A sixth concept is that of “exact” mass, also called accurate mass, illustrated in Fig. 5. The major utility of accurate mass is the determination of molecular formulae. In metabolomics, a given molecular formula usually corresponds to just a handful of known metabolites given the biological source; hence, an array of accurate mass information alone can enable assignment (tentative identities) to many hundreds, sometimes thousands, of peaks in a single spectrum. However, the attainment of sufficiently accurate mass data is technically demanding and not achieved by most MS instruments on the market today. One interesting fact shown in Fig. 5 is that very close (<1 m/z) peaks are not closely related structures; rather, they are substantially different in molecular formula.
Spectral simulations of protonated adducts of four diacyl glycerophospholipids (gPLs) illustrating an important use of accurate mass. In this case, accurate mass is used to distinguish, in a mixture, among these four different classes of lipids that have very similar m/z near 830.6. They differ in molecular formulae, and thus differ also in their accurate mass. The spectral range shown is 0.5 Da, and sufficient resolution was simulated to distinguish the four lipid species. The naming conventions are PS phosphatidyl serine; PC phosphatidylcholine; PE phosphatidyl ethanolamine; PCe phosphatidylcholine plasmalogen (ether linkage). The numerals denote the total [# C]; [# unsaturations] across the two acyl chains.
A seventh concept conveyed by the mass spectrum is that of resolving power or resolution. The resolution of an MS instrument is generally defined as m1/(m1 – m2), where m1 and m2 are two peaks of equal intensity with 10% overlap. A common practice is to set m1 = 400 m/z because the actual resolving power varies across the spectrum depending on the design of the instrument. By selecting a single m/z, specifications can be compared. Thus, resolving power = R(400 m/z, 10%) = 4,000 means that the MS instrument is able to discriminate unambiguously (with only 10% overlap) between two equally sized peaks at 400.0 and 400.1 m/z. It is important to note that such a specification is dependent on the m/z; thus, if R(400 m/z, 10%) = 4,000 but the analyte of interest is m1 = 800 m/z, then that same resolution specification is theoretically reduced to 2,000 for the analyte of interest. It is also important to note that a mass histogram contains no information regarding resolving power.
It turns out that “sufficient” resolution must be achieved in order to obtain useful accurate mass, as previously shown in Fig. 5 (where R = 40,000 was used) and further shown in Fig. 6 and its legend. In this illustration, the accuracy of the m/z (intensity-weighted center of an m/z peak) is affected by the “purity” of each m/z peak, which in turn is a function of the resolution. Note that insufficient resolution in this case led to both false positives and false negatives with regards to the mass spectral peaks, even in this simplest case of a pure standard. A further consequence of insufficient resolution is also seen in Fig. 6, namely, that intensities are nonrepresentative of the individual peaks, hampering quantification if these peaks had originated from different metabolites.
Negative ion mass spectra of monogalactosyl PL-A (InvivoGen) illustrating the influence of resolution on mass accuracy. Spectra were obtained using a Thermo LTQ-FT MS hybrid ion trap/FT-ICR-MS instrument. Under continuous spray, spectral acquisition was alternated between low resolution (ion trap MS) and high resolution (FT-ICR-MS) repeatedly so that the spectra shown are a true average over a 5-min acquisition span. Note that low resolution returns average values that are nonexistent m/z peaks (false positive), as shown by the red lines; here, as in other cases, the reader is reminded that averages are values that do not actually exist. At the same time, low resolution fails to reveal the presence of some peaks that actually do exist (false negative), as shown by the blue lines. Unpublished data by the author.
Thus, MS instruments sporting excellent accurate mass specifications cannot realize this advantage on real-life complex samples (or even the pure standard in Fig. 6) without sufficiently good resolving power. As is the case with accurate mass, the attainment of very high resolution is technically demanding and not achieved by most MS instruments. In fact, the need to achieve simultaneous accurate mass with high resolution only compounds the difficulty.
An eighth key concept is isotopic abundances. In Fig. 7, the theoretical natural-abundance (NA) 13C isotope peaks are shown as simulated mass spectra for increasing number of carbons per molecule. As is evident from Fig. 7, the 13C isotopic mass isomers (isotopologues) increase with the carbon-skeleton size of the molecule, which has two main advantages in metabolomics: (a) for the Biomarker approach, the relative intensities of the natural-abundance 13C isotopomer peaks can be used for structure confirmation; (b) for the Pathway approach, the isotopomer intensities that are in excess of natural abundance are useful in that they encode metabolic pathway information (3, 4).
Illustration of isotopic abundance dependence on number of carbons. The series of simulated spectra are of pure carbon compounds of the designated number of carbons. In the C1 spectrum, the 13C1 peak (m + 1 isotopologue) has the expected natural abundance (intensity) of approximately 1.1%. For the C10 spectrum, the m + 1 isotopologue has ten possible locations to populate, increasing by 10× the probability of the molecule harboring a single 13C isotope. Thus, the 12C 139 C1 peak is 1.1% × 10 = 11% the abundance. At C90, the m + 1 isotopologue is approximately the same abundance as the monoisotopic. Furthermore, the 12C 1389 C1 peak itself has a natural abundance peak of one more 13C (m + 2, or 12C 1388 C2) and so forth, forming a cluster of molecular ions. Note that at C200 the molecular ion cluster is dominated by the various 13C isotopologues.
Figure 6 illustrates “sufficient” resolution on a pure standard—so how much resolution is sufficient for real samples? Of course, this depends on the goal and particular sample, but the Pathway approach will be far more demanding than the Biomarker approach. For the latter, it is feasible to calculate the natural-abundance isotopologues for each known analyte, and subtract all the NA isotopologues using a “deisotoping” algorithm based on the known metabolite identity (5). However, deisotoping obviously cannot be done for unknowns, and also cannot be done in the Pathway approach, because the isotopic variation is the very experimental variable that is sought. Figure 8 and its legend illustrate the minimum resolution that is needed for assignments of molecular ions for the simplest case—that is, the Biomarker approach assuming NA isotopologues—for an idealized mixture of two lipid metabolites. Figure 9 and its legend illustrate the greater resolving power needed for the same two metabolites, by simply having a larger abundance of an (m + 2) isotopologue, a common occurrence in an actual experimental sample. Here, it is seen that the m + 2 isotopologue peak of one lipid overlaps considerably with the monoisotopic peak of another lipid species such that R = 200,000 must be used to distinguish the peaks. Figure 9 also shows the application to a real sample of the apparently “sufficient” resolution with accurate mass.
Illustration of “sufficient resolution” for the simplest case of two metabolites with natural-abundance isotopic distributions. Spectra were simulated at different resolutions of a mixture of PC34:3 and PC34:2, the former at tenfold the concentration of the latter. (a–c) each show mixture spectra at the top, PC34:3 alone, and PC34:2 alone, respectively, at R(400 m/z, 10%) = 1 K, 10 K, and 100 K. The insets to the right zoom in on the 0.15-Da spectral region corresponding to an overlap of PC34:3 (m + 2) isotopologue and PC34:2 monoisotopic peaks. Note that at least R = 100 K is required to begin to distinguish these two molecular species. In principle, for the Biomarker approach, it is possible to calculate the natural abundance contribution of the PC34:3 (m + 2) to “deisotope” the peak, if these two species were the only compounds in the mixture. However, in practice, it is common to have isotopologues of many molecular species crowded into even such a narrow region, making “deisotope” data analyses techniques extremely difficult.
Illustration of the greater demand for “sufficient resolution” in a 13C-labeled Pathway approach, 0.15-Da m/z window. The simulated spectral mixture from Fig. 8c is now tenfold enriched in [13C2] PC34:3 such that there is much more PC34:3 (m + 2) than PC34:2 (m + 0). As shown in (a), this simple change in only one of the isotopologues renders insufficient the R = 100 K spectrum that previously appeared to be adequate (Fig. 8c). (b) Illustrates that R = 200 K is now needed to cleanly resolve the PC34:3 (m + 2) isotopologue ion from the PC34:2 (m + 0) ion. (c) A spectrum from a methanolic extract of [U-13C] glucose-grown human lung adenocarcinoma (A549) cell culture illustrates that even the R = 200 K is barely adequate for real samples due to the presence of other metabolites, such as the much more abundant one at 758.5507 m/z. The concentrations of the gPLs were in the low picomolar range, analyzed using a continuous nanoelectrospray Fourier-transform ion-cyclotron resonance mass spectrometry (FT-ICR-MS) instrument. The assignments of corresponding peaks for PC34:3 (m + 2) and PC34:2 (m + 0) are shown by the arrows in (c) (the accurate mass is in slight error which is readily corrected by an internal mass accuracy standard). (c) Is unpublished data by the author.
The lessons from Figs. 8 and 9 point to at least three important ramifications of insufficient resolution: (a) in a 13C isotope-based study (Pathway approach), the MS would fail to distinguish among the 13C isotopologues of numerous compounds; (b) if metabolite X were known to be unlabeled (i.e., Biomarker approach) but at far higher in concentration than metabolite Y, the natural-abundance isotopologues of X will be sufficiently large to skew the quantification of the Y monoisotopic peak and its isotopologues (Fig. 9); (c) insufficient resolution would preclude the use of the extremely powerful multielement labeling approaches for Pathway metabolomics, an approach that is readily accommodated by NMR techniques. It turns out that problems (a) and (b) can be partly addressed by turning to tandem MS or coupling MS to separation instruments, which is discussed later. However, problem (c) has no such solution, and extremely high resolution with accurate mass is the only reasonable solution; this is discussed later in this chapter. Figure 10 illustrates how extremely high-resolution, accurate-mass MS can be used to distinguish among monoisotopic and element-specific (in this case, 13C) isotopologues, and more on this topic is discussed below.
Continuous nanoelectrospray FT-ICR-MS spectrum, obtained at an R(400 m/z, 10%) = 200,000, of a methanolic extract of human lung adenocarcinoma (A549) cell culture grown in [U-13C]-glucose. The top panel shows the region of 700–900 m/z which is dominated by gPLs, consisting of >1,500 distinct ion peaks, many of which is not visible at this graphical scale. The 5.0-Da window denoted by the rectangle is expanded into the middle panel, where gPLs of identical structures are indicated by the colored dots; they are isotopologues of each other. Due to the extremely high resolution, the peaks are widely separated; however, there is a tight clustering at the right. This 0.1-Da-wide region is expanded into the bottom panel that shows gPLs of three different structures that are nearly isobaric (same molecular mass). As illustrated by Fig. 9 and its legend, these would not have been distinguished at lower resolutions. The boxes below the bottom panel show the comparison of theoretical and measured m/z; recall from Fig. 6 that such small errors in mass accuracy require simultaneous extremely high resolution. These m/z errors are small enough to readily distinguish incremental addition of protons (+1.00782 m/z each) which denote different gPL structures, from among isotopologues achieved through incremental additions of neutrons in 13C (+1.00336 m/z each), deuterium (+1.00618 m/z each), 15N (+0.99703 m/z each), 18O (+1.99668 m/z each), 34S (+1.99580 m/z each), etc. Thus, the blue and yellow dot peaks in the bottom panel can be confidently assigned as 13C isotopologues (and not of other elements) of the same dot color in the middle panel, and clearly distinct from another, third monoisotopic gPL (purple dot). Unpublished data by the author.
In summary, the key concepts six through eight, respectively, accurate mass, resolution, and isotopologues, interplay to produce confounding interferences or rich information, or both, depending on the perspective and instrument capabilities. The latter aspect is now considered.
3 Types of Mass Spectrometers
As there are numerous excellent books on the subject of MS instrument types and design, only a vastly simplified overview, from the unique perspective of metabolomics, is presented here. Perhaps, the most significant classification of MS instruments in this regard is the “MS-in-space” versus the “MS-in-time” designs. Table 1 is divided along these lines, with attempts to clarify the utility of most instrument designs with regards to the Biomarker and Pathway approaches to metabolomics. For example, it indicates whether coupling to chromatography is generally necessary to achieve the tasks in the column headings; in this sense, chromatography is called upon to solve inadequate MS capability. The first six columns are the various modes of measurement supported by the instrument type, which are very useful to the Biomarker approach. The last three columns state additional requirements by the Pathway approach, indicating whether the instrument type supports sufficient resolution simultaneously with sufficient mass accuracy in order to distinguish isotopologues from nearly isobaric monoisotopic peaks, and whether isotopologues of multiple-element-labeled metabolites can be distinguished. The resolution required to achieve unambiguous distinction among combinations of 13C isotopologues in natural abundance (Biomarker) and enriched (Pathway) was illustrated earlier in Figs. 8–10.
3.1 MS-in-Space Designs
The top part of Table 1 lists the metabolomics capabilities of the MS-in-space designs. The MS-in-space designs comprise most of the instruments historically, and modern ones are at the state of the art in speed and sensitivity. They are characterized principally by reliance on spatial transport of ions to achieve fragmentation experiments and additional resolution of m/z; a considerable practical consequence of the MS-in-space designs is that additional physical sections are required in order to achieve MSn (illustrated in Fig. 4). They range from the historically common quadrupole MS instruments to high-end “hybrid” instruments combining quadrupole with time-of-flight (ToF) and other designs.
Many MS instruments in use are MS-in-space designs, such as those dedicated to GC, rapid very-high-sample throughput nucleotide sequencing, portable field instruments, etc. Often, the more affordable instruments are also of this category, which is a very important advantage, although there is considerable overlap of prices with MS-in-time designs. At the high end, the MS-in-space designs include some of the highest sensitivity “tandem” MS (MS/MS) instruments available for preselected analytes or structural classes, such as for drug analysis, their toxicodynamics, and abuse testing.
Tandem MS, or MSn, is an extremely powerful technique that greatly enhances the detection of target analytes. It is instructive to describe how the MSn concept (Fig. 4) is translated into hardware operation, which is illustrated by Fig. 11 and its legend. In the most basic use, Q1 is set to pass the target molecular ion (m/z corresponding to MOLECULE3 in the example) that is then fragmented by collision with Ar gas in the “q” section, from which the mass spectrum of the fragments is obtained by scanning Q3 (the operation is, thus, Q1fixed/q/Q3scan). An LC-triple-quad MS instrument will, thus, continuously acquire full spectra on only molecules that pass selection in Q1. This results in very good detection due to the specificity of Q1 while Q3 provides structural confirmation fragments. Figure 12 shows an operational mode for further s/n improvement, whereby Q1 and Q3 can both cycle through a list of preselected pairs of m/z (for each pair, Q1fixed/q/Q3fixed), which would continuously detect MS/MS products of multiple analytes in a technique called mutiple reaction monitoring (MRM). This second operational mode eschews confirmation spectra in order to gain the advantage of improved detection by higher throughput of only the target ion(s), since the instrument does not waste time collecting unwanted “noise” portions that can constitute the major portion of mass spectra.
Illustration of the basic mode of tandem MS for an MS-in-space instrument. The original design, schematized here, consisted of three physical quadrupole MS instrument sections, hence the traditional name of “triple quad.” See Table 1 for a listing of design variations. In comparison with Fig. 1, note that Q3 is the basic MS instrument, and the added front sections Q1 and q attain the reaction control for the tandem MS experiment. In this experiment, Q1 is set to pass a desired m/z, which is collided in q to generate fragments, which are sorted and detected in Q3. Since the operations for MSn occur in different physical sections of the MS instrument, this is called MS in space.
Illustration of a second mode of operation for tandem MS on the “triple0quad” MS-in-space instrument. Q1 and Q3 can be programmed to pass many such fixed pairs of m/z, which is called MRM as described in the text. Two such fixed pairs are shown here. Since all of the m/z values are fixed, there is no mass spectrum acquired in this mode of operation.
An interesting use of the high-end tandem MS-in-space instruments for metabolomics is substructure monitoring. The most common use of this capability is shown in Fig. 13, a simulation of an LC-“triple-quad” MS instrument. In this third mode of operation, which flips the operation from that shown in Fig. 11, Q1 is set to obtain full-scan spectra while Q3 is fixed on a given m/z (hence, Q1scan/q/Q3fixed). Operated in this manner, the LC-triple-quad MS will continuously detect a given fragment from any compound as it elutes from the LC, resulting in a chromatogram consisting of all compounds that contain that fragment (and thus presumably the desired substructure). In the example shown, if the interest was only in metabolites that contain MOLEC moieties, both known and unknown, then Q1 can be set to full scan while Q3 can be fixed on a particular fragment, in this case the m/z corresponding to the MOLEC substructure.
Illustration of a third mode of operation for tandem MS on the “triple-quad” MS-in-space instrument. In this mode, the MS instrument is used to detect any substructure of interest. Here, Q1 scans allow one m/z metabolite at a time, which undergoes the fragmentation process in q as usual; however, Q3 is set to pass m/z ions corresponding only to MOLEC. Thus, any metabolite—including unknowns—with m/z corresponding to MOLEC is detected while all others are rejected. In a sense, this is an “any-and-all” MRM mode that does not have a predefined molecular m/z list, which is useful for unknown or unexpected metabolites.
Table 1 also alludes to some of the metabolomic considerations of MS-in-space instruments. Note that most of the instrument configurations have a “0” designation, meaning that for extremely complex metabolomic samples it is possible to perform unassisted MS (e.g., without prior chromatographic separation). However, note that for the Pathway approach, which entails analysis of isotopologues, the designations mostly have a “1,” which means that some assistance such as prior chromatographic separation is needed in order to reduce overlaps in the spectra (see Fig. 9). Also note that none of these instruments are generally capable of multielement isotopologue analysis due to lack of sufficient resolution and mass accuracy.
3.2 MS-in-Time Designs
The bottom part of Table 1 lists some of the most common MS-in-time designs. MS-in-time instruments—at least ones available for purchase—appeared historically later than the MS-in-space instruments. This instrument strategy is based on first trapping ions in a confined space, and then mass spectrometry by manipulations (usually radio-frequency pulses) in the same physical device over time, rather than spatially across discrete devices. Thus, one of the reasons for the later historic appearance is a much greater need for integrated computer control, the technology for which greatly accelerated in the 1980s. An important practical consequence of the MS-in-time designs is that MSn is achieved in the same physical device, unlike MS-in-space designs. The MS-in-time instruments range from the early retail-available “ion trap” MS to high-end hybrid instruments combining ion traps or quadrupoles on the front end with ICR or even another ion trap and other MS designs at the back. In short, hybrid instruments can, in principle, utilize any combination MS-in-space and MS-in-time devices. The basic operation of an ion trap MS instrument is illustrated in Fig. 14.
Illustration of the basic mode of operation for “ion trap” MS-in-time instrument. Most MS-in-time instruments are ion traps of some sort. A major type of ion trap design features a two-dimensional cross section of four curved surfaces facing each other, schematized here as curved plates. Other types of ion traps can differ considerably in both hardware and principle of operation; see Table 1 for a listing of some of the designs. Compare to Fig. 1: ion traps first collect all of the ions with the assistance of He gas to “cool” (slow) down the ions, and then eject them by m/z over time. The time abscissa is calibrated to m/z to produce a mass spectrum.
Tandem MS, or MSn, is achieved in MS-in-time instruments in the manner shown in Fig. 15; compare this with MS2 on the MS-in-space instruments (Figs. 11–13). Achievement of multiple MS stages is, thus, a matter of repeating steps 2 and 3, for which the isolation and fragmentation steps over time (as explained in Fig. 15 legend) are readily controlled by the computer. A practical limitation of requesting many steps in MSn is that the population of available ions decreases dramatically at each MSn step; thus, analytical utilization of MS10, for example, is very rare while up to MS5 is not unusual. Another practical limitation is that many stages in MSn take considerable time, usually many milliseconds per step, thus limiting MSn stages in applications where the analytes are available to the MS instrument for only a short time, such as in LC-MS. This argument is often countered by pointing out that at least the option of many MSn stages is available in MS-in-time instruments while the option is almost always limited to just MS2 in the MS-in-space instruments.
Illustration of one round of tandem MS for the “ion trap” MS-in-time instrument. (1) The ions comprising the MS1 spectrum is stored in the ion trap, usually “cooled” by the presence of He gas; (2) the precursor ion of interest (MOLECULE3) is isolated in the trap by ejection of all other ions; (3) the precursor ion is fragmented by collision with He gas that is always present in the ion trap (does not have to be pumped in and out); (4) the fragments are scanned out of the trap as in Fig. 14. Steps 2–3 can be repeated for many cycles over time to obtain multistage MS or MSn. Hence, this type of MSn is called MS over time.
The straightforward implementation of MSn on MS-in-time instruments led to extensive use of “data-dependant” MSn (ddMSn). Figure 16 and its legend illustrate the ddMSn process, in which the computer makes on-the-fly decisions of the MSn acquisition based on predetermined user criteria. Note that ddMSn requires switching between different operational modes of an MS instrument, which can introduce significant time-consuming steps; in fact, a consequence of this time limitation is illustrated in Fig. 16, where the command stage “g” is never executed because the LC peak is no longer present. Generally, ddMS2 is performed less commonly in MS-in-space instruments. To see why, using a triple- quad MS as an example, a Q1scan first finds the target m/zs, followed by Q1fixed/q/Q3scan, and then cycling back to the Q1scan. This takes considerable time because the fragmentation is achieved in the collision cell by the physical presence of the collision gas, which must be introduced and stabilized between the two steps and then pumped out again, all of which take many seconds. Thus, a typical LC peak elution of 20 s duration misses much of data while the triple-quad MS is switching modes. In contrast, in, e.g., an ion trap, the collision gas is always present and fragmentation is induced by excitation of ions using an RF pulse. Although such switching modes by RF pulses is far faster, even the ion trap needs time to switch operational modes because ions still have a physical settling time after fragmentation, isolation, ejection, etc., all of which take many milliseconds. In summary, the advantages and disadvantages between MS-in-space and MS-in-time instruments for ddMS2 depend on the specific features of each model and how they are implemented. However, for ddMSn where n > 2, the MS-in-time instruments are usually the only choice because the MS-in-space instruments simply lack the required hardware sections.
The “data-dependent” MSn process which automates MSn acquisition without specifying a predetermined list of analyte m/z. The bars represent total intensities of spectra collected over time so that the bars comprise a chromatogram peak eluting from an LC. The table shows the preprogrammed commands that the data-dependent MSn uses, and the stages corresponding to the letters in the chromatogram. Stage a: Collection of MS1 spectra (black) remaining below a preprogrammed threshold intensity. Stage b: The MS1 spectra are detected to have exceeded the threshold. Stage c: The instrument performs MS2 just once on the most intense peak m/z 637.1 (blue), as programmed. Stage d: Returns to MS1 which shows that the intensity is still above threshold. Stage e: The instrument performs MS2 on the second most intense peak m/z 266.8 (orange) twice as programmed, and then returns to Stage d which finds the intensity still above threshold. Stage f: The instrument performs MS2 on the third most intense peak m/z 469.0 (green) once. Stage g: This is never executed because the MS1 has fallen below the threshold. This is a very rudimentary illustration and data-dependent acquisition can include many more conditional parameters than shown here.
An example of intensive ddMSn by MS-in-time instruments is the generation of an “ion tree.” Figure 17, which is effectively a multiple version of Fig. 4, illustrates how ddMSn can be used to automatically investigate multiple parallel fragmentation pathways of a substance. If the ion tree is generated at accurate mass (recall that this also requires sufficiently high resolution), the ion tree can reveal details regarding the substructures of a given molecular ion, even in the highly complex sample illustrated here. Ion tree generation can take many seconds to minutes, making it practical only for direct infusion that provides a continuous source of molecular ions, as in the case of Fig. 17.
The “data-dependent” MSn process as applied to ion trees. These spectra were obtained directly from a crude extract of wheat plant under continuous electrospray into an ion trap MSn instrument, total acquisition time was <30 s, and the precursor ion of 637.1 m/z is an unknown metabolite. The MS instrument was programmed to automatically obtain up to MS4 spectra for each of the two most intense ions in a spectrum, thus yielding an extremely detailed, sequentially controlled fragmentation for interpretation without prior knowledge of the structure or expected ions. Figure redrawn from 10 using unpublished data by the author.
The bottom part of Table 1 lists the metabolomic considerations of MS-in-time instrument types. Note that all of these are capable of MSn of n > 2, which as discussed previously is an intrinsic advantage of the MS-in-time design strategy. Because MSn implies advanced applications of MS, many instruments are generally richer in features (i.e., noncollision dissociation modes, variety of ion sources, and higher accuracy and resolution), at of course correspondingly greater cost. However, for metabolomics, the advantages of such capabilities can be considerable. Lastly, note that this category represents the only instruments that have sufficient resolution and mass accuracy to perform multielement isotopologue analysis for advanced Pathway approaches, discussed below.
3.3 Sensitivity
Sensitivity is a general term that can have many meanings to different users, but in MS analysis the main concept is that of signal to noise ratio (s/n). Most chemists are familiar with concepts and determination of s/n for data generated by instruments, which are usually well defined for a given type of instrument, and is also sample specific. One common determination of the analytical s/n of a mass spectrometer is the signal strength (i.e., in ion counts) of series of known constituent mass spectra that is acquired over time on an actual sample, in comparison to a blank sample. Thus, if the total spectral ion count for NADH from Fig. 2 averaged 12,000 and the blank under identical conditions averaged 200 counts, then s/n = 60. However, the total spectral ion count includes spectral noise in between fragment m/z peaks, so if only the actual fragment m/z peaks of NADH were used the s/n would very likely be far better. This, in turn, means that dozens to hundreds of combinations of m/z peaks can be custom specified to optimize the s/n of each metabolite such that the s/n is often determined post acquisition at the time of data processing, and can be a different specification for each sample. Such myriad s/n complications have led to much simpler, single-numerical value standards, such as minimum detectable limits (MDLs) or minimum quantifiable limits (MQLs), as the most widely used metrics for sensitivity in MS analysis.
However, for metabolomics, since structure-revealing spectral data are critical to the extremely complex sample analyses, MDL and MQL are not useful metrics of s/n because they are specified for a very limited set of analytes, and not readily applied to submolecular parts (fragments) which are sometimes the key metabolomic data sought. Furthermore, MDL and MQL cannot be specified for unknown analytes or cases, whereby authentic standards are not available, both of which are extremely important for metabolomics. Thus, for metabolomics, such simplified mass spectrometry conventions need to be disregarded.
The analytical s/n of MS is intrinsically the best among structure-revealing analytical techniques, but it depends heavily on the instrument design, operational mode, and analytical intention, such as having a preset list of analytes versus profiling of unknowns. In case of the latter, s/n is effectively impossible to determine since it is not known which m/z peaks “belong” to the unknown. Thus, a discussion of MS spectral s/n in metabolomic applications threatens to span several chapters of discussion, at the end of which no general conclusions can be drawn. Ultimately, the adequacy of s/n—and not s/n as some citable numerical standard—is what should be addressed. Adequate s/n must be couched in terms of supporting the findings of an experimental study, and is not a consideration of the analytical platform alone. For example, a very basic and common question such as “Will the MS have enough sensitivity to analyze NADH at picomolar levels in my samples?” may need much discussion and testing to answer because it depends on the intended interpretation of such data. For discovery or hypothesis-generating studies, the intended interpretation is not known prior to analysis.
From a hardware perspective, the discussion of s/n is simplified because in most cases the MS sensitivity is limited by the ion source, and not the design of the MS itself. The classical ion source is electron ionization, which is variously estimated to have <1% ionization efficiency (1). This is in contrast with the electrospray source, which for a species with permanent charge, such as quaternary ammonium compounds (e.g., cholines, betaines), has a theoretical ionization efficiency of 100%. Actual efficiency is less than 100% and depends on the particular design and operation of both the ion source and MS, as well as the general nature of the sample (e.g., 1, 5).
The foregoing discussion touched upon just a few of the numerous considerations of s/n in MS analysis, and a comprehensive coverage is far beyond the scope of this chapter. The important practical parameter for sensitivity in metabolomics is how much material is needed to obtain a spectrum of experimentally adequate s/n across numerous analytes in a given analysis time. Modern electrospray MS instruments can routinely obtain experimentally adequate spectra from subpicomoles of each substance in an extremely complex metabolomic extract (e.g., 6). In contrast, NMR requires much larger amounts of each analyte in a complex mixture, on the order of nanomoles of each metabolite. It must be kept in mind that the metabolomic information from these two techniques is highly complementary so that this large difference in sensitivity alone is not a useful criterion in choosing one technique over the other.
4 Chromatography-MS
Although it is well beyond the scope of this chapter to cover metabolomic uses of chromatography—a vast field by itself—some perspectives with specific regards to MS in metabolomics are warranted. As stated earlier, chromatography interfaced to MS is a result of inadequate MS capability. For example, there are many metabolites that are isobaric compounds (identical molecular formulae), principally structural isomers, which can give essentially the same fragmentation pattern, and thus are indistinguishable even by MSn; simple examples of this are leucine vs. isoleucine, Z vs. E fatty acids, and so on. The most difficult is distinguishing stereoisomers, which is usually best left to NMR, although there are chiral chromatography columns that are useful for resolving enantiomers and diastereomers, if the samples are of relatively simple composition. However, it should be kept in mind that, as stated at the onset of this chapter, the analytical needs of metabolomics usually entail highly complex samples driven by the need for very rich data sets.
A major contribution of interfacing chromatography to MS instruments is to reduce the complexity of the sample presented to the MS instrument at any given time, thus reducing spectral ambiguity and improving s/n (however, that is defined). For example, the significant spectral overlap problems illustrated by Fig. 8 could possibly be addressed by chromatography which can separate the PC34:3 from PC34:2. This would allow the use of low- resolution MS instruments, which is a considerable cost and possible ease-of-use advantage. Among the most affordable MS instruments is GC-MS. Figure 18 and its legend illustrate the use of GC-MS to analyze silyl-derivatized amino and organic acids for their isotopologues. It should be noted that, due to the nonvolatile nature of almost all of the metabolome, and even with derivatization, the limitations on molecular mass imposed by the requirement of volatility, GC-based techniques are intrinsically poorly suited for metabolomics. Their broad and decades-long usage—including by this author—is principally due to the low cost and practical robustness of the technique. Such factors should be weighed against the paucity of metabolomic coverage by GC-MS, which imposes severe limits on research progress or diagnostic potential. Thus, it is expected that GC-MS usage will be phased out from metabolomics as other MS techniques develop.
GC-MS analysis of silyl-derivatized organic and amino acids, revealing isotopologue patterns from crude extracts of rhabdomyosarcoma cells grown in the presence of [U-13C]-glucose. For each metabolite, the corresponding molecular ion region of the tert-butyldimethylsilyl derivative is shown for analyses of both natural isotopic abundance chemical standard and cellular extract (“Rh 30 cells”). The total ion chromatogram is shown for the Rh 30 sample while the chromatogram for the standards is not shown. By comparison of the spectra, it is immediately apparent that some metabolites are heavily and multiply labeled (red lettering) while others are not (blue lettering), rapidly providing a glimpse into the glucose-utilization networks by these cancer cell lines. This would not have been apparent using unlabeled cell cultures, which would instead require extensive experimentation, tedious quantification of concentrations, and modeling of pathways to interpret the concentrations, just to obtain a similar glimpse. Redrawn from 10, 11.
Chromatography can help, but not necessarily solve, the problems faced by many Pathway approach samples. Figure 9 presents a very elementary example to revisit in this regard. As there are potentially scores of isotopologues for each gPL, they overlap the entire m/z range of gPLs. Thus, there is no assurance that any of the peaks detected by insufficient-resolution MS are free of interference from isotopologues of other lipid species; this point was illustrated in Figs. 9 and 10. There is currently no practical chromatographic solution that can separate all of the thousands of possible gPLs. Additionally, chromatography cannot ensure adequate separation of metabolites that are unknown, unexpected, or unavailable as authentic standards; all of these categories are of key importance to metabolomics.
An example of combining chromatography with data-dependent acquisition is shown in Fig. 19 and its legend. Note that data-dependent acquisition is not limited to MS instruments; in this case, a UV-visible detector was used to ensure acquisition of mass spectra only when there was sufficient material eluting from the LC. It is important to keep in mind that, as discussed for Fig. 16, chromatography can also severely limit the utility of ddMSn due to the short time the analyte is present at the MS interface as well as the constantly changing concentration of analytes in the ion source.
Example of the “data-dependent” MSn process as applied to LC-MS. The chromatogram is LC-photodiodearray total spectrum detection (PDA) of monobromobimane-derivatized wheat root extracts. “PC” refers to phytochelatins and the numerals refer to various ϒ-glutamyl-cysteine extensions. The PDA was set to trigger the ion trap MSn instrument to acquire MS1 spectrum (not shown) upon rise in signal, illustrating how data-dependent acquisition is not limited to MS alone. The instrument then acquired a high-resolution “zoom-scan” MS1 spectrum (right side of each spectrum) in the region of the most intense ion in the MS1 spectrum, followed by MS2 spectrum (left side of each spectrum) of the putative monoisotopic ion in the zoom scan. Thus, three data-dependent acquisitions were linked to obtain four spectra from each chromatographic peak (UV–vis, low-resolution MS1, zoom-scan MS1, and MS2 spectra). Redrawn from ref. 19.
5 Biomarkers vs. Pathway Approaches for MS
5.1 Biochemical Information Content
Ultimately, the information generated from MS analysis—not the data per se—is the objective of any analysis. Thus, the analysis must go well beyone the past exercise of merely obtaining metabolite concentration data, to providing key information on metabolic response differences (e.g. control vs. treatment) of hundreds or even thousands of metabolites, and doing so in the presence of a large background of the very same metabolites, including unknown or unexpected metabolites. This is an unprecedented challenge, as these requirements are completely beyond all other current applications of MS, such as clinical diagnostics, drug analysis, biotechnology, security, and environmental pollution, to name just a few.
The brief discussion above regarding s/n in MS focused mainly on spectral data. But determining the s/n for information is even less straightforward. Assuming that the data s/n is adequate—which is the initial step—we quickly find that the really important task is to determine s/n of information content. To assess this, we must necessarily include metadata, such as the experimental or diagnostic purpose and biochemical interpretation, other MS data, as well as data from other types of measurements. For the Biomarker approach, adequate information s/n would mean that the data can be used to statistically detect differences between biological treatment and control; thus, in this case, information s/n is a direct function of data s/n. For the Pathway approach, the detection must be (at the least) for differences between biochemical pathways of treatment and control samples; thus, data s/n alone is insufficient to judge the adequacy of MS analysis for a given study.
Figure 18 can be revisited as a simple example to compare the potential information s/n for both Biomarker and Pathway approaches. For the purposes of this discussion, the data s/n is assumed to be very good and thus have no influence on this discussion. For the Biomarker approach, which uses no stable-isotope labeling, this GC-MS run would consist of only of spectra identical to the natural abundance standards (only the bottom spectra for each metabolite in Fig. 18). This figure would convey almost no information s/n for the Biomarker approach; that is, there is no statistical significance that can be gleaned from runs of one chemical standard versus one sample.
For the Pathway approach, the data set from the same two GC-MS runs would include the isotopologue data (top spectra for each metabolite in Fig. 18), from which we can readily obtain hints of glucose metabolism pathways. Of course, there is almost no scientific information in Fig. 18 alone (to obtain that, the corresponding control samples must be run), but there is clear evidence for the analytical approach to be able to distinguish biochemical pathways, which clearly conveys the potential to gain scientific information from continued experimentation and analysis. In summary, from two GC-MS runs—one standard and only one sample—the Biomarker approach would gain next to no information (conceptually, information s/n ≤ 1) while the Pathway approach already yields some useful information (conceptually, information s/n > 1).
Once it is established that the information s/n > 1 for a given analytical approach, then the next focus of MS analysis for metabolomics should be high information throughput (HIT), not per se high sample or data throughput. The reasoning behind HIT is simple: obviously, high data or sample throughput that yields data with information s/n < 1 is in essence useless. HIT is not a function of the analytical technique alone. HIT requires the interaction of MS considerations with experimental design, quality of its execution, sampling, sample preparation, other non-MS considerations, such as other analytical instruments, most notably NMR, and data reduction.
Although HIT might at first sound like huge sample numbers or large data sets, that is not necessarily true. For an illustration of this, we turn once again to Fig. 18 and its legend, where arguably the most powerful “analytical” tool is the 13C labeling of the experiment, and otherwise utilized the least-expensive and least-sophisticated MS instrument type. Compared with nonlabeled experimentation of the Biomarker approach, the Pathway approach of Fig. 18 illustrates far greater potential for HIT because of its much greater information s/n, without the need to improve the data s/n, analysis speed, cost, sophistication, sample preparation, or any other aspect of the instrumental analysis. The greater HIT from labeling experiments, in turn, provides a major basis for far less experimentation as compared to nonlabeled experiments, which in turn accelerates scientific discovery and hypothesis testing. In this way, pursuit of HIT can lead to experimental designs with far less samples than would be needed otherwise. The approach has been used for a number of publications in stable isotope-resolved metabolomics (SIRM) which involved this author (2–4, 7–13).
However, modern MS instruments can readily provide HIT for the Biomarker approach as well. An example of this is shown in Fig. 20 and its legend. This is a case, where the number of samples is necessarily limited (resected tissue from human subjects) which makes HIT approaches indispensable. The information content of lipid analysis by FT-ICR-MS is a very high data throughput technique, with several thousand lipid metabolites analyzed in 10 min (6); more explanation of such use of FT-ICR-MS follows below. This makes it a true HIT for the Biomarker approach. The very high data content combined with rapid sample preparation and comprehensiveness of gPL analysis supplies extremely detailed, very high coverage data for comparative statistical analyses. The key point conveyed by Fig. 20 is that such surgically obtained human samples will rarely reach large sample numbers for optimal statistical needs. Therefore, alternative high sample throughput techniques that sacrifice HIT (e.g., fast MSn analysis of only 50 gPLs) would not be desired in such cases.
Comparison of glycerophospholipid (gPL) profiles from cancerous and noncancerous lung tissues from the same human subject using the method similar to Fig. 10. (a) Shows FT-ICR-MS analyses of gPL extracts. Note that overall the spectra look similar to each other, which is expected since the most abundant gPL serves functional roles, such as membrane structure. (b) Shows the expansion of the region indicated in (a), illustrating the three general categories of results for the thousands of m/z ions obtained from each such 10-min sample run. As indicated, the most abundant lipids tend to be similar in profile while many of the less abundant lipids decrease or increase in lung cancer relative to the noncancerous lung tissue from the same human subject. In this particular example, there are >1,000% differences in the indicated lipids, which are only two out of many dozens of differences in this concentration range that are present. This is a high-information throughput (HIT) technique for the Biomarker approach; high sample throughput techniques at the expense of HIT are not needed, nor desired, nor even feasible in such a limited-subject study. Figure redrawn from 8.
Figure 21 illustrates HIT in the Pathway approach. Considerable information exists even in these spectra from just two extracts of A549 cancer cells grown on [U-13C]-glucose. As explained in more detail in the legend, 13C incorporation into fatty acids was inhibited without affecting whole gPL synthesis; thus, one possible explanation is inhibition of the fatty acid synthase system (FAS) that would utilize glucose-metabolism sources of carbon. Such information can already be discerned from analyzing just two samples! In fact, many more clues lie in the detailed pattern of labeling (not shown) of Fig. 21 and other spectra from related experiments so that it is possible that there are hundreds of pieces of biochemical information embedded in these spectra.
Continuous nanoelectrospray FT-ICR-MS analysis of methanolic extracts of human lung adenocarcinoma (A549) cells using the method similar to Fig. 10. The A549 cells were grown in [U-13C]-glucose in the absence (control, top panel) or presence of selenite (bottom panel). The accurate mass data (numerals on peaks) usually provides sufficient information for assignment of all gPLs to the head group and total acyl chain level of detail, including all of the corresponding isotopologues (6). Extensive incorporation of 13C from [U-13C]-glucose into the acyl chains of gPLs is evident from detailed data analysis of neutron mass incorporation (see Fig. 10 for explanation). The detailed data analysis (not shown) also revealed that the very substantial difference in the appearance of the two spectra was due to the lack of 13C labeling in the fatty acyl chains of gPLs in the selenite treatment. Since the gPL head groups were 13C labeled in similar fashion in the two treatments, whole gPLs were being synthesized at similar rates; yet the fatty acyl chain portions of the gPLs were not. Therefore, one plausible explanation is inhibition of fatty acid synthesis by selenite; since much of that occurs by the fatty acid synthase multienzyme complex (FAS) that utilizes glucose-metabolism sources of carbon (e.g., via acetyl-CoA), that could be one of the targets of selenite effects. Unpublished data from T. Fan, A.N. Lane, and the author.
5.2 The Natural Abundance Problem in Enrichment Quantification
Several of the key fundamental concepts above come into play when labeling patterns are complex, which as mentioned earlier can be rich in information but confounding if it cannot be dealt with. First and foremost, each isotopologue itself has contributions from natural-abundance isotopologues, which in turn means that each isotopologue must itself consist of a mixture of enriched and natural-abundance isotopes; the latter commensurate with the structure. Because the amount of enrichment is unknown, the amounts of the corresponding NA isotopologues are also unknown.
These statements are better understood by the example in Fig. 22 and its legend. A key concept from the figure is that these isotopologue mixtures are not resolvable by chromatography or mass spectrometry; thus, there is no measurement solution to this problem. In order to deal with this complexity, we first have to understand distributions at natural abundance, which can be calculated by the binomial theorem (1):
where p(m) is the probability of finding m isotopic atoms, n is the total number of elemental atoms in the molecule, and f is the fraction of isotopic atoms (e.g., 13C = 1.108% at natural abundance). For example, for carbon in PC 34:1, n = 42, and the relative abundances of m + 0, m + 1, m + 2, and m + 3 are 0.626, 0.295, 0.068, and 0.01, respectively. Thus, for even a modest dynamic range of 103 (i.e., the largest and smallest peaks have intensity differences of 1,000-fold), in this case at least four NA isotopologues must be taken into account for each of the enriched isotopologues. Because in real applications the dynamic range is even greater at 104 or more, this means that even a single lipid species that is enriched in 13C can have many dozens of NA isotopologues. Since there are hundreds of lipid species in a single spectrum, it follows that a single spectrum can easily consist of thousands of NA isotopologues. This is, in fact, what is observed (see Figs. 10, 20, 21 and refs. 3, 4, 6, 7).
Illustration of the complexity and problems regarding multiply 13C-enriched isotopologues of a single compound. Simulated spectra represent, from bottom, respectively, several isotopologues of PC 34:3 with their relative levels of enrichment: black = natural abundance 1.0×; green = 13C1 3.5×; blue = 13C2 0.5×; red = 13C3 0.75×. The spectrum at the top is the mixture of these four isotopologues, as would be acquired from an actual sample. The problem is, except for the monoisotopic m/z 829.5616, each of the exact masses in the top spectrum harbors multiple 13C isotopologues of the same metabolite. Moreover, neither the number of isotopologues present nor their intensities are known; in fact, this is precisely the experimental data that is desired from the analysis. Most frustratingly, because natural-abundance and enriched isotopologues are physically–chemically identical in every way, there will be no MS measurement solution to this problem; there will be no chromatographic, no mass spectrometric technique forthcoming, ever, that can distinguish between sources of these chemically identical isotopologues. However, there is an informatics solution to this as described in the text.
Now, to account for the enrichment—effectively the change in the NA profile upon incorporation of multiple unknown numbers of an isotope (e.g., 13C) into molecules—an exact solution was not available until recently, when an iterative NA isotope stripping approach was developed for pathway metabolomics (6, 14). This is based on eventually decaying binomial terms (2) that describe the multiple possible combinations of 13C incorporation for a particular molecule,
where C Max is the number of carbon atoms in the molecule and NA (0.01108) is the fractional natural abundance for 13C. In the presence of 13C enrichment, the effects of the natural abundance can be calculated using a series of decaying binomial terms as described by (3) representing the contributions from 13C natural abundance for each mass isotopologue.
starting with i = 0. I is the intensity of the ith species, with B(x, i) defined in (2).
In essence, then, the complex isotopologue pattern acquired in the MS (e.g., top spectrum of Fig. 22) contains sufficient information to strip the NA contributions and, thus, reveals the biologically enriched contributions of each isotopologue. This is an illustration of the crucial importance of chemical informatics as an integral part of MS for metabolomics.
5.3 A Model Approach for Metabolomic MS Analysis
Thus far, this chapter has addressed separately the various analytical concepts for the use of MS in metabolomics. So how does it all come together? One analytical approach is to exploit the simultaneous accurate mass, extremely high resolution capability of the FT-ICR-MS (Table 1) to perform a broad-coverage, isotopologue-resolving, metabolite-identifiable, rapid, and sensitive analysis of a crude biological extract (5, 6). Figures 10 and 21 illustrate just such analyses, each consisting of >1,500 analytes including isotopologues, at picomole amounts, acquired in just 10 min of FT-ICR-MS time from biological samples using a simple and rapid extraction. In other words, a true HIT analysis: note that the Fig. 21 legend is able to discuss possible biological interpretations from the general appearance of just two spectra.
Beyond general spectral appearance, it is completely feasible to obtain extremely detailed metabolite information by this approach. A hint of this data richness was presented in Figs. 10 and 20, showing specific lipid isotopologue assignments and specific lipid differences, respectively. From any of these spectra, a long list of the m/z values can be exported, assigned, and sorted, as illustrated by Fig. 23 and its legend. In this way, the thousands of metabolites from a single spectrum can be tentatively assigned by relatively simple matching of m/z values. Once assigned, the NA isotopic contributions can be stripped from each lipid as described in the preceding section to yield the isotopic enrichment of each of the thousands of gPL isotopologues present in the spectrum.
Illustration of data output from precalculated mass isotopologue search engine (PREMISE). The first two columns consist of a small portion of text output from the FT-ICR-MS (spectrum shown in Fig. 10), in this case a Thermo LTQ-FT using Xcalibur software. Then, PREMISE compared this accurate mass list with a simple spreadsheet containing theoretical m/z of every possible mammalian gPL (including those never before observed, but theoretically possible): all known head groups (PC, SM, PE, PS, PI, etc.) were combined with every possible pair of known saturated and unsaturated fatty acid chains, utilizing both ester and ether linkages. This results in a tentative assignment of the gPL adducts (columns 4–7) and their isotopologue identity (column 3) of the thousands of peaks in column 2. The last column shows the extremely small deviation from the theoretical m/z of each peak, giving some measure of the confidence of the assignment. A second analysis using FT-ICR tandem MS can confirm the assignments of a select number of these gPLs.
Because this method uses one-dimensional MS, it is at once sensitive, rapid, and of high coverage that includes unknown or unexpected metabolites, with the caveat of tentative assignments. It fully supports secondary analyses by MSn for confirmatory and more detailed structural studies of selected metabolites. Thus, the method is amenable to both Biomarker (Fig. 20) and Pathway (Fig. 21) approaches, but places an ever-greater burden on chemical informatics.
Figure 24 and its legend describe how advanced experimental design can be supported by FT-ICR-MS analysis. This spectrum clearly shows ATP pools that have eight different metabolic “histories,” seven of them unambiguously involving exogenous glutamine. The remainder of this spectrum (not shown) contains similar detail of information regarding hundreds of other metabolites so that it becomes apparent that far more than pathway information is encoded in the MS spectrum with this approach. The experimental design in combination with the FT-ICR-MS yields information on the kinetics of multiple pathways interacting in the metabolic network of the cells. The data processing of such complex MS spectra that is required to extract the metabolic network information, but the current state of the technology is well suited to fully harness parallel advances in SIRM bioinformatics (15–18).
Negative ion nanoelectrospray FT-ICR-MS spectrum, acquired at R = 200,000 in less than 10 min, showing the region of the ATP molecular ion cluster, analysis of an extract of A549 cancer cells grown with [U-13C][U-15N] glutamine in the vehicle. Note that the last two labeled peaks are expanded on the ordinate to make them visible. The spectral quality is sufficient to detect that, for example, the m + 5 peak (ATP 13C 151 N1) is not from ATP 13C5 (e.g., a fully labeled ribose ring); that is, the resolution and accuracy are sufficient to separately account for the different neutron masses in the nuclei of 13C and 15N. This result unambiguously shows that some of the carbon and most of the nitrogen of the existing ATP pool were synthesized from glutamine in this experiment. In summary, this figure illustrates that it is practical to obtain simultaneously ATP structures with eight different metabolic “histories”; note that this biochemical information is gleaned by inspection of one metabolite from just one sample. Similar information exists for hundreds of other metabolites in the same spectrum (data not shown). Thus, far more than just pathway information is encoded in the isotopologue pattern: this spectrum contains information on the kinetics of multiple pathways interacting in the intact metabolic network of the cells. Unpublished figure courtesy of P. Lorkiewicz and T. Fan.
6 Concluding Remarks
Mass spectrometry continues its breakneck pace of instrument development, for which metabolomics is a major beneficiary. For metabolomics, because of the extreme complexity of samples and need for very high metabolite coverage, there is a critical need to consider the difference between structural data content and biochemical information content in MS spectra. Figure 25 is a conceptual diagram that describes the relationship between spectral resolution and its analytical content for a ~400 m/z metabolite. The green line illustrates that, in terms of chemical structure data—principally molecular formulae information—gains are not realized until the m/z resolution surpasses the third decimal place. There is considerable gain in molecular formula confidence with the fifth decimal place due to achievement of complete molecular formula based on the full count of protons, neutrons, and electrons while the sixth decimal place does not gain any more structural information. This green line also happens to represent the information content of MS for the Biomarker approach.
The red line in Fig. 25 describes the curve for stable isotope-labeled samples in the Pathway approach. Because biochemical pathway and network information is encoded in the isotopologue pattern (as illustrated by Figs. 18, 21, 22, 24, and refs. 2–4, 6–13), the information content is vastly higher. It follows that the red curve also describes HIT once sufficient resolution is reached.
This chapter has touched upon the key concepts of MS as it applies to metabolomics. However, perhaps the most important concept conveyed is that useful MS analysis is largely dependent on non-MS aspects, such as experimental design, in particular stable isotope encoding of pathways. Aspects of the experimental design (such as the use of stable isotope labeling) must maximally exploit the available MS resources, and vice versa. For the biologist, the analytical aspects should not be an afterthought, but rather an integral part of experimental design. Conversely, for the analytical chemist, intellectual engagement in the experimental design is increasingly a requirement in metabolomics.
Abbreviations
- ddMSn:
-
Data-dependent MSn
- FT-ICR:
-
Fourier transform ion cyclotron resonance
- GC:
-
Gas chromatography
- HIT:
-
High information throughput
- ICP-MS:
-
Inductively coupled plasma MS
- IR:
-
Infrared
- LC:
-
Liquid chromatography
- MDLs:
-
Minimum detectable limits
- MQLs:
-
Minimum quantifiable limits
- MRM:
-
Multiple reaction monitoring
- MS:
-
Mass spectrometry
- MS2:
-
MS to the 2nd or tandem MS
- MSn:
-
MS to the nth
- NA:
-
Natural abundance
- NMR:
-
Nuclear magnetic resonance spectroscopy
- PC:
-
Phosphatidylcholine
- RF:
-
Radio frequency
- SIRM:
-
Stable isotope-resolved metabolomics
- ToF:
-
Time of flight
- UV:
-
Ultraviolet visible
References
Watson JT (1985) Introduction to mass spectrometry. Raven, New York
Fan TWM et al (1997) Anaerobic nitrate and ammonium metabolism in flood-tolerant rice coleoptiles. J Exp Bot 48(314): 1655–1666
Fan TW et al (2009) Altered regulation of metabolic pathways in human lung cancer discerned by (13)C stable isotope-resolved metabolomics (SIRM). Mol Cancer 8:41
Fan T et al (2005) Metabolomics-edited transcriptomics analysis of Se anticancer action in human lung cancer cells. Metabolomics J 1(4):325–339
Aharoni A et al (2002) Nontargeted metabolome analysis by use of fourier transform ion cyclotron mass spectrometry. OMICS 6(3):217–234
Lane AN et al (2009) Isotopomer analysis of lipid biosynthesis by high resolution mass spectrometry and NMR. Anal Chim Acta 651: 201–208
Fan TW-M et al (2010) Stable isotope-resolved metabolomic analysis of lithium effects on glial-neuronal metabolism and interactions. Metabolomics 6(2):165–179
Lane AN et al (2009) Prospects for clinical cancer metabolomics using stable isotope tracers. Exp Mol Pathol 86(3):165–173
Lane AN, Fan TW, Higashi RM (2008) Stable isotope-assisted metabolomics in cancer research. IUBMB Life 60(2):124–129
Lane AN, Fan TW, Higashi RM (2008) Isotopomer-based metabolomic analysis by NMR and mass spectrometry. Methods Cell Biol 84: 541–588
Fan T et al (2008) Rhabdomyosarcoma cells show an energy producing anabolic metabolic phenotype compared with primary myocytes. Mol Cancer 7(1):79
Fan TWM, Higashi RM, Lane AN (2006) Integrating metabolomics and transcriptomics for probing Se anticancer mechanisms. Drug Metab Rev 38(4):707–732
Fan TWM, Lane AN, Higashi RM (2003) In vivo and in vitro metabolomic analysis of anaerobic rice coleoptiles revealed unexpected pathways. Russ J Plant Physiol 50(6): 787–793
Moseley HN (2010) Correcting for the effects of natural abundance in stable isotope resolved metabolomics experiments involving ultra-high resolution mass spectrometry. BMC Bioinformatics 11:139
Arita M (2003) In silico atomic tracing by substrate-product relationships in Escherichia coli intermediary metabolism. Genome Res 13(11): 2455–2466
Arita M (2004) Computational resources for metabolomics. Brief Funct Genomic Proteomic 3(1):84–93
Arita M (2009) What can metabolomics learn from genomics and proteomics? Curr Opin Biotechnol 20(6):610–615
Cascante M et al (2010) Metabolic network adaptations in cancer as targets for novel therapies. Biochem Soc Trans 38(5): 1302–1306
Fan TWM, Lane AN, Higashi RM (2004) An electrophoretic profiling method for thiol-rich phytochelatins and metallothioneins. Phytochem Anal 15(3):175–183
Acknowledgments
The unpublished data shown here was supported in part by grants from NSF EPSCoR EPS-0447479, NIH 1R01CA118434-01A2, NCI R21CA133688, and the Susan G. Komen Foundation BCTR0503648 while the published data was supported by the grants listed in the cited publications. The author thanks T. Fan, A.N. Lane, H, Moseley, J. Winiike, P. Lorkiewicz, M. Arita, J. Goran, and T. Cassel, among numerous others, for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer Science+Business Media New York
About this protocol
Cite this protocol
Higashi, R.M. (2012). Structural Mass Spectrometry for Metabolomics. In: Fan, TM., Lane, A., Higashi, R. (eds) The Handbook of Metabolomics. Methods in Pharmacology and Toxicology. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-61779-618-0_4
Download citation
DOI: https://doi.org/10.1007/978-1-61779-618-0_4
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-61779-617-3
Online ISBN: 978-1-61779-618-0
eBook Packages: Springer Protocols