Abstract
Nodal is a TGF-β family member ligand that is critical for early embryonic patterning in vertebrates. Nodal signaling functions through core TGF-β receptors to activate a Smad transcription factor signaling cascade. However, unlike other TGF-β ligands, Nodal signaling requires an additional co-receptor of the EGF-CFC family to activate intracellular signaling. Nodal signaling is also subject to extensive negative regulation by Lefty and other factors. Work in numerous model organisms, including mouse, chicken, and zebrafish, established that Nodal signaling plays an essential role during germ layer formation, anterior-posterior axis patterning, and left-right axis determination. Incomplete or delayed loss of Nodal signaling results in defective organogenesis and birth defects, including congenital heart defects, and clinical studies have linked aberrant Nodal signaling in humans with many common congenital malformations, including congenital heart defects. Congenital heart defects associated with disrupted Nodal signaling in mammals include those that arise due to global defects in left-right patterning of the embryo, such as heterotaxy. Other Nodal-associated heart defects appear to occur as more subtle isolated malformations of the great arteries and atrioventricular septum, which may not be related to overall perturbations in laterality. A more detailed understanding of the Nodal signaling pathway and its targets in the heart is required to more fully understand the etiology of Nodal signaling pathway-associated congenital heart defects.
Note: This chapter is also related to Part II.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
The Nodal signaling pathway is critical for early embryonic patterning in all vertebrates. Nodal is a secreted signaling molecule that belongs to the transforming growth factor-β (TGF-β) family and functions primarily to activate downstream signaling through a receptor-mediated response [1–3]. The Nodal pathway plays a critical role in mesoderm specification and anterior-posterior patterning during gastrulation and is also essential for establishing the left-right axis in the developing embryo [4–7]. Changes in Nodal expression or dosage can disrupt left-right patterning and result in a range of congenital defects that affect development of the forebrain, the craniofacial skeleton, and several other organs, including the heart.
2 The Nodal Signaling Pathway
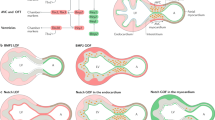
The Nodal signaling pathway shares many similarities with other TGF-β signaling pathways in that it utilizes core Smad-dependent signaling components (Fig. 24.1). Nodal is secreted extracellularly as a proprotein homodimer like other TGF-β ligands [8]. Once cleaved into the mature ligand, the Nodal homodimer binds tightly to a TGF-β receptor heterodimer consisting of both Type I and Type II receptors [9]. Nodal ligand binding causes the constitutively active Type II receptor serine/threonine kinases to associate with the inactive Type I receptor kinases and leads to phosphorylation and the subsequent dissociation of R-Smad from the TGF-β receptor [10]. Following formation of a trimeric complex composed of two R-Smads and the common partner Smad, Smad4, the Smad oligomer translocates to the nucleus where it regulates gene expression through direct and indirect DNA binding (Fig. 24.1) [1, 11, 12].
The Nodal signaling pathway. The Nodal ligand binds to a dimer of the TGF-β Type I and Type II receptors. In association with an EGF-CFC co-receptor (Cripto or Cryptic), Nodal activates the receptor complex causing the phosphorylation of either Smad2 or Smad3 followed by oligomerization with the common partner Smad4. The Smad2/Smad3-Smad4 complex then translocates to the nucleus where it binds to DNA with the transcriptional cofactor FoxH1 or Mixer, leading to transcription of downstream target genes. The Nodal signaling pathway is negatively regulated by proteins such as Lefty, which can bind to either EGF-CFC or the Nodal dimer to prevent activation of the receptor complex, or TGIF1, which recruits histone deacetylases to Smad2/Smad3 and represses transcriptional activation
The Nodal signaling pathway has important distinctions from other TGF-β-mediated signaling pathways. Nodal can only activate TGF-β receptor signaling in the presence of an EGF-CFC protein (Fig. 24.1). There are two EGF-CFC proteins, Cripto and Cryptic, which function as co-receptors for Nodal [13]. Cripto and Cryptic are extracellular proteins that contain an epidermal growth factor-like motif and a novel cystine-rich domain named the CFC [13]. EGF-CFC proteins function primarily by binding to the Type II TGF-β receptor thro ugh the CFC domain and by binding to Nodal through the EGF domain [14].
In response to Nodal signaling, the Smad complex cooperates specifically with FoxH1, a winged helix transcription factor, or Mixer, a member of the Mix subclass of homeodomain proteins [15, 16]. These cofactors are critical for Nodal-dependent downstream gene activation and act to stabilize Smad-DNA interactions, since Smads have relatively weak DNA-binding affinity [17, 18]. FoxH1 and Mixer recruit the Smad complex to promoter and enhancer elements and help to establish temporal and spatial regulation of Nodal-dependent target genes.
In addition, the Nodal signaling pathway is subject to specific negative regulation not found for other TGF-β family members. Proteins of the Lefty family, specifically Lefty1 and Lefty2, inhibit Nodal-dependent activation of TGF-β receptors by interacting directly with the Nodal ligand and preventing binding to the TGF-β receptor heterodimer and by binding to the EGF-CFC Nodal co-receptors Cripto and Cryptic and preventing their association with the TGF-β receptor complex (Fig. 24.1) [19–21].
3 Requirement for Nodal in Development
Nodal signaling is critical for the patterning of the developing embryo [4, 22]. Loss-of-function studies in vertebrate model organisms indicate that Nodal signaling is first required during gastrulation. Germline loss of Nodal in mice results in severe patterning and differentiation defects and embryonic lethality due to a failure to induce the primitive streak from the ectoderm and to disrupted specification of mesoderm and endoderm from the epiblast [5]. Additionally, loss of Nodal in mice results in impaired anterior-posterior axis formation due to the lack of anterior visceral endoderm formation [22]. The requirement of Nodal signaling for early embryonic pattern formation was further highlighted by loss-of-function studies of the EGF-CFC Nodal co-receptor Cripto. Inactivation of Tdgf1 , the gene encoding Cripto, in mice results in a phenocopy of early Nodal defects, including lethality shortly after gastrulation. Tdgf1 mutants lack a primitive streak, fail to form embryonic mesoderm, and exhibit anterior-posterior axis defects [6]. The one-eyed pinhead (oep) mutation in zebrafish results in a complete loss of function of the fish ortholog of the Nodal co-receptor Cripto [23]. These mutants exhibit a phenocopy of the early Nodal defects seen in mice, including an absence of mesoderm and anterior-posterior axis abnormalities [23]. Together, both animal models establish that EGF-CFC proteins are required for Nodal signaling and support an early requirement for the Nodal pathway in embryo morphogenesis.
Following gastrulation, Nodal signaling is indispensable for the establishment of left-right asymmetry. Conditional deletion of Nodal in the lateral plate mesoderm in mice circumvents the early requirement for Nodal signaling during gastrulation and results in heterotaxy, a condition characterized by left-right ambiguity of thoracic and abdominal visceral organs [24]. These mice exhibit transposition of the great arteries of the heart, right-sided isomerism of the lungs, and right-sided stomach [25]. Similarly, germline deletion of Cfc1, the gene encoding Cryptic, results in left-right laterality defects with mutants exhibiting heterotaxy [26].
In addition to playing a role in the establishment of laterality following gastrulation, Nodal signaling is also required for midline patterning of the ventral forebrain [27]. Zebrafish mutants for the Nodal ligands and for the ortholog of Cripto result in holoprosencephaly, a condition where bifurcation of the ventral forebrain fails to occur and results in fusion of the two brain hemispheres [28]. Similarly, mice heterozygous for germline knockout alleles of both Nodal and Smad2 have cyclopia, a rare and severe form of holoprosencephaly [29], further supporting the relationship between Nodal signaling and forebrain development. Mechanistically, this phenotype is thought to occur due to the patterning of Sonic Hedgehog (Shh) expression in the forebrain by Nodal [30, 31].
4 Congenital Heart Defects Associated with Perturbations in Nodal Signaling
It is perhaps not surprising to find that the Nodal signaling pathway is associated with pathogenesis in humans, given its critical role in patt erning during embryonic development. Nodal signaling was first linked to human congenital defects through the identification of mutations associated with left-right laterality defects. These include mutations in genes encoding Lefty family members, the EGF-CFC co-receptor Cryptic, and the Type II TGF-β receptors [32–34]. In addition, mutations in genes encoding transcriptional inhibitors of Smad2, such as TGIF1, are associated with holoprosencephaly, a defect strongly associated with disrupted Nodal signaling [30, 35]. Interestingly, these human pathologies are similar to the defects observed in animal models with defective (but not incomplete) Nodal signaling.
Congenital heart defects have also been linked to aberrant Nodal signaling (Table 24.1) [36]. Loss-of-function mutations in genes encoding numerous Nodal signaling components, including Nodal, Cripto, Cryptic, and FoxH1, have been identified in patients with heart defects [37, 38]. The spectrum of heart defects in these patients can be roughly grouped into two broadly defined classes: (1) those that occur as a result of overall isomerism or heterotaxy and (2) those that occur as isolated congenital heart defects. The isomerisms of the heart can be classified as situs inversus totalis, a complete mirror image of the visceral organs of the body including the heart, or situs inversus ambiguous, where the abdominal and visceral organs are distributed abnormally and randomly in a condition more commonly called heterotaxy [39]. Both of these conditions can be linked to aberrant Nodal signaling [40]. Heterotaxy often results in complex congenital heart defects [41, 42]. These defects include levo-transposition of the great arteries (l-TGA) and atrial isomerism [36, 39]. The feature characteristics of l-TGA are improper positioning of the aorta and the pulmonary artery such that the arteries are switched in conjunction with the ventricles such that they still have the normal relationship between the ventricles and the arteries [43].
Isolated congenital heart defects associated with Nodal mutations can result in structural and functional abnormalities that appear to be independent of overall left-right ambiguity. These isolated defects include dextro-transposition of the great arteries (d-TGA), double outlet right ventricle (DORV), tetralogy of Fallot, and isolated ventricular septal defects [37, 38]. Unlike l-TGA, which results in a proper alignment of the arteries with respect to the ventricles, d-TGA results in a switching of aorta and the pulmonary artery such that the pulmonary artery connects to the left ventricle and the aorta emanates from the right ventricle [43]. DORV, as the name implies, occurs when the aorta and the pulmonary artery connect to the right ventricle. Tetralogy of Fallot is essentially a milder form of DORV in which the aorta overrides the ventricular septum and empties blood from both ventricles [44]. There is also pulmonary artery stenosis and a hypertrophic right ventricle secondary to pulmonary artery blockage associated with tetralogy of Fallot [44]. These defects appear to be independent of overall laterality, although the mechanistic basis for why these defects occur when Nodal signaling is disrupted is unclear. Interestingly, Nodal expression occurs well prior to the patterning and final alignment of the aorta and pulmonary artery [40]. Furthermore, the temporal expression pattern of Nodal observed during development is tightly regulated within a narrow developmental window due, at least in part, to the extensive negative feedback by Lefty proteins and other factors. The diverse nature of heterotaxy in humans suggests that some isolated congenital heart defects associated with perturbed Nodal signaling may still be secondary to overall laterality defects.
A role for Nodal signaling in isolated congenital heart defects is also supported by studies in mice in which subtle perturbations of Nodal signaling result in less severe defects (Table 24.1), suggesting that the Nodal ligand functions in a dosage-dependent manner. For example, deletion of an intronic enhancer of Nodal resulted in decreased expression of Nodal in the lateral plate mesoderm. These mouse embryos developed laterality defects that were less severe than the heterotaxy observed in Nodal conditional knockouts where Nodal expression in lateral plate mesoderm was completely abolished [25, 45].
Consistent with Nodal function in left-right patterning, Nodal-dependent target genes are also critical for left-right patterning and heart development [21]. Pitx2 is perhaps the best-described target gene of Nodal signaling and is expressed asymmetrically on the left side after gastrulation [46]. Misexpression of Pitx2 on the embryonic right side in mice results in heterotaxia with conditions such as aberrant heart looping, cardiac isomerism, and visceral organ laterality defects [46]. In the mouse, germline loss of function of Pitx2 results in left-right asymmetry defects in specific organs, such as the lung [47]. Interestingly, Pitx2-null and isoform-specific Pitx2c-null embryos undergo normal heart looping but have a subset of congenital cardiovascular anomalies such as DORV and ventricular and atrial septal defects [47, 48]. These observations suggest that Pitx2 functions in heart development after left-right determination and that other Nodal-dependent target genes may be required for cardiac laterality. Together, these observations suggest that other unappreciated Nodal-dependent target genes are involved in the establishment of left-right identity and cardiac development. A more detailed elucidation of this fundamental pathway, including target genes in the cardiac mesoderm, is required to more fully understand the role of Nodal signaling in heart development and in congenital heart defects.
References
Massague J, Seoane J, Wotton D. Smad transcription factors. Genes Dev. 2005;19:2783–810.
Schier AF. Nodal morphogens. Cold Spring Harb Perspect Biol. 2009;1:a003459.
Shen MM. Nodal signaling: developmental roles and regulation. Development. 2007;134:1023–34.
Brennan J, Norris DP, Robertson EJ. Nodal activity in the node governs left-right asymmetry. Genes Dev. 2002;16:2339–44.
Conlon FL, Lyons KM, Takaesu N, Barth KS, Kispert A, Herrmann B, et al. A primary requirement for nodal in the formation and maintenance of the primitive streak in the mouse. Development. 1994;120:1919–28.
Ding J, Yang L, Yan YT, Chen A, Desai N, Wynshaw-Boris A, et al. Cripto is required for correct orientation of the anterior-posterior axis in the mouse embryo. Nature. 1998;395:702–7.
Perea-Gomez A, Vella FD, Shawlot W, Oulad-Abdelghani M, Chazaud C, Meno C, et al. Nodal antagonists in the anterior visceral endoderm prevent the formation of multiple primitive streaks. Dev Cell. 2002;3:745–56.
Walton KL, Makanji Y, Harrison CA. New insights into the mechanisms of activin action and inhibition. Mol Cell Endocrinol. 2012;359:2–12.
Reissmann E, Jornvall H, Blokzijl A, Andersson O, Chang C, Minchiotti G, et al. The orphan receptor ALK7 and the activin receptor ALK4 mediate signaling by nodal proteins during vertebrate development. Genes Dev. 2001;15:2010–22.
Moustakas A, Heldin CH. The regulation of TGFbeta signal transduction. Development. 2009;136:3699–714.
Ali MH, Imperiali B. Protein oligomerization: how and why. Bioorg Med Chem. 2005;13:5013–20.
Moustakas A. Smad signalling network. J Cell Sci. 2002;115:3355–6.
Shen MM, Schier AF. The EGF-CFC gene family in vertebrate development. Trends Genet. 2002;16:303–9.
Yeo C, Whitman M. Nodal signals to Smads through Cripto-dependent and Cripto-independent mechanisms. Mol Cell. 2001;7:949–57.
Kunwar PS, Zimmerman S, Bennett JT, Chen Y, Whitman M, Schier AF. Mixer/Bon and FoxH1/Sur have overlapping and divergent roles in Nodal signaling and mesendoderm induction. Development. 2003;130:5589–99.
Yamamoto M, Meno C, Sakai Y, Shiratori H, Mochida K, Ikawa Y, et al. The transcription factor FoxH1 (FAST) mediates Nodal signaling during anterior-posterior patterning and node formation in the mouse. Genes Dev. 2001;15:1242–56.
Germain S, Howell M, Esslemont GM, Hill CS. Homeodomain and winged-helix transcription factors recruit activated Smads to distinct promoter elements via a common Smad interaction motif. Genes Dev. 2000;14:435–51.
Osada SI, Saijoh Y, Frisch A, Yeo CY, Adachi H, Watanabe M, et al. Activin/nodal responsiveness and asymmetric expression of a Xenopus nodal-related gene converge on a FAST-regulated module in intron 1. Development. 2000;127:2503–14.
Chen C, Shen MM. Two modes by which Lefty proteins inhibit nodal signaling. Curr Biol. 2004;14:618–24.
Cheng SK, Olale F, Brivanlou AH, Schier AF. Lefty blocks a subset of TGFbeta signals by antagonizing EGF-CFC coreceptors. PLoS Biol. 2004;2:E30.
Shiratori H, Hamada H. The left-right axis in the mouse: from origin to morphology. Development. 2006;133:2095–104.
Brennan J, Lu CC, Norris DP, Rodriguez TA, Beddington RS, Robertson EJ. Nodal signalling in the epiblast patterns the early mouse embryo. Nature. 2001;411:965–9.
Gritsman K, Zhang J, Cheng S, Heckscher E, Talbot WS, Schier AF. The EGF-CFC protein one-eyed pinhead is essential for nodal signaling. Cell. 1999;97:121–32.
Lin AE, Ticho BS, Houde K, Westgate MN, Holmes LB. Heterotaxy: associated conditions and hospital-based prevalence in newborns. Genet Med. 2000;2:157–72.
Kumar A, Lualdi M, Lewandoski M, Kuehn MR. Broad mesodermal and endodermal deletion of Nodal at postgastrulation stages results solely in left/right axial defects. Dev Dyn. 2008;237:3591–601.
Gaio U, Schweickert A, Fischer A, Garratt AN, Muller T, Ozcelik C, et al. A role of the cryptic gene in the correct establishment of the left-right axis. Curr Biol. 1999;9:1339–42.
Lowe LA, Yamada S, Kuehn MR. Genetic dissection of nodal function in patterning the mouse embryo. Development. 2001;128:1831–43.
Hayhurst M, McConnell SK. Mouse models of holoprosencephaly. Curr Opin Neurol. 2003;16:135–41.
Nomura M, Li E. Smad2 role in mesoderm formation, left-right patterning and craniofacial development. Nature. 1998;393:786–90.
Gripp KW, Wotton D, Edwards MC, Roessler E, Ades L, Meinecke P, et al. Mutations in TGIF cause holoprosencephaly and link NODAL signalling to human neural axis determination. Nat Genet. 2000;25:205–8.
Rohr KB, Barth KA, Varga ZM, Wilson SW. The nodal pathway acts upstream of hedgehog signaling to specify ventral telencephalic identity. Neuron. 2001;29:341–51.
Bamford RN, Roessler E, Burdine RD, Saplakoglu U, dela Cruz J, Splitt M, et al. Loss-of-function mutations in the EGF-CFC gene CFC1 are associated with human left-right laterality defects. Nat Genet. 2000;26:365–9.
Kosaki K, Bassi MT, Kosaki R, Lewin M, Belmont J, Schauer G, et al. Characterization and mutation analysis of human LEFTY A and LEFTY B, homologues of murine genes implicated in left-right axis development. Am J Hum Genet. 1999;64:712–21.
Kosaki R, Gebbia M, Kosaki K, Lewin M, Bowers P, Towbin JA, et al. Left-right axis malformations associated with mutations in ACVR2B, the gene for human activin receptor type IIB. Am J Med Genet. 1999;82:70–6.
Massague J, Blain SW, Lo RS. TGFbeta signaling in growth control, cancer, and heritable disorders. Cell. 2000;103:295–309.
Mohapatra B, Casey B, Li H, Ho-Dawson T, Smith L, Fernbach SD, et al. Identification and functional characterization of NODAL rare variants in heterotaxy and isolated cardiovascular malformations. Hum Mol Genet. 2009;18:861–71.
Roessler E, Ouspenskaia MV, Karkera JD, Velez JI, Kantipong A, Lacbawan F, et al. Reduced NODAL signaling strength via mutation of several pathway members including FOXH1 is linked to human heart defects and holoprosencephaly. Am J Hum Genet. 2008;83:18–29.
Roessler E, Pei W, Ouspenskaia MV, Karkera JD, Velez JI, Banerjee-Basu S, et al. Cumulative ligand activity of NODAL mutations and modifiers are linked to human heart defects and holoprosencephaly. Mol Genet Metab. 2009;98:225–34.
Bisgrove BW, Morelli SH, Yost HJ. Genetics of human laterality disorders: insights from vertebrate model systems. Annu Rev Genomics Hum Genet. 2003;4:1–32.
Lowe LA, Supp DM, Sampath K, Yokoyama T, Wright CV, Potter SS, et al. Conserved left-right asymmetry of nodal expression and alterations in murine situs inversus. Nature. 1996;381:158–61.
Shiraishi I, Ichikawa H. Human heterotaxy syndrome - from molecular genetics to clinical features, management, and prognosis. Circ J. 2012;76:2066–75.
Bowers PN, Brueckner M, Yost HJ. The genetics of left-right development and heterotaxia. Semin Perinatol. 1996;20:577–88.
Warnes CA. Transposition of the great arteries. Circulation. 2006;114:2699–709.
Fox D, Devendra GP, Hart SA, Krasuski RA. When ‘blue babies’ grow up: what you need to know about tetralogy of Fallot. Cleve Clin J Med. 2010;77:821–8.
Norris DP, Brennan J, Bikoff EK, Robertson EJ. The Foxh1-dependent autoregulatory enhancer controls the level of Nodal signals in the mouse embryo. Development. 2002;129:3455–68.
Logan M, Pagan-Westphal SM, Smith DM, Paganessi L, Tabin CJ. The transcription factor Pitx2 mediates situs-specific morphogenesis in response to left-right asymmetric signals. Cell. 1998;94:307–17.
Kitamura K, Miura H, Miyagawa-Tomita S, Yanazawa M, Katoh-Fukui Y, Suzuki R, et al. Mouse Pitx2 deficiency leads to anomalies of the ventral body wall, heart, extra- and periocular mesoderm and right pulmonary isomerism. Development. 1999;126:5749–58.
Liu C, Liu W, Palie J, Lu MF, Brown NA, Martin JF. Pitx2c patterns anterior myocardium and aortic arch vessels and is required for local cell movement into atrioventricular cushions. Development. 2002;129:5081–91.
Goldmuntz E, Bamford R, Karkera JD, de la Cruz J, Roessler E, Muenke M. CFC1 mutations in patients with transposition of the great arteries and double-outlet right ventricle. Am J Hum Genet. 2002;70:776–80.
de la Cruz JM, Bamford RN, Burdine RD, Roessler E, Barkovich AJ, Donnai D, et al. A loss-of-function mutation in the CFC domain of TDGF1 is associated with human forebrain defects. Hum Genet. 2002;110:422–8.
Wang B, Yan J, Peng Z, Wang J, Liu S, Xie X, et al. Teratocarcinoma-derived growth factor 1 (TDGF1) sequence variants in patients with congenital heart defect. Int J Cardiol. 2011;146:225–7.
Chu J, Ding J, Jeays-Ward K, Price SM, Placzek M, Shen MM. Non-cell-autonomous role for Cripto in axial midline formation during vertebrate embryogenesis. Development. 2005;132:5539–51.
Zaidi S, Choi M, Wakimoto H, Ma L, Jiang J, Overton JD, et al. De novo mutations in histone-modifying genes in congenital heart disease. Nature. 2013;498:220–3.
Oh SP, Li E. The signaling pathway mediated by the type IIB activin receptor controls axial patterning and lateral asymmetry in the mouse. Genes Dev. 1997;11:1812–26.
Wang B, Yan J, Mi R, Zhou S, Xie X, Wang J, et al. Forkhead box H1 (FOXH1) sequence variants in ventricular septal defect. Int J Cardiol. 2010;145:83–5.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is distributed under the terms of the Creative Commons Attribution-Noncommercial 2.5 License (http://creativecommons.org/licenses/by-nc/2.5/), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited. The images or other third party material in this chapter are included in the work's Creative Commons license, unless indicated otherwise in the credit line; if such material is not included in the work's Creative Commons license and the respective action is not permitted by statutory regulation, users will need to obtain permission from the license holder to duplicate, adapt or reproduce the material.
Copyright information
© 2016 The Author(s)
About this chapter
Cite this chapter
Barnes, R.M., Black, B.L. (2016). Nodal Signaling and Congenital Heart Defects. In: Nakanishi, T., Markwald, R., Baldwin, H., Keller, B., Srivastava, D., Yamagishi, H. (eds) Etiology and Morphogenesis of Congenital Heart Disease. Springer, Tokyo. https://doi.org/10.1007/978-4-431-54628-3_24
Download citation
DOI: https://doi.org/10.1007/978-4-431-54628-3_24
Published:
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-54627-6
Online ISBN: 978-4-431-54628-3
eBook Packages: MedicineMedicine (R0)