Abstract
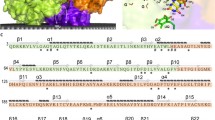
Reductive dehalogenases are a class of corrinoid and [4Fe–4S] cluster-dependent enzymes that are arguably key to organohalide respiration. Recently, the first crystal structures for these enzymes were reported, including one representative of both a respiratory as well as a nonrespiratory catabolic reductive dehalogenase. The comparison made between both structures establishes two highly conserved elements: the configuration of the redox chain within the protein and the Tyr–Lys/Arg active site dyad involved in proton transfer to the substrate. In contrast, the substrate binding elements are highly distinct. These insights serve to guide further study of RdhA structure–function relationships.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Adrian L, Szewzyk U, Wecke J, Gorisch H (2000) Bacterial dehalorespiration with chlorinated benzenes. Nature 408(6812):580–583. doi:10.1038/35046063
Banerjee R, Ragsdale SW (2003) The many faces of vitamin B-12: catalysis by cobalamin-dependent enzymes. Annu Rev Biochem 72:209–247. doi:10.1146/annurev.biochem.72.121801.161828
Bommer M, Kunze C, Fesseler J, Schubert T, Diekert G, Dobbek H (2014) Structural basis for organohalide respiration. Science 346(6208):455–458. doi:10.1126/science.1258118
Brown KL (2005) Chemistry and enzymology of vitamin B-12. Chem Rev 105(6):2075–2149
Bunge M, Adrian L, Kraus A, Opel M, Lorenz WG, Andreesen JR, Gorisch H, Lechner U (2003) Reductive dehalogenation of chlorinated dioxins by an anaerobic bacterium. Nature 421(6921):357–360. doi:10.1038/nature01237
Chen K, Huang LL, Xu CF, Liu XM, He J, Zinder SH, Li SP, Jiang JD (2013) Molecular characterization of the enzymes involved in the degradation of a brominated aromatic herbicide. Mol Microbiol 89(6):1121–1139
Futagami T, Goto M, Furukawa K (2008) Biochemical and genetic bases of dehalorespiration. Chem Rec 8(1):1–12. doi:10.1002/tcr.20134
Gribble GW (1998) Naturally occurring organohalogen compounds. Acc Chem Res 31(3):141–152. doi:10.1021/ar9701777
Holliger C, Wohlfarth G, Diekert G (1998) Reductive dechlorination in the energy metabolism of anaerobic bacteria. FEMS Microbiol Rev 22(5):383–398. doi:10.1111/j.1574-6976.1998.tb00377.x
Holscher T, Krajmalnik-Brown R, Ritalahti KM, Von Wintzingerode F, Gorisch H, Loffler FE, Adrian L (2004) Multiple nonidentical reductive-dehalogenase-homologous genes are common in dehalococcoides. Appl Environ Microbiol 70(9):5290–5297. doi:10.1128/AEM.70.9.5290-5297.2004
Hug LA, Edwards EA (2013) Diversity of reductive dehalogenase genes from environmental samples and enrichment cultures identified with degenerate primer PCR screens. Front Microbiol 4:341. doi:10.3389/fmicb.2013.00341
Hug LA, Maphosa F, Leys D, Loffler FE, Smidt H, Edwards EA, Adrian L (2013) Overview of organohalide-respiring bacteria and a proposal for a classification system for reductive dehalogenases. Philos Trans R Soc B 368(1616):2012. doi:10.1098/rstb.2012.0322 (ARTN0322)
John M, Schmitz RP, Westermann M, Richter W, Diekert G (2006) Growth substrate dependent localization of tetrachloroethene reductive dehalogenase in Sulfurospirillum multivorans. Arch Microbiol 186(2):99–106. doi:10.1007/s00203-006-0125-5
Koutmos M, Gherasim C, Smith JL, Banerjee R (2011) Structural basis of multifunctionality in a vitamin B-12-processing enzyme. J Biol Chem 286(34):29780–29787
Krasotkina J, Walters T, Maruya KA, Ragsdale SW (2001) Characterization of the B12- and iron–sulfur-containing reductive dehalogenase from desulfitobacterium chlororespirans. J Biol Chem 276(44):40991–40997. doi:10.1074/jbc.M106217200
Krautler B, Fieber W, Ostermann S, Fasching M, Ongania KH, Gruber K, Kratky C, Mikl C (2003) The cofactor of tetrachloroethene reductive dehalogenase of Dehalospirillum multivorans is norpseudo-B-12, a new type of a natural corrinoid. Helv Chim Acta 86(11):3698–3716. doi:10.1002/hlca.200390313
Lai QL, Li GZ, Shao ZZ (2012) Genome Sequence of Nitratireductor pacificus type strain pht-3B. J Bacteriol 194(24):6958
Mac Nelly A, Kai M, Svatos A, Diekert G, Schubert T (2014) Functional heterologous production of reductive dehalogenases from Desulfitobacterium hafniense strains. Appl Environ Microbiol 80(14):4313–4322. doi:10.1128/AEM.00881-14
Maillard J, Schumacher W, Vazquez F, Regeard C, Hagen WR, Holliger C (2003) Characterization of the corrinoid iron–sulfur protein tetrachloroethene reductive dehalogenase of Dehalobacter restrictus. Appl Environ Microbiol 69(8):4628–4638
Munro AW, Girvan HM, McLean KJ (2007) Variations on a (t)heme—novel mechanisms, redox partners and catalytic functions in the cytochrome P450 superfamily. Nat Prod Rep 24(3):585–609
Neumann A, Wohlfarth G, Diekert G (1996) Purification and characterization of tetrachloroethene reductive dehalogenase from Dehalospirillum multivorans. J Biol Chem 271(28):16515–16519
Neumann A, Siebert A, Trescher T, Reinhardt S, Wohlfarth G, Diekert G (2002) Tetrachloroethene reductive dehalogenase of Dehalospirillum multivorans: substrate specificity of the native enzyme and its corrinoid cofactor. Arch Microbiol 177(5):420–426. doi:10.1007/s00203-002-0409-3
Oberg G (2002) The natural chlorine cycle–fitting the scattered pieces. Appl Microbiol Biotechnol 58(5):565–581. doi:10.1007/s00253-001-0895-2
Parthasarathy A, Stich TA, Lohner ST, Lesnefsky A, Britt RD, Spormann AM (2015) Biochemical and EPR-spectroscopic investigation into heterologously expressed vinyl chloride reductive dehalogenase (VcrA) from Dehalococcoides mccartyi strain VS. J Am Chem Soc 137(10):3525–3532. doi:10.1021/ja511653d
Payne KA, Quezada CP, Fisher K, Dunstan MS, Collins FA, Sjuts H, Levy C, Hay S, Rigby SE, Leys D (2015) Reductive dehalogenase structure suggests a mechanism for B12-dependent dehalogenation. Nature 517(7535):513–516. doi:10.1038/nature13901
Richardson RE (2013) Genomic insights into organohalide respiration. Curr Opin Biotechnol 24(3):498–505. doi:10.1016/j.copbio.2013.02.014
Sjuts H, Fisher K, Dunstan MS, Rigby SE, Leys D (2012) Heterologous expression, purification and cofactor reconstitution of the reductive dehalogenase PceA from Dehalobacter restrictus. Protein Expr Purif 85(2):224–229. doi:10.1016/j.pep.2012.08.007
Smidt H, de Vos WM (2004) Anaerobic microbial dehalogenation. Annu Rev Microbiol 58:43–73
Studer A, Vuilleumier S, Leisinger T (1999) Properties of the methylcobalamin:H4folate methyltransferase involved in chloromethane utilization by Methylobacterium sp. strain CM4. Eur J Biochem 264(1):242–249
Studer A, Stupperich E, Vuilleumier S, Leisinger T (2001) Chloromethane: tetrahydrofolate methyl transfer by two proteins from Methylobacterium chloromethanicum strain CM4. Eur J Biochem 268(10):2931–2938. doi:10.1046/j.1432-1327.2001.02182.x
van de Pas BA, Smidt H, Hagen WR, van der Oost J, Schraa G, Stams AJ, de Vos WM (1999) Purification and molecular characterization of ortho-chlorophenol reductive dehalogenase, a key enzyme of halorespiration in Desulfitobacterium dehalogenans. J Biol Chem 274(29):20287–20292
Wohlfarth G, Diekert G (1997) Anaerobic dehalogenases. Curr Opin Biotechnol 8(3):290–295
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Dobbek, H., Leys, D. (2016). Insights into Reductive Dehalogenase Function Obtained from Crystal Structures. In: Adrian, L., Löffler, F. (eds) Organohalide-Respiring Bacteria. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-662-49875-0_20
Download citation
DOI: https://doi.org/10.1007/978-3-662-49875-0_20
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-662-49873-6
Online ISBN: 978-3-662-49875-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)