Abstract
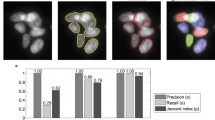
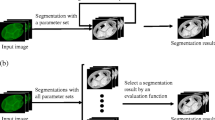
We address the problem of cell segmentation in confocal microscopy membrane volumes of the ascidian Ciona used in the study of morphogenesis. The primary challenges are non-uniform and patchy membrane staining and faint spurious boundaries from other organelles (e.g. nuclei). Traditional segmentation methods incorrectly attach to faint boundaries producing spurious edges. To address this problem, we propose a linear optimization framework for the joint correction of multiple over-segmentations obtained from different methods. The main idea motivating this approach is that multiple over-segmentations, resulting from a pool of methods with various parameters, are likely to agree on the correct segment boundaries, while spurious boundaries are method- or parameter-dependent. The challenge is to make an optimized decision on selecting the correct boundaries while discarding the spurious ones. The proposed unsupervised method achieves better performance than state of the art methods for cell segmentation from membrane images.
Chapter PDF
Similar content being viewed by others
References
Cohen, L., et al.: Fast marching the global min. of active contours. In: ICIP 1996 (1996)
Delibaltov, D., et al.: An automatic feature based model for cell segmentation from confocal microscopy volumes. In: IEEE ISBI (2011)
Delibaltov, D., et al.: Robust biological image sequence analysis using graph based approaches. In: Asilomar Conference on Signals, Systems and Computers (2012)
Grant, C.M., Boyd, S.P.: Graph implementations for nonsmooth convex programs. In: Recent Advances in Learning and Control. LNCIS, vol. 371, pp. 95–110. Springer, Heidelberg (2008)
Grant, M., et al.: CVX: Matlab software for disciplined convex (2011), http://cvxr.com/cvx
Meyer, F., et al.: Morphological segmentation. Journal of Visual Communication and Image Representation (1990)
Sabuncu, M.R., et al.: A generative model for image segmentation based on label fusion. IEEE Trans. on Medical Imaging (2010)
Chen, T., Vemuri, B.C., Rangarajan, A., Eisenschenk, S.J.: Mixture of segmenters with discriminative spatial regularization and sparse weight selection. In: Fichtinger, G., Martel, A., Peters, T. (eds.) MICCAI 2011, Part III. LNCS, vol. 6893, pp. 595–602. Springer, Heidelberg (2011)
Veeman, M., et al.: Chongmague reveals an essential role for laminin-mediated boundary formation in chordate convergence and extension movements. Development Genes and Evolution (2008)
Veeman, M.T., et al.: Whole-organ cell shape analysis reveals the developmental basis of ascidian notochord taper. Developmental Biology (2013)
Warfield, et al.: Simultaneous truth and performance level estimation (STAPLE): an algorithm for the validation of image segmentation. IEEE TMI (2004)
Artaechevarria, X., et al.: Combination strategies in multi-atlas image segmentation: Application to brain MR data. IEEE Trans. on Med. Img. (2009)
Zanella, C., et al.: Cells segmentation from 3-D confocal images Of early zebrafish embryogenesis. In: IEEE TIP 2010 (2010)
Vitaladevuni, S.N., Basri, R.: Co-clustering of image segments using convex optimization applied to EM neuronal reconstruction. In: CVPR 2010 (2010)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Delibaltov, D.L., Ghosh, P., Rodoplu, V., Veeman, M., Smith, W., Manjunath, B.S. (2013). A Linear Program Formulation for the Segmentation of Ciona Membrane Volumes. In: Mori, K., Sakuma, I., Sato, Y., Barillot, C., Navab, N. (eds) Medical Image Computing and Computer-Assisted Intervention – MICCAI 2013. MICCAI 2013. Lecture Notes in Computer Science, vol 8149. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-40811-3_56
Download citation
DOI: https://doi.org/10.1007/978-3-642-40811-3_56
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-40810-6
Online ISBN: 978-3-642-40811-3
eBook Packages: Computer ScienceComputer Science (R0)