Abstract
Quorum sensing signaling molecules also called auto-inducers secretes from bacteria into its immediate extracellular environment and the molecules are concentrated as their bacterial population increases. At certain threshold concentration, auto-inducers regulate the expression of different types of genes and phenotypes, which includes virulence and formation of bio-films. Bio-films are responsible for 65% of bacterial infections. Mycobacterium tuberculosis (Mtb) causes one of the infectious disease named as tuberculosis (TB), infected one-third of world’s population. In humans, following primary TB infection, Mtb enters into latent stage. Reactivation or re-infection by new Mtb soften and fragments the lung tissue leaving cavities. The success of Mtb comes from its ability to grow as pellicle, a bio-film like structure on the surface of such cavities. The Mtb bio-films are highly resistant to drugs and implicated in persistence. The presence of LuxR homologs and expression patterns of transcription regulator, WhiB3 suggests quorum sensing existence in Mtb. The involvement of nucleotide-based second messengers such as c-di-GMP in signal transduction gives another indirect evidence of quorum sensing mechanisms in Mtb. The present chapter reviews quorum sensing mechanisms and their importance in bio-film formation, regulation of gene expressions, virulence and pathogenesis of Mtb, which will provide basis for novel anti-tuberculosis therapy.
Similar content being viewed by others
Keywords
Introduction
Mycobacterium tuberculosis (Mtb) belongs to phylum actinobacteria, which is non-motile and rod shaped bacterium. It is also an acid-fast obligate aerobe which divides slowly for every 15–20 h. The Mtb cell wall contains three important components: arabinogalactan, peptidoglycan and mycolic acids. Based on the cell wall structure, Mtb considered as gram positive bacteria [2]. Tuberculosis (TB) is caused by Mtb, which affects one third of world’s population annually. The disease caused by Mtb in any of the following ways: (1) production of virulence factors, (2) Colonization of host body and its persistence, (3) Invasion of host cells, (4) Expression of immunosuppressive compounds and (5) Expression of toxins [1]. The increased emergence of multidrug–resistant TB and its co-infection with HIV has become a major problem in anti-TB therapy [3].
Quorum sensing (QS) is a bacterial community phenomenon, which is mediated by secretion of small signaling molecules. The small signaling molecules are divided into three major classes: (1) N-acyl homoserine lactones (AHLs), which are produced by many gram-negative bacteria, (2) Oligopeptides, which are employed by gram-positive bacteria, (3) Autoinducer-2, DPD (4,5-dihydroxy-2,3-pentanedione) are used by both gram-positive and negative bacteria [4]. The indiscriminate and over use of antibiotics increases the chances of drug resistance in bacteria, which has become a serious health concern globally. The emergence of drug-resistance bacteria decreases the effectiveness of current treatment modalities. Therefore, novel strategies or development of compounds against drug resistance bacteria is needed immediately. Various studies have been reported that QS is responsible for the regulation of biofilms formation and pathogenicity in both gram positive and negative bacteria. Since QS is linked in regulation of biofilm formation and virulence in many pathogenic bacteria, it has been proposed that QS will become a new potential target in the development of novel antibacterial therapies. The compounds with anti-QS activity are known as quorum quenching (QQ), in which the molecules do not kill, but attenuate the pathogenic bacteria. Some studies reported that anti-QS molecules increased the bacterial biofilms sensitivity to antibiotics in both in vitro and in vivo [5]. The present chapter describes about QS mechanisms in Mtb and its role in biofilms formation and pathogenesis.
The Concept of QS and Its Evidence in Mtb
Communal behaviors of various gram-positive and gram-negative bacteria are coordinated by the mechanism called QS. In the QS of different bacteria, many specific genes are involved in the regulation in response to bacterial population density. Coordination between the size of bacterial population and expression of specific genes are achieved by specific signaling molecules called auto-inducers. Bacteria sense their population density by releasing these diffusible auto-inducers. Small amounts of auto-inducers are released and diluted in the surrounding environment by basal level expression at low bacterial population density. These signaling molecules gradually accumulate as the bacterial population expands until a certain threshold concentration is reached. At beyond a threshold concentration, auto-inducers bind to receptors present on the cell membrane or cytoplasm [6].
Different QS signaling mechanisms have been reported in both gram-negative and positive bacteria. In gram-negative bacteria, regulatory proteins such as LuxI and LuxR are involved in QS. These regulatory proteins activate the synthesis of autoinducers such as acylated homoserine lactone (AHL). The secreted AHL binds to its cognate receptor and forms autoinducer-receptor complex, which in turn binds to promoter region and regulates the expression of genes. In gram-positive bacteria, modified oligopeptides as autoinducers activates the sensor kinase in the cytoplasm, which in turn phosphorylates the response regulator proteins. Finally, the response regulator protein binds to target promoter and regulates the expression of genes [7]. Bacteria exploit QS to exhibit certain phenotypes such as bio-film formation, extracellular matrix and virulence factors production. These phenotypes are important for establishment of strong associations between pathogenic bacteria and with their specific hosts [8].
In gram positive mycobacteria, there is no clear evidence of QS reported. However, some evidences of QS mechanism have been revealed in Mtb. In one bioinformatics study, they have shown that the presence of LuxR-like regulators in mycobacteria, which are important players in QS [9]. Some indirect evidences revealed that the putative transcription regulator called WhiB3 gene was linked in QS regulation of Mtb recently. The expression patterns of WhiB3 correlated with the changes in bacterial density, which suggests the existence of QS in Mtb [10]. The expression of WhiB3 will decrease with the increase in bacterial density after post-infection by Mtb (Fig. 1). Other indirect evidences of QS also shown that the nucleotide based second messengers are involved in the regulation of various physiological processes of mycobacterial species [11].
Role of QS in Mtb Biofilms Formation
Biofilms are surface adherent, multicelluar bacterial communities formed by different types of stresses such as nutrient limitation, heat shock and exposure to antibiotics. Biofilms are also described as structured aggregations of microorganisms develop on non-biological (medical devices) and biological surfaces. Biofilms are one of the survival strategies of bacteria, responsible for 65% or more of all infections and highly resistant to host immune mechanisms and also to other conventional anti-bacterial therapy [12].
Formations of biofilms are common in many different types of micro-organisms such as bacteria, archaea and fungi. Initially, bacteria attach firmly to surfaces and proliferate in their numbers. As the number increases to certain concentration triggers the activation of QS circuits, which endogenously regulates secretion of signal molecules. When secreted molecules reach the certain critical concentrations, they are taken up and activate regulatory mechanisms. The secreted signal molecules regulate the transition from susceptible planktonic to adaptively resistant multi-cellular biofilm formation. Some cells dissociate from mature biofilm structures and spread, colonize other new surfaces, where they develop very rapidly against to stress signals in order to avoid adverse effects of stress signals. This planktonic-biofilm transition is a highly regulated and complex process [13]. A biofilm program’is activated by differential expression of specific genes even before a structured biofilm is formed [14].
The infectious disease TB is not much virulent in humans when compared with animals. Primary TB infection regresses within a few weeks in humans due to immunity and enters into latent stage, where it is not completely sterilized. In Primary TB infection, granulomas are formed and spread systemically and also to lymph nodes. Reactivation of dormant (or) re-infection by new Mtb causes cavities on the soft lung tissues. In these cavities Mtb grows massively and form the structure called pellicle, which separates from host immune response to prevent from lymphocytes penetration and also increases drug tolerance like that of many other bacteria [15].
There are studies reported the formation of biofilms by Mtb H37Rv in the laboratory conditions [16]. But when, where and how Mtb forms biofilms in vivo remains to be determined. The existence of three genetic loci such as pks16, helY and pks1 suggested that their role in Mtb biofilm formation [17]. Among several protein kinases, PknG was shown that it regulates the formation of biofilms in Mtb by redox sensing pathways [18].Some studies shown that biofilms formation by Mtb depends on keto mycolic acids [19]. Another studies also reported that QS is responsible for the formation of biofilms by Mtb, where it is mediated through variety of small molecules such as c-di-GMP [11]. Bacterial biofilms are developed by many factors such as contact surface, pH, nutrient availability, contact time with surface, growth stage, surface hydrophobicity and textures of surface, temperature, humidity etc.,. Among these factors temperature, pH and nutrient composition are suggested to be important for the growth of mycobacterial biofilms [20].
Role of QS in the Pathogenesis of Mtb
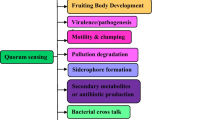
Biofilms formations as well as other physiological processes such as persistence and pathogenesis of Mtb are regulated by QS (Fig. 2). Many virulence factors of mycobacteria were identified such as: sigma factors, proteases, lipids, secretion systems, regulators etc. Depending on the function, the virulence factors have been classified into following groups: (1) Lipid metabolism, (2) cell envelop proteins, (3) protein kinases, (4) Proteases, (5) metal-transporter proteins, (6) proteins inhibits anti-microbial effectors of macrophages, (7) regulators of gene expression, (8) unknown function proteins, (9) other virulence proteins [21]. The Mtb cell envelope is characterized by complex lipids and glycolipids along with mycolic acids. Multiple methyl branched fatty acids are the components of Mtb cell wall lipids, which plays crucial role in pathogenesis [22]. The surface-exposed lipoglycan is lipoarabinomannan, suggested to be play as virulence factor. The Mtb cell envelope lipoprotein LprG also suggested that it is playing role in the normal expression of lipoarabinomannan, virulence and pathogenesis [23].
But, the clear signaling mechanism in pathogenesis (or) virulence of Mtb is largely unknown. The stationary phase is responsible for persistence and pathogenecity in mycobacteria. A second messenger such as cyclic-di-GMP is required for persistence in mycobacteria [24]. Some studies also suggested that high levels of cAMP in Mtb linked in the regulation of specific genes responsible for persistence and virulence [25]. Virulence factors are produced by QS in many pathogenic bacterial species [26]. The unusual cell wall of Mtb contains mycolic acid, which is a key virulence factor. Mutation in hadC, which encodes HadBC dehydratase results in dramatic change in mycolic acid structures, which causes loss of virulence in Mtb [27]. There are studies also reported that LuxR family regulator Rv0195 is responsible for the modulation of dormancy and virulence in Mtb [28].
Conclusion
Several studies reported that the involvement of QS in the regulation of biofilms formation and virulence in many gram positive and negative bacteria. However, some studies have been shown the indirect evidence of QS in Mtb biofilms formation and pathogenesis. In QS mechanisms multiple signaling molecules and their integration in the in vivo conditions produce various phenotypes by bacteria. But, till now only few molecules have been reported in the QS mechanisms of Mtb. Understanding of clear signaling networks in the QS of Mtb and its role in biofilms formation and pathogenesis helps us for the discovery of new QQ molecules, which in turn improve the treatment of tuberculosis.
References
Cambier, C. J., Falkow, S., & Ramakrishnan, L. (2014). Host evasion and exploitation schemes of Mycobacterium tuberculosis. Cell, 159(7), 1497–1509.
Kenneth, J. R., & Ray, C. G. (2004). Mycobacteria. Sherris medical microbiology: An introduction to infectious diseases (4th ed.p. 439). New York: McGraw-Hill ISBN 0-83-858529-9.
Mukherjee, K., Tribedi, P., Mukhopadhyay, B., & Sil, A. K. (2013). Antibacterial activity of long-chain fatty alcohols against mycobacteria. FEMS Microbiology Letters, 338(2), 177–183.
Kaufmann, G. F., Park, J., & Janda, K. D. (2008). Bacterial quorum sensing: A new target for anti-infective immunotherapy. Expert Opinion on Biological Therapy, 8(6), 719–724.
Brackman, G., Cos, P., Maes, L., Nelis, H. J., & Coenye, T. (2011). Quorum sensing inhibitors increase the susceptibility of bacterial biofilms to antibiotics in vitro and in vivo. Antimicrobial Agents and Chemotherapy, 55(6), 2655–2661.
Polkade, A. V., Mantri, S. S., Patwekar, U. J., & Jangid, K. (2016). Quorum sensing: An under-explored phenomenon in the phylum actinobacte. Frontiers in Microbiology, 7, 131.
LaSarre, B., & Federle, M. J. (2013). Exploiting quorum sensing to confuse bacterial pathogens. Microbiology and Molecular Biology Reviews, 77(1), 73–111.
Deep, A., Chaudhary, U., & Gupta, V. (2011). Quorum sensing and bacterial pathogenicity: From molecules to disease. Journal of Laboratory and Physicians, 3(1), 4–11.
Chen, J., & Xie, J. (2011). Role and regulation of bacterial LuxR-like regulators. Journal of Cellular Biochemistry, 112(10), 2694–2702.
Banaiee, N., Jacobs, W. R., Jr., & Ernst, J. D. (2006). Regulation of Mycobacterium tuberculosis whiB3 in the mouse lung and macrophages. Infection and Immunity, 74(11), 6449–6457.
Sharma, I. M., Petchiappan, A., & Chatterji, D. (2014). Quorum sensing and biofilm formation in mycobacteria: Role of c-di-GMP and methods to study this second messenger. IUBMB Life, 66(12), 823–834.
de la Fuente-Núñez, C., Reffuveille, F., Fernández, L., & Hancock, R. E. (2013). Bacterial biofilm development as a multicellular adaptation: Antibiotic resistance and new therapeutic strategies. Current Opinion in Microbiology, 16(5), 580–589.
O’Toole, G., Kaplan, H. B., & Kolter, R. (2000). Biofilm formation as microbial development. Annual Review of Microbiology, 54, 49–79.
Beloin, C., & Ghigo, J. M. (2005). Finding gene-expression patterns in bacterial biofilms. Trends in Microbiology, 13(1), 16–19.
Hunter, R. L., Actor, J. K., Hwang, S. A., Karev, V., & Jagannath, C. (2014). Pathogenesis of post primary tuberculosis: Immunity and hypersensitivity in the development of Cavities. Annals of Clinical and Laboratory Science, 44(4), 365–387.
Ojha, A. K., Baughn, A. D., Sambandan, D., Hsu, T., Trivelli, X., Guerardel, Y., Alahari, A., Kremer, L., Jacobs, W. R., Jr., & Hatfull, G. F. (2008). Growth of Mycobacterium tuberculosis biofilms containing free mycolic acids and harbouring drug-tolerant bacteria. Molecular Microbiology, 69(1), 164–174.
Pang, J. M., Layre, E., Sweet, L., Sherrid, A., Moody, D. B., Ojha, A., & Sherman, D. R. (2012). The polyketide Pks1 contributes to biofilm formation in Mycobacterium tuberculosis. Journal of Bacteriology, 194(3), 715–721.
Wolff, K. A., de la Peña, A. H., Nguyen, H. T., Pham, T. H., Amzel, L. M., Gabelli, S. B., & Nguyen, L. (2015). A redox regulatory system critical for mycobacterial survival in macrophages and biofilm development. PLoS Pathogens, 11(4), e1004839.
Sambandan, D., Dao, D. N., Weinrick, B. C., Vilchèze, C., Gurcha, S. S., Ojha, A., Kremer, L., Besra, G. S., Hatfull, G. F., & Jacobs, W. R., Jr. (2013). Keto-mycolic acid-dependent pellicle formation confers tolerance to drug-sensitive Mycobacterium tuberculosis. MBio, 4(3), e00222–e00213.
Johansen, T. B., Agdestein, A., Olsen, I., Nilsen, S. F., Holstad, G., & Djønne, B. (2009). Biofilm formation by Mycobacterium avium isolates originating from humans, swine and birds. BMC Microbiology, 9, 159.
Mwambete, K. D. (2015). Targeting microbial virulence factors: potential alternative to evade the antimicrobial resistance threats. ©Formatex. Strategies, 4, 5.
Glickman, M. S., & Jacobs, W. R. (2001). Microbial pathogenesis of Mycobacterium tuberculosis: Dawn of a discipline. Cell, 104(4), 477–485.
Manabe, Y. C., Saviola, B. J., Sun, L., Murphy, J. R., & Bishai, W. R. (1999.;26). Attenuation of virulence in Mycobacterium tuberculosis expressing a constitutively active iron repressor. Proceedings of the National Academy of Sciences of the United States of America, 96(22), 12844–12848.
Bharati, B. K., & Chatterji, D. (2013). Quorum sensing and pathogenesis: Role of small signalling molecules in bacterial persistence. Current Science, 105(5), 10 september.
Rickman, L., Scott, C., Hunt, D. M., Hutchinson, T., Menéndez, M. C., Whalan, R., Hinds, J., Colston, M. J., Green, J., & Buxton, R. S. (2005). A member of the cAMP receptor protein family of transcription regulators in Mycobacterium tuberculosis is required for virulence in mice and controls transcription of the rpfA gene coding for a resuscitation promoting factor. Molecular Microbiology, 56(5), 1274–1286.
Rutherford, S. T., & Bassler, B. L. (2012). Bacterial quorum sensing: Its role in virulence and possibilities for its control. Cold Spring Harbor Perspectives in Medicine, 2(11), a012427.
Slama, N., Jamet, S., Frigui, W., Pawlik, A., Bottai, D., Laval, F., Constant, P., Lemassu, A., Cam, K., Daffé, M., Brosch, R., Eynard, N., & Quémard, A. (2016). The changes in mycolic acid structures caused by hadC mutation have a dramatic effect on the virulence of Mycobacterium tuberculosis. Molecular Microbiology, 99(4), 794–807.
Fang, H., Yu, D., Hong, Y., Zhou, X., Li, C., & Sun, B. (2013). The LuxR family regulator Rv0195 modulates Mycobacterium tuberculosis dormancy and virulence. Tuberculosis (Edinburgh, Scotland), 93(4), 425–431.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Mallaiah, D., Pallaval Veera Bramhachari (2018). Quorum Sensing in Mycobacterium Tuberculosis: Its Role in Biofilms and Pathogenesis. In: Pallaval Veera Bramhachari (eds) Implication of Quorum Sensing System in Biofilm Formation and Virulence. Springer, Singapore. https://doi.org/10.1007/978-981-13-2429-1_22
Download citation
DOI: https://doi.org/10.1007/978-981-13-2429-1_22
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-2428-4
Online ISBN: 978-981-13-2429-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)




 Increase,
Increase,  decrease
decrease
 )
)