Abstract
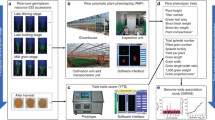
Asian rice is a staple food in Indonesia and worldwide, and its production is essential to food security. Cataloging and linking genetic variation in Asian rice to important traits, such as quality and yield, is needed in developing superior varieties of rice. We develop a bioinformatics workflow for quality control and data analysis of genetic and trait data for a diversity panel of 467 rice varieties found in Indonesia. The bioinformatics workflow operates using a back-end relational database for data storage and retrieval. Quality control and data analysis procedures are implemented and automated using the whole genome data analysis toolset, PLINK, and the [R] statistical computing language. The 467 rice varieties were genotyped using a custom array (717,312 genotypes total) and phenotyped for 12 traits in four locations in Indonesia across multiple seasons. We applied our bioinformatics workflow to these data and present prototype genome-wide association results for a continuous trait - days to flowering. Two genetic variants, located on chromosome 4 and 12 of the rice genome, showed evidence for association in these data. We conclude by outlining extensions to the workflow and plans for more sophisticated statistical analyses.
Chapter PDF
Similar content being viewed by others
Keywords
References
International Rice Genome Sequencing Project: The map-based sequence of the rice genome. Nature 436(7052), 793–800 (2005)
Zhao, K., Tung, C.W., Eizenga, G.C., Wright, M.H., Ali, M.L., Price, A.H., Norton, G.J., Islam, M.R., Reynolds, A., Mezey, J., McClung, A.M., Bustamante, C.D., McCouch, S.: Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat. Commun. 2, 467 (2011)
Purcell, S., Neale, B., Todd-Brown, K., Thomas, L., Ferreira, M.A., Bender, D., Maller, J., Sklar, P., de Bakker, P.I., Daly, M.J., Sham, P.: PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81(3), 559–575 (2007)
R Development Core Team.: R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria, http://www.R-project.org/ , ISBN 3-900051-07-0
Price, A.L., Patterson, N.J., Plenge, R.M., Weinblatt, M.E., Shadick, N.A., Reich, D.: Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38(8), 904–909 (2006)
Wright, M.H., Tung, C.W., Zhao, K., Reynolds, A., McCouch, S.R., Bustamante, C.: ALCHEMY: a reliable method for automated SNP genotype calling for small batch sizes and highly homozygous populations. Bioinformatics 26(23), 2952–2960 (2010)
Yu, J., Pressoir, G., Briggs, W.H., Vroh Bi, I., Yamasaki, M., Doebley, J.F., McMullen, M.D., Gaut, B.S., Nielsen, D.M., Holland, J.B., Kresovich, S., Buckler, E.S.: A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat. Genet. 38(2), 203–208 (2006)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 IFIP International Federation for Information Processing
About this paper
Cite this paper
Baurley, J.W., Pardamean, B., Perbangsa, A.S., Utami, D., Rijzaani, H., Satyawan, D. (2014). A Bioinformatics Workflow for Genetic Association Studies of Traits in Indonesian Rice. In: Linawati, Mahendra, M.S., Neuhold, E.J., Tjoa, A.M., You, I. (eds) Information and Communication Technology. ICT-EurAsia 2014. Lecture Notes in Computer Science, vol 8407. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-55032-4_35
Download citation
DOI: https://doi.org/10.1007/978-3-642-55032-4_35
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-55031-7
Online ISBN: 978-3-642-55032-4
eBook Packages: Computer ScienceComputer Science (R0)