Abstract
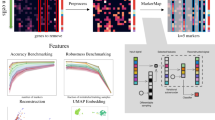
The development of high-throughput technology in genome sequencing provide a large amount of raw data to study the regulatory functions of transcription factors (TFs) on gene expression. It is possible to realize a classifier system in which the gene expression level, under a certain condition, is regarded as the response variable and features related to TFs are taken as predictive variables. In this paper we consider the families of Instance-Based (IB) classifiers, and in particular the Prototype exemplar learning classifier (PEL-C), because IB-classifiers can infer a mixture of representative instances, which can be used to discover the typical epigenetic patterns of transcription factors which explain the gene expression levels. We consider, as case study, the gene regulatory system in mouse embryonic stem cells (ESCs). Experimental results show IB-classifier systems can be effectively used for quantitative modelling of gene expression levels because more than 50% of variation in gene expression can be explained using binding signals of 12 TFs; moreover the PEL-C identifies nine typical patterns of transcription factors activation that provide new insights to understand the gene expression machinery of mouse ESCs.
Chapter PDF
Similar content being viewed by others
References
Soon, W.W., Hariharan, M., Snyder, M.P.: High-throughput sequencing for biology and medicine. Molecular Systems Biology 9, Article number:640 (2013)
Hawkins, R.D., Hon, G.C., Ren, B.: Next-generation genomics: an integrative approach. Nature Review Genetics 11(7), 476–486 (2010)
Ouyanga, Z., Zhoub, Q., Wongc, W.H.: ChIP-Seq of transcription factors predicts absolute and differential gene expression in embryonic stem cells. PNAS 106(51), 21521–21526 (2009)
Young, M.D., Willson, T.A., Wakefield, M.J., Trounson, E., Hilton, D.J., Blewitt, M.E., Oshlack, A., Majewski, I.J.: ChIP-seq analysis reveals distinct H3K27me3 profiles that correlate with transcriptional activity. Nucleic Acids Research 39(17), 7415–7427 (2011)
Witten, I.H., Frank, E.: Data Mining: Practical Machine Learning Tools and Techniques with Java Implementations, 2nd edn. Morgan Kaufmann, San Francisco (2005)
Hastie, T., Tibshirani, R., Friedman, J.: Prototype Methods and Nearest-Neighbors. In: The Elements of Statistical Learning. Data Mining; Inference; and Prediction, 2nd edn., pp. 459–484. Springer, New York (2009)
Fayyad, U., Piatetsky-Shapiro, G., Smyth, P., Uturusamy, R.: Advances in Knowledge Discovery and Data Mining. MIT Press, Cambridge (1996)
Gagliardi, F.: Instance-based classifiers applied to medical databases: diagnosis and knowledge extraction. Artificial Intelligence in Medicine 52(3), 123–139 (2011)
Nieddu, L., Patrizi, G.: Formal methods in pattern recognition: A review. European Journal of Operational Research 120, 459–495 (2000)
Gagliardi, F.: Instance-Based Classifiers to Discover the Gradient of Typicality in Data. In: Pirrone, R., Sorbello, F. (eds.) AI*IA 2011. LNCS, vol. 6934, pp. 457–462. Springer, Heidelberg (2011)
Pavlidis, P., Noble, W.S.: Matrix2png: A Utility for Visualizing Matrix Data. Bioinformatics 19(2), 295–296 (2003)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Gagliardi, F., Angelini, C. (2013). Discovering Typical Transcription-Factors Patterns in Gene Expression Levels of Mouse Embryonic Stem Cells by Instance-Based Classifiers. In: Petrosino, A., Maddalena, L., Pala, P. (eds) New Trends in Image Analysis and Processing – ICIAP 2013. ICIAP 2013. Lecture Notes in Computer Science, vol 8158. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-41190-8_41
Download citation
DOI: https://doi.org/10.1007/978-3-642-41190-8_41
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-41189-2
Online ISBN: 978-3-642-41190-8
eBook Packages: Computer ScienceComputer Science (R0)