Abstract
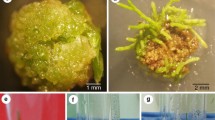
The soil bacteria Agrobacterium tumefaciens and A. rhizogenes are the etiological agents of the plant diseases crown gall and hairy root, respectively. They belong to the family of Rhizobiaceae, and thus are related to the nitrogen fixing rhizobia. Whereas crown gall is characterized by the presence of tumors on plants, the hairy root disease is so called because of a conspicuous proliferation of roots from infection sites (Fig. 1). Plant cells in crown galls and hairy roots have two features with which they are distinguished from normal plant cells: 1) they are tumorous i.e. they proliferate in the absence of added growth factors in in vitro culture, and 2) they produce and secrete specific compounds which have been given the generic name of opines. It is now known that these novel properties of crown gall and hairy root cells are a consequence of the presence of a segment of bacterial DNA, the T(ransferred)-DNA within these cells. This bacterial DNA forms part of a large (about 200 kbp) bacterial plasmid which is present in virulent strains of these Agrobacteria, and is known as Ti (tumor inducing) plasmid in the case of A. tumefaciens and Ri (root inducing) plasmid in the case of A. rhizogenes. The T-DNA of the Ti plasmid contains a number of genes which are expressed in the transformed plant cells. Some of these are onc-genes which encode enzymes involved in the production of plant growth regulators, viz. the phytohormones indole acetic acid (an auxin) and isopentenyl-AMP (a cytokinin). Other determine enzymes which catalyze the production of the opines. The presence of these genes in the T-DNA explains the novel properties of crown gall cells. Similarly, the T-DNA of Ri plasmids contains genes called rol genes which confer the capacity to differentiate into roots on transformed cells as well as opine synthase genes. There are several different classes of opines which may be present in the T-DNA transformed plant cells (Fig. 2). This depends entirely on the types of opine synthase genes present on the T-DNA. Opine producing overgrowths form a specific ecological niche for the infecting agrobacteria, since only these can utilize the opines, which are secreted from the tumor cells, for growth. Apparently, agrobacteria are metabolic engineers which apply genetic transformation to force plant cells into producing the required metabolites. For further detailed information on Agrobacterium the reader may consult a number of recent reviews1–4.
Chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
References
Binns, A.N. and Thomashow, M.F. (1988). Cell biology of Agrobacterium infection and transformation of plants. Ann. Rev. Microbiol. 42, 575–606.
Hooykaas, P.J.J. and Beijersbergen, A.G.M. (1994). The virulence system of Agrobacterium tumefaciens. Ann. Rev. Phytopathol. 32, 157–179.
Sheng, J. and Citovsky, V. (1996). Agrobacterium-plant cell DNA transport: have virulence proteins, will travel. Plant Cell 8, 1699–1710.
Dessaux, Y., Petit, A. and Tempé, J. (1993). Chemistry and biochemistry of opines, chemical mediators of parasitism. Phytochemistry 34, 31–38.
Stachel, S.E., Messens, E., Van Montagu, M., et al. (1985). Identification of the signal molecules produced by wounded plant cells that activate T-DNA transfer in Agrobacterium tumefaciens. Nature 318, 624–629.
Melchers, L.S., Regensburg-Tuïnk, A.J.G., Schilperoort, R.A. and Hooykaas, P.J.J. (1989). Specificity of signal molecules on the activation of Agrobacterium virulence gene expression. Mol. Microbiol. 3, 969–977.
Winans, S.C. (1992). Two-way chemical signaling in Agrobacterium -plant interactions. Microbiol. Rev. 56, 12–31.
Lessl, M. and Lanka, E. (1994). Common mechanisms in bacterial conjugation and Ti-mediated T-DNA transfer to plant cells. Cell 77, 321–324.
Zupan, J. and Zambryski, P. (1997). The Agrobacterium DNA transfer complex. Crit. Rev. Plant Sci. 16, 279–295.
Rossi, L., Hohn, B. and Tinland, B. (1996). Integration of complete transferred DNA units is dependent on the activity of virulence E2 protein of Agrobacterium tumefaciens. Proc. Natl. Acad. Sci USA 93, 126–130.
Ballas, N. and Citovsky, V. (1997). Nuclear localization signal binding protein from Arabidopsis mediates nuclear import of Agrobacterium VirD2 protein. Proc. Natl. Acad. Sci. USA 94, 10723–10728.
Regensburg-Tuïnk, A.J.G. and Hooykaas, P.J.J. (1993). Transgenic N.glauca plants expressing bacterial virulence gene virF are converted into hosts for nopaline strains of A.tumefaciens. Nature 363, 69–71.
Hooykaas-Van Slogteren, G.M., Hooykaas, P.J.J. and Schilperoort, R.A. (1984). Expression of Ti plasmid genes in monocotyledonous plants infected with Agrobacterium tumefaciens. Nature 311, 763–764.
Beijersbergen, A., Den Dulk-Ras, A., Schilperoort, R.A., et al. (1992). Conjugative transfer by the virulence system of Agrobacterium tumefaciens. Science 256, 1324–1327.
Bundock P., Den Dulk-Ras, A., Beijersbergen, A., et al. (1995). Trans-kingdom T-DNA transfer from Agrobacterium tumefaciens to Saccharomyces cerevisiae. EMBO J. 14, 3206–3214.
Yadav, N.S., Vanderleyden, J., Bennett, D.R., et al. Short direct repeats flank the T-DNA on a nopaline Ti plasmid. Proc. Natl. Acad. Sci. USA 79, 6322–6326.
Zambryski, P., Joos, H., Genetello, C., et al. (1983). Ti plasmid vector for the introduction of DNA into plant cells without alteration of their normal regeneration capacity. EMBO J. 2, 2143–2150.
Hoekema, A., Hirsch, P.R., Hooykaas, P.J.J., et al. (1983). A binary plant vector strategy based on separation of vir and T- region of the Agrobacterium tumefaciens Ti-plasmid. Nature 303, 179–180.
Bevan, M. (1984). Binary Agrobacterium vectors for plant transformation. Nucleic Acids Res. 12, 8711–8721.
Jefferson, R.A., Kavanagh, T.A. and Bevan, M.W. (1987). GUS fusions: 13-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 6, 3901–3907.
Vancanneyt, G., Schmidt, R., O’Connor-Sanchez, A., et al. (1990). Construction of an intron-containing marker gene: splicing of the intron in transgenic plants and its use in monitoring early events in Agrobacterium -mediated plant transformation. Mol. Gen. Genet. 220, 245–250.
Haseloff, J., Siemering, K.R., Prasher, D.C. and Hodge, S. (1997). Removal of a cryptic intron and subcellular localization of green fluorescent protein are required to mark transgenic Arabidopsis plants brightly. Proc. Natl. Acad. Sci. USA 94, 2122–2127.
Tinland, B. (1996). The integration of T-DNA into plant genomes. Trends Plant Science 1, 178–184.
Mayerhofer, R., Koncz-Kalman, Z., Nawrath, C., et al. (1991). T-DNA integration: a mode of illegitimate recombination in plants. EMBO J. 10, 697–704.
Hiei, Y., Ohta, S., Komari, T., et al. (1994). Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T- DNA. Plant J. 6, 271–282.
Bundock, P. and Hooykaas, P.J.J. (1996). Integration of Agrobacterium tumefaciens T-DNA in the Saccharomyces cerevisiae genome by illegitimate recombination. Proc. Natl. Acad. Sci. USA 93, 15272–15275.
Deroles, S.C. and Gardner, R.C. (1988). Analysis of the T-DNA structure in a large number of transgenic petunias generated by Agrobacterium -mediated transformation. Plant Mol. Biol. 11, 365–377.
Van der Graaff, E., Den Dulk-Ras, A. and Hooykaas, P.J.J. (1996). Deviating T-DNA transfer from Agrobacterium tumefaciens to plants. Plant Mol. Biol. 31, 677–681.
Koncz, C., Németh, K., Rédei, G.P., et al. (1992). T-DNA international mutagenesis in Arabidopsis. Plant Mol. Biol. 20, 963–976.
Forsthoefel, N.R., Wu, Y., Schulz, B., et al. (1992). T-DNA insertion mutagenesis in Arabidopsis: prospects and perspectives. Aust. J. Plant Physiol 19, 353–366.
Azpiroz-Leehan, R. and Feldmann, K.A. (1997). T-DNA insertion mutagenesis in Arabidopsis: going back and forth. TIG 13, 152–156.
Szekeres, M., Németh, K., Kauschmann, A., et al. (1998). Brassinosteroids rescue the deficiency of CYP90, a cytochrome P450, controlling cell elongation and de-etiolation in Arabidopsis. Cell 85, 171–182.
Lindsey, K., Wei, W., Clarke, M.C., et al.. (1993). Tagging genomic sequences that direct transgene expression by activation of a promoter trap in plants. Transgenic Res. 2, 33–47.
Walden, R., Fritze, K., Hayashi, H., et al. (1994). Activation tagging: a means of isolating genes implicated as playing a role in plant growth and development. Plant Mol. Biol. 26, 1521–1528.
Van der Graaff, E. and Hoogkaas, P.J.J. (1996) Improvements in the transformation of Arabidopsis thaliana C20. leaf-discs by Agrobacterium tumefaciens. Plant Cell Rep. 15, 572–577.
Kakimoto, T. (1996). CK11, a histidine kinase homolog implicated in cytokinin signal transduction. Science 274, 982–985.
Furini, A., Koncz, C., Salamini, F., et al. (1997). High level transcription of a member of a repeated gene family confers dehydration tolerance to callus tissue of Craterostigma plantagineum. EMBO J. 16, 3599–3608.
Hamilton, C.M., Frary, A., Lewis, C., et al. (1996). Stable transfer of intact high molecular weight DNA into plant chromosomes. Proc. Natl. Acad. Sci USA 93, 9975–9979.
Visser, R.G.F., Somhorst, I., Kuipers, G.J., et al. (1991). Inhibition of the expression of the gene for granule-bound starch synthase in potato by antisense constructs. Mol. Gen. Genet. 225, 289–296.
Smith, C.J.S., Watson, C.F., Ray, J., et al. (1988). Antisense RNA inhibition of polygalacturonase gene expression in transgenic tomatoes. Nature 334, 724–726.
Li, H.-Q., Sautter, C., Potrykus, I., et al. (1996). Genetic transformation of cassave (Manihot esculenta Crantz). Nature Biotechnology 14, 736–740.
Hiei, Y., Komari, T. and Kubo, T. (1997). Transformation of rice mediated by Agrobacterium tumefaciens. Plant Mol. Biol. 35, 205–218.
Ishida, Y., Saito, H., Ohta, S., et al. (1996). High efficiency transformation of maize (Zea mays L.) mediated by Agrobacterium tumefaciens. Nature Biotechnology 14, 745–750.
Cheng, M., Fry, J.E., Pang, S., Zhou, H., Hironaka, M., Duncan, D.R., Conner, T.W. and Wan, Y. (1997). Genetic transformation of wheat mediated by Agrobacterium tumefaciens. Plant Physiol. 115, 971–980.
Comai, L., Facciotti, D., Hiatt, W.R., Thompson, G., Rose, R.E. and Stalker, D.M. (1985). Expression in plants of a mutant aroA gene from Salmonella typhimurium confers tolerance to glyphosate. Nature 317, 741–744.
Abel, P.P., Nelson, R.S., De, B., Hoffmann, N., Rogers, S.G., Fraley, R.T. and Beachy, R.N. (1986). Delay of disease development in transgenic plants that express the tobacco mosaic virus coat protein gene. Science 232, 738–743.
Mariani, C., De Beuckeleer, M., Truettner, J., Leemans, J. and Goldberg, R.B. (1990). Induction of male sterility in plants by a chimaeric ribonuclease gene. Nature 347, 737–741.
Van der Krol, A.R., Lenting, P.E., Veenstra, J., Van der Meer, I.M., Koes, R.E., Gerats, A.G.M., Mol, J.N.M. and Stuitje, A.R. (1988). An anti-sense chalcone synthase gene in transgenic plants inhibits flower pigmentation. Nature 333, 866–869.
Vaeck, M., Reynaerts, A., Höfte, H., Jansens, S., De Beuckeleer, M., Dean, C., Zabeau, M., Van Montagu, M. and Leemans, J. (1987). Transgenic plants protected from insect attack. Nature 328, 33–37.
De Luca, V., Marineau, C. and Brisson, N. (1989). Molecular cloning and analysis of cDNA encoding a plant tryptophan decarboxylase: Comparison with animal dopa decarboxylases. Proc. Natl. Acad. Sci. USA 86, 2582–2586.
Pasquali, G., Goddijn, O.J.M., De Waal, A., Verpoorte, R., Schilperoort, R.A., Hoge, J.H.C. and Memelink, J. (1992). Coordinated regulation of two indole alkaloid biosynthetic genes from Catharanthus roseus by auxin and elicitors. Plant Mol. Biol. 18, 1121–1131.
Büschges, R., Hollrichter, K., Panstruga, R., Simons, G., Wolter, M., Frijters, A., Van Daelen, R., Van der Lee, T., Diergaarde, P., Groenendijk, J., Töpsch, S., Vos, P., Salamini, F. and Schulze-Lefert, P. (1997). The barley Mlo gene: a novel control element of plant pathogen resistance. Cell 88, 695–705.
Guivarc’h, A., Caissard, J.-C., Brown, S., Marie, D., Dewitte, W., Van Onckelen, H. and Chriqui, D. (1993). Localization of target cells and improvement of Agrobacterium-mediated transformation efficiency by direct acetosyringone pretreatment of carrot root discs. Protoplasma 174, 10–18.
Villemont, E., Dubois, F., Sangwan, R.S., Vasseur, G., Bourgeois, Y. and Sangwan-Norreel, B.S. (1997). Role of the host cell cycle in the Agrobacterium-mediated genetic transformation of Petunia: evidence of an S-phase control mechanism for T-DNA transfer. Planta 201, 160–172.
Perl, A., Lotan, O., Abu-Abied, M. and Holland, D. (1996). Establishment of an Agrobacteriummediated transformation system for grape (Vitis vinifera L.): The role of antioxidants during grapeAgrobacterium interactions. Nature Biotechnology 14, 624–628.
Chilton, M.-D., Tepfer, D.A., Petit, A., David, C., Casse-Delbart, F. and Tempé, J. (1982). Agrobacterium rhizogenes inserts T-DNA into plants roots. Nature 295, 432–434.
Tepfer, D. (1984). Transformation of several species of higher plants by Agrobacterium rhizogenes: sexual transmission of the transformed genotype and phenotype. Cell 37, 959–967.
Simpson, R.B., Spielmann, A., Margossian, L. and McKnight, T.D. (1986). A disarmed binary vector from Agrobacterium tumefaciens functions in Agrobacterium rhizogenes. Plant Mol. Biol. 6, 403–415.
Ooms, G., Hooykaas, P.J.J., Moolenaar, G. and Schilperoort, R.A. (1981). Crown gall plant tumors of abnormal morphology, induced by Agrobacterium tumefaciens carrying mutated octopine Ti plasmids: analysis of T-DNA functions. Gene 14, 33–50.
Meyer, P., Linn, F., Heidmann, I., Meyer, H., Niedenhof, I. and Saedler, H. (1992). Endogenous and environmental factors influence 35S promoter methylation of a maize Al gene construct in transgenic petunia and its colour phenotype. Mol. Gen. Genet. 231, 345–352.
Van der Graaff, E. and Hooykaas, P.J.J. (1996). Improvements in the transformation of Arabidopsis thaliana C24 leaf-discs by Agrobacterium tumefaciens. Plant Cell Rep. 15, 572–577.
Schmülling, T. and Schell, J. (1993). Transgenic tobacco plants regenerated from leaf disks can be periclinal chimeras. Plant Mol. Biol. 21, 705–708.
Ingelbrecht, I., Breyne, P., Vancompernolle, K., Jacobs, A., Van Montagu, M. and DePicker, A. (1991). Transcriptional interference in transgenic plants. Gene 109, 239–242.
Pröls, F. and Meyer, P. (1992). The methylation patterns of chromosomal integration regions influence gene activity of transferred DNA in Petunia hybrida. Plant J. 2, 465–475.
Breyne, P., Van Montagu, M. and Gheysen, G. (1994). The role of scaffold attachment regions in the structural and functional organization of plant chromatin. Transgenic Res. 3, 195–202.
Mlynarova, L., Jansen, R.C., Conner, A.J., Stiekema, W.J. and Nap, J.-P. (1995). The MAR-mediated reduction in position effect can be uncoupled from copy number-dependent expression in transgenic plants. Plant Cell 7, 599–609.
Kooter, J.M. and Mol, J.N.M. (1993). Trans-inactivation of gene expression in plants. Corr. Opinion Biotechnol. 4, 166–171.
Meyer, P. and Saedler, H. (1996). Homology-dependent gene silencing in plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 47, 23–48.
Haring, M.A., Rommens, C.M.T., Nijkamp, H.J.J. and Hille, J. (1991). The use of transgenic plants to understand transposition mechanisms and to develop transposon tagging strategies. Plant Mol. Biol. 16, 449–461.
Aarts, M.G.M., Dirkse, W.G., Stiekema, W.J. and Pereira, A. (1993). Transposon tagging of a male sterility gene in Arabidopsis. Nature 363, 715–717.
Koes, R., Souer, E., Van Houwelingen, A., Mur, L., Spelt, C., Quattrocchio, F., Wing, J., Oppedijk, B., Ahmed, S., Maes, T., Gerats, T., Hoogeveen, P., Meesters, M., Kloos, D. and Mol, J.N.M. (1995). Targeted gene inactivation in petunia by PCR-based selection of transposon insertion mutants. Proc. Natl. Acad. Sci. USA 92, 8149–8153.
Krysan, P.J., Young, J.C., Tax, F. and Sussman, M.R. (1996). Identification of transferred DNA insertions within Arabidopsis genes involved in signal transduction and ion transport. Proc. Natl. Acad. Sci. USA 93, 8145–8150.
McKinney, E.C., Ali, N., Traut, A., Feldmann, K.A., Belostotsky, D.A., McDowell, J.M. and Meagher, R.B. (1995). Sequence-based identification of T-DNA insertion mutations in Arabidopsis: actin mutants act2–1 and act4–1. Plant J. 8, 613–622.
Morton, R. and Hooykaas, P.J.J. (1995). Gene Replacement. Mol. Breeding 1, 123–132.
Reiss, B., Klemm, M., Kosak, H. and Schell, J. (1996). RecA protein stimulates homologous recombination in plants. Proc. Natl. Acad. Sci. USA 93, 3094–3098.
Schaefer, D.G. and Zrijd, J.-P. (1997). Efficient gene targeting in the moss Physcomitrella patens. Plant J. 11, 1195–1206.
Dale, E.C. and Ow, D.W. (1991). Gene transfer with subsequent removal of the selection gene from the host genome. Proc. Natl. Acad. Sci. USA 88, 10558–10562.
Ebinuma, H., Sugita, K., Matsunaga, E. and Yamakado, M. (1997). Selection of marker-free transgenic plants using the isopentenyl transferase gene. Proc. Natl. Acad. Sci. USA 94, 2117–2121.
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2000 Springer Science+Business Media Dordrecht
About this chapter
Cite this chapter
Hooykaas, P.J.J. (2000). Agrobacterium, A Natural Metabolic Engineer of Plants. In: Verpoorte, R., Alfermann, A.W. (eds) Metabolic Engineering of Plant Secondary Metabolism. Springer, Dordrecht. https://doi.org/10.1007/978-94-015-9423-3_3
Download citation
DOI: https://doi.org/10.1007/978-94-015-9423-3_3
Publisher Name: Springer, Dordrecht
Print ISBN: 978-90-481-5475-3
Online ISBN: 978-94-015-9423-3
eBook Packages: Springer Book Archive