Abstract
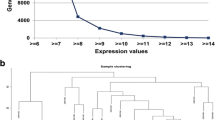
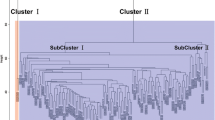
It is important to understand the interaction mechanism among co-expressed and co-regulated genes in stem cell to restrict the abnormal growth of cell tissues (tumor) which may lead to cancer. In this article, differentially co-expressed and co-regulated genes exist in normal stem cells and stem cell derived tumors are identified from sample Bone Marrow microarray data. By performing statistical t-test between sample groups, first we have identified differentially expressed genes (DEG). Then up-regulated (UR) and down-regulated (DR) genes are separated by setting a p-value cutoff at 0.001. After identifying the differentially expressed genes, distinguished co-expressed up-regulated and down-regulated genes are found. Subsequently, we have constructed pair-wise co-expression networks with the co-expressed genes. Finally, we have studied the significance of co-expressed genes with gene ontology (GO) and we have found significant GO-ids. This study is expected to lead to finding of pathways for diseases.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
William, et al, J.B.: Functional recovery of spinal cord injury following application of intralesional bone marrow mononuclear cells embedded in polymer scaffold-two year follow-up in a canine. J. Stem Cell Res. Ther. 1(3) 2011
Hanna, J., et al.: Treatment of sickle cell anemia mouse model with iPS cells generated from autologous skin. Science 318(5858), 1920–1923 (2007)
Saha, K., Jaenisch, R.: Technical challenges in using human induced pluripotent stem cells to model disease. Cell Stem Cell 5(6), 584–595 (2009)
Jiang, Y., et al.: Pluripotency of mesenchymal stem cells derived from adult marrow. Nature 418(6893), 41–49 (2002)
Vaishali, K., Vinayababu, A.: Application of microarray technology and soft computing in cancer biology: a review. Int. J. Biometr. Bioinform. 5(4), 225–233 (2011)
Chen, J.J.: Key aspects of analyzing microarray gene-expression data. Pharmacogenomics 8(5), 473–482 (2007)
Allen, et al. J.D.: Comparing statistical methods for constructing large scale gene networks. PLoS ONE 7(1) (2012)
Vardhanabhuti, S., et al.: A comparison of statistical tests for detecting differential expression using affymetrix oligonucleotide microarrays. OMICS 10(4), 555–566 (2006)
Benjamini, Y., Hochberg, Y.: Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. Roy. Stat. Soc. 57(1), 289–300 (1995)
Bandyopadhyay, S., Mallik, S., Mukhopadhyay, A.: A survey and comparative study of statistical tests for identifying differential expression from microarray data. IEEE/ACM Trans. Comput. Biol. Bioinform. 11(1), 95–115 (2014)
Storey, J.D., Tibshirani, R.: Statistical significance for genome-wide studies. Proc. Natl. Acad. Sci. 100(16), 9440–9445 (2003)
Storey, J.D., Taylor, J.E., Siegmund, D.: Strong control conservative point estimation and simultaneous conservative consistency of false discovery rates: a unified approach. J. Roy. Stat. Soc. 66(1), 187–205 (2004)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer India
About this paper
Cite this paper
Biswas, P., Barman, B., Mukhopadhyay, A. (2015). Construction of Co-expression and Co-regulation Network with Differentially Expressed Genes in Bone Marrow Stem Cell Microarray Data. In: Mandal, J., Satapathy, S., Kumar Sanyal, M., Sarkar, P., Mukhopadhyay, A. (eds) Information Systems Design and Intelligent Applications. Advances in Intelligent Systems and Computing, vol 339. Springer, New Delhi. https://doi.org/10.1007/978-81-322-2250-7_76
Download citation
DOI: https://doi.org/10.1007/978-81-322-2250-7_76
Published:
Publisher Name: Springer, New Delhi
Print ISBN: 978-81-322-2249-1
Online ISBN: 978-81-322-2250-7
eBook Packages: EngineeringEngineering (R0)